miRNomic Signature in Very Low Birth-Weight Neonates Discriminates Late-Onset Gram-Positive Sepsis from Controls
Abstract
1. Introduction
2. Materials and Methods
2.1. The Study Design and Patients’ Characteristics
2.2. RNA Extraction
2.3. Small Non-Coding RNAs and mRNA Expression Profiling
2.4. Data Analysis of Microarrays
2.5. Data Analysis of Clinical and Demographic Analysis
3. Results
3.1. Patients’ Characteristics
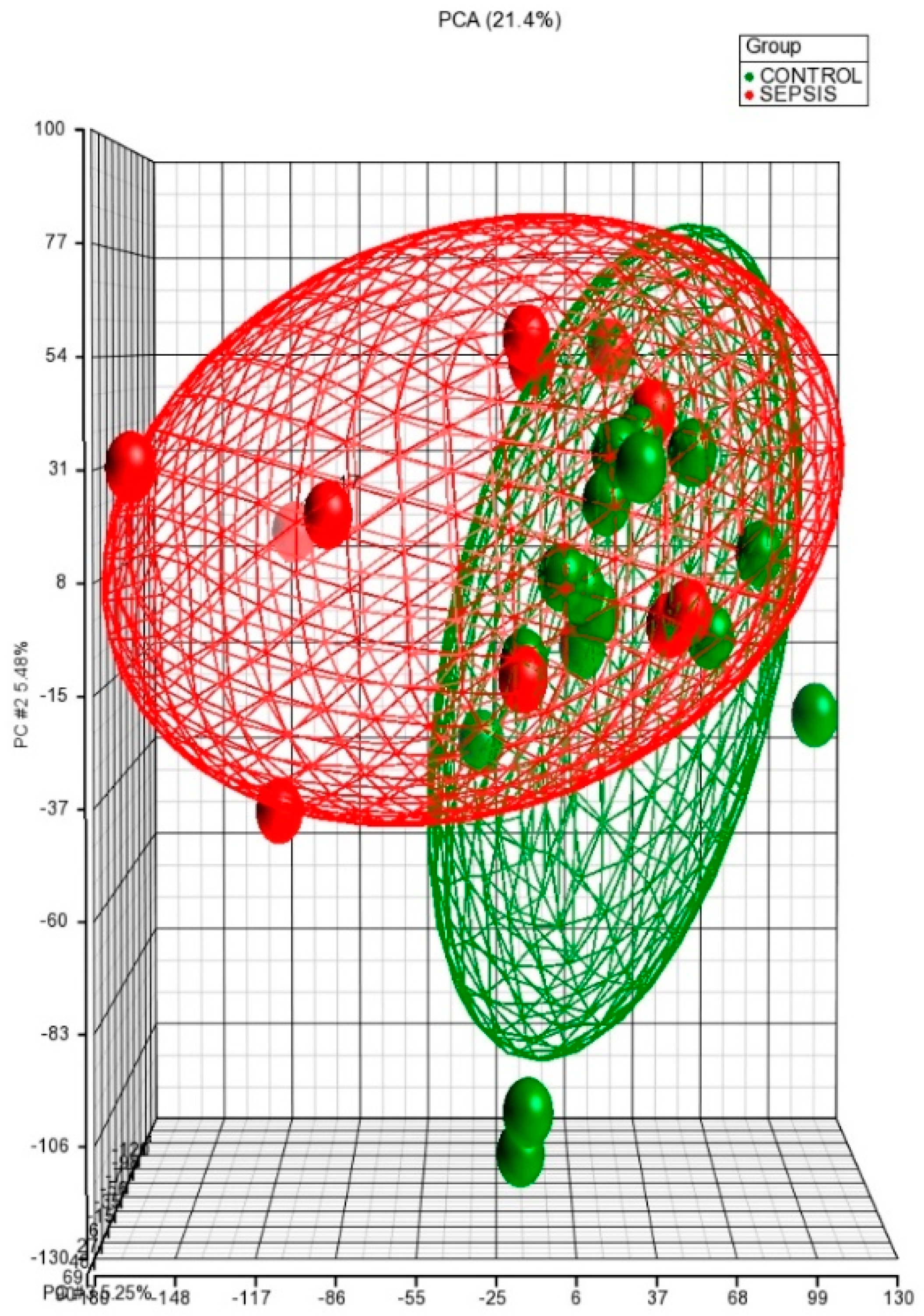
3.2. Principal Component Analysis (PCA) of Neonatal miRNome
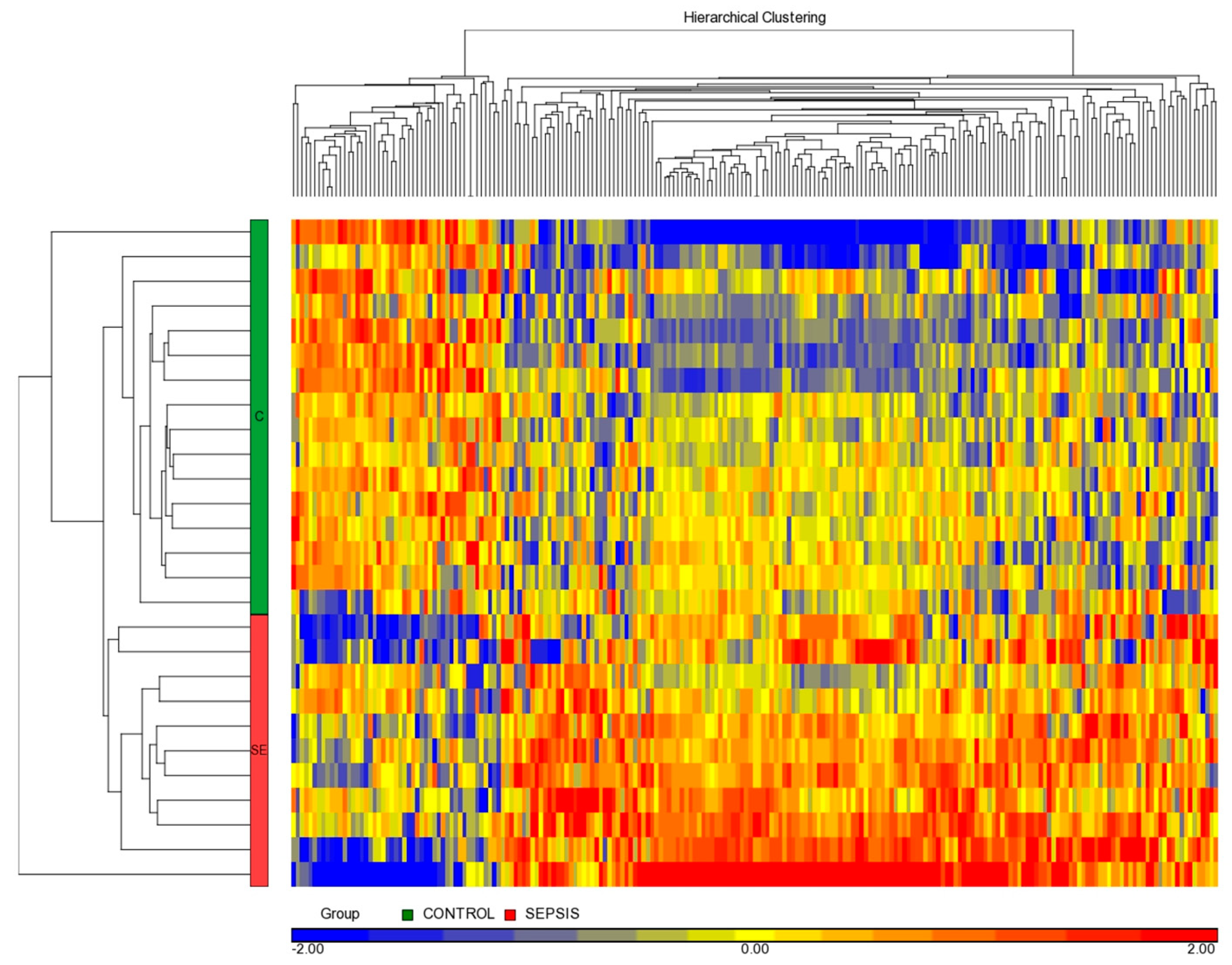
3.3. miRNome Differential Analysis
3.4. Combined Analysis: miRNAs Susceptible to Regulate mRNAs in Gram-Positive Sepsis versus Controls Neonates
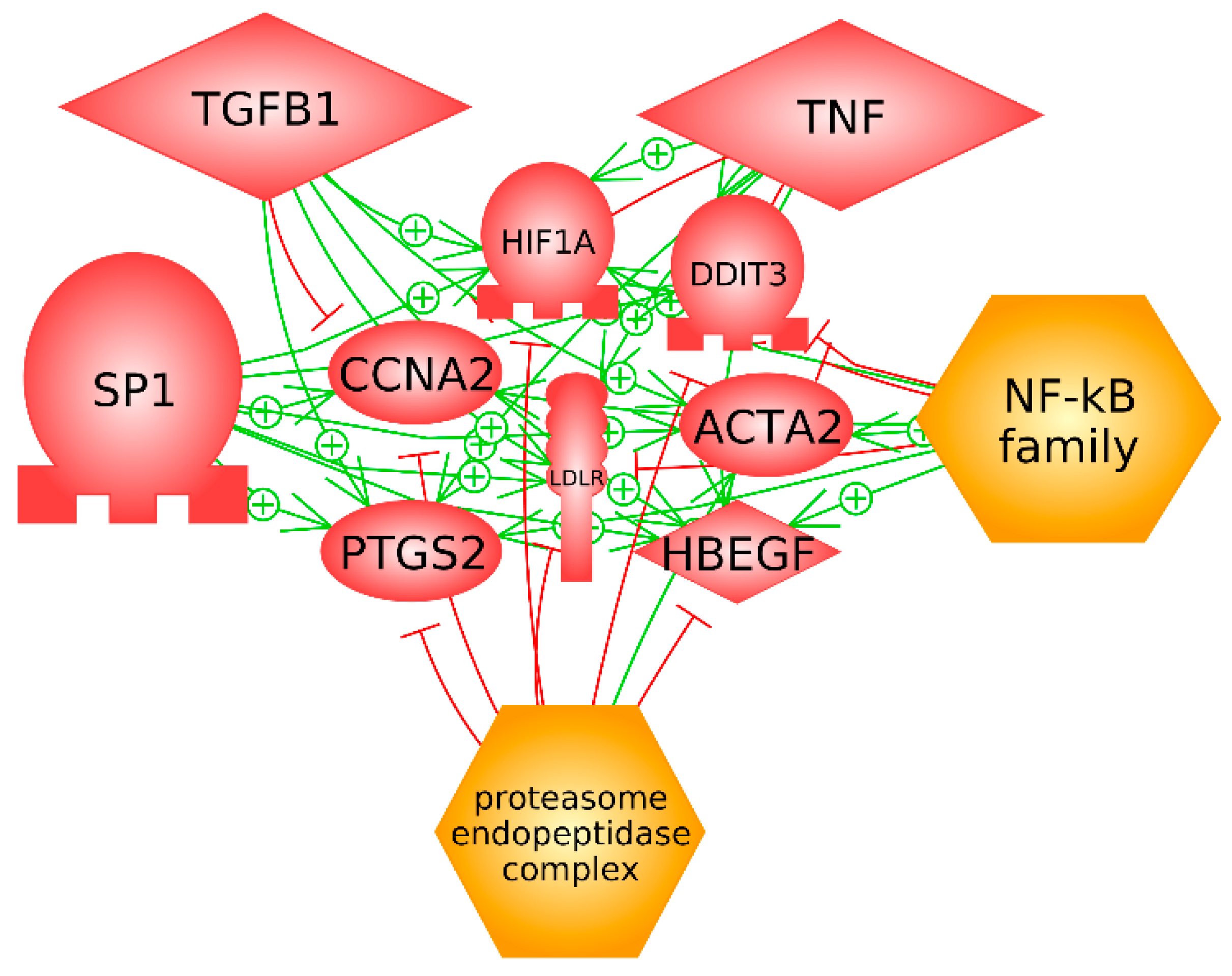
3.5. Biological Analysis from a Combined Analysis
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Shane, A.L.; Sánchez, P.J.; Stoll, B.J. Neonatal sepsis. Lancet 2017, 390, 1770–1780. [Google Scholar] [CrossRef]
- Schlapbach, L.J.; Aebischer, M.; Adams, M.; Natalucci, G.; Bonhoeffer, J.; Latzin, P.; Nelle, M.; Bucher, H.U.; Latal, B.; The Swiss Neonatal Network and Follow-Up Group. Impact of Sepsis on Neurodevelopmental Outcome in a Swiss National Cohort of Extremely Premature Infants. Pediatrics 2011, 128, e348–e357. [Google Scholar] [CrossRef]
- Goldstein, B.; Giroir, B.; Randolph, A.; International Consensus Conference on Pediatric Sepsis. International pediatric sepsis consensus conference: Definitions for sepsis and organ dysfunction in pediatrics. Pediatr. Crit. Care Med. 2005, 6, 2–8. [Google Scholar] [CrossRef]
- Kumar, Y.; Qunibi, M.; Neal, T.J.; Yoxall, C.W. Time to positivity of neonatal blood cultures. Arch. Dis. Child. Fetal Neonatal Ed. 2001, 85, F182–F186. [Google Scholar] [CrossRef]
- Connell, T.G.; Rele, M.; Cowley, D.; Buttery, J.P.; Curtis, N. How Reliable Is a Negative Blood Culture Result? Volume of Blood Submitted for Culture in Routine Practice in a Children’s Hospital. Pediatrics 2007, 119, 891–896. [Google Scholar] [CrossRef] [PubMed]
