1. Introduction
Alzheimer’s disease (AD) features can be analyzed to create more effective and accurate tools based on recent economical, and publicly available, technologies. Currently, there have been several approaches which can be applied to detect AD in its early phases, such as neuroimaging techniques [
1,
2,
3], behavior and emotion analysis [
4,
5], often referred to as cognitive approaches, and cognitive test. Behavioral analysis methods help to detect irregular reactions to frequent problems in daily living activities, and some of which involve the installation of sensors in the patient’s house. One of the main drawbacks of this strategy is that it comes with a lot of limitations, as it needs the patient’s permission to mount the sensors in his/her home. One of the signs of AD is a decline in social cognition, and some studies have focused on patients’ ability to interpret emotions using various data, such as eye-tracking data [
6], voice/speech recordings [
7], facial expressions [
8], and electroencephalograms (EEG) [
9,
10].
Neuroimaging techniques such as structural magnetic resonance imaging (sMRI) [
11,
12], fMRI [
13], fluorodeoxyglucose positron emission tomography (FDG-PET) imaging [
14], amyloid PET [
1], and diffusion tensor imaging (DTI) [
15]. These neuroimaging techniques have shown to be promising modalities to assess abnormal brain changes linked to AD, and they remain mainly used in the more advanced centers. In amyloid PET, diffuse amyloid deposits in the cortex are considered a measure of neurodegeneration, and a marker that binds to the Aβ protein is injected into the subject. Amyloid PET shows both quantitative information, that can be regionally based, and qualitative information about the topology of Aβ deposition in the brain. For fMRI, the alteration in blood flow and blood oxygen concentration measurement shows the brain’s metabolic activities [
16]. The amount of shrinkage in brain sub-regions, especially the hippocampus, corroborates the structural changes of the brain [
16]. FDG-PET provides quantitative measurements of the brain’s metabolic activity [
17].
Compared to other neuroimaging modalities, fMRI has helped AD analysts to assess functionally activated regions when conducting a task to diagnose AD early [
18]. When performing tasks, the frontal-subcortical-parietal regions, thalamus, striatum, and intraparietal cortex are all co-activated brain regions. Preliminary fMRI studies have shown a strong link between cognitive functions and sensorimotor eye movements, assisting in the development of a better understanding of neurodegenerative diseases’ network-level brain disruptions. Authors in [
19] investigated functional brain connectivity, and they concluded that new approaches are needed to comprehend the functional pattern alternation in the initial phase of AD.
The AD functional brain changes among mild cognitive impairment (MCI) stages are closely related and constant trait information from the features that represent each stage are often very complex to delineate. Based on this, it is difficult to accurately predict AD early. Distinguishing the functional brain pattern of MCI stages need accurate information and knowledge. In this paper, to diagnose AD early, authors created a DL algorithm to take out useful features from hippocampal fMRI data from the ADNI database. A clinician may use the proposed model to easily diagnose a patient with MCI and monitor their progress over time.
The contribution of this paper are as follows:
This study proposes a modified ResNet18 and performs binary classification of AD which include EMCI/LMCI, AD/CN, CN/EMCI, CN/LMCI, EMCI/AD, LMCI/AD, and MCI/EMCI.
To effectively identify the brain changes associated with each of the classes, we investigate fine tuning framework for classification of AD images based on seven binary classes.
To avoid over fitting and be able to generalize the data and reduce validation loss, dropout of 0.2 is introduced to the custom layer over fully connected layer to predict the best result on binary classification.
The following sections make up the remainder of this paper: a literature review about using deep learning (DL) in fMRI is stated in
Section 2.
