Knee Injury Detection Using Deep Learning on MRI Studies: A Systematic Review
Abstract
:1. Introduction
1.1. Backdrop
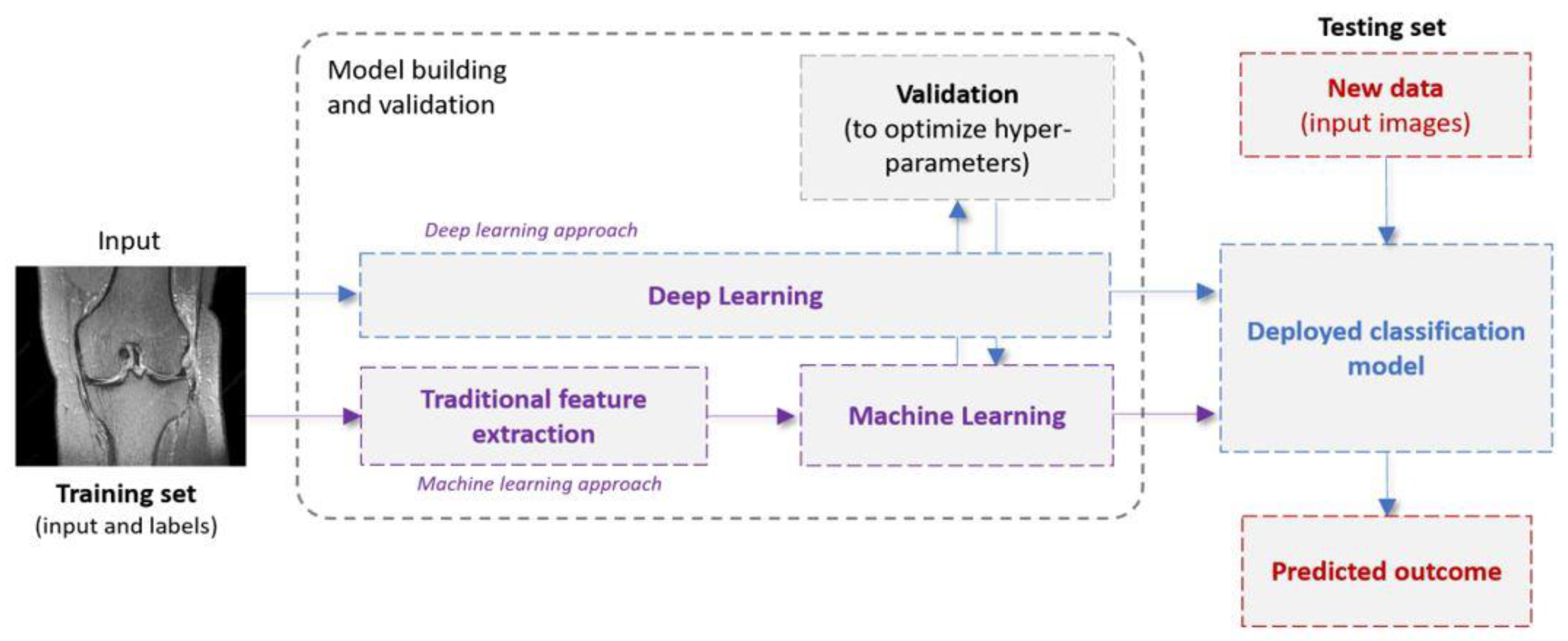
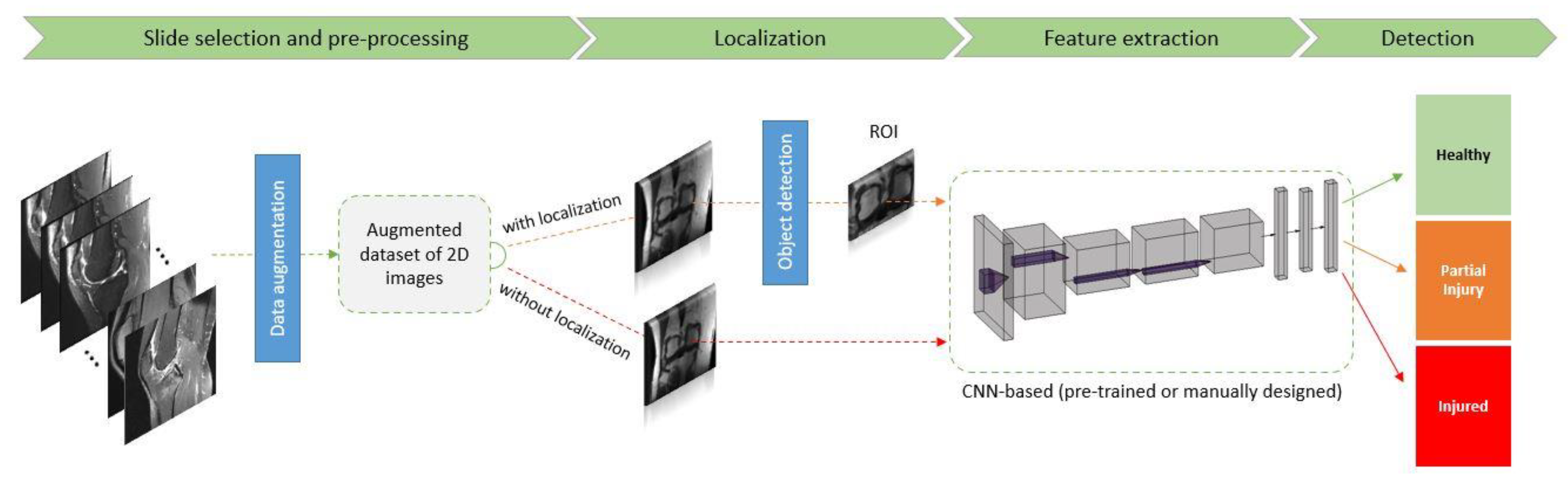
1.2. Machine Learning in a Nutshell: Definitions and Terminology
2. Materials and Methods
2.1. Reporting
2.2. Literature Search
2.3. Eligibility Criteria
2.4. Data Extraction
2.5. Statistical Analysis
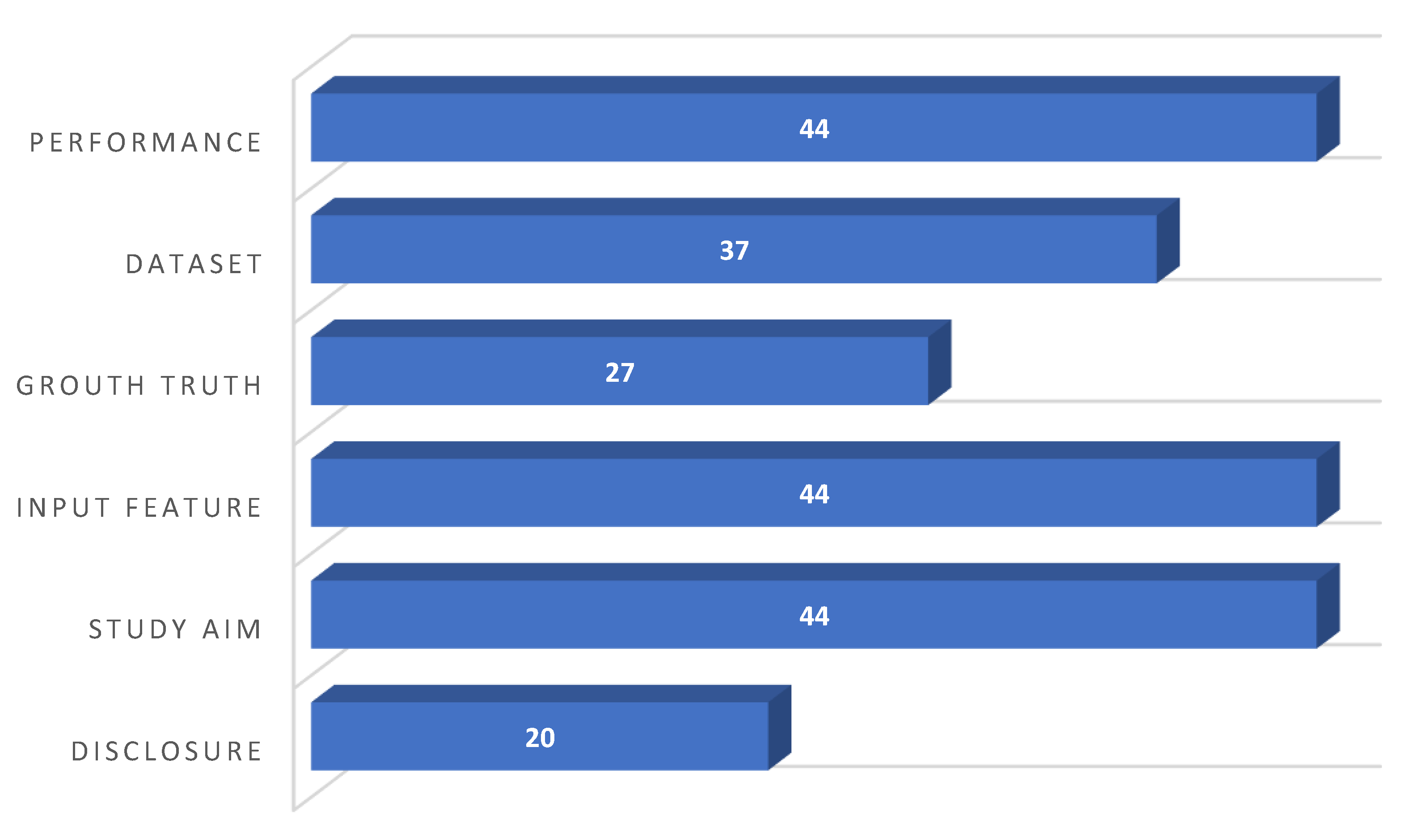
2.6. Quality Assessment
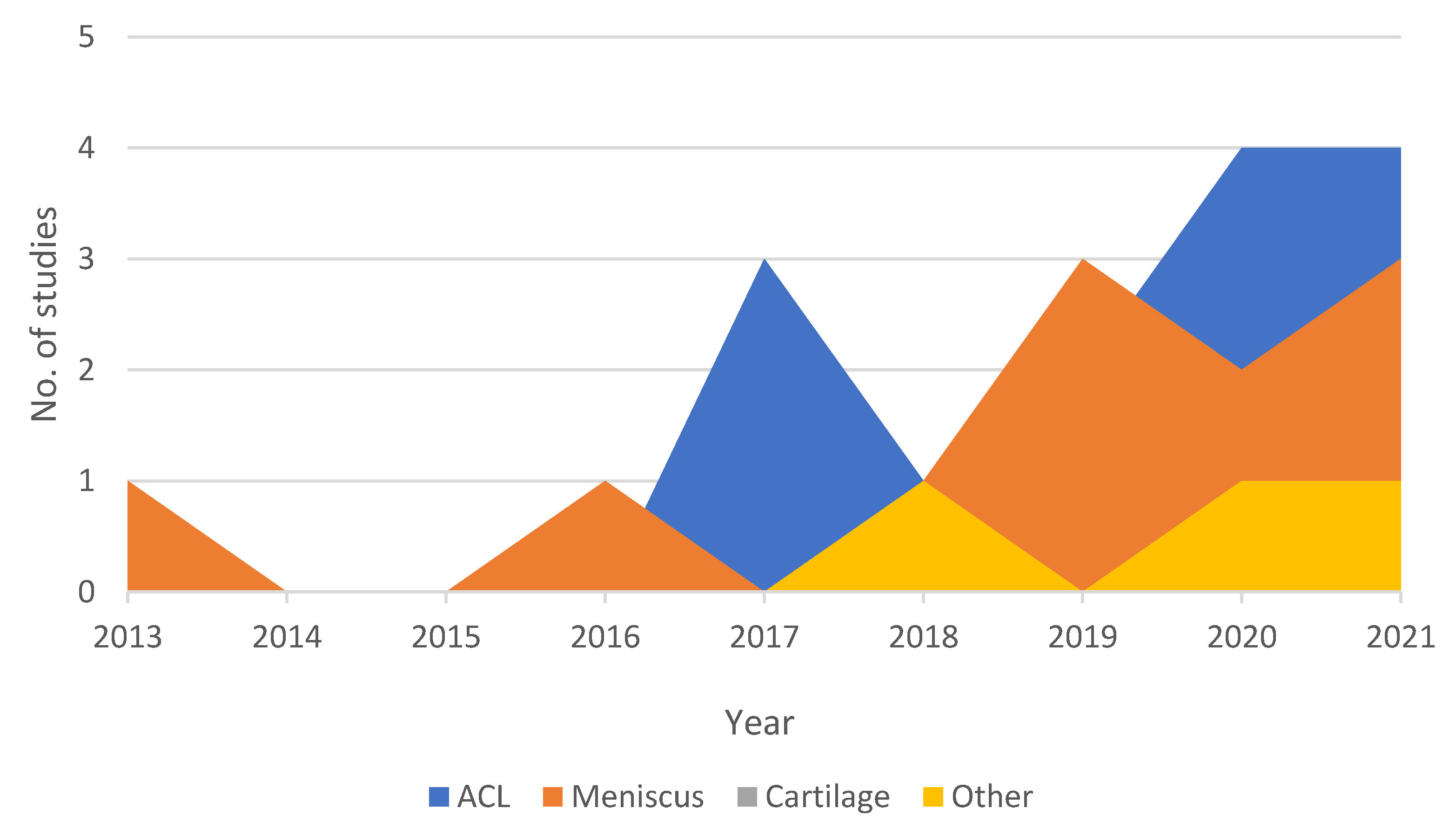
3. Results
3.1. ACL Injury Detection
3.1.1. Machine Learning
3.1.2. Deep Learning with Transfer Learning
3.1.3. Custom-Made Deep-Learning Networks
3.2. Meniscus Tear
3.2.1. Machine Learning
3.2.2. Deep Learning with Transfer Learning
3.2.3. Custom-Made Deep-Learning Networks
3.3. Cartilage Lesion and Other Abnormalities
4. Discussion and Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Musahl, V.; Karlsson, J. Anterior cruciate ligament tear. N. Engl. J. Med. 2019, 380, 2341–2348. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, I.; Bowes, M.; Hutchinson, C.E.; Parsons, N.; Staniszewska, S.; Price, A.J.; Metcalfe, A. Protocol: Meniscal tear outcome Study (METRO Study): A study protocol for a multicentre prospective cohort study exploring the factors which affect outcomes in patients with a meniscal tear. BMJ Open 2020, 10, e038681. [Google Scholar] [CrossRef] [PubMed]
- Darrow, C.J.; Collins, C.L.; Yard, E.E.; Comstock, R.D. Epidemiology of severe injuries among United States high school athletes: 2005–2007. Am. J. Sports Med. 2009, 37, 1798–1805. [Google Scholar] [CrossRef] [PubMed]
- Gage, B.E.; McIlvain, N.M.; Collins, C.L.; Fields, S.K.; Dawn Comstock, R. Epidemiology of 6.6 million knee injuries presenting to United States emergency departments from 1999 through 2008. Acad. Emerg. Med. 2012, 19, 378–385. [Google Scholar] [CrossRef]
- Merkely, G.; Ackermann, J.; Lattermann, C. Articular cartilage defects: Incidence, diagnosis, and natural history. Oper. Tech. Sports Med. 2018, 26, 156–161. [Google Scholar] [CrossRef]
- Zhang, L.; Li, M.; Zhou, Y.; Lu, G.; Zhou, Q. Deep Learning Approach for Anterior Cruciate Ligament Lesion Detection: Evaluation of Diagnostic Performance Using Arthroscopy as the Reference Standard. J. Magn. Reson. Imaging 2020, 52, 1745–1752. [Google Scholar] [CrossRef]
- Kaeding, C.C.; Léger-St-Jean, B.; Magnussen, R.A. Epidemiology and diagnosis of anterior cruciate ligament injuries. Clin. Sports Med. 2017, 36, 1–8. [Google Scholar] [CrossRef]
- Chang, P.D.; Wong, T.T.; Rasiej, M.J. Deep Learning for Detection of Complete Anterior Cruciate Ligament Tear. J. Digit. Imaging 2019, 32, 980–986. [Google Scholar] [CrossRef]
- Logerstedt, D.S.; Snyder-Mackler, L.; Ritter, R.C.; Axe, M.J.; Godges, J.; Altman, R.D.; Briggs, M.; Chu, C.; Delitto, A.; Ferland, A. Knee pain and mobility impairments: Meniscal and articular cartilage lesions: Clinical practice guidelines linked to the international classification of functioning, disability, and health from the orthopaedic section of the American Physical Therapy Association. J. Orthop. Sports Phys. Ther. 2010, 40, A1–A35. [Google Scholar]
- Mather, R.C., III; Koenig, L.; Kocher, M.S.; Dall, T.M.; Gallo, P.; Scott, D.J.; Bach Jr, B.R.; Spindler, K.P.; Group, M.K. Societal and economic impact of anterior cruciate ligament tears. J. Bone Jt. Surg. Am. Vol. 2013, 95, 1751. [Google Scholar] [CrossRef] [Green Version]
- Cameron, K.L.; Thompson, B.S.; Peck, K.Y.; Owens, B.D.; Marshall, S.W.; Svoboda, S.J. Normative values for the KOOS and WOMAC in a young athletic population: History of knee ligament injury is associated with lower scores. Am. J. Sports Med. 2013, 41, 582–589. [Google Scholar] [CrossRef] [PubMed]