- Ng, P.C.; Ma, T.P.Y.; Lam, H. The use of laboratory biomarkers for surveillance, diagnosis and prediction of clinical outcomes in neonatal sepsis and necrotising enterocolitis. Arch. Dis. Child. Fetal Neonatal Ed. 2015, 100, F448–F452. [Google Scholar] [CrossRef] [PubMed]
- Dellinger, R.P.; The Surviving Sepsis Campaign Guidelines Committee including The Pediatric Subgroup; Levy, M.M.; Rhodes, A.; Annane, D.; Gerlach, H.; Opal, S.M.; Sevransky, J.E.; Sprung, C.L.; Douglas, I.; et al. Surviving Sepsis Campaign: International Guidelines for Management of Severe Sepsis and Septic Shock, 2012. Intensiv. Care Med. 2013, 39, 165–228. [Google Scholar] [CrossRef]
- Simon, L.; Gauvin, F.; Amre, D.K.; Saint-Louis, P.; Lacroix, J. Serum Procalcitonin and C-Reactive Protein Levels as Markers of Bacterial Infection: A Systematic Review and Meta-Analysis. Clin. Infect. Dis. 2004, 39, 206–217. [Google Scholar] [CrossRef]
- Benz, F.; Roy, S.; Trautwein, C.; Roderburg, C.; Luedde, T. Circulating MicroRNAs as Biomarkers for Sepsis. Int. J. Mol. Sci. 2016, 17, 78. [Google Scholar] [CrossRef]
- Jawa, R.S.; Anillo, S.; Huntoon, K.; Baumann, H.; Kulaylat, M.N. Interleukin-6 in Surgery, Trauma, and Critical Care Part II: Clinical Implications. J. Intensiv. Care Med. 2010, 26, 73–87. [Google Scholar] [CrossRef]
- Cernada, M.; Badía, N.; Modesto, V.; Alonso, R.; Mejías, A.; Golombek, S.; Vento, M. Cord blood interleukin-6 as a predictor of early-onset neonatal sepsis. Acta Paediatr. 2012, 101, e203–e207. [Google Scholar] [CrossRef] [PubMed]
- Caldas, J.P.S.; Marba, S.T.M.; Blotta, M.H.S.L.; Calil, R.; Morais, S.S.; Oliveira, R.T.D. Accuracy of white blood cell count, C-reactive protein, interleukin-6 and tumor necrosis factor alpha for diagnosing late neonatal sepsis. J. Pediatr. 2008, 84, 536–542. [Google Scholar] [CrossRef]
- Groselj-Grenc, M.; Ihan, A.; Pavcnik-Arnol, M.; Kopitar, A.N.; Gmeiner-Stopar, T.; Derganc, M. Neutrophil and monocyte CD64 indexes, lipopolysaccharide-binding protein, procalcitonin and C-reactive protein in sepsis of critically ill neonates and children. Intensiv. Care Med. 2009, 35, 1950–1958. [Google Scholar] [CrossRef]
