Demarcation Line Determination for Diagnosis of Gastric Cancer Disease Range Using Unsupervised Machine Learning in Magnifying Narrow-Band Imaging
Abstract
1. Introduction
2. Methods
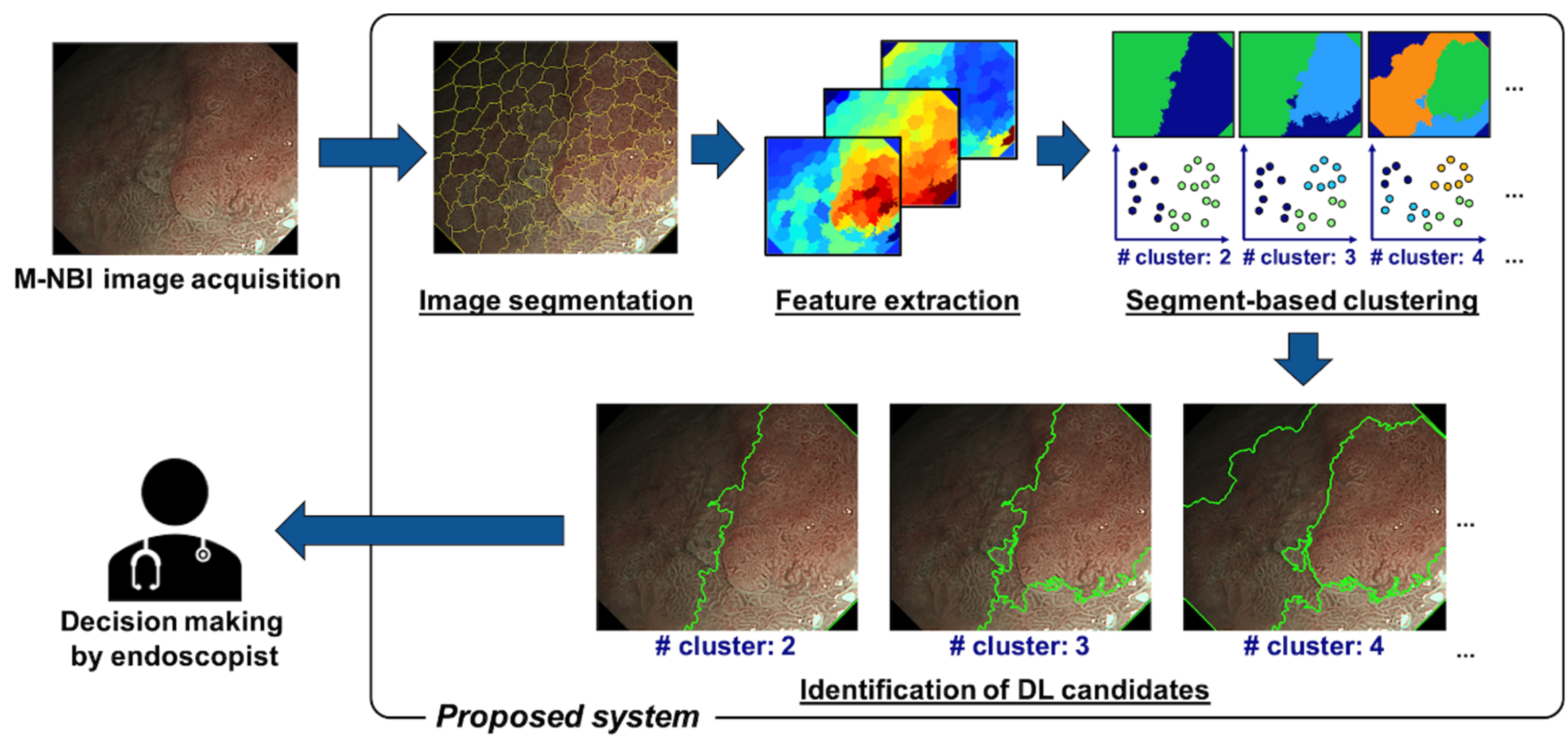
2.1. Proposed Method
2.1.1. Image Segmentation
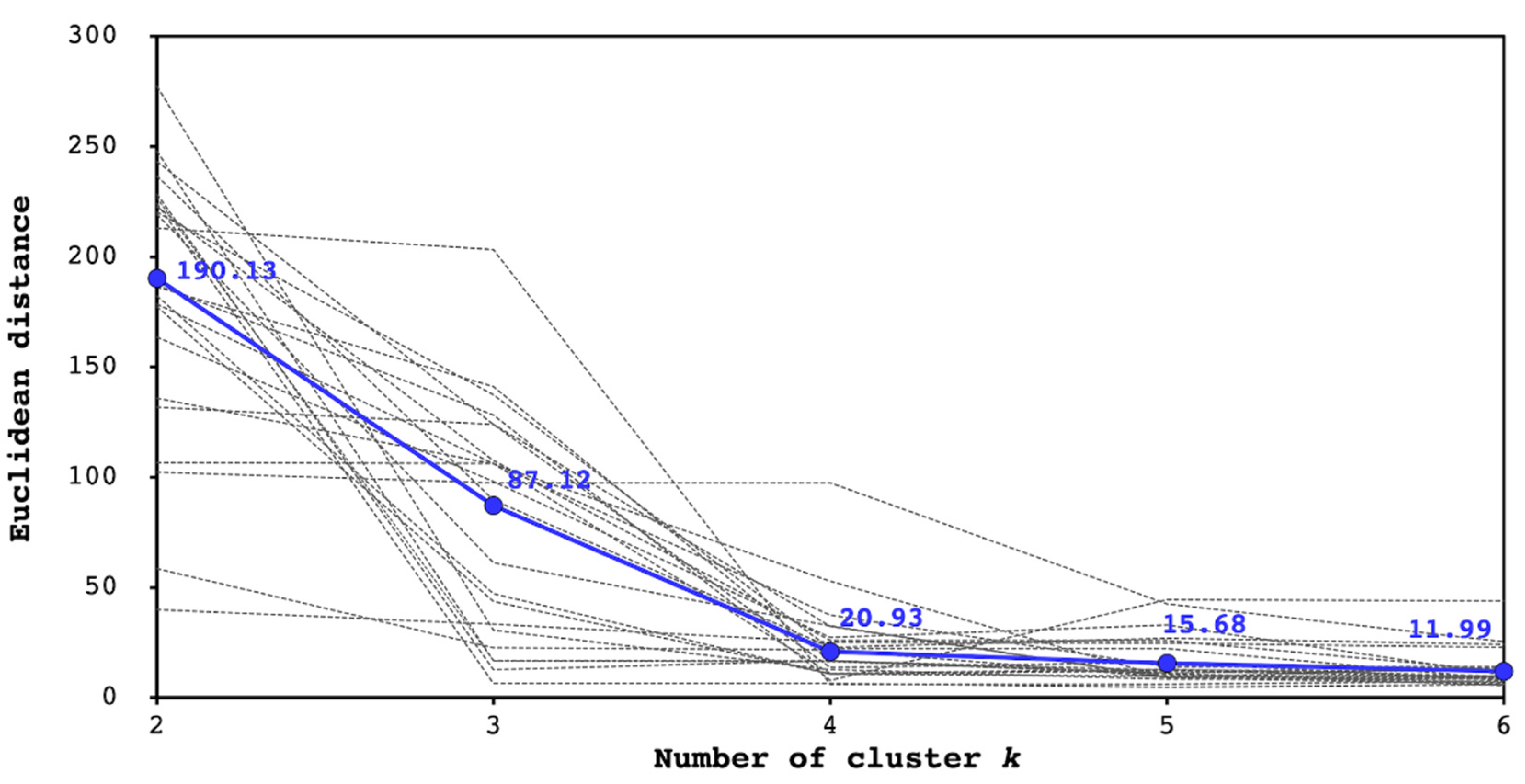
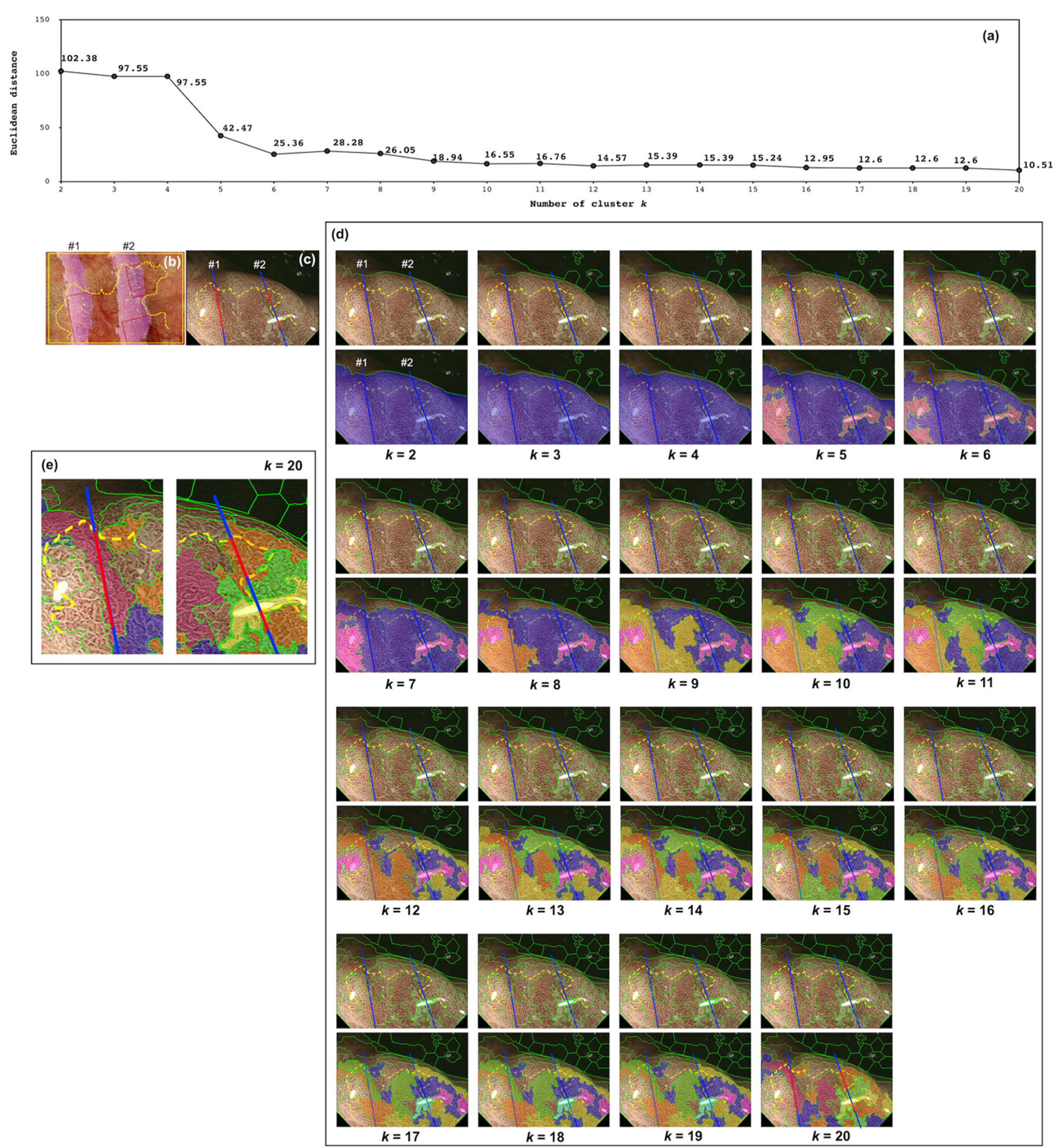
2.1.2. Feature Extraction and Clustering
2.1.3. Identification of DL
2.2. Experimental Setup
2.2.1. Test Images
2.2.2. Parameter Settings of the Proposed Method
2.2.3. Performance Metrics
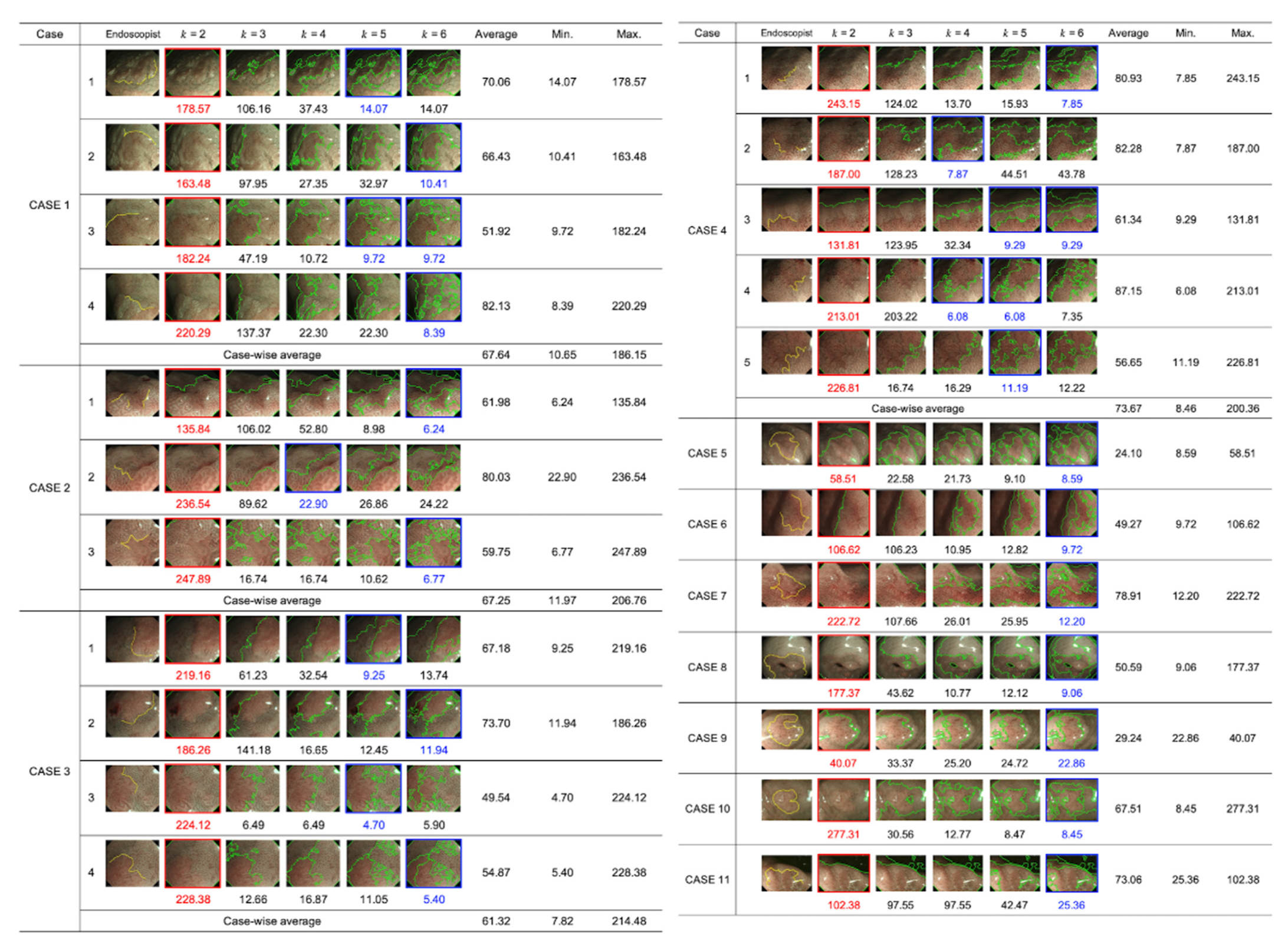
3. Results
4. Discussion
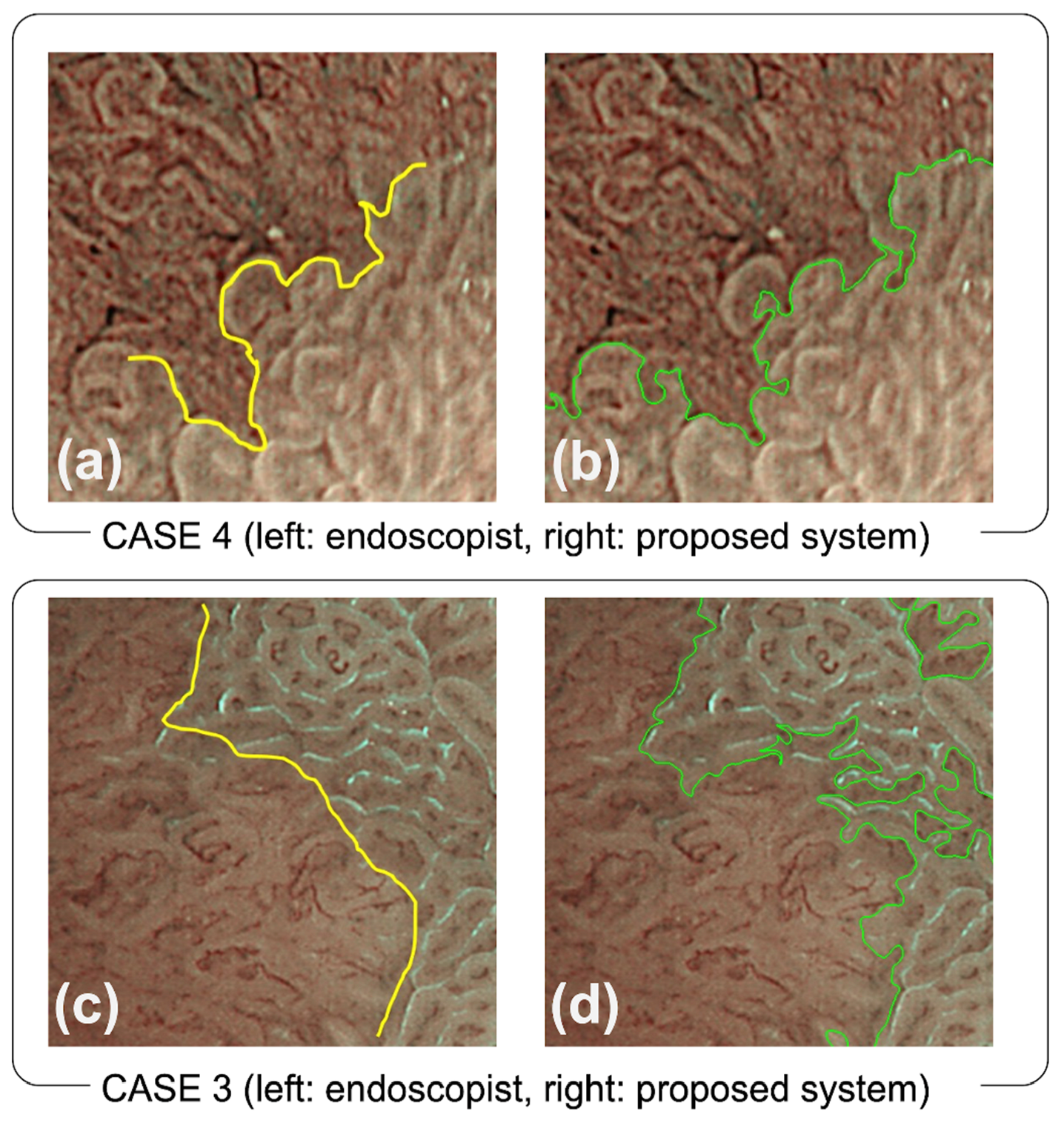
4.1. Comparison between the Endoscopists and the Proposed System
4.2. Pathological Evaluation of the Proposed System
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Yao, K. Clinical Application of Magnifying Endoscopy with Narrow-Band Imaging in the Stomach. Clin. Endosc. 2015, 48, 481–490. [Google Scholar] [CrossRef] [PubMed]
- Ezoe, Y.; Muto, M.; Uedo, N.; Doyama, H.; Yao, K.; Oda, I.; Kaneko, K.; Kawahara, Y.; Yokoi, C.; Sugiura, Y.; et al. Magnifying narrowband imaging is more accurate than con-ventional white-light imaging in diagnosis of gastric mucosal cancer. Gastroenterology 2011, 141, 2017–2025. [Google Scholar] [CrossRef] [PubMed]
