Proposal to Improve the Image Quality of Short-Acquisition Time-Dedicated Breast Positron Emission Tomography Using the Pix2pix Generative Adversarial Network
Abstract
:1. Introduction
2. Materials and Methods
2.1. Participants
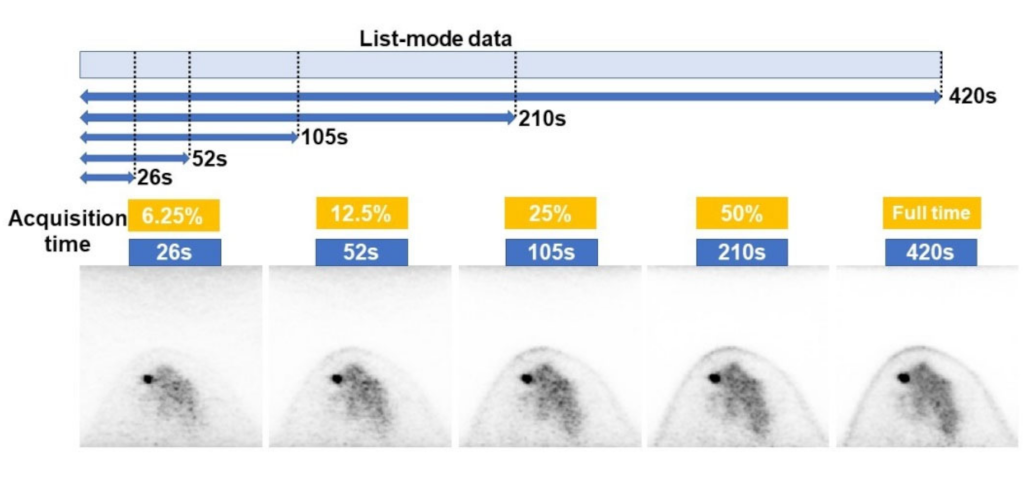
2.2. Imaging Protocol
2.3. Final Diagnoses
2.4. Data Set
2.5. Model Description and Training Protocol
2.6. Evaluation of Models
2.7. Statistical Analysis
3. Results
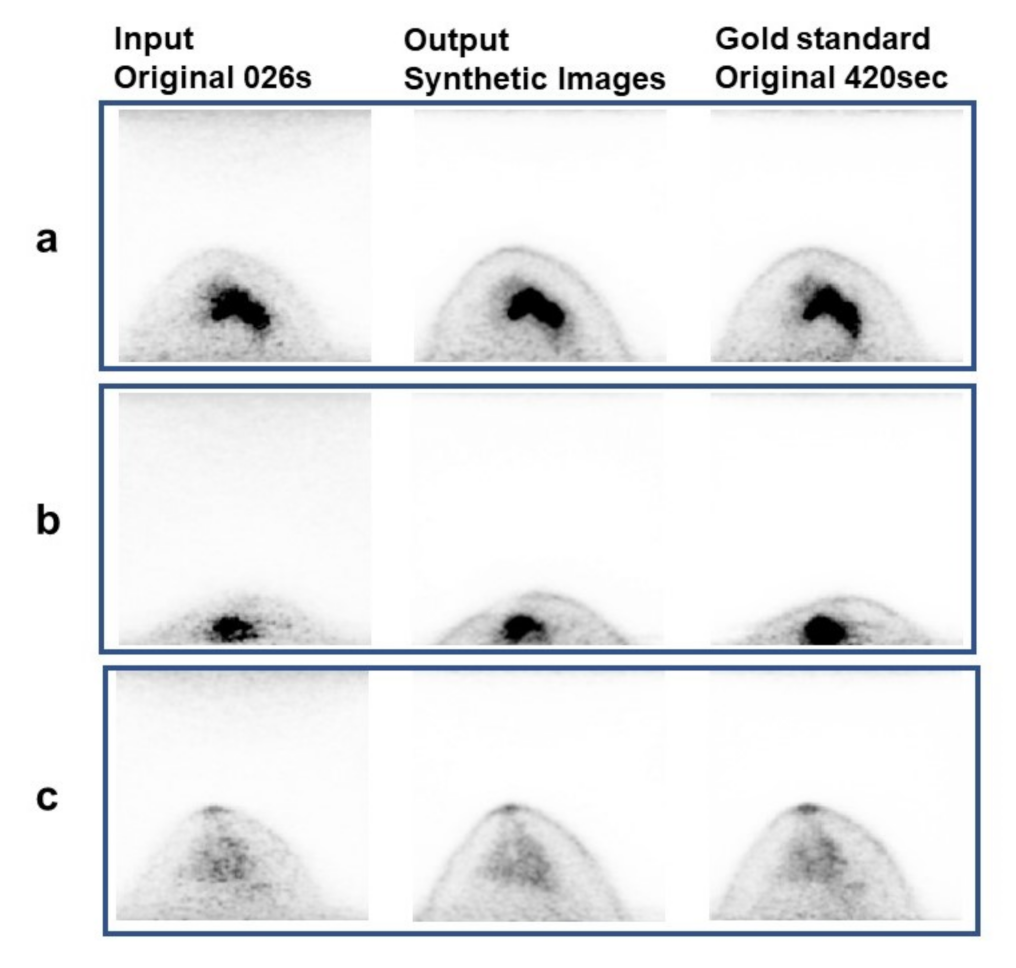
3.1. Visual Analysis Results
3.2. Quantitative Analysis Results
3.3. Detection Rate of Abnormal Accumulation
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2020. CA Cancer J. Clin. 2020, 70, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Mori, M.; Fujioka, T.; Katsuta, L.; Tsuchiya, J.; Kubota, K.; Kasahara, M.; Oda, G.; Nakagawa, T.; Onishi, I.; Tateishi, U. Diagnostic performance of time-of-flight PET/CT for evaluating nodal metastasis of the axilla in breast cancer. Nucl. Med. Commun. 2019, 40, 958–964. [Google Scholar] [CrossRef] [PubMed]
- Tateishi, U.; Gamez, C.; Dawood, S.; Yeung, H.W.; Cristofanilli, M.; Macapinlac, H.A. Bone metastases in patients with metastatic breast cancer: Morphologic and metabolic monitoring of response to systemic therapy with integrated PET/CT. Radiology 2008, 247, 189–196. [Google Scholar] [CrossRef] [PubMed]
- Kitajima, K.; Miyoshi, Y. Present and future role of FDG-PET/CT imaging in the management of breast cancer. Jpn. J. Radiol. 2016, 34, 167–180. [Google Scholar] [CrossRef]
- Mori, M.; Fujioka, T.; Kubota, K.; Katsuta, L.; Yashima, Y.; Nomura, K.; Yamaga, E.; Tsuchiya, J.; Hosoya, T.; Oda, G.; et al. Relationship between prognostic stage in breast cancer and fluorine-18 fluorodeoxyglucose positron emission tomography/computed tomography. J. Clin. Med. 2021, 10, 3173. [Google Scholar] [CrossRef] [PubMed]
- Satoh, Y.; Nambu, A.; Ichikawa, T.; Onishi, H. Whole-body total lesion glycolysis measured on fluorodeoxyglucose positron emission tomography/computed tomography as a prognostic variable in metastatic breast cancer. BMC Cancer 2014, 14, 525. [Google Scholar] [CrossRef] [Green Version]