- Viemann, D.; Dubbel, G.; Schleifenbaum, S.; Harms, E.; Sorg, C.; Roth, J. Expression of Toll-Like Receptors in Neonatal Sepsis. Pediatr. Res. 2005, 58, 654–659. [Google Scholar] [CrossRef]
- Lee, R.C.; Ambros, V. An Extensive Class of Small RNAs in Caenorhabditis elegans. Science 2001, 294, 862–864. [Google Scholar] [CrossRef]
- Hrach, H.C.; Mangone, M. miRNA Profiling for Early Detection and Treatment of Duchenne Muscular Dystrophy. Int. J. Mol. Sci. 2019, 20, 4638. [Google Scholar] [CrossRef]
- Pinilla, L.; Benitez, I.D.; González, J.; Torres, G.; Barbé, F.; de Gonzalo-Calvo, D. Peripheral blood microRNAs and the COVID-19 patient: Methodological considerations, technical challenges and practice points. RNA Biol. 2021, 18, 688–695. [Google Scholar] [CrossRef]
- Ho, J.; Chan, H.; Wong, S.H.; Wang, M.H.T.; Yu, J.; Xiao, Z.; Liu, X.; Choi, G.; Leung, C.C.H.; Wong, W.T.; et al. The involvement of regulatory non-coding RNAs in sepsis: A systematic review. Crit. Care 2016, 20, 383. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wang, X.; Liu, X.; Wang, X.; Xu, J.; Hou, S.; Zhang, X.; Ding, Y. miR-15a/16 are upreuglated in the serum of neonatal sepsis patients and inhibit the LPS-induced inflammatory pathway. Int. J. Clin. Exp. Med. 2015, 8, 5683–5690. [Google Scholar]
- Huang, L.; Qiao, L.; Zhu, H.; Jiang, L.; Yin, L. Genomics of neonatal sepsis: Has-miR-150 targeting BCL11B functions in disease progression. Ital. J. Pediatr. 2018, 44, 145. [Google Scholar] [CrossRef]
- Dhas, B.B.; Dirisala, V.R.; Bhat, B.V. Expression Levels of Candidate Circulating microRNAs in Early-Onset Neonatal Sepsis Compared with Healthy Newborns. Genom. Insights 2018, 11. [Google Scholar] [CrossRef]
- Cheng, Q.; Tang, L.; Wang, Y. Regulatory role of miRNA-26a in neonatal sepsis. Exp. Ther. Med. 2018, 16, 4836–4842. [Google Scholar] [CrossRef]
