Assay System for Simultaneous Detection of SARS-CoV-2 and Other Respiratory Viruses
Abstract
1. Introduction
2. Materials and Methods
2.1. Clinical Specimen Collection and Storage
2.2. Commercial Assay Kits for RV Detection
2.3. Analytical Performance of Commercial Kits
2.4. Statistical Analysis
3. Results
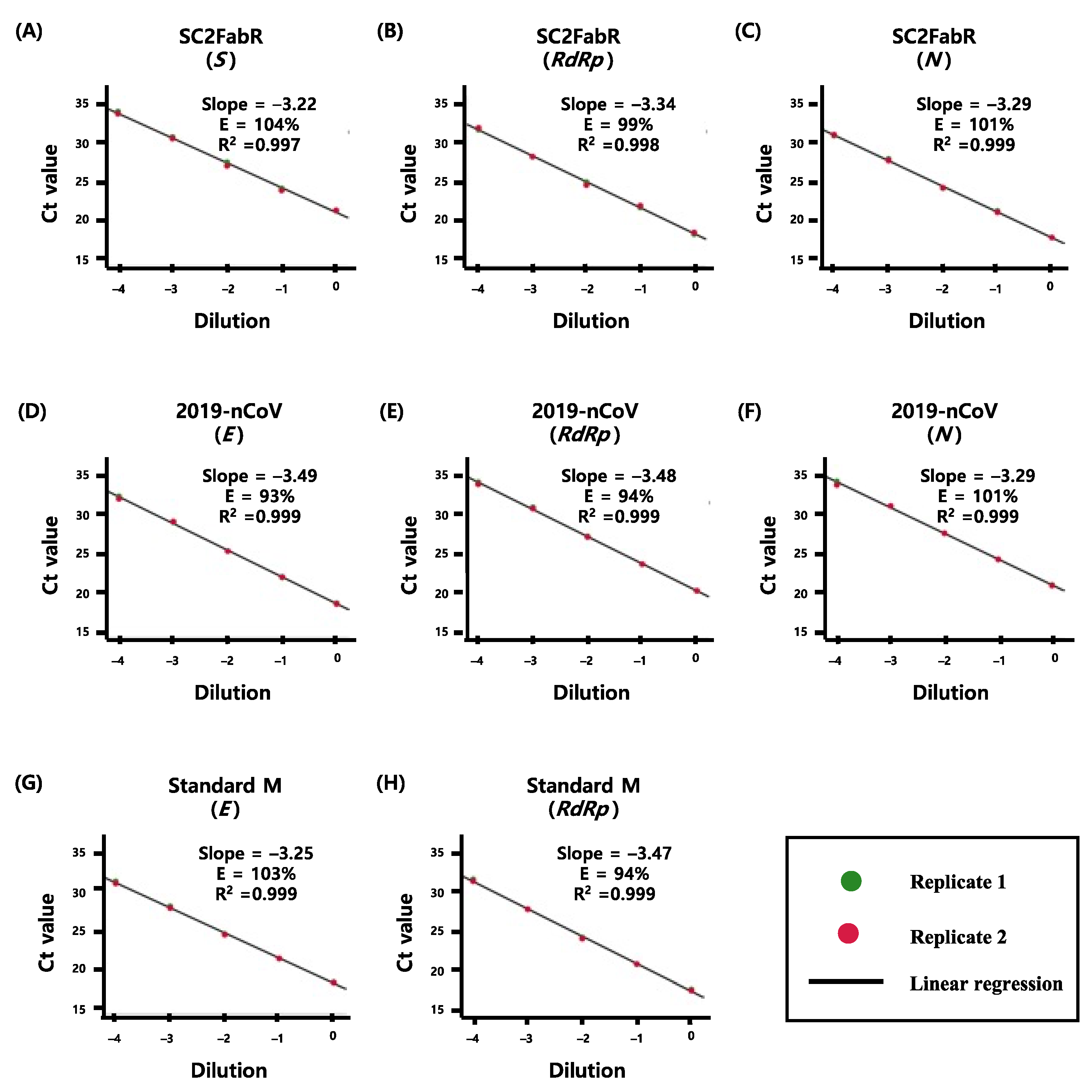
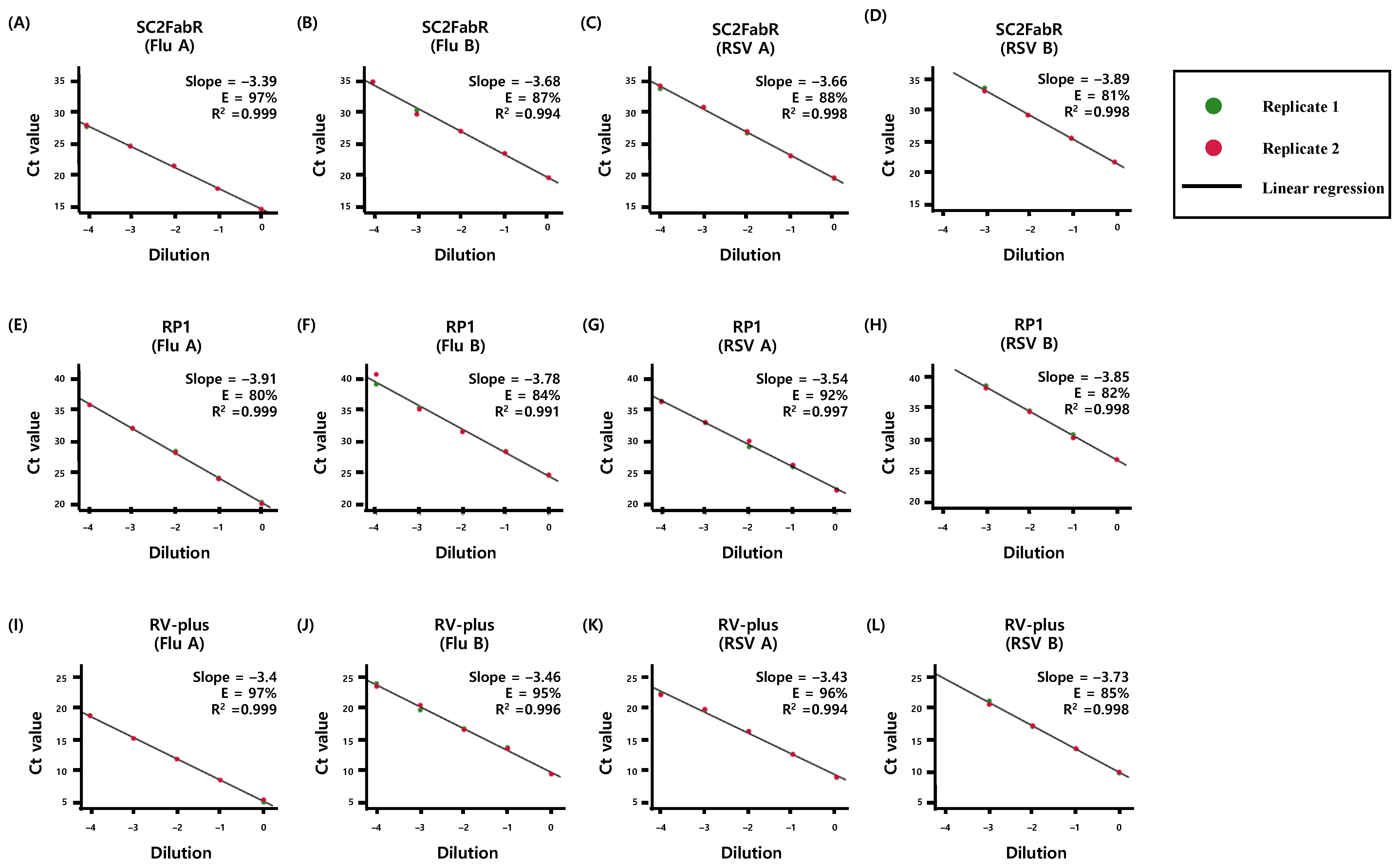
3.1. Comparison of PCR Efficiency of the Virus Detection Systems
3.2. RV Distribution
3.3. Clinical Performance of the Detection Systems