- Satoh, Y.; Imai, M.; Ikegawa, C.; Hirata, K.; Abo, N.; Kusuzaki, M.; Oyama-Manabe, N.; Onishi, H. Effect of radioactivity outside the field of view on image quality of dedicated breast positron emission tomography: Preliminary phantom and clinical studies. Ann. Nucl. Med. 2022, 36, 1010–1018. [Google Scholar] [CrossRef] [PubMed]
- Satoh, Y.; Imai, M.; Hirata, K.; Asakawa, Y.; Ikegawa, C.; Onishi, H. Optimal relaxation parameters of dynamic row-action maximum likelihood algorithm and post-smoothing filter for image reconstruction of dedicated breast PET. Ann. Nucl. Med. 2021, 35, 608–616. [Google Scholar] [CrossRef]
- Satoh, Y.; Motosugi, U.; Omiya, Y.; Onishi, H. Unexpected abnormal uptake in the breasts at dedicated breast PET: Incidentally detected small cancers or nonmalignant features? AJR Am. J. Roentgenol. 2019, 212, 443–449. [Google Scholar] [CrossRef]
- Satoh, Y.; Motosugi, U.; Imai, M.; Onishi, H. Comparison of dedicated breast positron emission tomography and whole-body positron emission tomography/computed tomography images: A common phantom study. Ann. Nucl. Med. 2020, 34, 119–127. [Google Scholar] [CrossRef]
- Miyake, K.; Kataoka, M.; Ishimori, T.; Matsumoto, Y.; Torii, M.; Takada, M.; Satoh, Y.; Kubota, K.; Satake, H.; Yakami, M.; et al. A proposed dedicated breast PET lexicon: Standardization of description and reporting of radiotracer uptake in the breast. Diagnostics 2021, 11, 1267. [Google Scholar] [CrossRef] [PubMed]
- Tokuda, Y.; Yanagawa, M.; Fujita, Y.; Honma, K.; Tanei, T.; Shimoda, M.; Miyake, T.; Naoi, Y.; Kim, S.J.; Shimazu, K.; et al. Prediction of pathological complete response after neoadjuvant chemotherapy in breast cancer: Comparison of diagnostic performances of dedicated breast PET, whole-body PET, and dynamic contrast-enhanced MRI. Breast Cancer Res. Treat. 2021, 188, 107–115. [Google Scholar] [CrossRef] [PubMed]
- Hathi, D.K.; Li, W.; Seo, Y.; Flavell, R.R.; Kornak, J.; Franc, B.L.; Joe, B.N.; Esserman, L.J.; Hylton, N.M.; Jones, E.F. Evaluation of primary breast cancers using dedicated breast PET and whole-body PET. Sci. Rep. 2020, 10, 21930. [Google Scholar] [CrossRef] [PubMed]
- Brenner, D.J.; Elliston, C.D. Estimated radiation risks potentially associated with full-body CT screening. Radiology 2004, 232, 735–738. [Google Scholar] [CrossRef] [Green Version]
