A New Gold Rush: A Review of Current and Developing Diagnostic Tools for Urinary Tract Infections
Abstract
1. Introduction
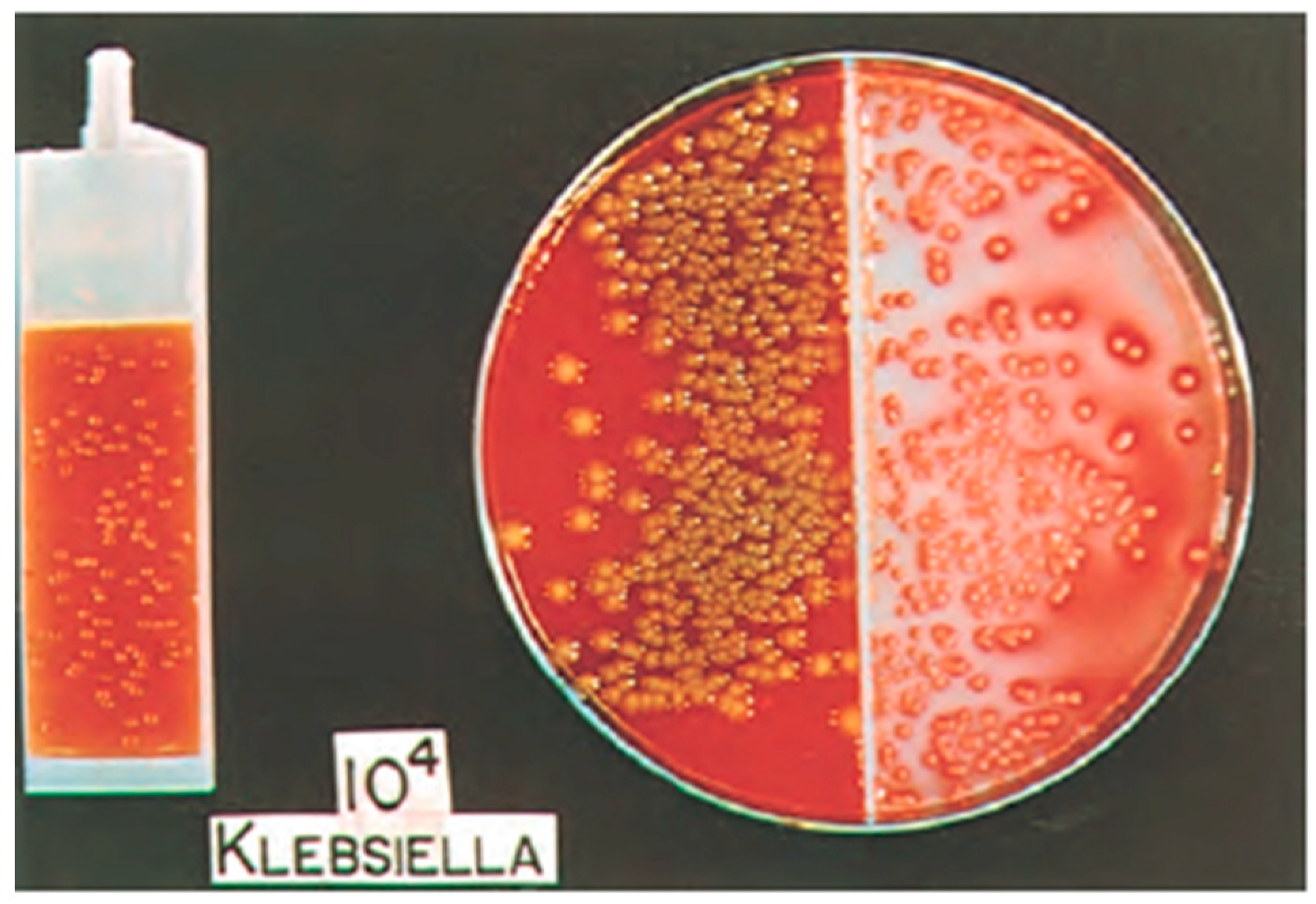
2. Standard Urine Culture with Sensitivity
3. Enhanced Culture for Atypical Organisms
4. Polymerase Chain Reaction (PCR)
5. Expanded Quantitative Urine Culture (EQUC)
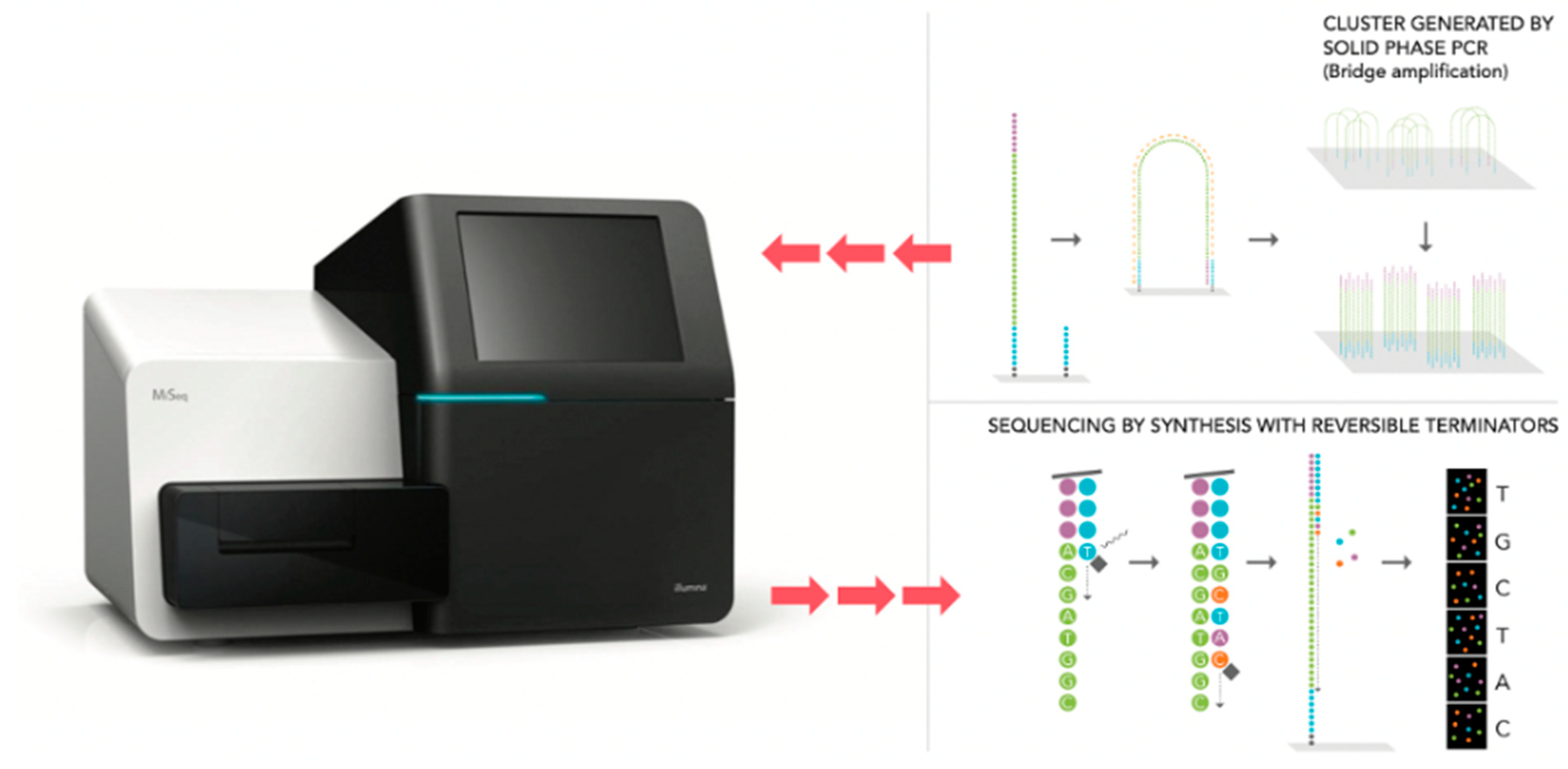
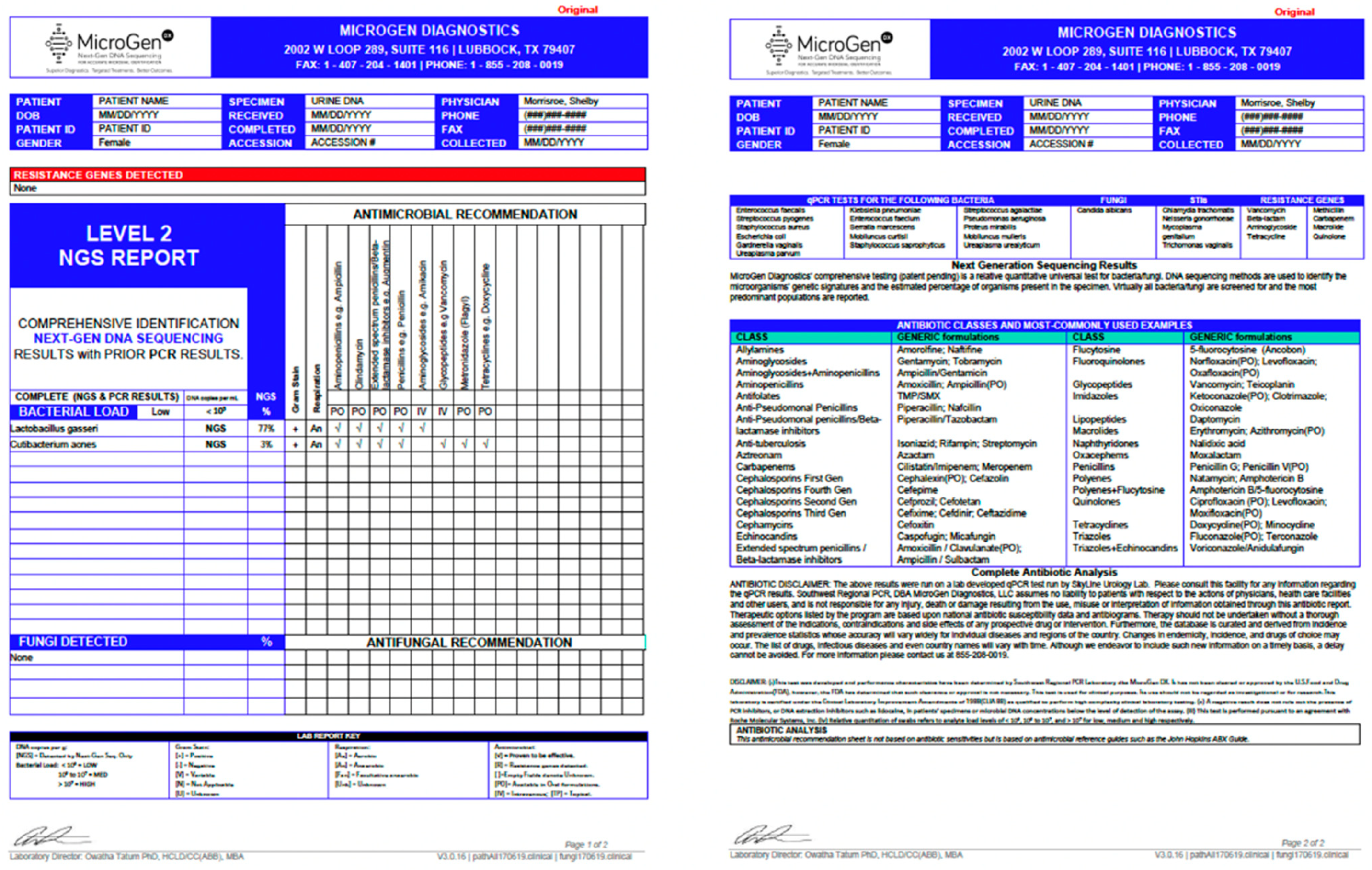
6. Next Generation Sequencing (NGS)
7. Discussion
8. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
| Study | Patient Population | Pertinent Findings | Limitations |
|---|---|---|---|
| Liss et al., 2019 [65] | -Patients scheduled for ureteroscopy -Compared urine from 20 patients tested with NGS versus SUC | -SUC: 2/20 positive -NGS: 12/20 positive; 10 of which were negative with SUC | -Low power -No antibiotic resistance data -Despite NGS data identifying a dominant bacteria, difficult to tell which organism to tailor therapy toward |
| Siddiqui et al., 2011 and 2012 [66,67] | -Examined the urinary microbiome of healthy females (HF) compared to 8 patients with interstitial cystitis (IC) | -Decreased microbiome diversity in IC vs. HF -31 vs. 45 genera respectively -Significant increase in Lactobacillus species in IC compared to HF -17 genera identified in HF urine only | -Low power -The role of the urinary microbiome in IC pathophysiology is unknown -Discrepancies in the literature regarding Lactobacillus predominance [68] |
| Karstens et al., 2016 [69] | -Urine collected from 10 women with urgency urinary incontinence (UUI) and 10 control patients -Performed NGS | -As microbial diversity decreases, UUI severity increases -Differences in abundance profiles of bacteria between groups | -Low Power -Discrepancies in the literature regarding correlation between diversity and symptom severity [54] |