- Chen, J.; Jiang, S.; Cao, Y.; Yang, Y. Altered miRNAs Expression Profiles and Modulation of Immune Response Genes and Proteins During Neonatal Sepsis. J. Clin. Immunol. 2014, 34, 340–348. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Dalli, J.; Chiang, N.; Baron, R.M.; Quintana, C.; Serhan, C.N. Plasticity of Leukocytic Exudates in Resolving Acute Inflammation Is Regulated by MicroRNA and Proresolving Mediators. Immunity 2013, 39, 885–898. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.-R.; Huang, L.-H.; Li, S.-C. Roles of microRNA in the immature immune system of neonates. Cancer Lett. 2018, 433, 99–106. [Google Scholar] [CrossRef] [PubMed]
- Zaas, A.K.; Chen, M.; Varkey, J.; Veldman, T.; Hero, A.O.; Lucas, J.; Huang, Y.; Turner, R.; Gilbert, A.; Lambkin-Williams, R.; et al. Gene Expression Signatures Diagnose Influenza and Other Symptomatic Respiratory Viral Infections in Humans. Cell Host Microbe 2009, 6, 207–217. [Google Scholar] [CrossRef]
- Sharma, P.; Sharma, A.; Sodhi, M.; Verma, P.; Parvesh, K.; Swami, S.K.; Jast, A.; Shandilya, U.K.; Mukesh, M. Characterizing binding sites of heat responsive microRNAs and their expression pattern in heat stressed PBMCs of native cattle, exotic cattle and riverine buffaloes. Mol. Biol. Rep. 2019, 46, 6513–6524. [Google Scholar] [CrossRef]
- Suzuki, Y.; Kim, H.W.; Ashraf, M.; Haider, H.K. Diazoxide potentiates mesenchymal stem cell survival via NF-kappaB-dependent miR-146a expression by targeting Fas. Am. J. Physiol. Heart. Circ. Physiol. 2010, 299, H1077–H1082. [Google Scholar] [CrossRef]
- Lu, L.-F.; Boldin, M.; Chaudhry, A.; Lin, L.-L.; Taganov, K.D.; Hanada, T.; Yoshimura, A.; Baltimore, D.; Rudensky, A.Y. Function of miR-146a in Controlling Treg Cell-Mediated Regulation of Th1 Responses. Cell 2010, 142, 914–929. [Google Scholar] [CrossRef]
- Petrocca, F.; Vecchione, A.; Croce, C.M. Emerging role of miR-106b-25/miR-17-92 clusters in the control of transforming growth factor beta signaling. Cancer Res. 2008, 68, 8191–8194. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Liu, X.; Wang, W.; Cai, Y.; Li, S.; Chen, Q.; Liao, M.; Zhang, M.; Zeng, G.; Zhou, B.; et al. Down-regulation of miR-20a-5p triggers cell apoptosis to facilitate mycobacterial clearance through targeting JNK2 in human macrophages. Cell Cycle 2016, 15, 2527–2538. [Google Scholar] [CrossRef]
