Breast Cancer Heterogeneity
Abstract
:1. Introduction
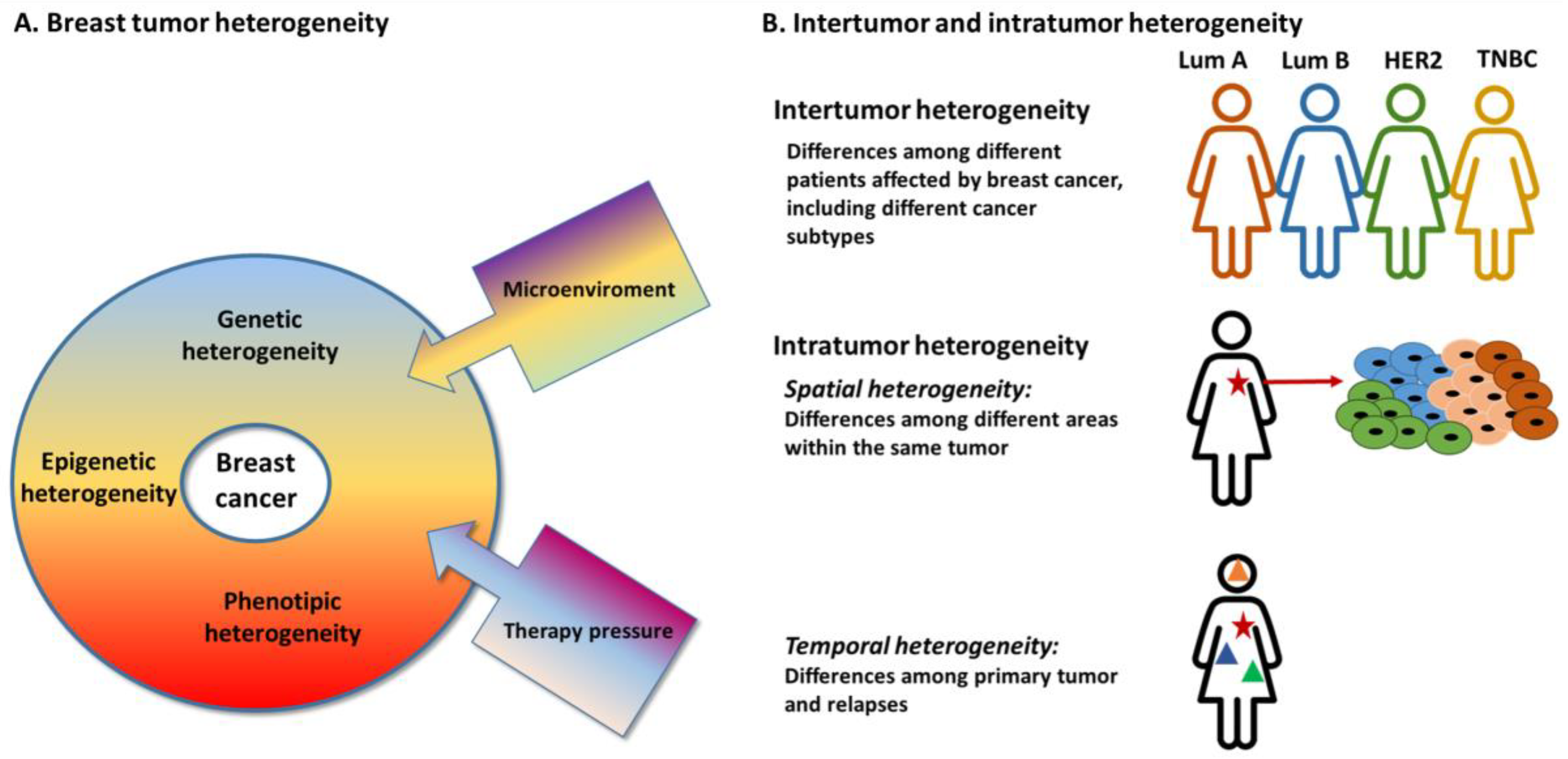
2. Inter-Tumor Heterogeneity
3. Intra-Tumor Heterogeneity
3.1. Spatial Heterogeneity
3.2. Temporal Heterogeneity
4. Clinical Implications of Tumor Heterogeneity
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: Globocan Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef]
- Young, R.H.; Louis, D.N. The Warrens and Other Pioneering Clinician Pathologists of the Massachusetts General Hospital during Its Early Years: An Appreciation on the 200th Anniversary of the Hospital Founding. Mod. Pathol. 2011, 24, 1285–1294. [Google Scholar] [CrossRef]
- Hawkins, R.A.; Killen, E.; Tesdale, A.L.; Sangster, K.; Thomson, M.; Steele, R.J.C.; Blackie, R.A.S. Oestrogen Receptors, Lactate Dehydrogenase and Cellularity in Human Breast Cancer. Clin. Chim. Acta 1988, 175, 89–96. [Google Scholar] [CrossRef]
- Turashvili, G.; Brogi, E. Tumor Heterogeneity in Breast Cancer. Front. Med. 2017, 4, 227. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vogelstein, B.; Papadopoulos, N.; Velculescu, V.E.; Zhou, S.; Diaz, L.A.; Kinzler, K.W. Cancer Genome Landscapes. Science 2013, 339, 1546–1558. [Google Scholar] [CrossRef]
- Marusyk, A.; Almendro, V.; Polyak, K. Intra-Tumour Heterogeneity: A Looking Glass for Cancer? Nat. Rev. Cancer 2012, 12, 323–334. [Google Scholar] [CrossRef] [PubMed]
- Fane, M.; Weeraratna, A.T. How the Ageing Microenvironment Influences Tumour Progression. Nat. Rev. Cancer 2020, 20, 89–106. [Google Scholar] [CrossRef] [PubMed]
- Dunnwald, L.K.; Rossing, M.A.; Li, C.I. Hormone Receptor Status, Tumor Characteristics, and Prognosis: A Prospective Cohort of Breast Cancer Patients. Breast Cancer Res. 2007, 9, R6. [Google Scholar] [CrossRef]
- Davis, S.L.; Eckhardt, S.G.; Tentler, J.J.; Diamond, J.R. Triple-Negative Breast Cancer: Bridging the Gap from Cancer Genomics to Predictive Biomarkers. Ther. Adv. Med. Oncol. 2014, 6, 88–100. [Google Scholar] [CrossRef] [Green Version]
- Vogel, C.L.; Cobleigh, M.A.; Tripathy, D.; Gutheil, J.C.; Harris, L.N.; Fehrenbacher, L.; Slamon, D.J.; Murphy, M.; Novotny, W.F.; Burchmore, M.; et al. Efficacy and Safety of Trastuzumab as a Single Agent in First-Line Treatment of HER2-Overexpressing Metastatic Breast Cancer. JCO 2002, 20, 719–726. [Google Scholar] [CrossRef] [PubMed]
- Early Breast Cancer Trialists’ Collaborative Group. Effects of Chemotherapy and Hormonal Therapy for Early Breast Cancer on Recurrence and 15-Year Survival: An Overview of the Randomised Trials. Lancet 2005, 365, 1687–1717. [Google Scholar] [CrossRef]
- Nanda, R.; Chow, L.Q.M.; Dees, E.C.; Berger, R.; Gupta, S.; Geva, R.; Pusztai, L.; Pathiraja, K.; Aktan, G.; Cheng, J.D.; et al. Pembrolizumab in Patients With Advanced Triple-Negative Breast Cancer: Phase Ib KEYNOTE-012 Study. JCO 2016, 34, 2460–2467. [Google Scholar] [CrossRef]
- Emens, L.A.; Adams, S.; Barrios, C.H.; Diéras, V.; Iwata, H.; Loi, S.; Rugo, H.S.; Schneeweiss, A.; Winer, E.P.; Patel, S.; et al. First-Line Atezolizumab plus Nab-Paclitaxel for Unresectable, Locally Advanced, or Metastatic Triple-Negative Breast Cancer: IMpassion130 Final Overall Survival Analysis. Ann. Oncol. 2021, 32, 983–993. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Bao, H.; Wu, X.; Wang, X.; Shao, Y.; Sun, T. Elevated Tumor Mutation Burden and Immunogenic Activity in Patients with Hormone Receptor-negative or Human Epidermal Growth Factor Receptor 2-positive Breast Cancer. Oncol. Lett. 2019, 18, 449–455. [Google Scholar] [CrossRef] [Green Version]
- Ebili, H.O.; Oluwasola, A.O.; Olopade, O.I. Molecular subtypes and prognosis of breast cancer. In Personalized Management of Breast Cancer; Jatoi, I., Holloway, T.L., Eds.; Future Medicine Ltd.: London, UK, 2014; pp. 21–33. [Google Scholar] [CrossRef]
- Loibl, S.; Treue, D.; Budczies, J.; Weber, K.; Stenzinger, A.; Schmitt, W.D.; Weichert, W.; Jank, P.; Furlanetto, J.; Klauschen, F.; et al. Mutational Diversity and Therapy Response in Breast Cancer: A Sequencing Analysis in the Neoadjuvant GeparSepto Trial. Clin. Cancer Res. 2019, 25, 3986–3995. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fumagalli, C.; Ranghiero, A.; Gandini, S.; Corso, F.; Taormina, S.; De Camilli, E.; Rappa, A.; Vacirca, D.; Viale, G.; Guerini-Rocco, E.; et al. Inter-Tumor Genomic Heterogeneity of Breast Cancers: Comprehensive Genomic Profile of Primary Early Breast Cancers and Relapses. Breast Cancer Res. 2020, 22, 107. [Google Scholar] [CrossRef]
- Denkert, C.; Liedtke, C.; Tutt, A.; von Minckwitz, G. Molecular Alterations in Triple-Negative Breast Cancer—The Road to New Treatment Strategies. Lancet 2017, 389, 2430–2442. [Google Scholar] [CrossRef] [Green Version]
- CBIOPORTAL DATABASE. Available online: http://cbioportal.org (accessed on 17 July 2021).
