1. Introduction
The number of patients who are on dialysis due to end-stage renal failure is increasing worldwide, which has become a major health economic problem. According to a recent report [
1], the number of patients undergoing chronic dialysis worldwide exceeded two million in 2010, and this number may double by 2030. The early detection and management of chronic kidney disease (CKD) is important in order to prevent its progression to end-stage renal failure. Immunoglobulin A nephropathy (IgAN) is the leading cause of CKD worldwide. It typically progresses to end-stage renal failure in 15–20% of patients after 10 years, and approximately 40% of patients after around 20 years [
2,
3]. Using evidence-based clinical practice guidelines in Japan [
4], the clinical predictors for the progression of IgAN at the time of the initial renal biopsy include the following: (1) the presence of hypertension; (2) the amount of proteinuria with a usual cut-off of >1 g/day; (3) the degree of renal dysfunction; and (4) the histopathological grade, based on renal pathology. Of these predictors, histopathological findings play a key role but require observation by experts under a microscope. Patients with IgAN have varied histopathological lesions, ranging from mild mesangial proliferation, endocapillary hypercellularity, and crescentic glomerulonephritis to global and segmental sclerosis. For example, sclerosis represents the final appearance of glomerular injury that is caused by various diseases. When sclerosis occurs globally, determining the cause of the injury can be difficult.
Two histopathological grading systems are referred to in the clinical guidelines. The first system is the Oxford classification [
5,
6], which is based on the score of mesangial hypercellularity (M: M0, ≤0.5; M1, >0.5), endocapillary hypercellularity (E: E0, absent; E1, present), segmental sclerosis (S: S0, absent; S1, present), tubular atrophy or interstitial fibrosis (T: T0, 0–25%; T1, 26–50%; T2, >50%), and cellular or fibrocellular crescents (C: C0, absent; C1, 0–25%; C2, >25%). The second system is the Japanese histological grade classification (H-Grade) [
7,
8], which is based on the presence of acute lesions (i.e., cellular crescent, tuft necrosis, and fibrocellular crescent) and chronic lesions (i.e., global sclerosis, segmental sclerosis, and fibrous crescent). Detecting these complex findings among all of the glomeruli in whole slide images (WSIs) is laborious and time consuming, even for highly trained pathologists or nephrologists. Furthermore, the assessment is not always consistent [
9,
10]. Suppose the findings of all of the glomeruli on a WSI could be quantified with a computer, it may lead to a more thorough investigation of their impact on the prognosis of immunoglobulin A nephropathy (IgAN) and accelerate such research.
In the past decade, the number of studies aiming to develop deep learning applications for nephropathology has increased rapidly. Computational image recognition focusing on the glomerulus is generally classified into the following three types: the detection of glomeruli [
11,
12,
13,
14], the classification of the glomeruli [
10,
15], and the segmentation of the glomeruli [
16,
17,
18,
19,
20,
21,
22,
23,
24]. The glomeruli that are detected in the WSI are localized by drawing bounding boxes. This approach would be a good application of automation because detecting glomeruli is simple but tedious for humans. Additionally, the development of such tools is realistic, as previously reported [
13]. The classification of glomeruli, such as the presence or the absence of certain pathological findings, is more challenging because it requires the interpretation of quantitative histopathological lesions into qualitative expressions, for which expert assessment is not always consistent [
9,
10]. The segmentation of glomeruli localizes and quantifies every glomerulus by identifying the regions of each glomerulus in the pixels. Several studies have attempted to distinguish between the entire glomerulus and the background [
16,
17] or to distinguish between the normal and the sclerotic glomeruli [
20,
21,
24]. Other studies have focused on the tubules, the blood vessels, and the interstitium, in addition to the glomerulus [
19,
23], or on the components inside of the glomerulus [
18,
22]. Segmenting the glomerulus and its components would be more helpful for a better understanding of kidney disease because it will be applied in the classification of pathological findings and to develop a prognostic model by utilizing quantified histopathological regions.
Table 1 shows the previous studies for glomerular segmentation from WSI.