Section 3 outlines the proposed approach in detail and is divided into 6 subsections, the first of which is
Section 3.1, which describes data that was used in the procedure for evaluating. In
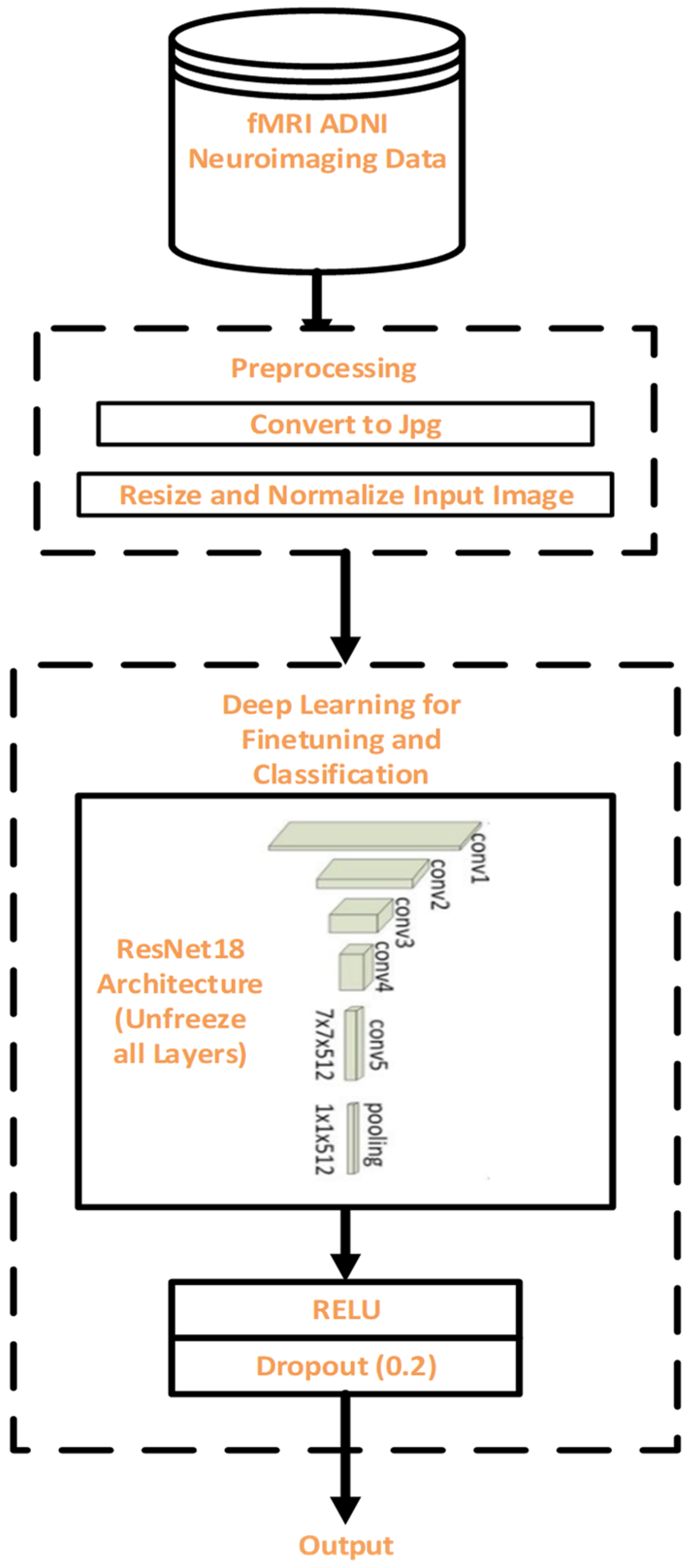
Section 3.2, a description of DL for finetuning and classification is discussed.