- Guerini-Rocco, E.; Gray, K.P.; Fumagalli, C.; Reforgiato, M.R.; Leone, I.; Rafaniello Raviele, P.; Munzone, E.; Kammler, R.; Neven, P.; Hitre, E.; et al. Genomic Aberrations and Late Recurrence in Postmenopausal Women with Hormone Receptor–Positive Early Breast Cancer: Results from the SOLE Trial. Clin. Cancer Res. 2021, 27, 504–512. [Google Scholar] [CrossRef] [PubMed]
- Adams, S.; Gray, R.J.; Demaria, S.; Goldstein, L.; Perez, E.A.; Shulman, L.N.; Martino, S.; Wang, M.; Jones, V.E.; Saphner, T.J.; et al. Prognostic Value of Tumor-Infiltrating Lymphocytes in Triple-Negative Breast Cancers from Two Phase III Randomized Adjuvant Breast Cancer Trials: ECOG 2197 and ECOG 1199. JCO 2014, 32, 2959–2966. [Google Scholar] [CrossRef] [PubMed]
- Mao, Y.; Qu, Q.; Chen, X.; Huang, O.; Wu, J.; Shen, K. The Prognostic Value of Tumor-Infiltrating Lymphocytes in Breast Cancer: A Systematic Review and Meta-Analysis. PLoS ONE 2016, 11, e0152500. [Google Scholar] [CrossRef]
- Lotfinejad, P.; Asghari Jafarabadi, M.; Abdoli Shadbad, M.; Kazemi, T.; Pashazadeh, F.; Sandoghchian Shotorbani, S.; Jadidi Niaragh, F.; Baghbanzadeh, A.; Vahed, N.; Silvestris, N.; et al. Prognostic Role and Clinical Significance of Tumor-Infiltrating Lymphocyte (TIL) and Programmed Death Ligand 1 (PD-L1) Expression in Triple-Negative Breast Cancer (TNBC): A Systematic Review and Meta-Analysis Study. Diagnostics 2020, 10, 704. [Google Scholar] [CrossRef]
- Lu, P.; Weaver, V.M.; Werb, Z. The extracellular matrix: A dynamic niche in cancer progression. J. Cell Biol. 2012, 196, 395–406. [Google Scholar] [CrossRef]
- Winkler, J.; Abisoye-Ogunniyan, A.; Metcalf, K.J.; Werb, Z. Concepts of extracellular matrix remodelling in tumour progression and metastasis. Nat. Commun. 2020, 11, 5120. [Google Scholar] [CrossRef]
- Conklin, M.W.; Eickhoff, J.; Riching, K.M.; Pehlke, C.A.; Eliceiri, K.W.; Provenzano, P.P.; Friedl, A.; Keely, P.J. Aligned collagen is a prognostic signature for survival in human breast carcinoma. Am. J. Pathol. 2011, 178, 1221–1232. [Google Scholar] [CrossRef]
- Sachs, N.; de Ligt, J.; Kopper, O.; Gogola, E.; Bounova, G.; Weeber, F.; Balgobind, A.V.; Wind, K.; Gracanin, A.; Begthel, H.; et al. Living Biobank of Breast Cancer Organoids Captures Disease Heterogeneity. Cell 2018, 172, 373–386.e10. [Google Scholar] [CrossRef] [Green Version]
- Keren, L.; Bosse, M.; Marquez, D.; Angoshtari, R.; Jain, S.; Varma, S.; Yang, S.R.; Kurian, A.; Van Valen, D.; West, R.; et al. Structured Tumor-Immune Microenvironment in Triple Negative Breast Cancer Revealed by Multiplexed Ion Beam Imaging. Cell 2018, 174, 1373–1387.e19. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yates, L.R. Intratumoral Heterogeneity and Subclonal Diversification of Early Breast Cancer. Breast 2017, 34, S36–S42. [Google Scholar] [CrossRef]
- Patani, N.; Barbashina, V.; Lambros, M.B.K.; Gauthier, A.; Mansour, M.; Mackay, A.; Reis-Filho, J.S. Direct Evidence for Concurrent Morphological and Genetic Heterogeneity in an Invasive Ductal Carcinoma of Triple-Negative Phenotype. J. Clin. Pathol. 2011, 64, 822–828. [Google Scholar] [CrossRef] [PubMed]
- Nik-Zainal, S.; Alexandrov, L.B.; Wedge, D.C.; Van Loo, P.; Greenman, C.D.; Raine, K.; Jones, D.; Hinton, J.; Marshall, J.; Stebbings, L.A.; et al. Mutational Processes Molding the Genomes of 21 Breast Cancers. Cell 2012, 149, 979–993. [Google Scholar] [CrossRef] [Green Version]