| Fouts et al., 2012 [70] | -26 controls versus 27 patients with spinal cord injury (SCI) and neurogenic bladder (NB) -NGS performed on urine | -Healthy patients had more Lactobacillus and Corynebacterium relative to NB patients -NB bladder patients had higher levels of Enterococcus, Escherichia, Klebsiella, and Proteus sp. -The NB microbiome changes with duration from SCI event | -Limited power for differentiating bacterial species within groups (ex. time since SCI in NB patients) -Variation in methods for urine collection (clean catch versus catheterized specimen) |
| Mouraviev et al., 2018 [71] | -13 patients with NB and chronic urinary retention exposed to culture and NGS -Resistance genes were also examined | -Predominant organisms included Escherichia, Staphylococcus sp., Enterobacter, Pseudomonas, Klebsiella -69.2% (n = 9) of samples carried antibiotic resistance genes and 46.2% (n = 6) had multidrug resistance genes | -Lower power for NB group -Despite NGS data identifying a dominant bacteria, difficult to tell which organism to tailor therapy toward |
References
- Weiss, R.A. Robert Koch: The grandfather of cloning? Cell 2005, 123, 539–542. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Foxman, B.; Barlow, R.; D’Arcy, H.; Gillespie, B.; Sobel, J.D. Urinary tract infection: Self-reported incidence and associated costs. Ann. Epidemiol. 2000, 10, 509–515. [Google Scholar] [CrossRef]
- Stamm, W.E.; Hooton, T.M. Management of urinary tract infections in adults. N. Engl. J. Med. 1993, 329, 1328–1334. [Google Scholar]
- Foxman, B. Epidemiology of urinary tract infections: Incidence, morbidity, and economic costs. Am. J. Med. 2002, 113, 5S–13S. [Google Scholar] [CrossRef]
- Dixon, M.; Sha, S.; Stefil, M.; McDonald, M. Is it Time to Say Goodbye to Culture and Sensitivity? The Case for Culture-independent Urology. Urology 2020, 136, 112–118. [Google Scholar] [CrossRef]
- Kallen, A.J.; Welch, H.G.; Sirovich, B.E. Current antibiotic therapy for isolated urinary tract infections in women. Arch. Intern. Med. 2006, 166, 635–639. [Google Scholar] [CrossRef] [PubMed]