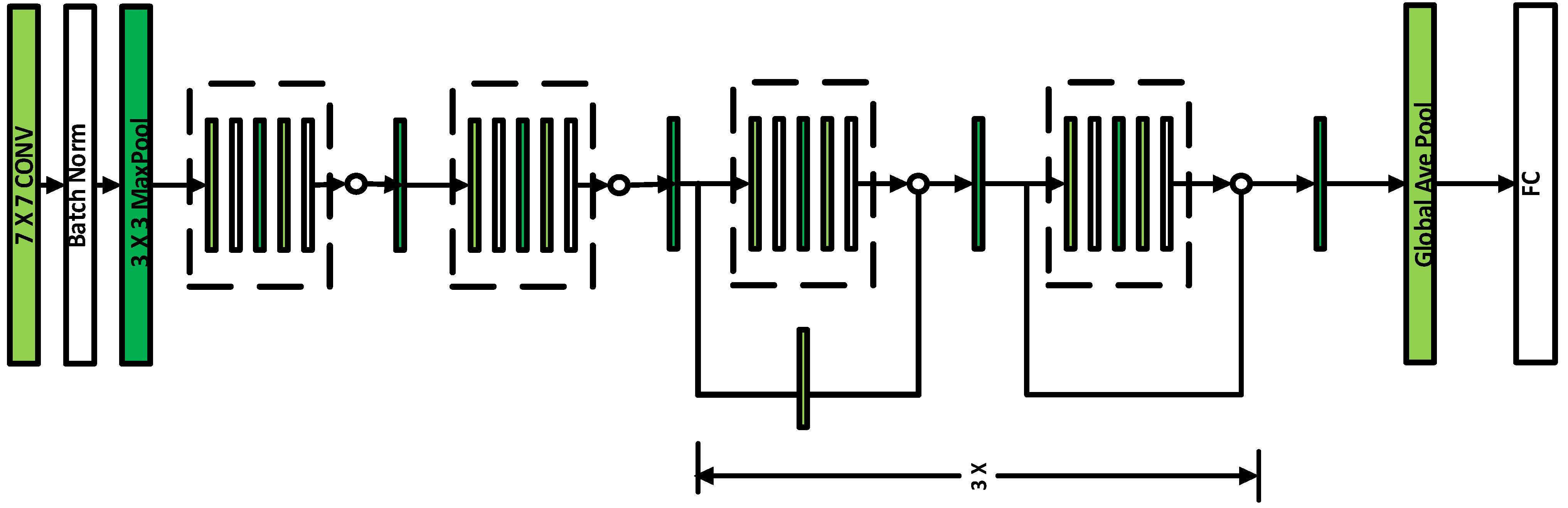
Section 3.3 gives the detailed preprocessing steps while the description of CNN architecture is presented in
Section 3.4. In
Section 3.5, the description of our proposed fine-tuning model using ResNet18 is explained, and
Section 3.6 gives the evaluation measures used to assess the proposed model. The experimental findings are summarized in
Section 4, and the discussion is presented in
Section 5. The comparison of the proposed model, with existing studies, is presented in
Section 6. The paper concludes in
Section 7 with the discussion on future research.
2. Related Work
The DL algorithms for extracting latent features of neuroimaging data, for early detection of Alzheimer’s disease, have piqued the interest of researchers. To distinguish an Alzheimer’s disease affected brain from a normal (healthy) brain, authors in [
19] used CNN to successfully identify functional MRI data of Alzheimer’s patients from standard controls. The model achieved an accuracy of 96.85%. However, more complicated network architecture is required to handle complicated problems. Centered on graph theory and machine learning (ML), authors in [
20] created a novel framework for the classification of MCI. The areas of the brain that changed significantly in the MCI groups were correctly described. The proposed model only showed the progression of MCI, differentiating the intermediate stages of the MCI was not considered. Authors in [
21] suggested a machine learning-based computer-assisted diagnostic approach that can automatically differentiate Alzheimer’s patients from safe controls. Although the proposed model gave a high prediction accuracy bur it is tough to say exactly which components had an impact on the overall neural network decision. Authors in [
22] suggested using fMRI to identify subjects with MCI or AD, incorporating CNN and Ensemble Learning to construct a classifier ensemble (EL). A combined CNN and EL method can find the brain regions that the qualified ensemble model suggests are the most discriminable. The authors concluded that using optimization techniques or other DL approaches, classification accuracy could be improved. To improve the EMCI detection, the Authors in [
23] proposed a multi-scale enhanced GCN (MSE-GCN) to investigate individual differences and knowledge association among various subjects. Using image and population phenotypic data, the proposed model was able to learn rich features. With LMCI vs. NC, the accuracy of 93.46% was achieved. More efficient network models for accurate brain region location were suggested by the authors. Authors in [
24] proposed a method based on autoencoders for distinguishing between natural aging and disease progression. The proposed approach makes use of effectively biased neural network functionality to accurately diagnose Alzheimer’s disease. Authors in [
25] used a 3D CNN to construct a binary classifier that could distinguish between AD and CN resting-state fMRI results. The proposed model used three binary classifications, with AD vs. CN achieving the highest validity accuracy of 97.77%, but there was high computational complexity. Authors in [
26] introduced a CNN DL algorithm that predicts who will develop Alzheimer’s disease and who will develop MCI. The proposed model was found to be extremely effective at distinguishing AD and MCI patients from healthy controls, as well as predicting AD conversion. The proposed model did not consider the heterogenous nature of AD. Authors in [
27] extracted spatial features from each volume of a 3D static image in an fMRI image sequence. The feature maps were fed into a long short-term memory (LSTM) network to capture the data’s time-varying details. The proposed model had a classification accuracy of 92.11% for AD vs. MCI and 88.12% for MCI vs. NC. However, the multiclass classification accuracy is very low.