- Nik-Zainal, S.; Van Loo, P.; Wedge, D.C.; Alexandrov, L.B.; Greenman, C.D.; Lau, K.W.; Raine, K.; Jones, D.; Marshall, J.; Ramakrishna, M.; et al. The Life History of 21 Breast Cancers. Cell 2012, 149, 994–1007. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yates, L.R.; Gerstung, M.; Knappskog, S.; Desmedt, C.; Gundem, G.; Van Loo, P.; Aas, T.; Alexandrov, L.B.; Larsimont, D.; Davies, H.; et al. Subclonal Diversification of Primary Breast Cancer Revealed by Multiregion Sequencing. Nat. Med. 2015, 21, 751–759. [Google Scholar] [CrossRef]
- Karthikeyan, S.; Waters, I.G.; Dennison, L.; Chu, D.; Donaldson, J.; Shin, D.H.; Rosen, D.M.; Gonzalez-Ericsson, P.I.; Sanchez, V.; Sanders, M.E.; et al. Hierarchical Tumor Heterogeneity Mediated by Cell Contact between Distinct Genetic Subclones. J. Clin. Investig. 2021, 131, e143557. [Google Scholar] [CrossRef] [PubMed]
- Salgado, R.; Denkert, C.; Demaria, S.; Sirtaine, N.; Klauschen, F.; Pruneri, G.; Wienert, S.; Van den Eynden, G.; Baehner, F.L.; Penault-Llorca, F.; et al. The Evaluation of Tumor-Infiltrating Lymphocytes (TILs) in Breast Cancer: Recommendations by an International TILs Working Group 2014. Ann. Oncol. 2015, 26, 259–271. [Google Scholar] [CrossRef]
- Kato, T.; Park, J.-H.; Kiyotani, K.; Ikeda, Y.; Miyoshi, Y.; Nakamura, Y. Integrated Analysis of Somatic Mutations and Immune Microenvironment of Multiple Regions in Breast Cancers. Oncotarget 2017, 8, 62029–62038. [Google Scholar] [CrossRef] [Green Version]
- Berben, L.; Wildiers, H.; Marcelis, L.; Antoranz, A.; Bosisio, F.; Hatse, S.; Floris, G. Computerised Scoring Protocol for Identification and Quantification of Different Immune Cell Populations in Breast Tumour Regions by the Use of QuPath Software. Histopathology 2020, 77, 79–91. [Google Scholar] [CrossRef]
- Fusco, N.; Lopez, G.; Corti, C.; Pesenti, C.; Colapietro, P.; Ercoli, G.; Gaudioso, G.; Faversani, A.; Gambini, D.; Michelotti, A.; et al. Mismatch Repair Protein Loss as a Prognostic and Predictive Biomarker in Breast Cancers Regardless of Microsatellite Instability. JNCI Cancer Spectr. 2018, 2, pky056. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, M.; Li, A.; Zhou, S.; Xu, Y.; Xiao, Y.; Bi, R.; Yang, W. Heterogeneity of PD-L1 Expression in Primary Tumors and Paired Lymph Node Metastases of Triple Negative Breast Cancer. BMC Cancer 2018, 18, 4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gyanchandani, R.; Lin, Y.; Lin, H.M.; Cooper, K.; Normolle, D.P.; Brufsky, A.; Fastuca, M.; Crosson, W.; Oesterreich, S.; Davidson, N.E.; et al. Intratumor Heterogeneity Affects Gene Expression Profile Test Prognostic Risk Stratification in Early Breast Cancer. Clin. Cancer Res. 2016, 22, 5362–5369. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Seol, H.; Lee, H.J.; Choi, Y.; Lee, H.E.; Kim, Y.J.; Kim, J.H.; Kang, E.; Kim, S.-W.; Park, S.Y. Intratumoral Heterogeneity of HER2 Gene Amplification in Breast Cancer: Its Clinicopathological Significance. Mod. Pathol. 2012, 25, 938–948. [Google Scholar] [CrossRef] [Green Version]
- Lee, H.J.; Seo, A.N.; Kim, E.J.; Jang, M.H.; Suh, K.J.; Ryu, H.S.; Kim, Y.J.; Kim, J.H.; Im, S.-A.; Gong, G.; et al. HER2 Heterogeneity Affects Trastuzumab Responses and Survival in Patients With HER2-Positive Metastatic Breast Cancer. Am. J. Clin. Pathol. 2014, 142, 755–766. [Google Scholar] [CrossRef] [Green Version]
- Hou, Y.; Nitta, H.; Wei, L.; Banks, P.M.; Portier, B.; Parwani, A.V.; Li, Z. HER2 Intratumoral Heterogeneity Is Independently Associated with Incomplete Response to Anti-HER2 Neoadjuvant Chemotherapy in HER2-Positive Breast Carcinoma. Breast Cancer Res. Treat. 2017, 166, 447–457. [Google Scholar] [CrossRef]