- Wein, A.J.; Kavoussi, L.R.; Patrin, A.W.; Peters, C.A. Campbell-Walsh Urology, 11th ed.; Elsevier Saunders: Philadelphia, PA, USA, 2015; p. 376. [Google Scholar]
- Giesen, L.G.; Cousins, G.; Dimitrov, B.D.; van de Laar, F.A.; Fahey, T. Predicting acute uncomplicated urinary tract infection in women: A systematic review of the diagnostic accuracy of symptoms and signs. BMC Fam. Pract. 2010, 11, 78. [Google Scholar] [CrossRef]
- Grigoryan, L.; Trautner, B.W.; Gupta, K. Diagnosis and management of urinary tract infections in the outpatient setting: A review. JAMA 2014, 312, 1677–1684. [Google Scholar] [CrossRef]
- Stamm, W.E.; Counts, G.W.; Running, K.R.; Fihn, S.; Turck, M.; Holmes, K.K. Diagnosis of coliform infection in acutely dysuric women. N. Engl. J. Med. 1982, 307, 463–468. [Google Scholar] [CrossRef] [PubMed]
- Fihn, S.D. Clinical practice. Acute uncomplicated urinary tract infection in women. N. Engl. J. Med. 2003, 349, 259–266. [Google Scholar] [CrossRef] [PubMed]
- Hooton, T.M.; Roberts, P.L.; Cox, M.E.; Stapleton, A.E. Voided midstream urine culture and acute cystitis in premenopausal women. N. Engl. J. Med. 2013, 369, 1883–1891. [Google Scholar] [CrossRef]
- Silver, S.A.; Baillie, L.; Simor, A.E. Positive urine cultures: A major cause of inappropriate antimicrobial use in hospitals? Can. J. Infect. Dis Med. Microbiol. 2009, 20, 107–111. [Google Scholar] [CrossRef]
- Bekeris, L.G.; Jones, B.A.; Walsh, M.K.; Wagar, E.A. Urine culture contamination: A College of American Pathologists Q-Probes study of 127 laboratories. Arch. Pathol. Lab. Med. 2008, 132, 913–917. [Google Scholar] [CrossRef]
- LaRocco, M.T.; Franek, J.; Leibach EK et, a.l. Effectiveness of Preanalytic Practices on Contamination and Diagnostic Accuracy of Urine Cultures: A Laboratory Medicine Best Practices Systematic Review and Meta-analysis. Clin. Microbiol. Rev. 2016, 29, 105–147. [Google Scholar] [CrossRef]