For EMCI classification, the authors in [
28] proposed a new 3D CNN for removing features that are deeply rooted from dynamic as well as static fMRI brain functional networks with an accuracy of 76.07%. Multi-model, multi-channel, and time-consuming, on the other hand, did not improve classification accuracy. On fMRI, Authors in [
29], used a 2D CNN model in conjunction with a transfer learning technique to correctly identify AD, EMCI, and NC. with a 98.41% accuracy. Although the proposed method performed well, it did not deal with the issue of EMCI vs. NC binary classification. Authors in [
30] have used a hybrid ML method that utilized bidirectional long short term memory (LSTM) network for identifying discriminative features among AD binary classification from multimodal neuroimaging data. The proposed model had a high run-time complexity. Authors in [
31] further develop a novel unified CNN framework using 3D CNN. A 3D Convolutional LSTM (CLSTM) is then applied to extract features and efficiently classified AD binary classes. The proposed model gave an improved classification accuracy, but the Prodromal stages of AD were not considered. Authors in [
32] also proposed using CNN fMRI data for early AD classification. Authors in [
33] also presented CNN architecture to diagnose AD early using fMRI with an accuracy of 96.7%, but the power of the method to diagnose the disease severity is low. Similarly, the authors in [
34] used 3D-CNN on MRI images to obtain high-level features for AD binary classification task with 87.2% accuracy for AD/CN. Authors in [
35] further utilized VoxCNN and ResNet for early AD diagnosis and had an accuracy of 80% for AD vs. CN classification. Authors in [
36] presented a simple 3D CNN framework, based on the transfer learning strategy for MCI classification, with an accuracy of 94.1%. The proposed model gave a low binary classification accuracy when compared to existing methods.
Furthermore, for six classification tasks, the authors in [
37] proposed a layer-wise transfer learning method using VGG 19. The experiments were conducted on 300 ADNI subjects who were divided into six binary groups. With an accuracy of 98.73% on AD vs. NC and 83.72% on EMCI vs. LMCI, the proposed model obtained the best performance but gave a high computational complexity. Authors in [
38] further utilized VGG 16 on fMRI dataset for two binary classification tasks. The proposed model was effective in achieving a classification accuracy of 99.27% for AD vs. MCI. The authors recommended other pre-trained networks, such as the Inception Network and the Residual Network for building a better classifier for binary AD classification. Authors in [
39] suggested a CNN-based technique for extracting discriminative features from structural MRI with the goal of diagnosing EMCI and LMCI, as well as classifying these two groups from healthy people. Authors in [
35] suggested two distinct 3D convolutional network topologies for brain MRI classification and demonstrated the performance of the suggested methodology on the ADNI for the classification of Alzheimer’s disease vs moderate cognitive impairment. Authors in [
40] used multimodal data for AD stage classification. Stacked denoising auto-encoders extracted features from genetic and clinical records, whereas 3D CNNs analyzed MRI data to recognize AD vs MCI and healthy controls. Classification of different stages of AD was performed on fMRI dataset, authors in [
41] used the architecture of a CNN AlexNet for efficient classification of AD with 97.64% average accuracy. The authors concluded that the use of other pre-trained models, and transfer learning, could improve classification accuracy. Authors in [
42] presented an approach for early detection of AD by fine-tuning CaffeNet and GoogLeNet models on 2D MRI images. On an fMRI dataset with AD and NC groups, authors in [
43] investigated the performance of ResNet18 based on transfer learning for AD detection. Experiments reported that the proposed model had a 96.88% accuracy. Authors in [
44] examined the usefulness of rs-fMRI for multi-class classification of AD and its stages. The classification task was performed using residual neural networks, and the findings showed a wide variety of outcomes depending on the stage of the disease.