- Yao, K.; Anagnostopoulos, G.; Ragunath, K. Magnifying endoscopy for diagnosing and delineating early gastric cancer. Endoscopy 2009, 41, 462–467. [Google Scholar] [CrossRef] [PubMed]
- Gotoda, T.; Yamamoto, H.; Soetikno, R.M. Endoscopic submucosal dissection of early gastric cancer. J. Gastroenterol. 2006, 41, 929–942. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, M.; Ono, H.; Hasuike, N.; Takizawa, K. Endoscopic submucosal dissection of early gastric cancer. Digestion 2008, 77, 23–28. [Google Scholar] [CrossRef]
- Kakushima, N.; Ono, H.; Tanaka, M.; Takizawa, K.; Yamaguchi, Y.; Matsubayashi, H. Factors related to lateral margin positivity for cancer in gastric specimens of endoscopic submucosal dissection. Dig. Endosc. 2011, 23, 227–232. [Google Scholar] [CrossRef]
- Sekiguchi, M.; Suzuki, H.; Oda, I.; Abe, S.; Nonaka, S.; Yoshinaga, S.; Taniguchi, H.; Sekine, S.; Kushima, R.; Saito, Y. Risk of recurrent gastric cancer after endoscopic resection with a positive lateral margin. Endoscopy 2014, 46, 273–278. [Google Scholar] [CrossRef]