- Bjerklund Johansen, T.E.; Bonkat, G.; Cai, T.; Tandogdu, Z.; Wagenlehner, F.; Grabe, M. Grey Zones in the Field of Urinary Tract Infections. Eur. Urol. Focus 2016, 2, 460–462. [Google Scholar] [CrossRef]
- Tomas, M.E.; Getman, D.; Donskey, C.J.; Hecker, M.T. Overdiagnosis of Urinary Tract Infection and Underdiagnosis of Sexually Transmitted Infection in Adult Women Presenting to an Emergency Department. J. Clin. Microbiol. 2015, 53, 2686–2692. [Google Scholar] [CrossRef]
- Mabeck, C.E. Studies in urinary tract infections. I. The diagnosis of bacteriuria in women. Acta Med. Scand. 1969, 186, 35–38. [Google Scholar] [CrossRef]
- Álvarez-Lerma, F.; Nolla-Salas, J.; León, C.; Palomar, M.; Jordá, R.; Carrasco, N.; Bobillo, F. Candiduria in critically ill patients admitted to intensive care medical units. Intensiv. Care Med. 2003, 29, 1069–1076. [Google Scholar] [CrossRef]
- Paul, N.; Mathai, E.; Abraham, O.C.; Michael, J.S.; Mathai, D. Factors associated with candiduria and related mortality. J. Infect. 2007, 55, 450–455. [Google Scholar] [CrossRef]
- Slavin, M.; Fastenau, J.; Sukarom, I.; Mavros, P.; Crowley, S.; Gerth, W.C. Burden of hospitalization of patients with Candida and Aspergillus infections in Australia. Int. J. Infect. Dis 2004, 8, 111–120. [Google Scholar] [CrossRef][Green Version]
- Nickel, J.C. Management of urinary tract infections: Historical perspective and current strategies: Part 2—Modern management. J. Urol. 2005, 173, 27–32. [Google Scholar] [CrossRef]
- Sobel, J.D.; Fisher, J.F.; Kauffman, C.A.; Newman, C.A. Candida urinary tract infections—Epidemiology. Clin. Infect. Dis. 2011, 52, S433–S436. [Google Scholar] [CrossRef]
- Toner, L.; Papa, N.; Aliyu, S.H.; Dev, H.; Lawrentschuk, N.; Al-Hayek, S. Candida growth in urine cultures: A contemporary analysis of species and antifungal susceptibility profiles. QJM 2016, 109, 325–329. [Google Scholar] [CrossRef]