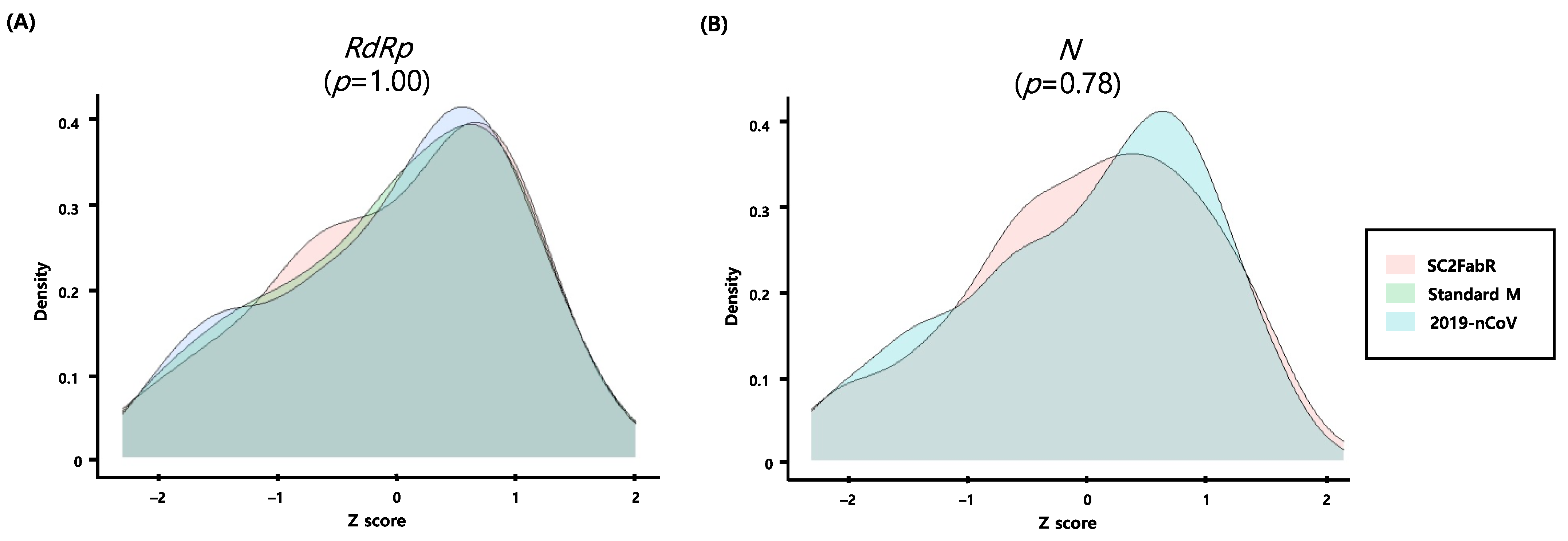
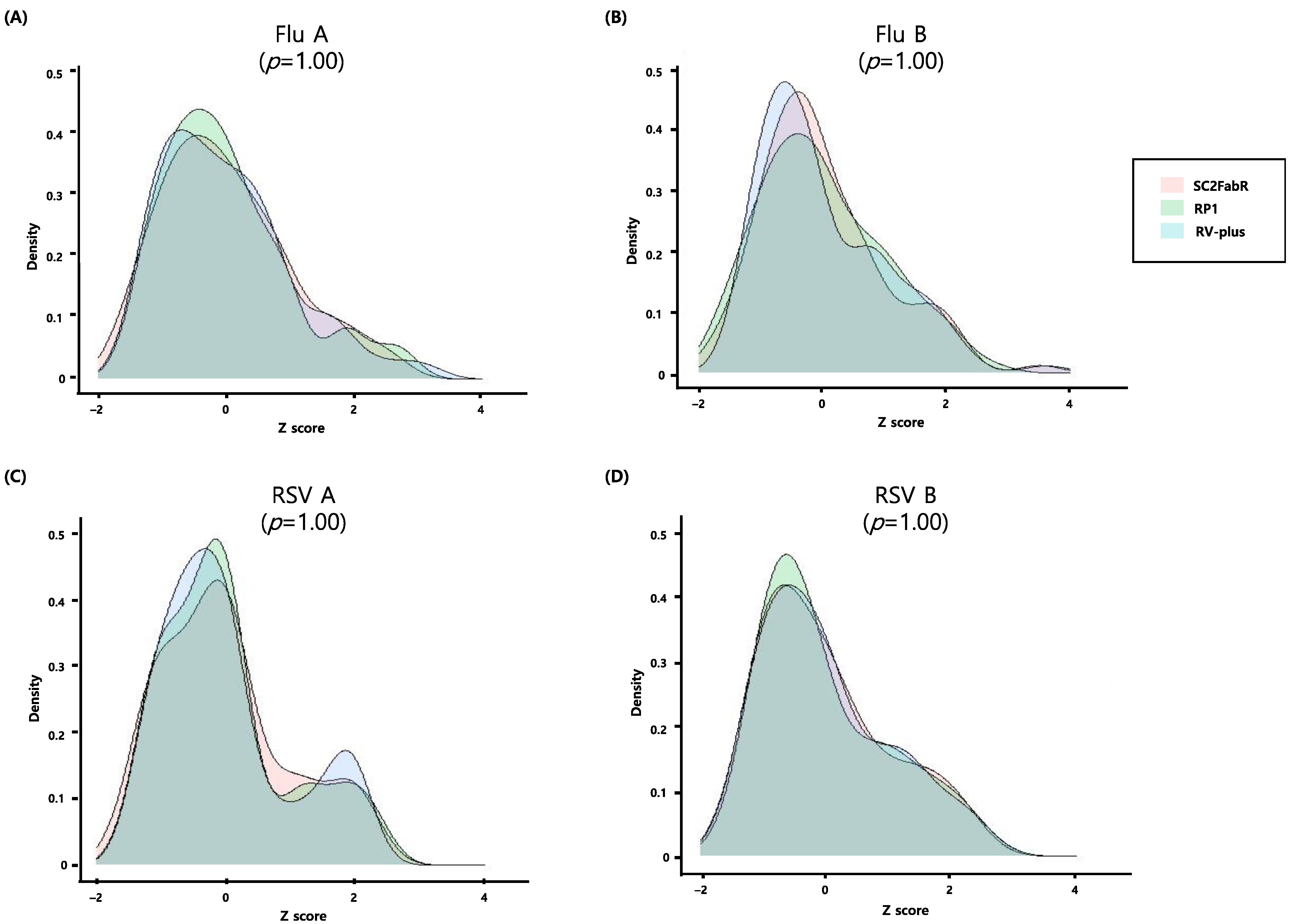
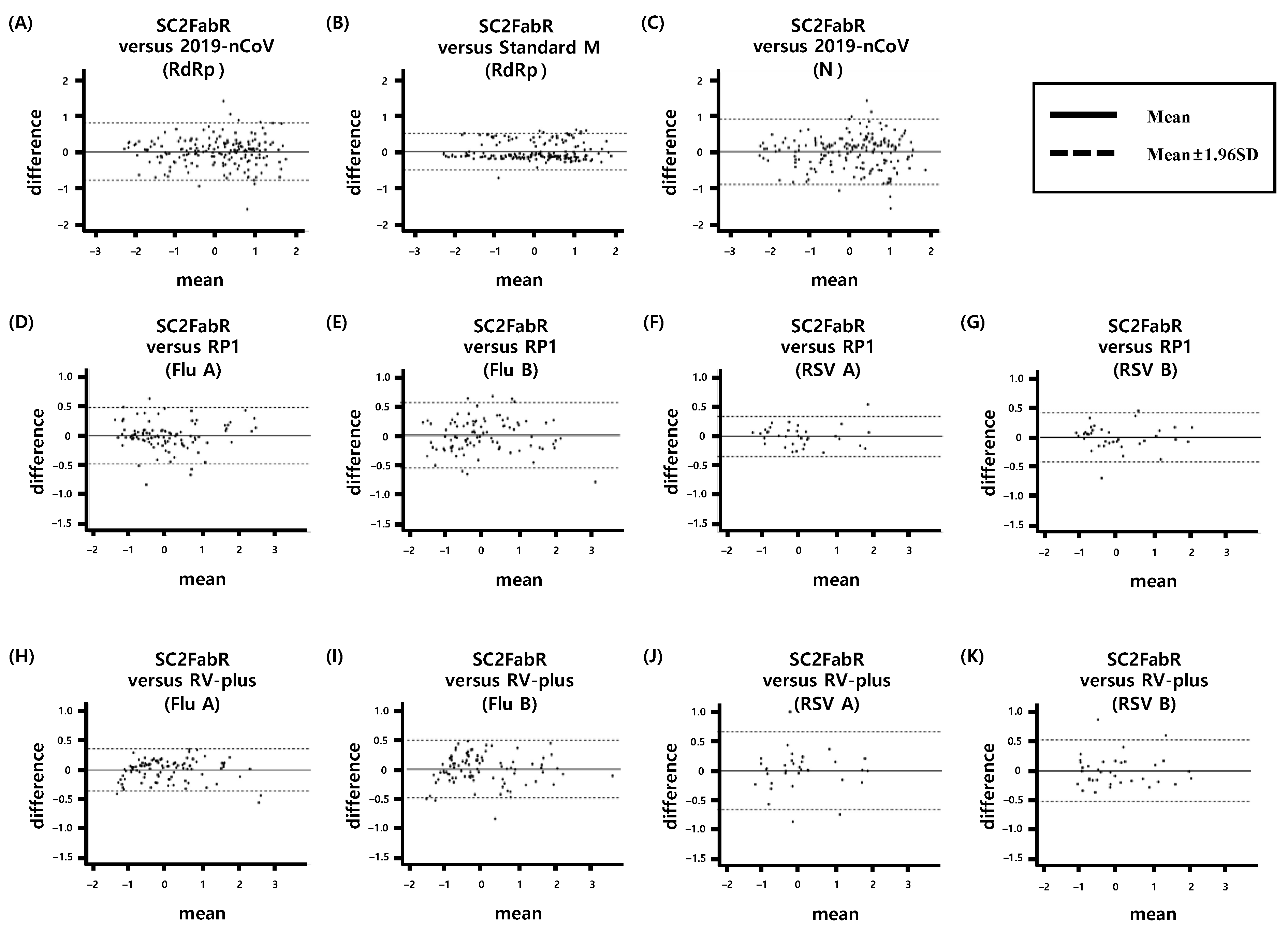
3.4. Intersystem Comparison of the z-Scores Calculated from the Ct Values
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Folgueira, L.; Moral, N.; Pascual, C.; Delgado, R. Comparison of the Panther Fusion and Allplex assays for the detection of respiratory viruses in clinical samples. PLoS ONE 2019, 14, e0226403. [Google Scholar] [CrossRef]