- Kim, T.K.; Kim, G.H.; Park, D.Y.; Lee, B.E.; Jeon, T.Y.; Kim, D.H.; Jo, H.J.; Song, G.A. Risk factors for local recurrence in patients with positive lateral resection margins after endoscopic submucosal dissection for early gastric cancer. Surg. Endosc. 2015, 29, 2891–2898. [Google Scholar] [CrossRef]
- Imagawa, A.; Okada, H.; Kawahara, Y.; Takenaka, R.; Kato, J.; Kawamoto, H.; Fujiki, S.; Takata, R.; Yoshino, T.; Shiratori, Y. Endoscopic submucosal dissection for early gastric cancer: Results and degrees of technical difficulty as well as success. Endoscopy 2006, 38, 987–990. [Google Scholar] [CrossRef]
- Sugimoto, T.; Okamoto, M.; Mitsuno, Y.; Kondo, S.; Ogura, K.; Ohmae, T.; Mizuno, H.; Yoshida, S.; Isomura, Y.; Yamaji, Y.; et al. Endoscopic submucosal dissection is an effective and safe therapy for early gastric neoplasms: A multicenter feasible study. J. Clin. Gastroenterol. 2012, 46, 124–129. [Google Scholar] [CrossRef]
- Iizuka, H.; Kakizaki, S.; Sohara, N.; Onozato, Y.; Ishihara, H.; Okamura, S.; Itoh, H.; Mori, M. Stricture after endoscopic submucosal dissection for early gastric cancers and adenomas. Dig. Endosc. 2010, 22, 282–288. [Google Scholar] [CrossRef] [PubMed]