- Satoh, Y.; Imai, M.; Ikegawa, C.; Onishi, H. Image quality evaluation of real low-dose breast PET. Jpn. J. Radiol. 2022, 40, 1186–1193. [Google Scholar] [CrossRef]
- Barat, M.; Chassagnon, G.; Dohan, A.; Gaujoux, S.; Coriat, R.; Hoeffel, C.; Cassinotto, C.; Soyer, P. Artificial intelligence: A critical review of current applications in pancreatic imaging. Jpn. J. Radiol. 2021, 39, 514–523. [Google Scholar] [CrossRef]
- Nakao, T.; Hanaoka, S.; Nomura, Y.; Hayashi, N.; Abe, O. Anomaly detection in chest 18F-FDG PET/CT by Bayesian deep learning. Jpn. J. Radiol. 2022, 40, 730–739. [Google Scholar] [CrossRef]
- NNakai, H.; Fujimoto, K.; Yamashita, R.; Sato, T.; Someya, Y.; Taura, K.; Isoda, H.; Nakamoto, Y. Convolutional neural network for classifying primary liver cancer based on triple-phase CT and tumor marker information: A pilot study. Jpn. J. Radiol. 2021, 39, 690–702. [Google Scholar] [CrossRef]
- Okuma, T.; Hamamoto, S.; Maebayashi, T.; Taniguchi, A.; Hirakawa, K.; Matsushita, S.; Matsushita, K.; Murata, K.; Manabe, T.; Miki, Y. Quantitative evaluation of COVID-19 pneumonia severity by CT pneumonia analysis algorithm using deep learning technology and blood test results. Jpn. J. Radiol. 2021, 39, 956–965. [Google Scholar] [CrossRef]
- Fujioka, T.; Yashima, Y.; Oyama, J.; Mori, M.; Kubota, K.; Katsuta, L.; Kimura, K.; Yamaga, E.; Oda, G.; Nakagawa, T.; et al. Deep-learning approach with convolutional neural network for classification of maximum intensity projections of dynamic contrast-enhanced breast magnetic resonance imaging. Magn. Reson. Imaging 2021, 75, 1–8. [Google Scholar] [CrossRef]
- Ozaki, J.; Fujioka, T.; Yamaga, E.; Hayashi, A.; Kujiraoka, Y.; Imokawa, T.; Takahashi, K.; Okawa, S.; Yashima, Y.; Mori, M.; et al. Deep learning method with a convolutional neural network for image classification of normal and metastatic axillary lymph nodes on breast ultrasonography. Jpn. J. Radiol. 2022, 40, 814–822. [Google Scholar] [CrossRef] [PubMed]
- Fujioka, T.; Kubota, K.; Mori, M.; Kikuchi, Y.; Katsuta, L.; Kimura, M.; Yamaga, E.; Adachi, M.; Oda, G.; Nakagawa, T.; et al. Efficient anomaly detection with generative adversarial network for breast ultrasound imaging. Diagnostics 2020, 10, 456. [Google Scholar] [CrossRef] [PubMed]