- Dabisch-Ruthe, M.; Vollmer, T.; Adams, O.; Knabbe, C.; Dreier, J. Comparison of three multiplex PCR assays for the detection of respiratory viral infections: Evaluation of xTAG respiratory virus panel fast assay, RespiFinder 19 assay and RespiFinder SMART 22 assay. BMC Infect. Dis. 2012, 12, 163. [Google Scholar] [CrossRef]
- Keske, S.; Ergonul, O.; Tutucu, F.; Karaaslan, D.; Palaoglu, E.; Can, F. The rapid diagnosis of viral respiratory tract infections and its impact on antimicrobial stewardship programs. Eur. J. Clin. Microbiol. Infect. Dis. 2018, 37, 779–783. [Google Scholar] [CrossRef]
- Rogan, D.T.; Kochar, M.S.; Yang, S.; Quinn, J.V. Impact of rapid molecular respiratory virus testing on real-time decision making in a pediatric emergency department. J. Mol. Diagn. 2017, 19, 460–467. [Google Scholar] [CrossRef]
- van Kasteren, P.B.; van der Veer, B.; van den Brink, S.; Wijsman, L.; de Jonge, J.; van den Brandt, A.; Molenkamp, R.; Reusken, C.; Meijer, A. Comparison of seven commercial RT-PCR diagnostic kits for COVID-19. J. Clin. Virol. 2020, 128, 104412. [Google Scholar] [CrossRef]
- Gatti, P.; Ilamathi, H.S.; Todkar, K.; Germain, M. Mitochondria targeted viral replication and survival strategies-prospective on SARS-CoV-2. Front. Pharmacol. 2020, 11, 578599. [Google Scholar] [CrossRef]
- World Health Organization. Weekly Epidemiological Update—9 February 2021. Available online: https://www.who.int/publications/m/item/weekly-epidemiological-update---9-february-2021 (accessed on 9 February 2021).