- Horie, Y.; Yoshio, T.; Aoyama, K.; Yoshimizu, S.; Horiuchi, Y.; Ishiyama, A.; Hirasawa, T.; Tsuchida, T.; Ozawa, T.; Ishihara, S.; et al. Diagnostic outcomes of esophageal cancer by artificial intelligence using convolutional neural networks. Gastrointest. Endosc. 2018, 89, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Hirasawa, T.; Aoyama, K.; Tanimoto, T.; Ishihara, S.; Shichijo, S.; Ozawa, T.; Ohnishi, T.; Fujishiro, M.; Matsuo, K.; Fujisaki, J.; et al. Application of artificial intelligence using a convolutional neural network for detecting gastric cancer in endoscopic images. Gastric Cancer 2018, 21, 653–660. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, T.; Hiroyasu, T.; Hiwa, S.; Okada, Y.; Hayashi, S.; Nakahata, Y.; Yasuda, Y.; Omatsu, T.; Obora, A.; Kojima, T.; et al. Potential of automatic diagnosis system with linked color imaging for diagnosis of Helicobacter pylori infection. Dig. Endosc. 2020, 32, 373–381. [Google Scholar] [CrossRef]
- Misawa, M.; Kudo, S.-E.; Mori, Y.; Nakamura, H.; Kataoka, S.; Maeda, Y.; Kudo, T.; Hayashi, T.; Wakamura, K.; Miyachi, H.; et al. Characterization of Colorectal Lesions Using a Computer-Aided Diagnostic System for Narrow-Band Imaging Endocytoscopy. Gastroenterology 2016, 150, 1531–1532. [Google Scholar] [CrossRef]