- Santoro, M.; Nociti, V.; Lucchini, M.; Loiodice, M.; Centofanti, F.; Botta, A.; Losavio, F.A.; De Fino, C.; Mirabella, M. A pilot study of lncRNAs expression profile in serum of progressive multiple sclerosis patients. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 3267–3273. [Google Scholar]
- Scalavino, V.; Liso, M.; Serino, G. Role of microRNAs in the Regulation of Dendritic Cell Generation and Function. Int. J. Mol. Sci. 2020, 21, 1319. [Google Scholar] [CrossRef]
- Yang, J.; Chen, Y.; Jiang, K.; Yang, Y.; Zhao, G.; Guo, S.; Deng, G. MicroRNA-106a Provides Negative Feedback Regulation in Lipopolysaccharide-Induced Inflammation by targeting TLR4. Int. J. Biol. Sci. 2019, 15, 2308–2319. [Google Scholar] [CrossRef]
- Galbraith, N.J.; O’Brien, S.J.; Walker, S.P.; Gardner, S.A.; Polk, H.C.; Barnes, S.L. Temporal expression of circulating miRNA after severe injury. Surgery 2018, 164, 665–672. [Google Scholar] [CrossRef]
- Anwar, M.A.; Basith, S.; Choi, S. Negative regulatory approaches to the attenuation of Toll-like receptor signaling. Exp. Mol. Med. 2013, 45, e11. [Google Scholar] [CrossRef]
- Zhang, W.; Lu, F.; Xie, Y.; Lin, Y.; Zhao, T.; Tao, S.; Lai, Z.; Wei, N.; Yang, R.; Shao, Y.; et al. miR-23b Negatively Regulates Sepsis-Induced Inflammatory Responses by Targeting ADAM10 in Human THP-1 Monocytes. Mediat. Inflamm. 2019, 2019, 1–13. [Google Scholar] [CrossRef]
- Fatmi, A.; Rebiahi, S.A.; Chabni, N.; Zerrouki, H.; Azzaoui, H.; Elhabiri, Y.; Benmansour, S.; Ibáñez-Cabellos, J.S.; Smahi, M.C.; Aribi, M.; et al. miRNA-23b as a biomarker of culture-positive neonatal sepsis. Mol. Med. Camb. Mass. 2020, 26, 94. [Google Scholar] [CrossRef] [PubMed]
- Li, J.-F.; Wang, P.-M.; Zhang, N.-N.; Zhang, X.; Guan, J.; Ye, Z.; Yan, J.-Y. Hypoxia-induced miR-17-5p ameliorates the viability reduction of astrocytes via targeting p21. Eur. Rev. Med. Pharmacol. Sci. 2016, 20, 3051–3059. [Google Scholar]
- Podshivalova, K.; Salomon, D.R. MicroRNA Regulation of T-Lymphocyte Immunity: Modulation of Molecular Networks Responsible for T-Cell Activation, Differentiation, and Development. Crit. Rev. Immunol. 2013, 33, 435–476. [Google Scholar] [CrossRef]