- Metzger Filho, O.; Viale, G.; Stein, S.; Trippa, L.; Yardley, D.A.; Mayer, I.A.; Abramson, V.G.; Arteaga, C.L.; Spring, L.M.; Waks, A.G.; et al. Impact of HER2 Heterogeneity on Treatment Response of Early-Stage HER2-Positive Breast Cancer: Phase II Neoadjuvant Clinical Trial of T-DM1 Combined with Pertuzumab. Cancer Discov. 2021. [Google Scholar] [CrossRef]
- Brouckaert, O.; Paridaens, R.; Floris, G.; Rakha, E.; Osborne, K.; Neven, P. A Critical Review Why Assessment of Steroid Hormone Receptors in Breast Cancer Should Be Quantitative. Ann. Oncol. 2013, 24, 47–53. [Google Scholar] [CrossRef]
- Clarke, R.; Liu, M.C.; Bouker, K.B.; Gu, Z.; Lee, R.Y.; Zhu, Y.; Skaar, T.C.; Gomez, B.; O’Brien, K.; Wang, Y.; et al. Antiestrogen Resistance in Breast Cancer and the Role of Estrogen Receptor Signaling. Oncogene 2003, 22, 7316–7339. [Google Scholar] [CrossRef] [Green Version]
- Allegra, J.C.; Lippman, M.E.; Thompson, E.B.; Simon, R.; Barlock, A.; Green, L.; Huff, K.K.; Do, H.M.Y.T.; Aitken, S.C.; Warren, R. Estrogen Receptor Status: An Important Variable in Predicting Response to Endocrine Therapy in Metastatic Breast Cancer. Eur. J. Cancer 1980, 16, 323–331. [Google Scholar] [CrossRef]
- Schrijver, W.A.M.E.; Suijkerbuijk, K.P.M.; van Gils, C.H.; van der Wall, E.; Moelans, C.B.; van Diest, P.J. Receptor Conversion in Distant Breast Cancer Metastases: A Systematic Review and Meta-Analysis. JNCI J. Natl. Cancer Inst. 2018, 110, 568–580. [Google Scholar] [CrossRef] [Green Version]
- Grinda, T.; Joyon, N.; Lusque, A.; Lefèvre, S.; Arnould, L.; Penault-Llorca, F.; Macgrogan, G.; Treilleux, I.; Vincent-Salomon, A.; Haudebourg, J.; et al. Phenotypic Discordance between Primary and Metastatic Breast Cancer in the Large-Scale Real-Life Multicenter French ESME Cohort. NPJ Breast Cancer 2021, 7, 41. [Google Scholar] [CrossRef]
- Gomez-Fernandez, C.; Daneshbod, Y.; Nassiri, M.; Milikowski, C.; Alvarez, C.; Nadji, M. Immunohistochemically Determined Estrogen Receptor Phenotype Remains Stable in Recurrent and Metastatic Breast Cancer. Am. J. Clin. Pathol. 2008, 130, 879–882. [Google Scholar] [CrossRef] [Green Version]
- Gerlinger, M.; Swanton, C. How Darwinian Models Inform Therapeutic Failure Initiated by Clonal Heterogeneity in Cancer Medicine. Br. J. Cancer 2010, 103, 1139–1143. [Google Scholar] [CrossRef] [Green Version]
- Carlsson, J.; Nordgren, H.; Sjöström, J.; Wester, K.; Villman, K.; Bengtsson, N.O.; Ostenstad, B.; Lundqvist, H.; Blomqvist, C. HER2 Expression in Breast Cancer Primary Tumours and Corresponding Metastases. Original Data and Literature Review. Br. J. Cancer 2004, 90, 2344–2348. [Google Scholar] [CrossRef]
- van Agthoven, T.; Timmermans, M.; Dorssers, L.C.J.; Henzen-Logmans, S.C. Expression of Estrogen, Progesterone and Epidermal Growth Factor Receptors in Primary and Metastatic Breast Cancer. Int. J. Cancer 1995, 63, 790–793. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.-F.; Liao, Y.-Y.; Yang, M.; Peng, N.-F.; Xie, S.-R.; Xie, Y.-F. Discordances in ER, PR and HER2 Receptors between Primary and Recurrent/Metastatic Lesions and Their Impact on Survival in Breast Cancer Patients. Med. Oncol. 2014, 31, 214. [Google Scholar] [CrossRef]
- Peng, L.; Zhang, Z.; Zhao, D.; Zhao, J.; Mao, F.; Sun, Q. Discordance in ER, PR, HER2, and Ki-67 Expression Between Primary and Recurrent/Metastatic Lesions in Patients with Primary Early Stage Breast Cancer and the Clinical Significance: Retrospective Analysis of 75 Cases. Pathol. Oncol. Res. 2021, 27, 599894. [Google Scholar] [CrossRef] [PubMed]