As the configuration of segmentation tasks varies from researcher to researcher, the high performance of a machine learning model does not necessarily indicate its usefulness for subsequent analyses. Previous studies [
18,
19,
22] have assessed the usefulness of segmentation results in subsequent analyses, whereas other studies have only assessed the performance of machine learning models. In addition, these previous studies have only [
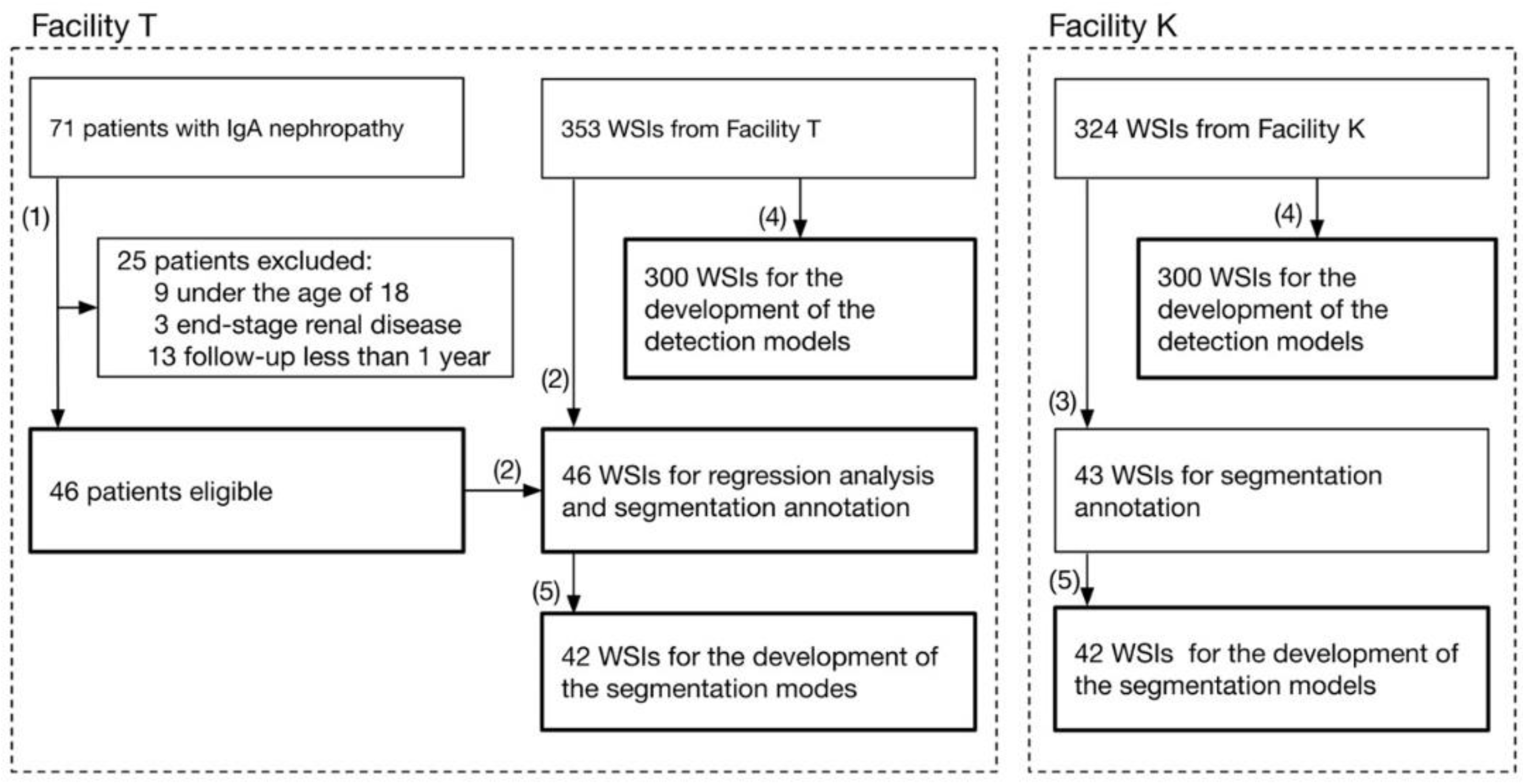
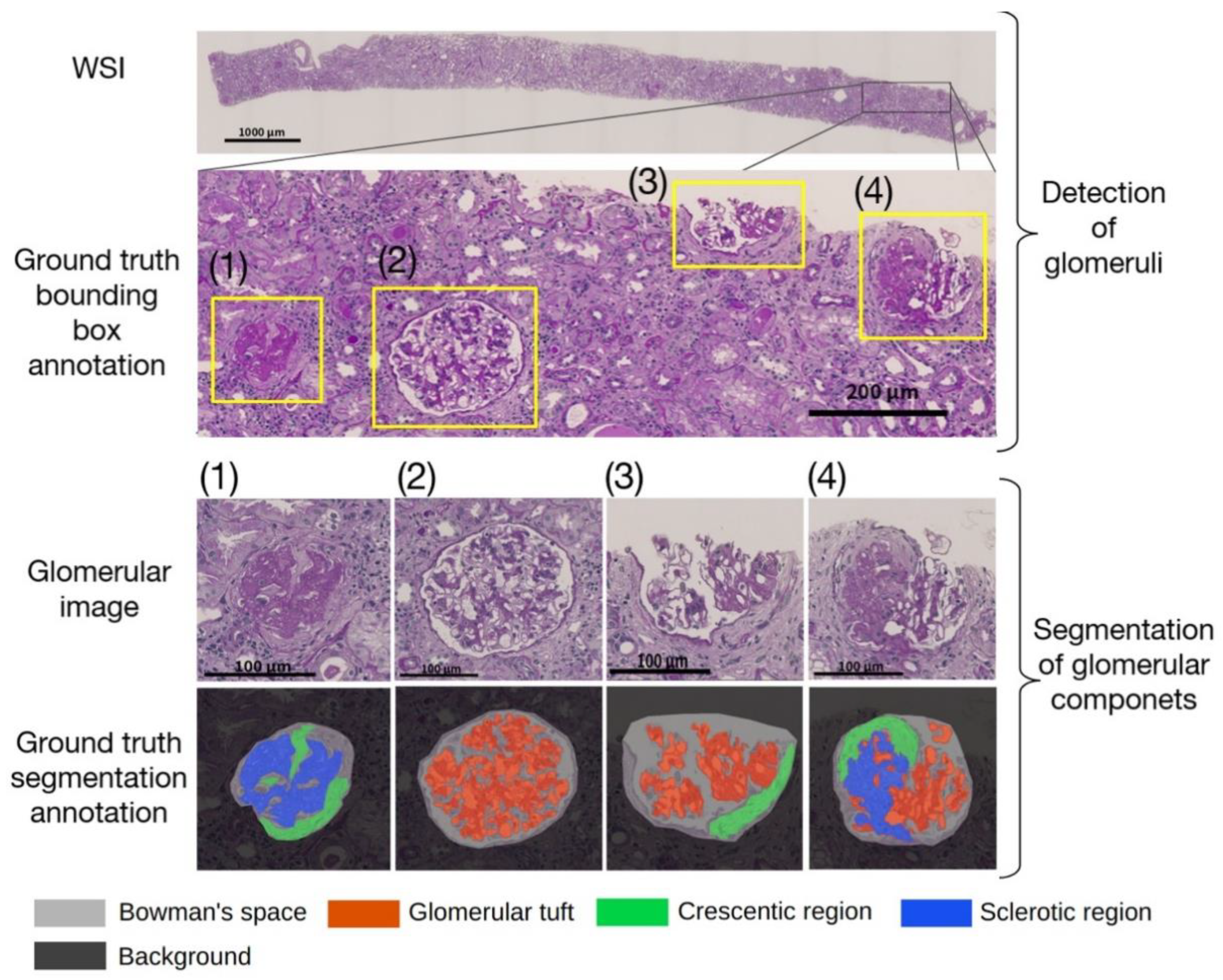
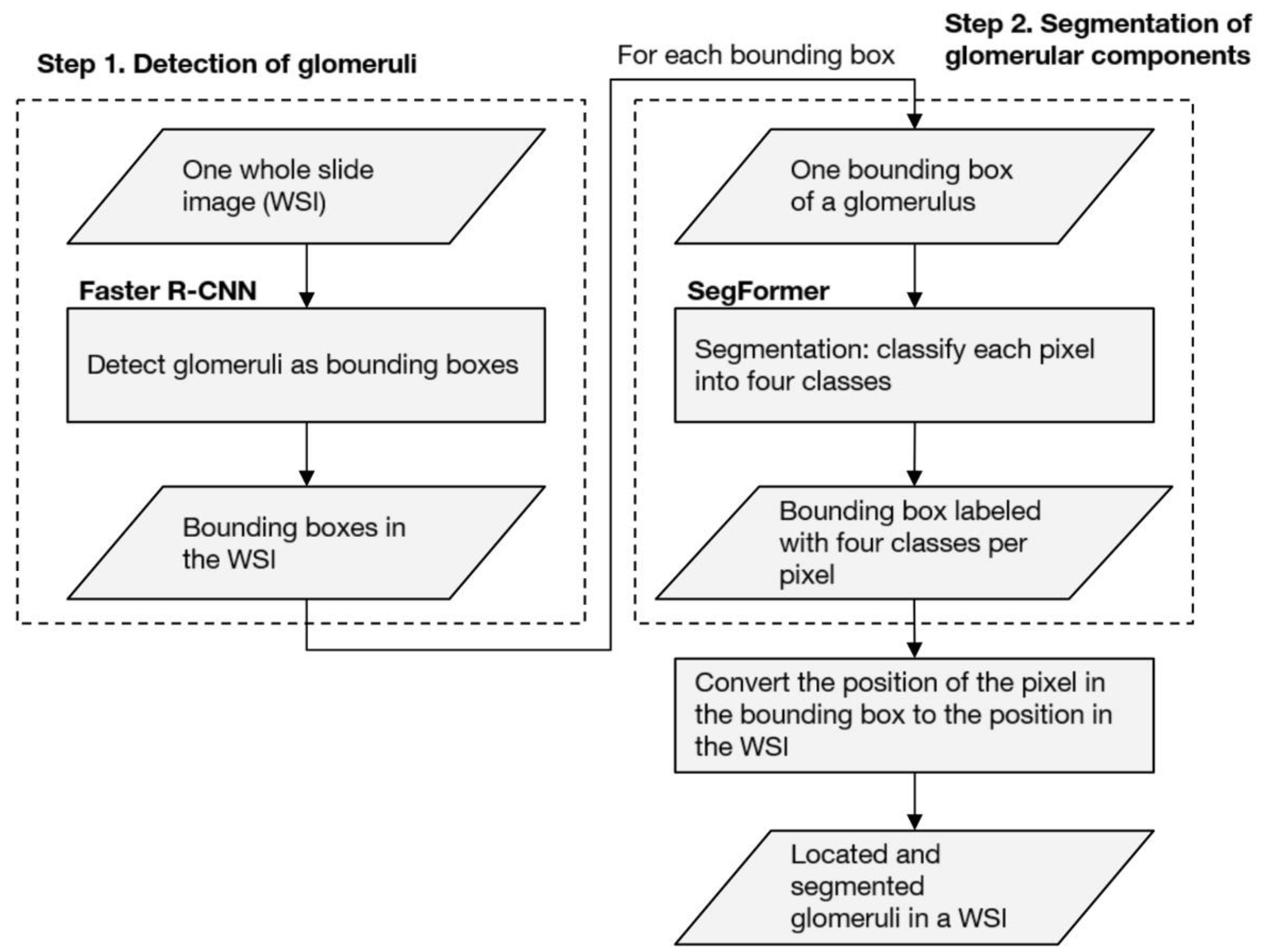
19] evaluated the performance of machine learning models against external WSI, whereas the other studies have evaluated a single facility. Due to their high performance, deep neural networks (DNNs) tend to overfit to minute differences in the images that are used for training. Furthermore, the pathological specimens differ between facilities due to the differences in the preparation protocols. These factors have a non-negligible impact on the generalizability of studies dealing with WSI in DNNs. Therefore, in assessing the performance of the developed DNNs, an internal evaluation using only the WSIs of a single facility is not sufficient; external evaluations of the WSIs of different facilities are also important. Based on these two points, we propose an automated computational pipeline to detect the glomeruli from periodic acid-Schiff (PAS)-stained WSI and to segment the Bowman’s space, the glomerular tuft, and the histopathological components of crescentic and sclerotic regions. The pipelines were developed using the WSIs of two facilities independently, and the performances across the facilities were evaluated. In order to assess the significance of the quantified histopathological regions, we conducted a multivariate regression analysis to determine whether the proportion of the sclerotic regions was significantly associated with the prognosis of kidney function in patients with IgAN.
4. Discussion
In this paper, we describe an automated computational pipeline that can detect glomeruli in PAS-stained WSI and segment the histopathological components inside of the glomerulus. Based on multivariate analysis, the predicted sclerotic regions, even the regions that were predicted by the external model, had a significant negative impact on the eGFR slope within two years after biopsy. We believe that this study is the first to demonstrate the usefulness of an automated computational pipeline for segmenting the histopathological glomerular components on WSIs and demonstrate that quantified sclerotic regions impact the prognosis of the kidney function in patients with IgAN.