- Shichijo, S.; Nomura, S.; Aoyama, K.; Nishikawa, Y.; Miura, M.; Shinagawa, T.; Takiyama, H.; Tanimoto, T.; Ishihara, S.; Matsuo, K.; et al. Application of Convolutional Neural Networks in the Diagnosis of Helicobacter pylori Infection Based on Endoscopic Images. eBioMedicine 2017, 25, 106–111. [Google Scholar] [CrossRef]
- Mori, Y.; Kudo, S.; Mohmed, H.E.N.; Misawa, M.; Ogata, N.; Itoh, H.; Oda, M.; Mori, K. Artificial intelligence and upper gastrointestinal endoscopy: Current status and future perspective. Dig. Endosc. 2018, 31, 378–388. [Google Scholar] [CrossRef]
- Achanta, R.; Shaji, A.; Smith, K.; Lucchi, A.; Fua, P.; Süsstrunk, S. SLIC Superpixels Compared to State-of-the-Art Superpixel Methods. IEEE Trans. Pattern Anal. Mach. Intell. 2012, 34, 2274–2282.e3. [Google Scholar] [CrossRef]
- Wyszecki, G.; Stiles, W.S. Color Science; Wiley: New York, NY, USA, 1982; Volume 8. [Google Scholar]
- Agoston, M.K.; Agoston, M.K. Computer Graphics and Geometric Modeling; Springer: London, UK, 2005; Volume 1. [Google Scholar]