- Adachi, M.; Fujioka, T.; Mori, M.; Kubota, K.; Kikuchi, Y.; Xiaotong, W.; Oyama, J.; Kimura, K.; Oda, G.; Nakagawa, T.; et al. Detection and diagnosis of breast cancer using artificial intelligence based assessment of maximum intensity projection dynamic contrast-enhanced magnetic resonance images. Diagnostics 2020, 10, 330. [Google Scholar] [CrossRef] [PubMed]
- Fujioka, T.; Kubota, K.; Mori, M.; Kikuchi, Y.; Katsuta, L.; Kasahara, M.; Oda, G.; Ishiba, T.; Nakagawa, T.; Tateishi, U. Distinction between benign and malignant breast masses at breast ultrasound using deep learning method with convolutional neural network. Jpn. J. Radiol. 2019, 37, 466–472. [Google Scholar] [CrossRef] [PubMed]
- Satoh, Y.; Tamada, D.; Omiya, Y.; Onishi, H.; Motosugi, U. Diagnostic performance of the support vector machine model for breast cancer on ring-shaped dedicated breast positron emission tomography images. J. Comput. Assist. Tomogr. 2020, 44, 413–418. [Google Scholar] [CrossRef] [PubMed]
- Satoh, Y.; Imokawa, T.; Fujioka, T.; Mori, M.; Yamaga, E.; Takahashi, K.; Takahashi, K.; Kawase, T.; Kubota, K.; Tateishi, U.; et al. Deep learning for image classification in dedicated breast positron emission tomography (dbPET). Ann. Nucl. Med. 2022, 36, 401–410. [Google Scholar] [CrossRef]
- Takahashi, K.; Fujioka, T.; Oyama, J.; Mori, M.; Yamaga, E.; Yashima, Y.; Imokawa, T.; Hayashi, A.; Kujiraoka, Y.; Tsuchiya, J.; et al. Deep learning using multiple degrees of maximum-intensity projection for PET/CT image classification in breast cancer. Tomography 2022, 8, 131–141. [Google Scholar] [CrossRef]
- Hirata, K.; Sugimori, H.; Fujima, N.; Toyonaga, T.; Kudo, K. Artificial intelligence for nuclear medicine in oncology. Ann. Nucl. Med. 2022, 36, 123–132. [Google Scholar] [CrossRef]
- Goodfellow, I.J.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative adversarial networks. arXiv 2014, arXiv:1406.2661. [Google Scholar] [CrossRef]
- Isola, P.; Zhu, J.Y.; Zhou, T.; Efros, A.A. Image-to-image translation with conditional adversarial networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 1125–1134. [Google Scholar]