- Huffman, G.R.; Park, J.; Roser-Jones, C.; Sennett, B.J.; Yagnik, G.; Webner, D. Normative SF-36 values in competing NCAA intercollegiate athletes differ from values in the general population. JBJS 2008, 90, 471–476. [Google Scholar] [CrossRef] [PubMed]
- Lam, K.C.; Thomas, S.S.; Valier, A.R.S.; McLeod, T.C.V.; Bay, R.C. Previous knee injury and health-related quality of life in collegiate athletes. J. Athl. Train. 2017, 52, 534–540. [Google Scholar] [CrossRef] [PubMed]
- Pouly, M.; Koller, T.; Gottfrois, P.; Lionetti, S. Artificial intelligence in image analysis-fundamentals and new developments. Der Hautarzt Z. Fur Dermatol. Venerol. Und Verwandte Geb. 2020, 71, 660–668. [Google Scholar] [CrossRef] [PubMed]
- Fritz, B.; Marbach, G.; Civardi, F.; Fucentese, S.F.; Pfirrmann, C.W. Deep convolutional neural network-based detection of meniscus tears: Comparison with radiologists and surgery as standard of reference. Skelet. Radiol. 2020, 49, 1207–1217. [Google Scholar] [CrossRef]
- Garwood, E.R.; Tai, R.; Joshi, G. The Use of Artificial Intelligence in the Evaluation of Knee Pathology. In Seminars in Musculoskeletal Radiology; Thieme Medical Publishers: New York, NY, USA, 2020; pp. 21–29. [Google Scholar]
- Palermi, S.; Massa, B.; Vecchiato, M.; Mazza, F.; De Blasiis, P.; Romano, A.M.; Di Salvatore, M.G.; Della Valle, E.; Tarantino, D.; Ruosi, C.J.J.O.F.M.; et al. Indirect Structural Muscle Injuries of Lower Limb: Rehabilitation and Therapeutic Exercise. J. Funct. Morphol. Kinesiol. 2021, 6, 75. [Google Scholar] [CrossRef]
- Sirico, F.; Palermi, S.; Massa, B.; Corrado, B. Tendinopathies of the hip and pelvis in athletes: A narrative review. J. Hum. Sports Exerc. 2020, 15, S748–S762. [Google Scholar]
- Hetsroni, I.; Lyman, S.; Do, H.; Mann, G.; Marx, R. Symptomatic pulmonary embolism after outpatient arthroscopic procedures of the knee: The incidence and risk factors in 418 323 arthroscopies. J. Bone Jt. Surg. Br. Vol. 2011, 93, 47–51. [Google Scholar] [CrossRef] [Green Version]
- Alanazi, H.O.; Abdullah, A.H.; Qureshi, K.N. A critical review for developing accurate and dynamic predictive models using machine learning methods in medicine and health care. J. Med. Syst. 2017, 41, 69. [Google Scholar] [CrossRef]
- Prickett, W.D.; Ward, S.I.; Matava, M.J. Magnetic resonance imaging of the knee. Sports Med. 2001, 31, 997–1019. [Google Scholar] [CrossRef]
- Krampla, W.; Roesel, M.; Svoboda, K.; Nachbagauer, A.; Gschwantler, M.; Hruby, W. MRI of the knee: How do field strength and radiologist’s experience influence diagnostic accuracy and interobserver correlation in assessing chondral and meniscal lesions and the integrity of the anterior cruciate ligament? Eur. Radiol. 2009, 19, 1519–1528. [Google Scholar] [CrossRef] [PubMed]
- Mohankumar, R.; White, L.M.; Naraghi, A. Pitfalls and pearls in MRI of the knee. Am. J. Roentgenol. 2014, 203, 516–530. [Google Scholar] [CrossRef] [PubMed]
- Germann, C.; Marbach, G.; Civardi, F.; Fucentese, S.F.; Fritz, J.; Sutter, R.; Pfirrmann, C.W.; Fritz, B. Deep Convolutional Neural Network–Based Diagnosis of Anterior Cruciate Ligament Tears: Performance Comparison of Homogenous Versus Heterogeneous Knee MRI Cohorts With Different Pulse Sequence Protocols and 1.5-T and 3-T Magnetic Field Strengths. Investig. Radiol. 2020, 55, 499. [Google Scholar] [CrossRef] [PubMed]
- Gyftopoulos, S.; Lin, D.; Knoll, F.; Doshi, A.M.; Rodrigues, T.C.; Recht, M.P. Artificial intelligence in musculoskeletal imaging: Current status and future directions. Am. J. Roentgenol. 2019, 213, 506–513. [Google Scholar] [CrossRef] [PubMed]
- Shen, D.; Wu, G.; Suk, H.-I. Deep learning in medical image analysis. Annu. Rev. Biomed. Eng. 2017, 19, 221–248. [Google Scholar] [CrossRef] [Green Version]