- Helbig, S.; Achkar, J.M.; Jain, N.; Wang, X.; Gialanella, P.; Levi, M.; Fries, B.C. Diagnosis and inflammatory response of patients with candiduria. Mycoses 2013, 56, 61–69. [Google Scholar] [CrossRef]
- Merchant, S.; Bharati, A.; Merchant, N. Tuberculosis of the genitourinary system-Urinary tract tuberculosis: Renal tuberculosis-Part I. Indian J. Radiol. Imaging 2013, 23, 46–63. [Google Scholar] [CrossRef] [PubMed]
- Ribot, S.; Gal, K.; Goldblat, M.V.; Eslami, H.H. The role of anaerobic bacteria in the pathogenesis of urinary tract infections. J. Urol. 1981, 126, 852–853. [Google Scholar] [CrossRef]
- Headington, J.T.; Beyerlein, B. Anaerobic bacteria in routine urine culture. J. Clin. Pathol. 1966, 19, 573–576. [Google Scholar] [CrossRef]
- Shepard, M.C.; Lunceford, C.D. Differential agar medium (A7) for identification of Ureaplasma urealyticum (human T mycoplasmas) in primary cultures of clinical material. J. Clin. Microbiol. 1976, 3, 613–625. [Google Scholar]
- Stellrecht, K.A.; Woron, A.M.; Mishrik, N.G.; Venezia, R.A. Comparison of multiplex PCR assay with culture for detection of genital mycoplasmas. J. Clin. Microbiol. 2004, 42, 1528–1533. [Google Scholar] [CrossRef] [PubMed]
- Valentine-King, M.A.; Brown, M.B. Antibacterial Resistance in Ureaplasma Species and Mycoplasma hominis Isolates from Urine Cultures in College-Aged Females. Antimicrob. Agents Chemother. 2017, 61, e01104–e01117. [Google Scholar] [CrossRef]
- Elnifro, E.M.; Ashshi, A.M.; Cooper, R.J.; Klapper, P.E. Multiplex PCR: Optimization and application in diagnostic virology. Clin. Microbiol. Rev. 2000, 13, 559–570. [Google Scholar] [CrossRef]
- Wojno, K.J.; Baunoch, D.; Luke, N.; Opel, M.; Korman, H.; Kelly, C.; Jafri, S.M.A.; Keating, P.; Hazelton, D.; Hindu, S.; et al. Multiplex PCR Based Urinary Tract Infection (UTI) Analysis Compared to Traditional Urine Culture in Identifying Significant Pathogens in Symptomatic Patients. Urology 2020, 136, 119–126. [Google Scholar] [CrossRef]