Several studies [
18,
19,
20,
21] aiming for pixel-level semantic segmentation for WSI of renal tissue sections have set the task of distinguishing between nonsclerotic and sclerotic glomeruli. Bueno et al. [
20] sequentially applied SegNet-VGG19 [
34] in order to segment glomeruli and applied AlexNet to classify them as nonsclerotic or sclerotic glomeruli. The segmentation accuracies for the nonsclerotic and the sclerotic were 96.06% and 83.22%, respectively. Hermsen et al. [
19] evaluated U-Net-based 11 class segmentation, as described by Ronneberger et al. [
35]. The normal glomeruli, sclerotic glomeruli, empty Bowman’s capsules, tubules, arteries, interstitium, and the capsules were fully annotated. The Dice coefficients of the normal and the sclerotic glomeruli were 0.95 and 0.62, respectively. Altini et al. [
21] conducted SegNet-based semantic segmentation of nonsclerotic and sclerotic glomeruli; their IoUs were 0.66546 and 0.49215, respectively. Jiang et al. [
24] conducted a mask region-based convolutional neural network (R-CNN)-based semantic segmentation for classifying glomeruli with a normal structure, an abnormal structure, and global sclerosis; the mean IoU for PAS-stained WSIs were 0.697, 0.544, and 0.646, respectively. The results of these previous studies could help us to quantify global glomerulosclerosis, the ratio between sclerotic glomeruli, and the overall number of glomeruli. However, because glomerular sclerosis does not always occur globally, pixel-level segmentation for partially sclerosed regions is required for detailed quantification. Such quantification should have an essential role in understanding kidney diseases.
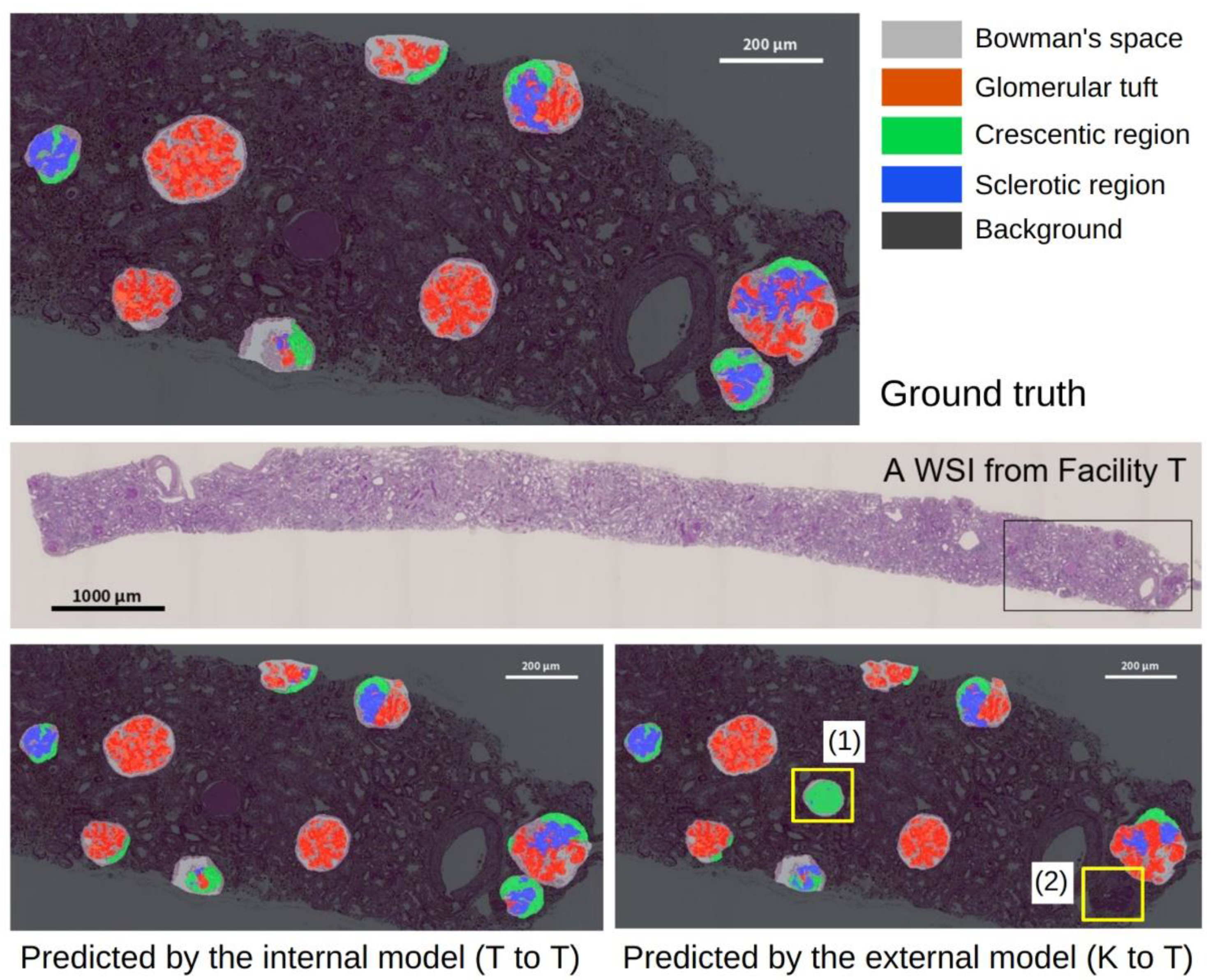
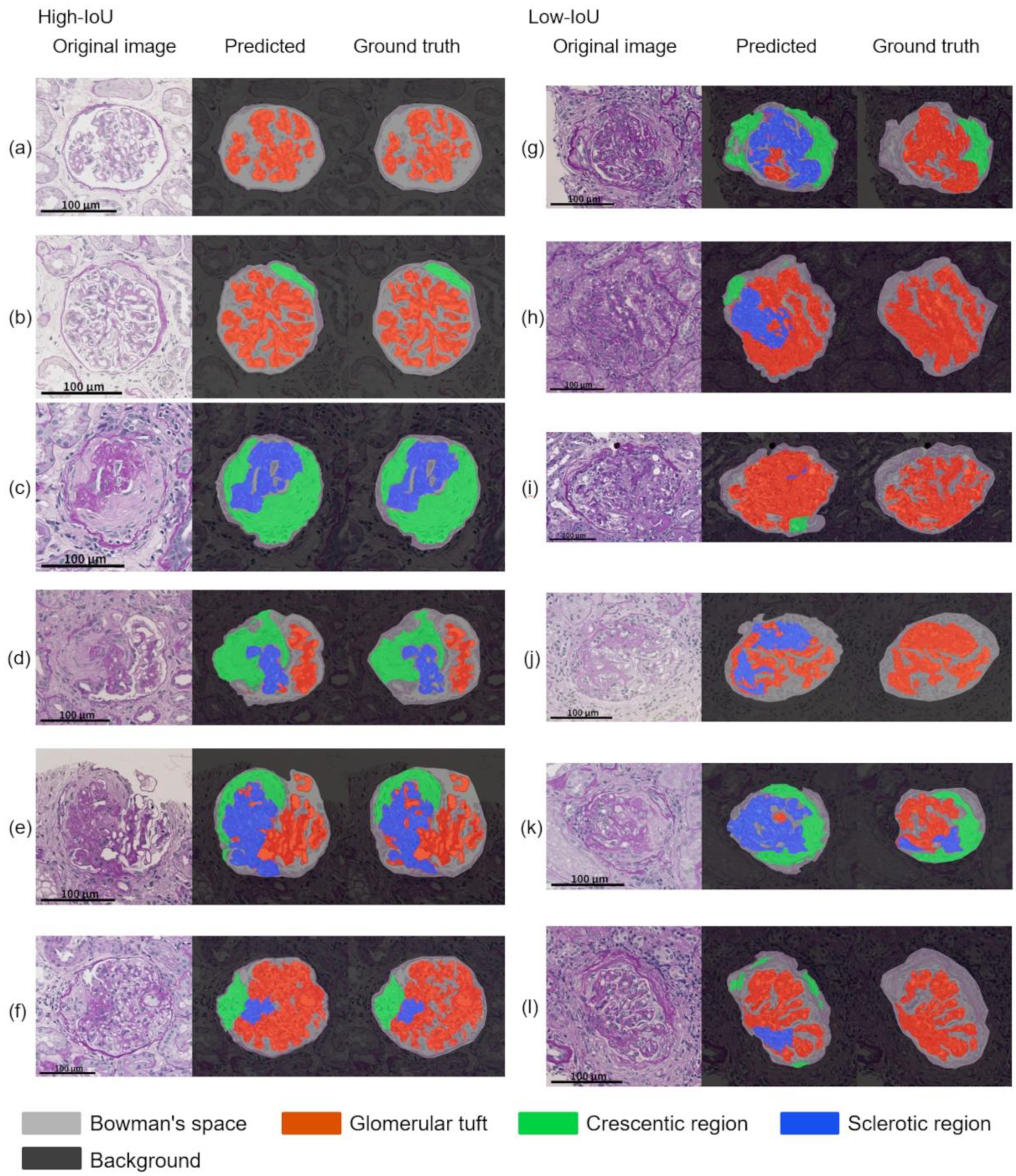
As shown in