- Bien, N.; Rajpurkar, P.; Ball, R.L.; Irvin, J.; Park, A.; Jones, E.; Bereket, M.; Patel, B.N.; Yeom, K.W.; Shpanskaya, K. Deep-learning-assisted diagnosis for knee magnetic resonance imaging: Development and retrospective validation of MRNet. PLoS Med. 2018, 15, e1002699. [Google Scholar] [CrossRef]
- Langerhuizen, D.W.; Janssen, S.J.; Mallee, W.H.; Van Den Bekerom, M.P.; Ring, D.; Kerkhoffs, G.M.; Jaarsma, R.L.; Doornberg, J.N. What are the applications and limitations of artificial intelligence for fracture detection and classification in orthopaedic trauma imaging? A systematic review. Clin. Orthop. Relat. Res. 2019, 477, 2482. [Google Scholar] [CrossRef]
- Kunze, K.N.; Rossi, D.M.; White, G.M.; Karhade, A.V.; Deng, J.; Williams, B.T.; Chahla, J. Diagnostic Performance of Artificial Intelligence for Detection of Anterior Cruciate Ligament and Meniscus Tears: A Systematic Review. Arthrosc. J. Arthrosc. Relat. Surg. 2020, 37, 771–781. [Google Scholar] [CrossRef]
- Hellerstein, J.M.; Naughton, J.F.; Pfeffer, A. Generalized Search Trees for Database Systems. In Proceedings of the 21st VLDB Conference, Zurich, Switzerland, 11–15 September 1995. [Google Scholar]
- Liu, F. Susan: Segment unannotated image structure using adversarial network. Magn. Reson. Med. 2019, 81, 3330–3345. [Google Scholar] [CrossRef]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 4700–4708. [Google Scholar]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. Imagenet classification with deep convolutional neural networks. Adv. Neural Inf. Processing Syst. 2012, 25, 1097–1105. [Google Scholar] [CrossRef]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Freeman, W.T.; Roth, M. Orientation histograms for hand gesture recognition. In Proceedings of the International Workshop on Automatic Face and Gesture Recognition, Zurich, Switzerland, 26–28 June 1995; pp. 296–301. [Google Scholar]
- Sebastian, V.B.; Unnikrishnan, A.; Balakrishnan, K. Gray level co-occurrence matrices: Generalisation and some new features. arXiv 2012, arXiv:1205.4831. [Google Scholar]
- Peterson, L.E. K-nearest neighbor. Scholarpedia 2009, 4, 1883. [Google Scholar] [CrossRef]
- Hearst, M.A.; Dumais, S.T.; Osuna, E.; Platt, J.; Scholkopf, B. Support vector machines. IEEE Intell. Syst. Appl. 1998, 13, 18–28. [Google Scholar] [CrossRef] [Green Version]
- Hassoun, M.H. Fundamentals of Artificial Neural Networks; MIT Press: Cambridge, MA, USA, 1995. [Google Scholar]
- O’Shea, K.; Nash, R. An introduction to convolutional neural networks. arXiv 2015, arXiv:1511.08458. [Google Scholar]
- Girshick, R.; Donahue, J.; Darrell, T.; Malik, J. Rich feature hierarchies for accurate object detection and semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Columbus, OH, USA, 23–28 June 2014; pp. 580–587. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Kamnitsas, K.; Ledig, C.; Newcombe, V.F.; Simpson, J.P.; Kane, A.D.; Menon, D.K.; Rueckert, D.; Glocker, B. Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med. Image Anal. 2017, 36, 61–78. [Google Scholar] [CrossRef]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S.J.A.P.A. An image is worth 16×16 words: Transformers for image recognition at scale. arXiv 2020, arXiv:2010.11929. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- Moher, D.; Liberati, A.; Tetzlaff, J.; Altman, D.G.; Altman, D.; Antes, G.; Atkins, D.; Barbour, V.; Barrowman, N.; Berlin, J.A. Preferred reporting items for systematic reviews and meta-analyses: The PRISMA statement (Chinese edition). J. Chin. Integr. Med. 2009, 7, 889–896. [Google Scholar] [CrossRef]