- Wu, Y.; Zhou, Y.; Saveriades, G.; Agaian, S.; Noonan, J.P.; Natarajan, P. Local Shannon entropy measure with statistical tests for image randomness. Inf. Sci. 2013, 222, 323–342. [Google Scholar] [CrossRef]
- Li, L.; Chen, Y.; Shen, Z.; Zhang, X.; Sang, J.; Ding, Y.; Yang, X.; Li, J.; Chen, M.; Jin, C.; et al. Convolutional neural network for the diagnosis of early gastric cancer based on magnifying narrow band imaging. Gastric Cancer 2019, 23, 126–132. [Google Scholar] [CrossRef]
- Kanesaka, T.; Lee, T.-C.; Uedo, N.; Lin, K.-P.; Chen, H.-Z.; Lee, J.-Y.; Wang, H.-P.; Chang, H.-T. Computer-aided diagnosis for identifying and delineating early gastric cancers in magnifying narrow-band imaging. Gastrointest. Endosc. 2018, 87, 1339–1344. [Google Scholar] [CrossRef] [PubMed]




| Characteristic | Value |
|---|---|
| Male/female, n | 6/5 |
| Age, mean, (range), years | 74 (59–92) |
| Tumor size, Average, (range), mm | |
| Major axis | 14.5 (5–30) |
| Minor axis | 8.8 (4–15) |
| Tumor location, n | |
| Upper | 1 |
| Middle | 3 |
| Lower | 7 |
| Morphologic types, n | |
| Type 0–I | 0 |
| Type 0–IIa | 3 |
| Type 0–IIb | 2 |
| Type 0–IIc | 6 |
| Type 1–4 | 0 |
| Depth of tumor, n | |