Table 4, the performance of the segmentation alone and the pipeline showed no significant differences in the mean IoU between T to T and K to K. This finding indicated that their internal performances were comparable. This finding supports that the annotation for glomerular detection and segmentation was conducted with a constant quality. Compared to the performance of the models that were trained with internal WSIs, the performance of the models that were trained with external WSIs tended to decrease in the segmentation alone and the pipeline. One of the reasons for this finding may be due to differences in the slide preparation to the digitization process between the facilities. The differences in the staining protocols, the manufacturing processes, and the digital scanner processing between the laboratories caused minute differences in the WSIs; however, the pathological samples were stained similarly. This difference is imperceptible to the human eye, but it is sufficient to affect deep learning-based applications [
36,
37,
38]. We applied color normalization in the preprocessing step and Gaussian blurring, sharpening, and contrast changes during the data augmentation. However, extended methods are required in order to compensate for the minute differences in WSIs between the facilities, which increases the robustness against external WSI. The successful adaptation of WSI in deep neural network-based applications depends on each step of high-quality pathology slide preparation, such as embedding, cutting, staining, and scanning [
39,
40], as well as color variations. Using precise and homogeneous WSIs is desirable; however, such a model may not necessarily be robust against external WSIs that have more diversity. Improving the interfacility applicability of the developed model is an important issue for