- Mori, M.; Fujioka, T.; Katsuta, L.; Kikuchi, Y.; Oda, G.; Nakagawa, T.; Kitazume, Y.; Kubota, K.; Tateishi, U. Feasibility of new fat suppression for breast MRI using pix2pix. Jpn. J. Radiol. 2020, 38, 1075–1081. [Google Scholar] [CrossRef]
- Ueda, D.; Katayama, Y.; Yamamoto, A.; Ichinose, T.; Arima, H.; Watanabe, Y.; Walston, S.L.; Tatekawa, H.; Takita, H.; Honjo, T.; et al. Deep learning-based angiogram generation model for cerebral angiography without misregistration artifacts. Radiology 2021, 299, 675–681. [Google Scholar] [CrossRef]
- Tsuda, T.; Murayama, H.; Kitamura, K.; Yamaya, T.; Yoshida, E.; Omura, T.; Kawai, H.; Inadama, N.; Orita, N. A four-layer depth of interaction detector block for small animal PET. IEEE Nucl. Sci. Symp. Conf. Rec. 2003, 3, 1789–1793. [Google Scholar]
- Yi, X.; Walia, E.; Babyn, P. Generative adversarial network in medical imaging: A review. Med. Image Anal. 2019, 58, 101552. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Landis, J.R.; Koch, G.G. The measurement of observer agreement for categorical data. Biometrics 1977, 33, 159–174. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yoshida, N.; Kageyama, H.; Akai, H.; Yasaka, K.; Sugawara, H.; Okada, Y.; Kunimatsu, A. Motion correction in MR image for analysis of VSRAD using generative adversarial network. PLoS ONE 2022, 17, e0274576. [Google Scholar] [CrossRef]
- Chen, H.; Yan, S.; Xie, M.; Huang, J. Application of cascaded GAN based on CT scan in the diagnosis of aortic dissection. Comput. Methods Programs Biomed. 2022, 226, 107130. [Google Scholar] [CrossRef]
- Ichikawa, Y.; Kanii, Y.; Yamazaki, A.; Nagasawa, N.; Nagata, M.; Ishida, M.; Kitagawa, K.; Sakuma, H. Deep learning image reconstruction for improvement of image quality of abdominal computed tomography: Comparison with hybrid iterative reconstruction. Jpn. J. Radiol. 2021, 39, 598–604. [Google Scholar] [CrossRef]
- Yasaka, K.; Akai, H.; Sugawara, H.; Tajima, T.; Akahane, M.; Yoshioka, N.; Kabasawa, H.; Miyo, R.; Ohtomo, K.; Abe, O.; et al. Impact of deep learning reconstruction on intracranial 1.5 T magnetic resonance angiography. Jpn. J. Radiol. 2022, 40, 476–483. [Google Scholar] [CrossRef] [PubMed]