- Slim, K.; Nini, E.; Forestier, D.; Kwiatkowski, F.; Panis, Y.; Chipponi, J.J.A.J.O.S. Methodological index for non-randomized studies (MINORS): Development and validation of a new instrument. ANZ J. Surg. 2003, 73, 712–716. [Google Scholar] [CrossRef]
- Štajduhar, I.; Mamula, M.; Miletić, D.; Ünal, G. Semi-automated detection of anterior cruciate ligament injury from MRI. Comput. Methods Programs Biomed. 2017, 140, 151–164. [Google Scholar] [CrossRef] [PubMed]
- Pal, M. Random forest classifier for remote sensing classification. Int. J. Remote Sens. 2005, 26, 217–222. [Google Scholar] [CrossRef]
- Abdullah, A.A.; Azz-Zahra Md Som, N.S.F. Design of an Intelligent Diagnostic System for Detection of Knee Injuries. Appl. Mech. Mater. 2013, 399, 219–224. [Google Scholar] [CrossRef]
- Mazlan, S.S.; Ayob, M.; Bakti, Z.K. Anterior cruciate ligament (ACL) injury classification system using support vector machine (SVM). In Proceedings of the 2017 International Conference on Engineering Technology and Technopreneurship (ICE2T), Kuala Lumpur, Malaysia, 18–20 September 2017; pp. 1–5. [Google Scholar]
- Azcona, D.; McGuinness, K.; Smeaton, A.F. A Comparative Study of Existing and New Deep Learning Methods for Detecting Knee Injuries using the MRNet Dataset. In Proceedings of the 2020 International Conference on Intelligent Data Science Technologies and Applications (IDSTA), Kuala Lumpur, Malaysia, 18–20 September 2020; pp. 149–155. [Google Scholar]
- Liu, F.; Guan, B.; Zhou, Z.; Samsonov, A.; Rosas, H.; Lian, K.; Sharma, R.; Kanarek, A.; Kim, J.; Guermazi, A. Fully automated diagnosis of anterior cruciate ligament tears on knee MR images by using deep learning. Radiol. Artif. Intell. 2019, 1, 180091. [Google Scholar] [CrossRef]
- Namiri, N.K.; Flament, I.; Astuto, B.; Shah, R.; Tibrewala, R.; Caliva, F.; Link, T.M.; Pedoia, V.; Majumdar, S. Deep Learning for Hierarchical Severity Staging of Anterior Cruciate Ligament Injuries from MRI. Radiol. Artif. Intell. 2020, 2, e190207. [Google Scholar] [CrossRef]
- Awan, M.J.; Rahim, M.S.M.; Salim, N.; Mohammed, M.A.; Garcia-Zapirain, B.; Abdulkareem, K.H. Efficient detection of knee anterior cruciate ligament from magnetic resonance imaging using deep learning approach. Diagnostics 2021, 11, 105. [Google Scholar] [CrossRef]
- Jeon, Y.S.; Yoshino, K.; Hagiwara, S.; Watanabe, A.; Quek, S.T.; Yoshioka, H.; Feng, M.J.I.J.O.B.; Informatics, H. Interpretable and lightweight 3-D deep learning model for automated ACL diagnosis. IEEE J. Biomed. Health Inform. 2021, 25, 2388–2397. [Google Scholar] [CrossRef]
- Dai, Y.; Gao, Y.; Liu, F.J.D. Transmed: Transformers advance multi-modal medical image classification. Diagnostics 2021, 11, 1384. [Google Scholar] [CrossRef]
- Astuto, B.; Flament, I.K.; Namiri, N.; Shah, R.; Bharadwaj, U.; M. Link, T.; D. Bucknor, M.; Pedoia, V.; Majumdar, S.J.R.A.I. Automatic Deep Learning–assisted Detection and Grading of Abnormalities in Knee MRI Studies. Radiol. Artif. Intell. 2021, 3, e200165. [Google Scholar] [CrossRef]
- Fu, J.-C.; Lin, C.-C.; Wang, C.-N.; Ou, Y.-K. Computer-aided diagnosis for knee meniscus tears in magnetic resonance imaging. J. Ind. Prod. Eng. 2013, 30, 67–77. [Google Scholar] [CrossRef]
- Zarandi, M.F.; Khadangi, A.; Karimi, F.; Turksen, I. A computer-aided type-II fuzzy image processing for diagnosis of meniscus tear. J. Digit. Imaging 2016, 29, 677–695. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Couteaux, V.; Si-Mohamed, S.; Nempont, O.; Lefevre, T.; Popoff, A.; Pizaine, G.; Villain, N.; Bloch, I.; Cotten, A.; Boussel, L. Automatic knee meniscus tear detection and orientation classification with Mask-RCNN. Diagn. Interv. Imaging 2019, 100, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Roblot, V.; Giret, Y.; Antoun, M.B.; Morillot, C.; Chassin, X.; Cotten, A.; Zerbib, J.; Fournier, L. Artificial intelligence to diagnose meniscus tears on MRI. Diagn. Interv. Imaging 2019, 100, 243–249. [Google Scholar] [CrossRef] [PubMed]