the success of deep learning applications in digital pathology. In addition, the performance of K to T is significantly lower for both the segmentation alone and the pipeline, while the performance degradations of T to K are not significant. This may be because a small number of glomerular images (1011) were used to develop the segmentation in model K, compared to the number of glomerular images that were used to develop model T (1713). We used the same number of WSIs from both of the facilities for the segmentation task. However, the number of images differed because of the different number of glomeruli that were contained in each WSI. The relatively small number of glomerular images in the training data for model K may have resulted in less diversity, leading to the significant performance degradation of K to T.
As shown in
Table 5, the manually quantified (ground truth) sclerotic regions were associated with negatively impacting the eGFR slope in the multivariate analysis. Segmental sclerosis, which is defined by the Oxford Classification [
5,
6], or the chronic lesions including segmental sclerosis and global sclerosis, which are defined by the H-Grade [
7,
8] have a negative impact on the poor prognosis of IgAN; however, the current study showed that the quantified sclerotic regions also have a negative impact on the eGFR slope within two years after biopsy. In our analysis, the effect of the post-biopsy treatment on eGFR was not adjusted because of the retrospective design, which is a limitation of this analysis. In addition, other limitations of this analysis were that the 2-year period was relatively short and the number of IgAN cases (
n = 46) was also limited; these may have affected the relatively low coefficients of determination (0.18 in the ground truth model).