- Abdelrahman, Z.; Li, M.; Wang, X. Comparative Review of SARS-CoV-2, SARS-CoV, MERS-CoV, and Influenza A Respiratory Viruses. Front. Immunol. 2020, 11, 552909. [Google Scholar] [CrossRef]
- Miners, S.; Kehoe, P.G.; Love, S. Cognitive impact of COVID-19: Looking beyond the short term. Alzheimer’s Res. Ther. 2020, 12, 170. [Google Scholar] [CrossRef]
- Zhen, W.; Manji, R.; Smith, E.; Berry, G.J. Comparison of four molecular in vitro diagnostic assays for the detection of SARS-CoV-2 in nasopharyngeal specimens. J. Clin. Microbiol. 2020, 58, 8. [Google Scholar] [CrossRef]
- China-WHO Expert Team. Report of the WHO-China Joint Mission on Coronavirus Disease 2019 (COVID-19); China-WHO Expert Team: Beijing, China, 2020. [Google Scholar]
- Khorramdelazad, H.; Kazemi, M.H.; Najafi, A.; Keykhaee, M.; Zolfaghari Emameh, R.; Falak, R. Immunopathological similarities between COVID-19 and influenza: Investigating the consequences of co-infection. Microb. Pathog. 2021, 152, 104554. [Google Scholar] [CrossRef]
- Bai, L.; Zhao, Y.; Dong, J.; Liang, S.; Guo, M.; Liu, X.; Wang, X.; Huang, Z.; Sun, X.; Zhang, Z.; et al. Coinfection with influenza A virus enhances SARS-CoV-2 infectivity. Cell Res. 2021, 31, 395–403. [Google Scholar] [CrossRef]
- Lansbury, L.; Lim, B.; Baskaran, V.; Lim, W.S. Co-infections in people with COVID-19: A systematic review and meta-analysis. J. Infect. 2020, 81, 266–275. [Google Scholar] [CrossRef]
- Yue, H.; Zhang, M.; Xing, L.; Wang, K.; Rao, X.; Liu, H.; Tian, J.; Zhou, P.; Deng, Y.; Shang, J. The epidemiology and clinical characteristics of co-infection of SARS-CoV-2 and influenza viruses in patients during COVID-19 outbreak. J. Med. Virol. 2020, 92, 2870–2873. [Google Scholar] [CrossRef]
- Wu, X.; Cai, Y.; Huang, X.; Yu, X.; Zhao, L.; Wang, F.; Li, Q.; Gu, S.; Xu, T.; Li, Y.; et al. Co-infection with SARS-CoV-2 and influenza A virus in patient with pneumonia, China. Emerg. Infect. Dis. 2020, 26, 1324–1326. [Google Scholar] [CrossRef]
- Hashemi, S.A.; Safamanesh, S.; Ghasemzadeh-Moghaddam, H.; Ghafouri, M.; Azimian, A. High prevalence of SARS-CoV-2 and influenza A virus (H1N1) coinfection in dead patients in Northeastern Iran. J. Med. Virol. 2021, 93, 1008–1012. [Google Scholar] [CrossRef]
- Hur, K.H.; Park, K.; Lim, Y.; Jeong, Y.S.; Sung, H.; Kim, M.N. Evaluation of four commercial kits for SARS-CoV-2 real-time reverse-transcription polymerase chain reaction approved by emergency-use-authorization in Korea. Front. Med. 2020, 7, 521. [Google Scholar] [CrossRef]
- Huh, H.J.; Kim, J.Y.; Kwon, H.J.; Yun, S.A.; Lee, M.K.; Lee, N.Y.; Kim, J.W.; Ki, C.S. Performance evaluation of Allplex respiratory panels 1, 2, and 3 for detection of respiratory viruses and influenza A virus subtypes. J. Clin. Microbiol. 2017, 55, 479–484. [Google Scholar] [CrossRef]
- Ko, D.H.; Kim, H.S.; Hyun, J.; Kim, H.S.; Kim, J.S.; Park, K.U.; Song, W. Comparison of the Luminex xTAG respiratory viral panel fast v2 assay with Anyplex II RV16 detection kit and AdvanSure RV real-time RT-PCR assay for the detection of respiratory viruses. Ann. Lab. Med. 2017, 37, 408–414. [Google Scholar] [CrossRef][Green Version]