- Kaga, T.; Noda, Y.; Mori, T.; Kawai, N.; Miyoshi, T.; Hyodo, F.; Kato, H.; Matsuo, M. Unenhanced abdominal low-dose CT reconstructed with deep learning-based image reconstruction: Image quality and anatomical structure depiction. Jpn. J. Radiol. 2022, 40, 703–711. [Google Scholar] [CrossRef]
- Chen, K.T.; Gong, E.; de Carvalho Macruz, F.B.; Xu, J.; Boumis, A.; Khalighi, M.; Poston, K.L.; Sha, S.J.; Greicius, M.D.; Mormino, E.; et al. Ultralow-dose 18F-florbetaben amyloid PET imaging using deep learning with multi-contrast MRI inputs. Radiology 2018, 290, 649–656, Erratum in Radiology 2020, 296, E195. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.-R.; Baratto, L.; Hawk, K.E.; Theruvath, A.J.; Pribnow, A.; Thakor, A.S.; Gatidis, S.; Lu, R.; Gummidipundi, S.E.; Garcia-Diaz, J.; et al. Artificial intelligence enables whole-body positron emission tomography scans with minimal radiation exposure. Eur. J. Nucl. Med. Mol. Imaging 2021, 48, 2771–2781. [Google Scholar] [CrossRef] [PubMed]

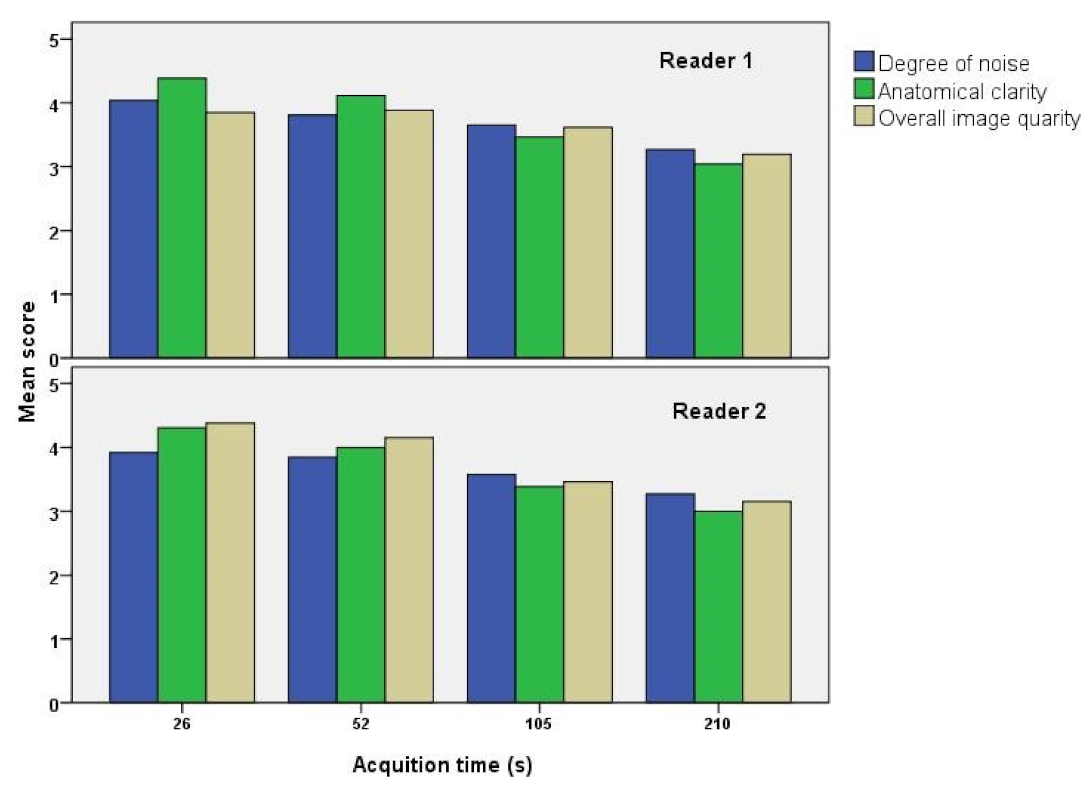

| Degree of Noise (Mean ± SD) | Anatomical Clarity (Mean ± SD) | Overall Image Quality (Mean ± SD) | |
|---|---|---|---|
| Reader 1 | |||
| 26 s | 4.04 ± 0.71 | 4.38 ± 0.74 | 3.85 ± 1.23 |
| 52 s | 3.81 ± 0.73 | 4.12 ± 0.85 | 3.88 ± 0.97 |
| 105 s | 3.65 ± 0.55 | 3.46 ± 0.69 | 3.62 ± 0.68 |
| 210 s | 3.27 ± 0.44 | 3.04 ± 0.34 | 3.19 ± 0.39 |
| Reader 2 | |||
| 26 s | 3.90 ± 0.62 | 4.31 ± 0.72 | 4.38 ± 0.74 |
| 52 s | 3.85 ± 0.53 | 4.00 ± 0.68 | 4.15 ± 0.66 |
| 105 s | 3.58 ± 0.49 | 3.38 ± 0.49 | 3.46 ± 0.50 |
| 210 s | 3.27 ± 0.44 | 3.00 ± 0.00 | 3.15 ± 0.36 |
| Acquisition Time | Image Type | SSIM (Mean ± SD) | p-Value | RSNR (mean ± SD) | p-Value |
|---|---|---|---|---|---|
| 026 s | Original | 0.83 ± 0.08 | <0.01 | 23.9 ± 2.4 | <0.01 |
| Synthetic | 0.85 ± 0.07 | 27.2 ± 3.6 | |||
| 52 s | Original | 0.88 ± 0.06 | 0.16 | 26.8 ± 2.8 | <0.01 |
| Synthetic | 0.87 ± 0.07 | 28.3 ± 4.0 | |||
| 105 s | Original | 0.92 ± 0.04 | <0.01 | 30.2 ± 3.2 | 0.62 |
| Synthetic | 0.90 ± 0.05 | 30.0 ± 4.3 | |||
| 210 s | Original | 0.96 ± 0.02 | <0.01 | 35.0 ± 2.3 | <0.01 |
| Synthetic | 0.93 ± 0.03 | 32.4 ± 3.9 |
| Original Images | Synthetic Images | |
|---|---|---|
| Reader 1 | ||
| 26 s | n = 10/15 (66.7%) | n = 9/15 (60.0%) |