The summary of some of the related work that applied DL algorithms on fMRI for early detection of AD is presented in
Table 1.
The existing studies suffered from some serious limitations such as low classification accuracy for MCI intermediate classes and non-consideration of binary classes such as EMCI vs. LMCI, EMCI vs. NC. However, there is still a need for more efficient network models for accurate brain region location to aid early detection of AD [
23]. Other CNN pre-trained models and more recent cutting-edge networks should be explored as the base model to build an efficient classifier for AD classification [
37].
5. Discussion
In this study, we have analyzed the effect of dropout on a fine-tuned pretrained model to classify fMR images from the ADNI database. This study’s findings revealed that fine-tuning the entire network gave high classification accuracy on all binary classification scenarios except AD vs. CN and CN vs. LMCI. Without dropout the best performance was achieved by EMCI vs. AD with an accuracy of 99.99% (
Table 7).
Table 8 shows the effect of dropout on the binary classification. We can see that the proposed model has yielded positive results on EMCI vs. AD, EMCI vs. LMCI, LMCI vs. AD, EMCI vs. MCI classification and achieved 99.99%, 99.76%, 99.95%, and 99.95% accuracy respectively. In terms of sensitivity, however, the AD vs. CN classification performance was superior. in the case of the model, without dropout, with a value of 97.8%. Regarding the confusion matrices in
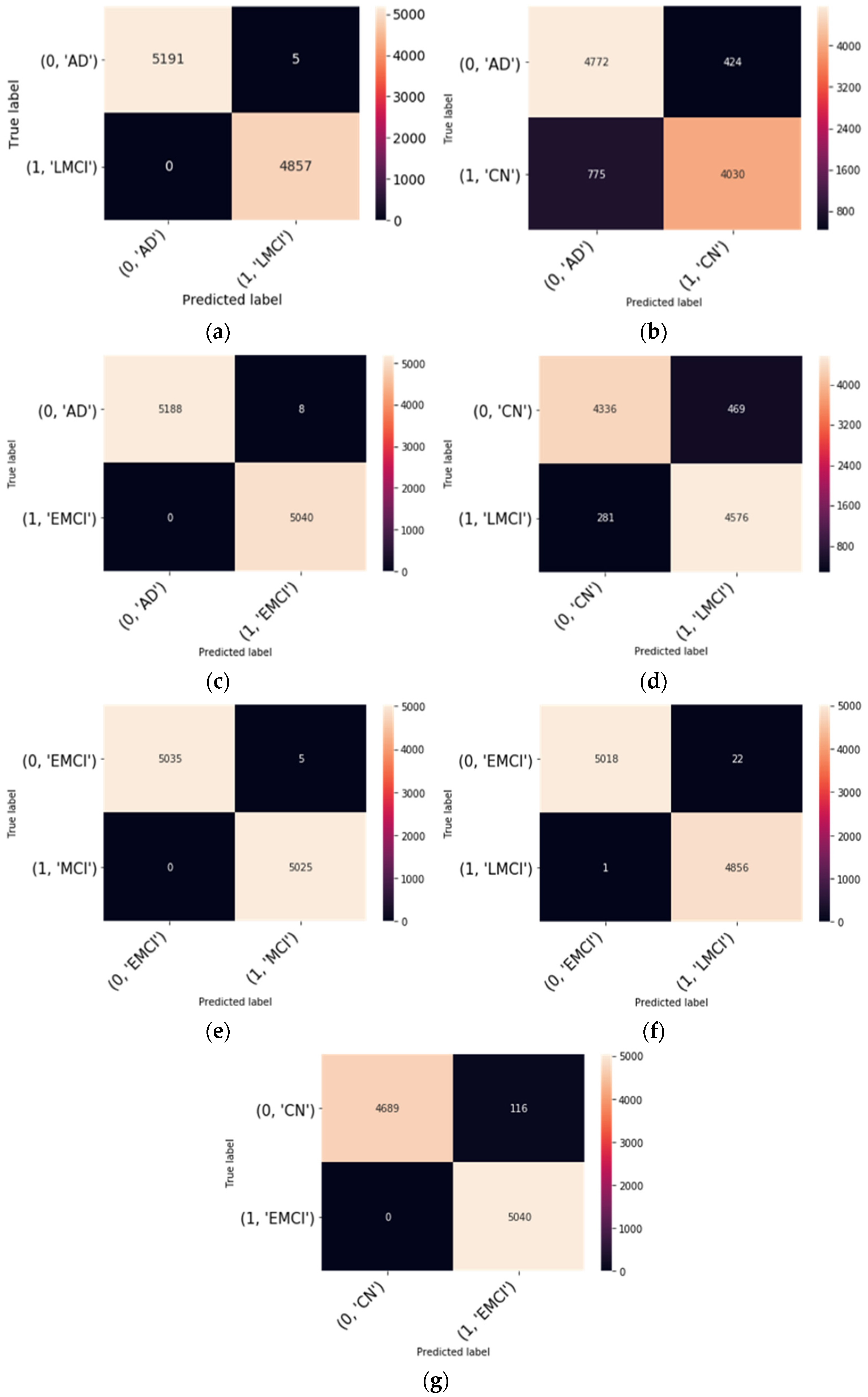
Figure 3, no subjects are misdiagnosed as AD as seen from the binary classification of AD vs. LMCI and AD vs. EMCI as shown in
Figure 3. Likewise, no subjects are misdiagnosed as EMCI and CN, respectively. However, few subjects are misdiagnosed as EMCI as in the case of EMCI vs. LMCI. This suggests that the proposed model is feasible and can correctly classify the intermediate stages of MCI, and using useful features derived from functional brain networks, the proposed model could effectively differentiate EMCI from LMCI.
With the high classification accuracy performance, the fine-tuning model produced some overfitting. To elucidate the overfitting hurdle on a noisy dataset that causes the model to learn patterns from the training data that do not generalize to the validation data, regularization technique such as dropout plays a major role. We observed that the dropout does not help in alleviating the overfitting. This finding indicates that the proposed model recognized the pattern in differentiating between the intermediary stages of MCI with the regularization technique. This corroborates the idea that the proposed network gave high precision in most of the binary classification, as shown in