- Simpson, L.J.; Patel, S.; Bhakta, N.R.; Choy, D.; Brightbill, H.D.; Ren, X.; Wang, Y.; Pua, H.H.; Baumjohann, D.; Montoya, M.M.; et al. A microRNA upregulated in asthma airway T cells promotes TH2 cytokine production. Nat. Immunol. 2014, 15, 1162–1170. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Gupta, S.; Du, W.W.; Yang, B.B. MicroRNA-17 inhibits tumor growth by stimulating T-cell mediated host immune response. Oncoscience 2014, 1, 531–539. [Google Scholar] [CrossRef]
- Cernada, M.; Pinilla-González, A.; Kuligowski, J.; Morales, J.M.; Lorente-Pozo, S.; Piñeiro-Ramos, J.D.; Parra-Llorca, A.; Lara-Cantón, I.; Vento, M.; Serna, E. Transcriptome profiles discriminate between Gram-positive and Gram-negative sepsis in preterm neonates. Pediatr. Res. 2021, 1–9. [Google Scholar] [CrossRef]
- Li, M.; Huang, X.; Zhuo, Q.; Zhang, J.; Ju, X. Clinical significance of miR-129-5p in patients with neonatal sepsis and its regulatory role in the LPS-induced inflammatory response. Bosn. J. Basic Med. Sci. 2020. [Google Scholar] [CrossRef] [PubMed]
- Cernada, M.; Serna, E.; Bäuerl, C.; Collado, M.C.; Martinez, G.P.; Vento, M. Genome-Wide Expression Profiles in Very Low Birth Weight Infants with Neonatal Sepsis. Pediatrics 2014, 133, e1203–e1211. [Google Scholar] [CrossRef] [PubMed]
- Wynn, J.; Cvijanovich, N.Z.; Allen, G.L.; Thomas, N.; Freishtat, R.; Anas, N.; Meyer, K.; Checchia, P.A.; Lin, R.; Shanley, T.; et al. The Influence of Developmental Age on the Early Transcriptomic Response of Children with Septic Shock. Mol. Med. 2011, 17, 1146–1156. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.L.; A Dickinson, P.; Forster, T.; Craigon, M.; Ross, A.; Khondoker, M.; France, R.; Ivens, A.; Lynn, D.; Orme, J.; et al. Identification of a human neonatal immune-metabolic network associated with bacterial infection. Nat. Commun. 2014, 5, 4649. [Google Scholar] [CrossRef]
| Risk Factors of Sepsis |
|
| Clinical Signs of Sepsis |
|
| Gram-Positive Sepsis (n = 11) | Non-Septic Controls (n = 16) | p-Value | |
|---|---|---|---|
| Gestational age (weeks) | 26 (26–29) | 29 (27–30) | 0.22 a |