| T1a (mucosa) | 10 |
| T1b (submucosa) | 1 |
| T2 (muscularis propria) or more | 0 |
| Histological classification, n | |
| Differentiated type | 11 |
| Undifferentiated type | 0 |
| Helicobacter pylori (H. pylori) status | |
| Current infection | 3 |
| Past infection | 8 |
| Atrophic border | |
| Closed type | 2 |
| Open type | 9 |
| CASE | Image Size (Pixels) Width (μ ± σ)/Height (μ ± σ) |
|---|---|
| 1 (n = 4) | 886.8 ± 2.2/767.8 ± 1.7 |
| 2 (n = 3) | 887.0 ± 1.7/768.0 ± 1.7 |
| 3 (n = 4) | 1041.0 ± 0.0/901.8 ± 0.5 |
| 4 (n = 5) | 887.0 ± 1.2/767.6 ± 1.3 |
| 5 (n = 1) | 816.0 ± 0.0/692.0 ± 0.0 |
| 6 (n = 1) | 832.0 ± 0.0/760.0 ± 0.0 |
| 7 (n = 1) | 799.0 ± 0.0/691.0 ± 0.0 |
| 8 (n = 1) | 839.0 ± 0.0/720.0 ± 0.0 |
| 9 (n = 1) | 752.0 ± 0.0/602.0 ± 0.0 |
| 10 (n = 1) | 800.0 ± 0.0/650.0 ± 0.0 |
| 11 (n = 1) | 717.0 ± 0.0/515.0 ± 0.0 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Okumura, S.; Goudo, M.; Hiwa, S.; Yasuda, T.; Kitae, H.; Yasuda, Y.; Tomie, A.; Omatsu, T.; Ichikawa, H.; Yagi, N.; et al. Demarcation Line Determination for Diagnosis of Gastric Cancer Disease Range Using Unsupervised Machine Learning in Magnifying Narrow-Band Imaging. Diagnostics 2022, 12, 2491. https://doi.org/10.3390/diagnostics12102491