- Sari, E.; Guler, G.; Hayran, M.; Gullu, I.; Altundag, K.; Ozisik, Y. Comparative Study of the Immunohistochemical Detection of Hormone Receptor Status and HER-2 Expression in Primary and Paired Recurrent/Metastatic Lesions of Patients with Breast Cancer. Med. Oncol. 2011, 28, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Aurilio, G.; Disalvatore, D.; Pruneri, G.; Bagnardi, V.; Viale, G.; Curigliano, G.; Adamoli, L.; Munzone, E.; Sciandivasci, A.; De Vita, F.; et al. A Meta-Analysis of Oestrogen Receptor, Progesterone Receptor and Human Epidermal Growth Factor Receptor 2 Discordance between Primary Breast Cancer and Metastases. Eur. J. Cancer 2014, 50, 277–289. [Google Scholar] [CrossRef] [Green Version]
- Jørgensen, C.L.T.; Larsson, A.-M.; Forsare, C.; Aaltonen, K.; Jansson, S.; Bradshaw, R.; Bendahl, P.-O.; Rydén, L. PAM50 Intrinsic Subtype Profiles in Primary and Metastatic Breast Cancer Show a Significant Shift toward More Aggressive Subtypes with Prognostic Implications. Cancers 2021, 13, 1592. [Google Scholar] [CrossRef]
- Rossi, L.; Verrico, M.; Tomao, S.; Ricci, F.; Fontana, A.; Spinelli, G.P.; Colonna, M.; Vici, P.-; Tomao, F. Expression of ER, PgR, HER-2, and Ki-67 in core biopsies and in definitive histological specimens in patients with locally advanced breast cancer treated with neoadjuvant chemotherapy. Cancer Chemother. Pharmacol. 2020, 85, 105–111. [Google Scholar] [CrossRef]
- Mohan, S.C.; Walcott-Sapp, S.; Lee, M.K.; Srour, M.K.; Kim, S.; Amersi, F.F.; Giuliano, A.E.; Chung, A.P. Alterations in Breast Cancer Biomarkers Following Neoadjuvant Therapy. Ann. Surg. Oncol. 2021. [Google Scholar] [CrossRef]
- Boman, C.; Zerdes, I.; Mårtensson, K.; Bergh, J.; Foukakis, T.; Valachis, A.; Matikas, A. Discordance of PD-L1 Status between Primary and Metastatic Breast Cancer: A Systematic Review and Meta-Analysis. Cancer Treat. Rev. 2021, 99, 102257. [Google Scholar] [CrossRef]
- Aftimos, P.; Oliveira, M.; Irrthum, A.; Fumagalli, D.; Sotiriou, C.; Nili Gal-Yam, E.; Robson, M.E.; Ndozeng, J.; Di Leo, A.; Ciruelos, E.M.; et al. Genomic and Transcriptomic Analyses of Breast Cancer Primaries and Matched Metastases in AURORA, the Breast International Group (BIG) Molecular Screening Initiative. Cancer Discov. 2021. [Google Scholar] [CrossRef]
- Agahozo, M.C.; Sieuwerts, A.M.; Doebar, S.C.; Verhoef, E.I.; Beaufort, C.M.; Ruigrok-Ritstier, K.; de Weerd, V.; Sleddens, H.F.B.M.; Dinjens, W.N.M.; Martens, J.W.M.; et al. PIK3CA Mutations in Ductal Carcinoma in Situ and Adjacent Invasive Breast Cancer. Endocr.-Relat. Cancer 2019, 26, 471–482. [Google Scholar] [CrossRef] [PubMed]
- Heselmeyer-Haddad, K.; Berroa Garcia, L.Y.; Bradley, A.; Ortiz-Melendez, C.; Lee, W.-J.; Christensen, R.; Prindiville, S.A.; Calzone, K.A.; Soballe, P.W.; Hu, Y.; et al. Single-Cell Genetic Analysis of Ductal Carcinoma in Situ and Invasive Breast Cancer Reveals Enormous Tumor Heterogeneity yet Conserved Genomic Imbalances and Gain of MYC during Progression. Am. J. Pathol. 2012, 181, 1807–1822. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hernandez, L.; Wilkerson, P.M.; Lambros, M.B.; Campion-Flora, A.; Rodrigues, D.N.; Gauthier, A.; Cabral, C.; Pawar, V.; Mackay, A.; A’Hern, R.; et al. Genomic and Mutational Profiling of Ductal Carcinomas in Situ and Matched Adjacent Invasive Breast Cancers Reveals Intra-Tumour Genetic Heterogeneity and Clonal Selection. J. Pathol. 2012, 227, 42–52. [Google Scholar] [CrossRef] [Green Version]
- Available online: http://oncokb.org (accessed on 17 July 2021).