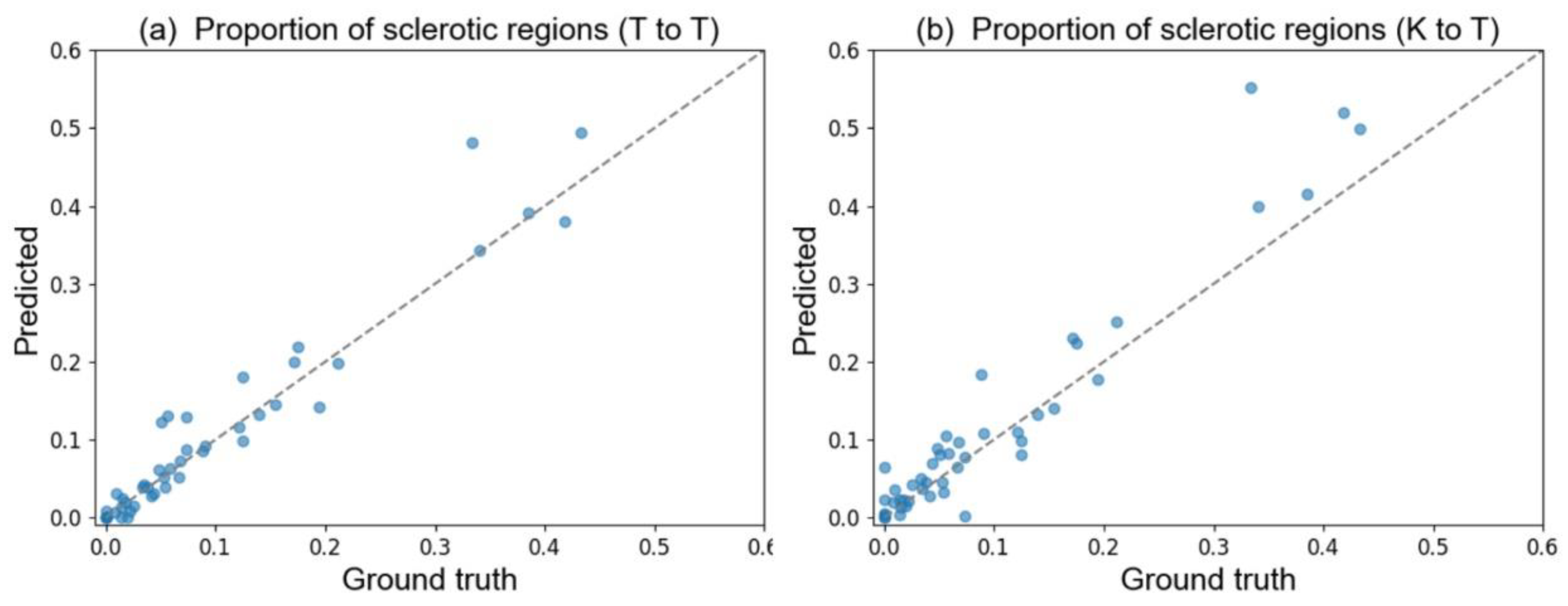
Table 5 also shows the same tendency in the standardized partial regression coefficients among the ground truth, the T to T (i.e., internal model), and the K to T (i.e., external model) models. The correlation between the ground truth regions and the predicted regions in each WSI aids in the understanding of their impact in the regression model. In
Table A3 in
Appendix E, the correlation coefficient for the sclerotic regions exceeded 0.96, even when using the external model. This finding indicated that the estimation of the total amount of sclerotic and glomerular tuft regions in each WSI was approximately correct. In light of the previous results, our developed pipeline shows a certain level of robustness for quantifying the glomerular tuft and sclerotic regions from WSI, even if the model is applied to the WSI of external facilities.
Another limitation of this study is that the concordance of the ground truth labels that have been used for developing glomerular detection and segmentation was not evaluated; however, the experts provided them. Surrounding the glomeruli with bounding boxes and drawing their histopathological components required distinguishing unclear boundaries with an understanding of pathology. Such labeling could vary among experts. Well-annotated examples are important in supervised learning; the main challenge in deep neural network-based applications for digital histopathology is obtaining high-quality labels. We carefully conducted the annotation with multiple experts, including a nephrologist and a pathologist, however the possibility of errors does exist. Nonetheless, annotation errors are not specific to this research; however, they should be kept in mind in studies on supervised learning.
5. Conclusions
We developed an automated computational pipeline for detecting glomeruli on PAS-stained WSIs, followed by segmenting the Bowman’s space, the glomerular tuft, the crescentic, and the sclerotic region inside of the glomeruli. The internal and external evaluation of the pipeline using WSIs from two facilities showed that the mean IoU of five regions, including the background, was 0.670 (T to T) and 0.693 (K to K) in the internal evaluation, and 0.609 (K to T) and 0.678 (T to K) in the external evaluation. The multivariate analysis for eGFR prognosis in cases of IgAN showed that the proportion of sclerotic regions that were quantified by the pipelines, even those that were quantified by the external model, had a significant negative impact on the eGFR slope, while five other clinical prognostic factors (i.e., age, sex, hypertension, eGFR at biopsy, and UPCR at biopsy) had no significant impact. These findings suggest the importance of quantifying the sclerotic region, as well as the usefulness and the robustness of the developed pipeline, for the purpose of predicting eGFR in cases of IgAN. The developed pipeline could aid in diagnosing renal pathology by visualizing and quantifying the histopathological feature of glomerulus. In addition, this high-throughput approach could potentially accelerate research in order to better understand the prognosis of IgAN.