Okumura S, Goudo M, Hiwa S, Yasuda T, Kitae H, Yasuda Y, Tomie A, Omatsu T, Ichikawa H, Yagi N, et al. Demarcation Line Determination for Diagnosis of Gastric Cancer Disease Range Using Unsupervised Machine Learning in Magnifying Narrow-Band Imaging. Diagnostics. 2022; 12(10):2491. https://doi.org/10.3390/diagnostics12102491
Chicago/Turabian StyleOkumura, Shunsuke, Misa Goudo, Satoru Hiwa, Takeshi Yasuda, Hiroaki Kitae, Yuriko Yasuda, Akira Tomie, Tatsushi Omatsu, Hiroshi Ichikawa, Nobuaki Yagi, and et al. 2022. "Demarcation Line Determination for Diagnosis of Gastric Cancer Disease Range Using Unsupervised Machine Learning in Magnifying Narrow-Band Imaging" Diagnostics 12, no. 10: 2491. https://doi.org/10.3390/diagnostics12102491
APA StyleOkumura, S., Goudo, M., Hiwa, S., Yasuda, T., Kitae, H., Yasuda, Y., Tomie, A., Omatsu, T., Ichikawa, H., Yagi, N., & Hiroyasu, T. (2022). Demarcation Line Determination for Diagnosis of Gastric Cancer Disease Range Using Unsupervised Machine Learning in Magnifying Narrow-Band Imaging. Diagnostics, 12(10), 2491. https://doi.org/10.3390/diagnostics12102491