- Balko, J.M.; Giltnane, J.M.; Wang, K.; Schwarz, L.J.; Young, C.D.; Cook, R.S.; Owens, P.; Sanders, M.E.; Kuba, M.G.; Sánchez, V.; et al. Molecular Profiling of the Residual Disease of Triple-Negative Breast Cancers after Neoadjuvant Chemotherapy Identifies Actionable Therapeutic Targets. Cancer Discov. 2014, 4, 232–245. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Waks, A.G.; Cohen, O.; Kochupurakkal, B.; Kim, D.; Dunn, C.E.; Buendia Buendia, J.; Wander, S.; Helvie, K.; Lloyd, M.R.; Marini, L.; et al. Reversion and Non-Reversion Mechanisms of Resistance to PARP Inhibitor or Platinum Chemotherapy in BRCA1/2-Mutant Metastatic Breast Cancer. Ann. Oncol. 2020, 31, 590–598. [Google Scholar] [CrossRef]
- De Santo, I.; McCartney, A.; Migliaccio, I.; Di Leo, A.; Malorni, L. The Emerging Role of ESR1 Mutations in Luminal Breast Cancer as a Prognostic and Predictive Biomarker of Response to Endocrine Therapy. Cancers 2019, 11, 1894. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jeselsohn, R.; Buchwalter, G.; De Angelis, C.; Brown, M.; Schiff, R. ESR1 Mutations—A Mechanism for Acquired Endocrine Resistance in Breast Cancer. Nat. Rev. Clin. Oncol 2015, 12, 573–583. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Turner, N.C.; Swift, C.; Kilburn, L.; Fribbens, C.; Beaney, M.; Garcia-Murillas, I.; Budzar, A.U.; Robertson, J.F.R.; Gradishar, W.; Piccart, M.; et al. ESR1 Mutations and Overall Survival on Fulvestrant versus Exemestane in Advanced Hormone Receptor–Positive Breast Cancer: A Combined Analysis of the Phase III SoFEA and EFECT Trials. Clin. Cancer Res. 2020, 26, 5172–5177. [Google Scholar] [CrossRef] [PubMed]
- Turner, N.C.; Reis-Filho, J.S. Genetic Heterogeneity and Cancer Drug Resistance. Lancet Oncol. 2012, 13, e178–e185. [Google Scholar] [CrossRef]
- Dagogo-Jack, I.; Shaw, A.T. Tumour Heterogeneity and Resistance to Cancer Therapies. Nat. Rev. Clin. Oncol. 2018, 15, 81–94. [Google Scholar] [CrossRef]
- Yates, L.R.; Campbell, P.J. Evolution of the Cancer Genome. Nat. Rev. Genet. 2012, 13, 795–806. [Google Scholar] [CrossRef] [Green Version]
- Swanton, C.; Burrell, R.A.; Futreal, P.A. Breast Cancer Genome Heterogeneity: A Challenge to Personalized Medicine? Breast Cancer Res. 2011, 13, 104. [Google Scholar] [CrossRef] [Green Version]
- Gao, H.; Korn, J.M.; Ferretti, S.; Monahan, J.E.; Wang, Y.; Singh, M.; Zhang, C.; Schnell, C.; Yang, G.; Zhang, Y.; et al. High-throughput screening using patient-derived tumor xenografts to predict clinical trial drug response. Nat. Med. 2015, 21, 1318–1325. [Google Scholar] [CrossRef] [PubMed]
- Bruna, A.; Rueda, O.M.; Greenwood, W.; Batra, A.S.; Callari, M.; Batra, R.N.; Pogrebniak, K.; Sandoval, J.; Cassidy, J.W.; Tufegdzic-Vidakovic, A.; et al. Biobank of Breast Cancer Explants with Preserved Intra-tumor Heterogeneity to Screen Anticancer Compounds. Cell 2016, 167, 260–274.e22. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Georgopoulou, D.; Callari, M.; Rueda, O.M.; Shea, A.; Martin, A.; Giovannetti, A.; Qosaj, F.; Dariush, A.; Chin, S.F.; Carnevalli, L.S.; et al. Landscapes of cellular phenotypic diversity in breast cancer xenografts and their impact on drug response. Nat. Commun. 2021, 12, 1998. [Google Scholar] [CrossRef]
- Zhong, Q.; Rüschoff, J.H.; Guo, T.; Gabrani, M.; Schüffler, P.J.; Rechsteiner, M.; Liu, Y.; Fuchs, T.J.; Rupp, N.J.; Fankhauser, C.; et al. Image-Based Computational Quantification and Visualization of Genetic Alterations and Tumour Heterogeneity. Sci. Rep. 2016, 6, 24146. [Google Scholar] [CrossRef] [Green Version]
- Heindl, A.; Nawaz, S.; Yuan, Y. Mapping Spatial Heterogeneity in the Tumor Microenvironment: A New Era for Digital Pathology. Lab. Investig. 2015, 95, 377–384. [Google Scholar] [CrossRef] [Green Version]
- Crowley, E.; Di Nicolantonio, F.; Loupakis, F.; Bardelli, A. Liquid Biopsy: Monitoring Cancer-Genetics in the Blood. Nat. Rev. Clin. Oncol. 2013, 10, 472–484. [Google Scholar] [CrossRef]