| 52 s | n = 14/15 (93.3%) | n = 13/15 (86.7%) |
| 105 s | n = 14/15 (93.3%) | n = 13/15 (86.7%) |
| 210 s | n = 15/15 (100%) | n = 15/15 (100%) |
| Total | n = 53/60 (88.3%) | n = 50/60 (83.3%) |
| Reader 2 | ||
| 26 s | n = 7/15 (46.7%) | n = 9/15 (60.0%) |
| 52 s | n = 12/15 (80.0%) | n = 12/15 (80.8%) |
| 105 s | n = 14/15 (93.3%) | n = 14/15 (93.3%) |
| 210 s | n = 15/15 (100%) | n = 15/15 (100%) |
| Total | n = 48/60 (80.0%) | n = 50 (83.3%) |
| Reader 1 and 2 (mean) | ||
| 26 s | n = 8.5/15 (56.6%) | n = 9/15 (60.0%) |
| 52 s | n = 13/15 (86.7%) | n = 12.5/15 (83.3%) |
| 105 s | n = 14/15 (93.3%) | n = 13.5/15 (90%) |
| 210 s | n = 15/15 (100%) | n = 15/15 (100%) |
| Total | n = 50.5/60 (84.2%) | n = 50/60 (83.3%) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fujioka, T.; Satoh, Y.; Imokawa, T.; Mori, M.; Yamaga, E.; Takahashi, K.; Kubota, K.; Onishi, H.; Tateishi, U. Proposal to Improve the Image Quality of Short-Acquisition Time-Dedicated Breast Positron Emission Tomography Using the Pix2pix Generative Adversarial Network. Diagnostics 2022, 12, 3114. https://doi.org/10.3390/diagnostics12123114
Fujioka T, Satoh Y, Imokawa T, Mori M, Yamaga E, Takahashi K, Kubota K, Onishi H, Tateishi U. Proposal to Improve the Image Quality of Short-Acquisition Time-Dedicated Breast Positron Emission Tomography Using the Pix2pix Generative Adversarial Network. Diagnostics. 2022; 12(12):3114. https://doi.org/10.3390/diagnostics12123114
Chicago/Turabian StyleFujioka, Tomoyuki, Yoko Satoh, Tomoki Imokawa, Mio Mori, Emi Yamaga, Kanae Takahashi, Kazunori Kubota, Hiroshi Onishi, and Ukihide Tateishi. 2022. "Proposal to Improve the Image Quality of Short-Acquisition Time-Dedicated Breast Positron Emission Tomography Using the Pix2pix Generative Adversarial Network" Diagnostics 12, no. 12: 3114. https://doi.org/10.3390/diagnostics12123114
APA StyleFujioka, T., Satoh, Y., Imokawa, T., Mori, M., Yamaga, E., Takahashi, K., Kubota, K., Onishi, H., & Tateishi, U. (2022). Proposal to Improve the Image Quality of Short-Acquisition Time-Dedicated Breast Positron Emission Tomography Using the Pix2pix Generative Adversarial Network. Diagnostics, 12(12), 3114. https://doi.org/10.3390/diagnostics12123114