- Pedoia, V.; Norman, B.; Mehany, S.N.; Bucknor, M.D.; Link, T.M.; Majumdar, S. 3D convolutional neural networks for detection and severity staging of meniscus and PFJ cartilage morphological degenerative changes in osteoarthritis and anterior cruciate ligament subjects. J. Magn. Reson. Imaging 2019, 49, 400–410. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Munich, Germany, 5–9 October 2015; pp. 234–241. [Google Scholar]
- Rizk, B.; Brat, H.; Zille, P.; Guillin, R.; Pouchy, C.; Adam, C.; Ardon, R.; d’Assignies, G.J.P.M. Meniscal lesion detection and characterization in adult knee MRI: A deep learning model approach with external validation. Phys. Med. 2021, 83, 64–71. [Google Scholar] [CrossRef]
- Liu, F.; Zhou, Z.; Samsonov, A.; Blankenbaker, D.; Larison, W.; Kanarek, A.; Lian, K.; Kambhampati, S.; Kijowski, R. Deep learning approach for evaluating knee MR images: Achieving high diagnostic performance for cartilage lesion detection. Radiology 2018, 289, 160–169. [Google Scholar] [CrossRef]
- Targ, S.; Almeida, D.; Lyman, K. Resnet in resnet: Generalizing residual architectures. arXiv 2016, arXiv:1603.08029. [Google Scholar]
- LeCun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proc. IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef] [Green Version]
- Zheng, S.; Lu, J.; Zhao, H.; Zhu, X.; Luo, Z.; Wang, Y.; Fu, Y.; Feng, J.; Xiang, T.; Torr, P.H. Rethinking semantic segmentation from a sequence-to-sequence perspective with transformers. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Online, 22–25 November 2021; pp. 6881–6890. [Google Scholar]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention is all you need. In Proceedings of the Advances in Neural Information Processing Systems, Long Beach, CA, USA, 4–9 December 2017; pp. 5998–6008. [Google Scholar]
- Devlin, J.; Chang, M.-W.; Lee, K.; Toutanova, K.J.A.P.A. Bert: Pre-training of deep bidirectional transformers for language understanding. arXiv 2018, arXiv:1810.04805. [Google Scholar]
- Brown, T.B.; Mann, B.; Ryder, N.; Subbiah, M.; Kaplan, J.; Dhariwal, P.; Neelakantan, A.; Shyam, P.; Sastry, G.; Askell, A.J.A.P.A. Language models are few-shot learners. Adv. Neural Inf. Process. Syst. 2020, 33, 1877–1901. [Google Scholar]
- Kouw, W.M.; Loog, M.; Bartels, L.W.; Mendrik, A.M.J.A.P.A. MR acquisition-invariant representation learning. arXiv 2017, arXiv:1709.07944. [Google Scholar]
- Billot, B.; Greve, D.; Van Leemput, K.; Fischl, B.; Iglesias, J.E.; Dalca, A.V.J.A.P.A. A learning strategy for contrast-agnostic MRI segmentation. arXiv 2020, arXiv:2003.01995. [Google Scholar]

| Category | Models | Description |
|---|---|---|
| Feature extraction | Histogram of oriented gradient (HOG) [35] | This is a feature descriptor used in computer vision and image processing for the purpose of object detection. The technique counts occurrences of gradient orientation in localized portions of an image. |
| Generalized search tree (GIST) [30] | GIST descriptor represents holistic spatial scene properties (spatial envelope) of an image. It summarizes gradient information on different spatial scales and orientations by splitting the image into a grid of cells on several scales and convolving each cell using a Gabor filter bank from different perspectives. | |
| Gray-level co-occurrence matrix (GLCM) [36] | GLCM is a way of extracting second-order statistical texture features. In particular, the texture of an image is estimated by calculating how often pairs of pixels with specific values and a certain spatial relationship occur. | |