- Smit, D.J.; Pantel, K.; Jücker, M. Circulating Tumor Cells as a Promising Target for Individualized Drug Susceptibility Tests in Cancer Therapy. Biochem. Pharmacol. 2021, 188, 114589. [Google Scholar] [CrossRef] [PubMed]
- Forshew, T.; Murtaza, M.; Parkinson, C.; Gale, D.; Tsui, D.W.Y.; Kaper, F.; Dawson, S.-J.; Piskorz, A.M.; Jimenez-Linan, M.; Bentley, D.; et al. Noninvasive Identification and Monitoring of Cancer Mutations by Targeted Deep Sequencing of Plasma DNA. Sci. Transl. Med. 2012, 4, 136ra68. [Google Scholar] [CrossRef] [PubMed]
- Onstenk, W.; Gratama, J.W.; Foekens, J.A.; Sleijfer, S. Towards a Personalized Breast Cancer Treatment Approach Guided by Circulating Tumor Cell (CTC) Characteristics. Cancer Treat. Rev. 2013, 39, 691–700. [Google Scholar] [CrossRef] [PubMed]
- Schiavon, G.; Hrebien, S.; Garcia-Murillas, I.; Cutts, R.J.; Pearson, A.; Tarazona, N.; Fenwick, K.; Kozarewa, I.; Lopez-Knowles, E.; Ribas, R.; et al. Analysis of ESR1 Mutation in Circulating Tumor DNA Demonstrates Evolution during Therapy for Metastatic Breast Cancer. Sci. Transl. Med. 2015, 7, 313ra182. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lambin, P.; Leijenaar, R.T.H.; Deist, T.M.; Peerlings, J.; de Jong, E.E.C.; van Timmeren, J.; Sanduleanu, S.; Larue, R.T.H.M.; Even, A.J.G.; Jochems, A.; et al. Radiomics: The Bridge between Medical Imaging and Personalized Medicine. Nat. Rev. Clin. Oncol. 2017, 14, 749–762. [Google Scholar] [CrossRef] [PubMed]
- Aerts, H.J.W.L.; Velazquez, E.R.; Leijenaar, R.T.H.; Parmar, C.; Grossmann, P.; Carvalho, S.; Bussink, J.; Monshouwer, R.; Haibe-Kains, B.; Rietveld, D.; et al. Decoding Tumour Phenotype by Noninvasive Imaging Using a Quantitative Radiomics Approach. Nat. Commun. 2014, 5, 4006. [Google Scholar] [CrossRef]
- Fan, M.; Zhang, P.; Wang, Y.; Peng, W.; Wang, S.; Gao, X.; Xu, M.; Li, L. Radiomic Analysis of Imaging Heterogeneity in Tumours and the Surrounding Parenchyma Based on Unsupervised Decomposition of DCE-MRI for Predicting Molecular Subtypes of Breast Cancer. Eur. Radiol. 2019, 29, 4456–4467. [Google Scholar] [CrossRef] [Green Version]
- Fan, M.; Xia, P.; Clarke, R.; Wang, Y.; Li, L. Radiogenomic Signatures Reveal Multiscale Intratumour Heterogeneity Associated with Biological Functions and Survival in Breast Cancer. Nat. Commun. 2020, 11, 4861. [Google Scholar] [CrossRef]
| Luminal-A | Luminal-B | HER2+ Non-Luminal | Triple Negative | |
|---|---|---|---|---|
| Hormone receptors | ER+ and/or PR+ | ER+ and/or PR+ | ER−, PR− | ER−, PR− |
| HER2 | HER2− | HER2+/− | HER2+ | HER2− |
| Ki67 | Ki67 < 20 | Ki67 ≥ 20 | ||
| Prevalence among populations [15] | Non Africans 47–61% | All 8–18% | Asians 19–36% | Africans 27–37% |
| Africans 26–27% | Caucasians 13.7–19% | Others 8–20% | ||
| Africans 15–23% | ||||
| Most frequently/peculiar mutated genes [19] | PIK3CA (47.5%) | TP53 (36.0%) | TP53 (70.5%) | TP53 (89.5%) |
| CDH1 (21.8%) | PIK3CA (29.9%) | PIK3CA (33.3%) | TTN (24.0%) | |
| GATA3 (15.4%) | TTN (20.3%) | TTN (28.6%) | BRCA1 (7.6%) | |
| MAP3K1 (14.2%) | GATA3 (20.3%) | MUC1 (19.2%) | NF1 (4.7%) | |
| Most frequently/peculiar CNA [19] | CCND1 (17.0%) | CCND1 (25.9%) | ERBB2 (70.5%) | MYC (35.7%) |
| FGF19 (16.8%) | FGF4 (24.4%) | |||
| FGF3 (16.6%) | FGF3 (23.9%) | |||
| FGF4 (16.6%) | FGFR1 (19.3%) | |||
| Biologic pathway [15] | ER signaling | ER signaling | HER-2 signaling | Immune response |
| ECM | ECM | Proliferation | ECM | |
| Proliferation | Immune response | Proliferation | ||
| Tumor invasion |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fumagalli, C.; Barberis, M. Breast Cancer Heterogeneity. Diagnostics 2021, 11, 1555. https://doi.org/10.3390/diagnostics11091555
Fumagalli C, Barberis M. Breast Cancer Heterogeneity. Diagnostics. 2021; 11(9):1555. https://doi.org/10.3390/diagnostics11091555
Chicago/Turabian StyleFumagalli, Caterina, and Massimo Barberis. 2021. "Breast Cancer Heterogeneity" Diagnostics 11, no. 9: 1555. https://doi.org/10.3390/diagnostics11091555
APA StyleFumagalli, C., & Barberis, M. (2021). Breast Cancer Heterogeneity. Diagnostics, 11(9), 1555. https://doi.org/10.3390/diagnostics11091555