Table 8. The use of the regularization technique for training allowed for the obtaining of better models, thus increasing the classification accuracy.
6. Comparison with Existing Studies
To validate our proposed approach, we compared our findings to previous studies that investigated the early diagnosis of AD using binary classification, as shown in
Table 9,
Table 10 and
Table 11. The proposed model gives better result in terms of accuracy, sensitivity, and specificity with 98.74% accuracy, 97.24% sensitivity, and 100% specificity on CN vs. EMCI classification scenario, and 99.76% accuracy, 99.56% sensitivity, and 99.97% specificity on EMCI vs. LMCI classification scenario.
The study [
23] achieved 93.46%, 94.03%, and 92.50% in terms of accuracy, sensitivity, and specificity respectively for CN vs. LMCI binary classification and thereby outperformed our proposed method in terms of accuracy and sensitivity. Likewise, the study [
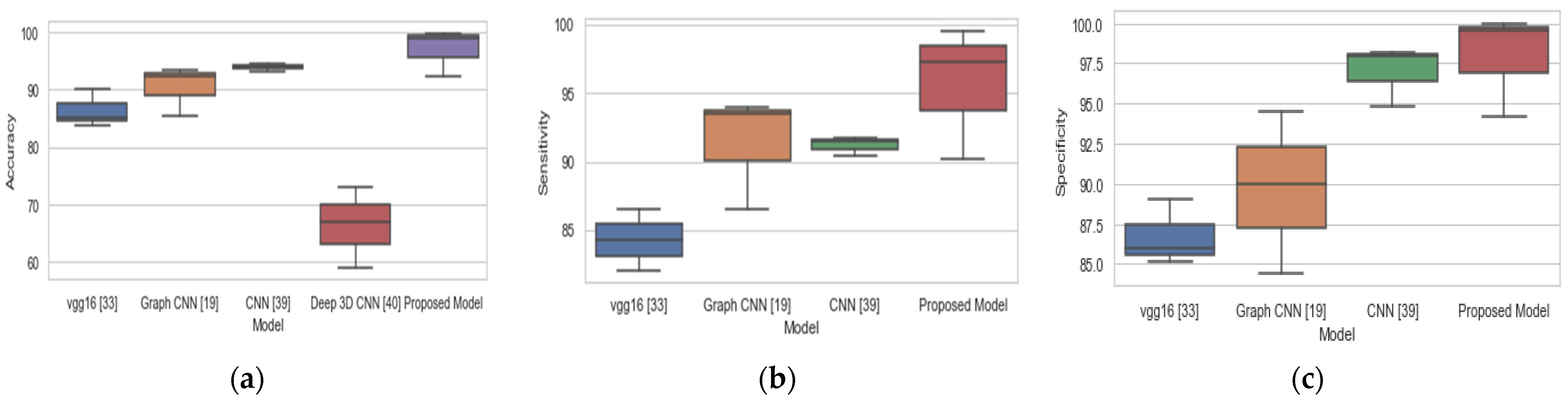
39] outperformed our proposed model in all the three-performance metrics for CN vs LMCI. The overall performance of our proposed model on the three binary classification tasks is based on accuracy, sensitivity, and specificity, and this is compared with existing methods as represented with box plots in
Figure 4. Our proposed model achieved the best performance with a median accuracy of 98.9%, median sensitivity of 98%, and median specificity of 99.9% over three binary classification tasks. Our proposed model achieved the highest accuracy, sensitivity, and specificity of 98.74%, 97.24%, and 100%, respectively, for CN vs EMCI binary classification as compared to other existing methods. In LMCI vs EMCI binary classification, our proposed method achieved a highest accuracy, sensitivity, and specificity of 99.76%, 99.56%, and 99.97%, respectively. By comparing the findings of this study to the findings of other research, we may conclude that our proposed system is a more trustworthy and accurate method.
For the clinical applicability of the proposed model in diseases such as stroke, most stroke survivors suffered from some cognitive functions, such as such as attention, concentration, memory, social cognition, language, spatial, and perceptual skills. The cognitive impairments for stroke survivors have not been addressed adequately. However, findings show that patients with neurocognitive disorders caused by AD had a higher level of affective suffering than those with neurocognitive disorders caused by stroke [