| Traditional Machine Learning | k-nearest neighbor (K-NN) [37] | KNN algorithm is a simple, easy-to-implement supervised ML algorithm that can be used to solve both classification and regression problems. It works by (i) finding the distances between a query and all the examples in the data, (ii) selecting the K nearest neighbors of the query, and (iii) voting for the most frequent label (in the case of classification) or averaging the labels (in the case of regression). |
| Support vector machines (SVMs) [38] | SVMs is a supervised method that identifies a hyperplane that best divides the data into two classes. To separate the two clouds of data points, there are many possible hyperplanes that could be chosen. The objective of the SVM algorithm is to find a slab that has the maximum thickness, i.e., the maximum distance between data points of the different classes. | |
| Shallow artificial neural networks (ANNs) [39] | The ANN vaguely simulates the way the human brain analyzes and processes information. They consist of sequential layers: input, hidden and output layers. The hidden layer processes and transmits the input information to the output layer. | |
| Deep Learning | Convolutional neural networks (CNNs) [40] | This is a class of DL algorithms commonly used in computer vision and pattern recognition. CNNs are a specific type of neural networks that are generally composed of the following layers: (i) input layer, (ii) convolution layers, (iii) pooling layers and (iv) fully connected layers. The convolution layers use filters that perform convolution operations as they are scanning the input with respect to its dimensions. Pooling is a down-sampling operation, which is typically applied after a convolution layer. The fully connected layers operate on a flattened input where each input is connected to all neurons in the next layer and are usually found towards the end of CNN architectures to optimize objectives such as class scores. |
| Region based convolutional neural networks (R-CNNs) [41] | The method of detecting and classifying objects in an image is known as object detection. R-CNN (regions with convolutional neural networks) is a deep learning technique that blends rectangular area proposals with convolutional neural network functionality. The R-CNN algorithm is a two-stage detection method. | |
| Deep residual networks [42] | A residual neural network (ResNet) is an ANN variant that uses residual mapping and shortcut connections to tackle the problem of vanishing and exploding gradients that is characteristic of deep CNNs. As a consequence of this, deep residual networks achieve better performance when compared to plain very deep networks, whereas their training is easier as well. Typical ResNet models are implemented with double- or triple-layer skips that contain nonlinearities such as rectified linear unit (ReLUs) and batch normalization in between. | |
| 3D-CNNs [43] | A 3D CNN is simply the 3D generalization of 2D CNNs. It takes as input a 3D volume or a sequence of 2D frames (e.g., slices in an MRI scan). Then kernels move through 3 dimensions of data producing 3D activation maps. Overall, they learn powerful representations of volumetric data. | |
| Computer Vision Transformers [44] | When data is modelized as a sequence of embeddings, the Transformer model is a basic yet scalable technique that can be used for any type of data. Even without typical convolutional pipelines, transformers can be utilized to provide SOTA results in Computer Vision. It is a DL network that extracts inherent properties of the interest domain via the self-attention technique. | |