| Gender: male | 5 (45.5) | 10 (62.5) | 0.38 b |
| Birth weight (g) | 940 (868–1175) | 1270 (879–1445) | 0.29 a |
| Ethnicity: Caucasian | 7 (63.6) | 15 (93.7) | 0.12 b |
| Delivery: | 0.053 b | ||
| Vaginal | 6 (54.5) | 3 (18.7) | |
| C-Section | 5 (45.5) | 13 (81.2) | |
| Apgar 1 min | 8 (5–9) | 6 (5–7) | 0.18 a |
| Apgar 5 min | 9 (9–10) | 9 (8–9) | 0.63 a |
| Nutrition: Breastmilk | 11 (100) | 15 (93.7) | 1 b |
| Central line | 8 (72.7) | 8 (50) | 0.24 b |
| Days after birth at sample collection | 11 (7–19) | 14 (3–26) | 0.72 a |
| Weight at sample collection (g) | 1020 (875–1220) | 1210 (888–1473) | 0.23 a |
| Transcript ID (Array Design) | p-Value (SEPSIS vs. CONTROL) | Fold Change (SEPSIS vs. CONTROL) |
|---|---|---|
| hsa-miR-31-5p | 0.0017 | −3.05 |
| hsa-miR-1271-5p | 0.0089 | −2.77 |
| hsa-miR-326 | 0.0013 | −2.39 |
| hsa-miR-146b-5p | 0.0050 | −2.38 |
| hsa-miR-140-5p | 0.0091 | −2.26 |
| hsa-miR-409-5p | 0.0084 | −2.24 |
| hsa-miR-668-3p | 0.0038 | −2.21 |
| hsa-miR-27b-3p | 0.0028 | −2.12 |
| hsa-miR-28-5p | 0.0019 | −2.06 |
| hsa-miR-152-3p | 0.0080 | −1.95 |
| hsa-miR-431-5p | 0.0094 | −1.95 |
| hsa-miR-106b-5p | 0.0063 | −1.73 |
| hsa-miR-151a-3p | 0.0002 | −1.72 |
| hsa-miR-15a-5p | 0.0046 | −1.71 |
| hsa-miR-339-5p | 0.0068 | −1.70 |
| hsa-miR-30b-5p | 0.0079 | −1.68 |
| hsa-miR-146a-5p | 0.0024 | −1.68 |
| hsa-miR-20a-5p | 0.0008 | −1.64 |
| hsa-miR-20b-5p | 0.0005 | −1.60 |
| hsa-miR-532-5p | 0.0059 | −1.51 |
| hsa-miR-30c-5p | 0.0044 | −1.47 |
| hsa-miR-106a-5p | 0.0006 | −1.43 |
| hsa-miR-17-5p | 0.0007 | −1.39 |
| hsa-miR-23b-3p | 0.0017 | −1.37 |
| hsa-miR-93-5p | 0.0088 | −1.22 |
| hsa-miR-425-5p | 0.0083 | −1.21 |
| hsa-miR-1298-5p | 0.0054 | −1.19 |
| hsa-miR-107 | 0.0009 | −1.14 |
| hsa-miR-103a-3p | 0.0004 | −1.14 |
| hsa-miR-372-3p | 0.0054 | 1.11 |
| hsa-miR-760 | 0.0063 | 1.34 |
| hsa-miR-6088 | 0.0011 | 1.68 |
| hsa-let-7c-5p | 0.0089 | 2.09 |
| Biological Processes | p-Value | Number of Genes |
|---|---|---|
| Treg-Cell Differentiation | 4.92 × 10−9 | 27 |
| Th2-Cell Differentiation | 1.84 × 10−8 | 26 |
| Nociception Expression Targets Signaling | 5.62 × 10−8 | 43 |