52].
7. Conclusions
AD is a debilitating brain disease that cannot be cured, and it impacts a large portion of the aging world’s population. The need to diagnose this disease early to establish effective care and enhance patients’ lives cannot be over-emphasized. This study proposed a modified ResNet18 fine-tuning approach for accurately classifying fMRI brain slices among seven binary classification tasks: CN vs. AD, CN vs. EMCI, CN vs. LMCI, EMCI vs. LMCI, EMCI vs. AD, LMCI vs. AD, and EMCI vs. MCI. The training data contained information about 61,502 images and the validation samples contained 30,095 samples. This study was able to address the problem of overfitting by finetuning all the convolutional layers and regularizing using a dropout of 0.2. This paper investigated the performance of two deep learning models (ResNet18 model without dropout and ResNet18 model with dropout) on the seven binary classification tasks. We demonstrated that finetuning ResNet18, and training it from scratch, was able to extract meaningful features for the seven binary classification tasks. The analysis results for our proposed model shows that, for regularizing with 0.2 dropout, the model was able to effectively diagnose AD early without any false positive but with very low false negative on the seven binary classification tasks. Our model achieved the best classification accuracy of 99.99%, 99.95%, and 99.95% for EMCI vs. AD, LMCI vs. AD, and MCI vs. EMCI, respectively. Additionally, for the proposed model, for EMCI vs. AD, LMCI vs. AD, and MCI vs. EMCI, the sensitivity is 99.84%, 99.90%, and 99.90%, respectively. The comparison of our method with existing methods shows that finetuning with regularization not only reduced overfitting but was also able to improve classification accuracy with low misclassification error.
For ascertaining and explaining the model decision, the use of visualization techniques (such as based on the neural network activations) will be considered in the future. We will also investigate other, recently proposed, neural network models for building classifiers in future studies. We will also explore a hybrid model, based on the pre-trained CNN, to achieve a better classification model with fewer false negatives. The performance of the model on a multiclass classification case also will be investigated in the future.