| Procedure | Training | The standard procedure involves a dataset of paired images and labels (x, y) for training and testing, an optimizer (e.g., stochastic gradient descent, Adam [45]), and a loss function to update the model parameters. The aim of the training is to find the optimal values for the network parameters so that the loss function is minimized. |
| Data augmentation | Data augmentation is a strategy that artificially generates more training samples to increase the diversity of the training data. This can be done via applying affine transformations (e.g., rotation, scaling), flipping or cropping to original labeled samples. | |
| Dropout | Dropout is a regularization method that randomly drops some units from the neural network during training, encouraging the network to learn a sparse representation. It is used to reduce overfitting. | |
| Loss function | The metric to assess the discrepancy between model predictions and labels is called loss function. The gradients of the loss function are used to update the weights of the neural networks. | |
| Transfer learning | This aims to transfer knowledge from one task to another different but related target task. This is often achieved by reusing the weights of a pre-trained model, to initialize the weights in a new model for the target task. Transfer learning can help to decrease the training time and achieve lower generalization error. |
| No. | Author | Year | AI Model Used | Pretrained CNN | MRI (T) | Localization Technique | Validation | Performance (Accuracy/AUC) | Application Domain |
|---|---|---|---|---|---|---|---|---|---|
| 1 | Awan et al. [55] | 2021 | CNN | ResNet-14 | 1.5 T | They applied normal approach to localize based upon region of interest (ROI) | 5-fold cross-validation | 92%/(healthy tear = 0.98, partial tear = 0.97 and fully ruptured tear = 0.99) | ACL tear |
| 2 | Jeon et al. [56] | 2021 | 3D CNN | VGGNet, AlexNet, and SqueezeNet | 3 T & 1.5 T | Custom localization technique | 5-fold cross-validation | N/A/0.983 and 0.980 on the Chiba and Stanford knee datasets, respectively | ACL tear |
| 3 | Rizk et al. [65] | 2021 | 3D CNN | CNN-based localization model | 1 T (54%)–1.5 T (9.7%)–3 T (36.3%) | Custom localization technique | ten-fold cross validation | Meidal = N/A/0.93, Lateral = N/A/0.84 | Meniscus tear |
| 4 | Dai et al. [57] | 2021 | TransMed | N/A | 3 T & 1.5 T | N/A | 120 exams | ACL tear = 94.9%/0.98, Abnormality = 91.8%/0.976, Meniscus tear = 85.3%/0.95 | ACL tear—Meniscus tear—Abnormalities |
| 5 | Astuto et al. [58] | 2021 | 3D CNN | N/A | 3 T | V-Net | Hold out (15% of sample) | N/A/from 0.83 to 0.93 | ACL tear—Meniscus tear—Cartilage Lession |
| 6 | Fritz et al. [15] | 2020 | DCNN | N/A | 1.5 T (64%)–3 T (36%) | To visually localize the tear, the software computes the class activation map (CAM) of the last convolution layer in the CNN and maps it to an axial knee image | Hold out (10% of sample) | Medial = (86%/0.88), Lateral = (84%/0.78), Overall = (N/A/0.96) | Meniscus tear |
| 7 | Namiri et al. [54] | 2020 | CNN | N/A | 3 T | three-dimensional V-Net | Hold out (10% of sample) | 3D-model = (89%/sensitivity of 89% and specificity of 88%), 2D-model = (92%/sensitivity of 93% and specificity of 90%) | ACL tear |
| 8 | Zhang et al. [6] | 2020 | CNN | 3D DenseNet, VGG16, ResNet | 1.5 T (74%)–3 T (26%) | - | Hold out (20% of sample) | Custom = (95.7%/0.96), ResNet = (NA/0.95), VGG16 = (NA/0.86) | ACL tear |
| 9 | Germann et al. [24] | 2020 | DCNN | N/A | 1.5 T–3 T | They cropped manually | Out of the 5802 MRI studies, 4802 were used for training, 500 for validation, and 500 for initial testing | N/A/0.94 | ACL tear |
| 10 | Azcona et al. [52] | 2020 | CNN | MRNet, ResNet18, Resnet50 and ResNet152, ImageNet | 3 T (56.6%)–1.5 T (43.4%) | - | N/A | NA/0.96–N/A/0.91–N/A/0.94 | ACL tear—Meniscus tear—Abnormalities |