- Mengelle, C.; Mansuy, J.M.; Sandres-Saune, K.; Barthe, C.; Boineau, J.; Izopet, J. Prospective evaluation of a new automated nucleic acid extraction system using routine clinical respiratory specimens. J. Med. Virol. 2012, 84, 906–911. [Google Scholar] [CrossRef]
- Cheadle, C.; Vawter, M.P.; Freed, W.J.; Becker, K.G. Analysis of microarray data using Z score transformation. J. Mol. Diagn. 2003, 5, 73–81. [Google Scholar] [CrossRef]
- Hindiyeh, M.; Mor, O.; Pando, R.; Mannasse, B.; Kabat, A.; Assraf-Zarfati, H.; Mendelson, E.; Sofer, D.; Mandelboim, M. Comparison of the new fully automated extraction platform eMAG to the MagNA PURE 96 and the well-established easyMAG for detection of common human respiratory viruses. PLoS ONE 2019, 14, e0211079. [Google Scholar] [CrossRef]
- Leung, E.C.-M.; Chow, V.C.-Y.; Lee, M.K.-P.; Tang, K.P.-S.; Li, D.K.-C.; Lai, R.W.-M. Evaluation of the xpert xpress SARS-CoV-2/Flu/RSV assay for simultaneous detection of SARS-CoV-2, influenza A and B viruses, and respiratory syncytial virus in nasopharyngeal specimens. J. Clin. Microbiol. 2021, 59, e02965–20. [Google Scholar] [CrossRef]
- Leung, N.H.L. Transmissibility and transmission of respiratory viruses. Nat. Rev. Microbiol. 2021, 1–18. [Google Scholar] [CrossRef]
- Sun, S.; Xie, Z.; Yu, K.; Jiang, B.; Zheng, S.; Pan, X. COVID-19 and healthcare system in China: Challenges and progression for a sustainable future. Glob. Health 2021, 17, 14–18. [Google Scholar] [CrossRef]
- Drame, M.; Tabue Teguo, M.; Proye, E.; Hequet, F.; Hentzien, M.; Kanagaratnam, L.; Godaert, L. Should RT-PCR be considered a gold standard in the diagnosis of COVID-19? J. Med. Virol. 2020, 92, 2312–2313. [Google Scholar] [CrossRef]
- Roh, K.H.; Kim, Y.K.; Kim, S.W.; Kang, E.R.; Yang, Y.J.; Jung, S.K.; Lee, S.H.; Sung, N. Coinfections with respiratory pathogens among COVID-19 patients in Korea. Can. J. Infect. Dis. Med. Microbiol. 2021, 2021, 6651045. [Google Scholar] [CrossRef]
- Wang, J.; Cai, K.; Zhang, R.; He, X.; Shen, X.; Liu, J.; Xu, J.; Qiu, F.; Lei, W.; Wang, J.; et al. Novel one-step single-tube nested quantitative real-time PCR assay for highly sensitive detection of SARS-CoV-2. Anal. Chem. 2020, 92, 9399–9404. [Google Scholar] [CrossRef]
- Kumar, A.; Chordia, N. In Silico PCR primer designing and validation. Methods Mol. Biol. 2015, 1275, 143–151. [Google Scholar] [CrossRef]
- Green, M.R.; Sambrook, J. Polymerase chain reaction (PCR) amplification of GC-rich templates. Cold Spring Harb. Protoc. 2019, 2019. [Google Scholar] [CrossRef]
- Mitropapas, G.; Nezos, A.; Halapas, A.; Pissimissis, N.; Lembessis, P.; Sourla, A.; Vassilopoulos, P.; Koutsilieris, M. Molecular detection of tyrosinase transcripts in peripheral blood from patients with malignant melanoma: Correlation of PCR sensitivity threshold with clinical and pathologic disease characteristics. Clin. Chem. Lab. Med. 2006, 44, 1403–1409. [Google Scholar] [CrossRef]
| Detection Kit (Release Year) | Manufacturer | Sample Type | Platform | Running Time | Target Regions of SARS-CoV2 | Target of Respiratory Virus | Maximum Capacity (96 Well Plate) | TAT (76,000 Samples) |
|---|---|---|---|---|---|---|---|---|
| SC2FabR (2020) | Seegene, Inc. | NPS, NPA, BAL, TS, Sputum, Saliva | CFX96 | ~115 min | S, RdRp, N | Flu A, Flu B, RSV | 94 samples | 1551 hr |