| mTOR Signaling | 2.80 × 10−7 | 42 |
| Serotonin Receptors Signaling | 3.06 × 10−7 | 23 |
| Catecholamines Secretion from Adrenal Gland | 5.00 × 10−7 | 26 |
| Peripheral T-Cell Tolerance | 6.81 × 10−7 | 23 |
| Vascular Motility | 7.59 × 10−7 | 28 |
| Th1-Cell Differentiation | 1.06 × 10−6 | 24 |
| Thrombopoietin Receptors Signaling in Platelet Maturation | 1.71 × 10−6 | 16 |
| Subnetworks | p-Value | Number of Genes |
|---|---|---|
| Protein targets of proteasome endopeptidase complex | 1.4 × 10−213 | 266 |
| Protein targets of TNF | 5.6 × 10−213 | 315 |
| Protein targets of TGFB1 | 4.4 × 10−212 | 308 |
| Protein targets of SP1 | 5.7 × 10−193 | 263 |
| Protein targets of NF-kB family | 4.8 × 10−171 | 247 |
| Protein targets of mitogen-activated protein kinase | 2.2 × 10−154 | 205 |
| Protein targets of TP53 | 3.3 × 10−145 | 200 |
| Protein targets of ubiquitin | 1.2 × 10−139 | 172 |
| Protein targets of INS | 1.0 × 10−134 | 193 |
| Protein targets of MAPK1 | 1.5 × 10−134 | 186 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Serna, E.; Parra-Llorca, A.; Panadero, J.; Vento, M.; Cernada, M. miRNomic Signature in Very Low Birth-Weight Neonates Discriminates Late-Onset Gram-Positive Sepsis from Controls. Diagnostics 2021, 11, 1389. https://doi.org/10.3390/diagnostics11081389
Serna E, Parra-Llorca A, Panadero J, Vento M, Cernada M. miRNomic Signature in Very Low Birth-Weight Neonates Discriminates Late-Onset Gram-Positive Sepsis from Controls. Diagnostics. 2021; 11(8):1389. https://doi.org/10.3390/diagnostics11081389
Chicago/Turabian StyleSerna, Eva, Anna Parra-Llorca, Joaquín Panadero, Máximo Vento, and María Cernada. 2021. "miRNomic Signature in Very Low Birth-Weight Neonates Discriminates Late-Onset Gram-Positive Sepsis from Controls" Diagnostics 11, no. 8: 1389. https://doi.org/10.3390/diagnostics11081389
APA StyleSerna, E., Parra-Llorca, A., Panadero, J., Vento, M., & Cernada, M. (2021). miRNomic Signature in Very Low Birth-Weight Neonates Discriminates Late-Onset Gram-Positive Sepsis from Controls. Diagnostics, 11(8), 1389. https://doi.org/10.3390/diagnostics11081389