| 11 | Chang et al. [8] | 2019 | CNN | ResNet | 1.5 T–3 T | The object localization CNN was implemented as a fully convolutional network based on U-net architecture | 5-fold-cross-validation | 96.7%/0.97 | ACL tear |
| 12 | Liu et al. [53] | 2019 | CNN | LeNet-5, DenseNet, VGG16, AlexNet | N/A | They used object detection technique YOLO | 50 subjects test set (14% of the sample) | N/A/0.98 | ACL tear |
| 13 | Couteaux et al. [61] | 2019 | CNN | ResNet-101, ConvNet, R-CNN | N/A | To localize both menisci and identify tears in each meniscus, they used the Mask R-CNN framework | 54 cases and the model with the highest validation accuracy was selected | N/A/0.90 | Meniscus tear |
| 14 | Pedoia et al. [63] | 2019 | 2D U-Net, CNN | N/A | 3 T | - | Hold out (20% of sample) | Sensitivity of 89.81% and specificity of 81.98% | Meniscus tear |
| 15 | Roblot et al. [62] | 2019 | CNN | AlexNet, MRNet | N/A | They used object detection technique Fast RCNN & Faster RCNN | The algorithm was thus used on a test dataset composed of 700 images for external validation | 72.5%/0.85 | Meniscus tear |
| 16 | Nicholas Bien et al. [27] | 2018 | CNN | AlexNET, MRNet | 3 T (56.6%)–1.5 T (43.4%) | - | 120 exams | 86.7%/0.97–72.5%/0.85–N/A/0.94 | ACL tear—Meniscus tear—Abnormalities |
| 17 | Liu et al. [66] | 2018 | CNN | VGG16 | 3 T | - | fellowship trained musculoskeletal radiologist (R.K., with 15 years of clinical experience) | N/A/0.92 | Cartilage lesion |
| 18 | Stajduhar et al. [48] | 2017 | HOG + linSVM, HOG + RF, GIST + rbfSVM, GIST + RF | N/A | 1.5 T | Manual extraction of a rectangular ROI | 10-fold cross validation | (Injury detection problem, complete rupture) = (N/A/0.89, N/A/0.94), (N/A/0.88, N/A/0.94), (N/A/0.889, N/A/0.91), (N/A/0.88, N/A/0.90) respectively with the models | ACL tear |
| 19 | Mazlan et al. [51] | 2017 | SVM | N/A | N/A | They use cropping technique | Hold out (10% of sample) | 100%/N/A | ACL tear |
| 20 | Zarandi et al. [60] | 2016 | IT2FCM, PNN | N/A | N/A | - | Hold out (20% of sample) | 0 and 1 mode: 90%/N/A Binary mode: 78%/N/A | Meniscus tear |
| 21 | Fu et al. [59] | 2013 | SVM | N/A | N/A | Active Contours without Edges method. This method combines Active Contours with Level Sets and is called ACLS | 5-Fold cross validation | SVM model: N/A/0.73 SFFS + SVM: N/A/0.91 | Meniscus tear |
| 22 | Abdullah et al. [50] | 2013 | BP ANN, K-NN | N/A | N/A | - | 5-fold and 6-fold | BP ANN: 94.44%/N/A k-NN: 87.83%/N/A | ACL tear |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Siouras, A.; Moustakidis, S.; Giannakidis, A.; Chalatsis, G.; Liampas, I.; Vlychou, M.; Hantes, M.; Tasoulis, S.; Tsaopoulos, D. Knee Injury Detection Using Deep Learning on MRI Studies: A Systematic Review. Diagnostics 2022, 12, 537. https://doi.org/10.3390/diagnostics12020537
Siouras A, Moustakidis S, Giannakidis A, Chalatsis G, Liampas I, Vlychou M, Hantes M, Tasoulis S, Tsaopoulos D. Knee Injury Detection Using Deep Learning on MRI Studies: A Systematic Review. Diagnostics. 2022; 12(2):537. https://doi.org/10.3390/diagnostics12020537
Chicago/Turabian StyleSiouras, Athanasios, Serafeim Moustakidis, Archontis Giannakidis, Georgios Chalatsis, Ioannis Liampas, Marianna Vlychou, Michael Hantes, Sotiris Tasoulis, and Dimitrios Tsaopoulos. 2022. "Knee Injury Detection Using Deep Learning on MRI Studies: A Systematic Review" Diagnostics 12, no. 2: 537. https://doi.org/10.3390/diagnostics12020537
APA StyleSiouras, A., Moustakidis, S., Giannakidis, A., Chalatsis, G., Liampas, I., Vlychou, M., Hantes, M., Tasoulis, S., & Tsaopoulos, D. (2022). Knee Injury Detection Using Deep Learning on MRI Studies: A Systematic Review. Diagnostics, 12(2), 537. https://doi.org/10.3390/diagnostics12020537