| 2019-nCoV (2020) | Seegene, Inc. | NPS, NPA, BAL, TS, Sputum | CFX96, ABI 7500 | ~110 min | E, RdRp, N | - | 94 samples | 1483 hr |
| STANDARD M (2020) | SD BIOSENSOR, Inc. | NPS, OS, Sputum | CFX96, ABI 7500 | ~98 min | E, RdRp | - | 94 samples | 1321 hr |
| RP1 (2016) | Seegene, Inc. | NPS, NPA, BAL | CFX96 | ~150 min | - | Flu A (H1, H1-pdm09, H3), Flu B, RSV A, RSV B | 94 samples | 2023 hr |
| RV-plus (2017) | LG Chem, Ltd. | NPS, NS, TS | SLAN-96P | ~100 min | - | Flu A, Flu B, RSV A, RSV B, OC43, HEV, BoV, | 22 samples | 5758 hr |
| RV-16 (2013) | Seegene, Inc. | NPS, NPA, BAL | CFX96 | ~230 min | - | Flu A, Flu B, RSV A, RSV B, AdV, MPV, BoV, 229E, NL63, OC43, HRV, HEV, PIV (1,2,3,4) | 46 samples | 6337 hr |
| Assay | Target | SC2FabR Agreement | Sen (%) | Spe (%) | DA (%) | ||||
|---|---|---|---|---|---|---|---|---|---|
| TP | FP | TN | FN | Total (n) | |||||
| 2019-nCoV | SARS-CoV2 | 180 | 0 | 200 | 0 | 380 | 100 | 100 | 100 |
| Standard M | SARS-CoV2 | 180 | 0 | 200 | 0 | 380 | 100 | 100 | 100 |
| RP1 | Flu A | 99 | 0 | 391 | 0 | 490 | 100 | 100 | 100 |
| Flu B | 91 | 0 | 399 | 0 | 490 | 100 | 100 | 100 | |
| RSV A/B | 74 | 0 | 415 | 1 | 490 | 98.7 | 100 | 99.8 | |
| RV-plus | Flu A | 98 | 2 | 389 | 1 | 490 | 99.0 | 99.5 | 99.4 |
| Flu B | 91 | 1 | 398 | 0 | 490 | 100 | 99.7 | 99.8 | |
| RSV A/B | 69 | 2 | 413 | 6 | 490 | 92.0 | 99.5 | 98.4 | |
| Sample ID | Discordance Type | SC2FabR Result | RP1 Result | RV-Plus Result | RV-16 Result | Final Determination |
|---|---|---|---|---|---|---|
| A | Flu A | Flu A | Flu A | Neg. | Neg. | Flu A |
| B | Neg. | Neg. | Flu A | Neg. | Neg. | |
| C | Neg. | Neg. | Flu A | Neg. | Neg. | |
| D | Flu B | Neg. | Neg. | Flu B | Neg. | Neg. |
| E | RSV | RSV | RSV A | Neg. | RSV A | RSV |
| F | RSV | Neg. | RSV A | RSV A | RSV | |
| G | RSV | RSV A | Neg. | RSV A | RSV | |
| H | RSV | RSV B | Neg. | Neg. | RSV | |
| I | RSV | RSV B | Neg. | RSV B | RSV | |
| J | RSV | RSV A | Neg. | Neg. | RSV | |
| K | RSV | RSV A | Neg. | Neg. | RSV | |
| L | Neg. | Neg. | RSV A | Neg. | Neg. | |
| M | Neg. | Neg. | RSV A | Neg. | Neg. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lim, H.-J.; Park, J.-E.; Park, M.-Y.; Baek, J.-H.; Jung, S.; Sung, N.; Yang, J.-H.; Lee, M.-W.; Lee, S.-H.; Yang, Y.-J. Assay System for Simultaneous Detection of SARS-CoV-2 and Other Respiratory Viruses. Diagnostics 2021, 11, 1084. https://doi.org/10.3390/diagnostics11061084
Lim H-J, Park J-E, Park M-Y, Baek J-H, Jung S, Sung N, Yang J-H, Lee M-W, Lee S-H, Yang Y-J. Assay System for Simultaneous Detection of SARS-CoV-2 and Other Respiratory Viruses. Diagnostics. 2021; 11(6):1084. https://doi.org/10.3390/diagnostics11061084
Chicago/Turabian StyleLim, Ho-Jae, Jung-Eun Park, Min-Young Park, Joo-Hwan Baek, Sunkyung Jung, Nackmoon Sung, Jae-Hyun Yang, Min-Woo Lee, Sun-Hwa Lee, and Yong-Jin Yang. 2021. "Assay System for Simultaneous Detection of SARS-CoV-2 and Other Respiratory Viruses" Diagnostics 11, no. 6: 1084. https://doi.org/10.3390/diagnostics11061084
APA StyleLim, H.-J., Park, J.-E., Park, M.-Y., Baek, J.-H., Jung, S., Sung, N., Yang, J.-H., Lee, M.-W., Lee, S.-H., & Yang, Y.-J. (2021). Assay System for Simultaneous Detection of SARS-CoV-2 and Other Respiratory Viruses. Diagnostics, 11(6), 1084. https://doi.org/10.3390/diagnostics11061084