- Scerbo, M.H.; Kaplan, H.B.; Dua, A.; Litwin, D.B.; Ambrose, C.G.; Moore, L.J.; Murray, C.C.K.; Wade, C.E.; Holcomb, J.B. Beyond Blood Culture and Gram Stain Analysis: A Review of Molecular Techniques for the Early Detection of Bacteremia in Surgical Patients. Surg. Infect. 2016, 17, 294–302. [Google Scholar] [CrossRef]
- Gayet-Ageron, A.; Combescure, C.; Lautenschlager, S.; Ninet, B.; Perneger, T.V. Comparison of Diagnostic Accuracy of PCR Targeting the 47-Kilodalton Protein Membrane Gene of Treponema pallidum and PCR Targeting the DNA Polymerase I Gene: Systematic Review and Meta-analysis. J. Clin. Microbiol. 2015, 53, 3522–3529. [Google Scholar] [CrossRef]
- Zboromyrska, Y.; Vila, J. Advanced PCR-based molecular diagnosis of gastrointestinal infections: Challenges and opportunities. Expert Rev. Mol. Diagn. 2016, 16, 631–640. [Google Scholar] [CrossRef]
- Preiser, W.; Bräuninger, S.; Schwerdtfeger, R.; Ayliffe, U.; A Garson, J.; Brink, N.S.; Franck, S.; Doerr, H.W.; Rabenau, H.F. Evaluation of diagnostic methods for the detection of cytomegalovirus in recipients of allogeneic stem cell transplants. J. Clin. Virol. 2001, 20, 59–70. [Google Scholar] [CrossRef]
- Gaydos, C.A.; Theodore, M.; Dalesio, N.; Wood, B.J.; Quinn, T.C. Comparison of three nucleic acid amplification tests for detection of Chlamydia trachomatis in urine specimens. J. Clin. Microbiol. 2004, 42, 3041–3045. [Google Scholar] [CrossRef]
- Pillay, A.; Radebe, F.; Fehler, G.; Htun, Y.; Ballard, R.C. Comparison of a TaqMan-based real-time polymerase chain reaction with conventional tests for the detection of Trichomonas vaginalis. Sex. Transm. Infect. 2007, 83, 126–129. [Google Scholar] [CrossRef]
- Price, T.K.; Dune, T.; Hilt, E.E.; Thomas-White, K.J.; Kliethermes, S.; Brincat, C.; Brubaker, L.; Wolfe, A.J.; Mueller, E.R.; Schreckenberger, P.C. The Clinical Urine Culture: Enhanced Techniques Improve Detection of Clinically Relevant Microorganisms. J. Clin. Microbiol. 2016, 54, 1216–1222. [Google Scholar] [CrossRef] [PubMed]
- Hilt, E.E.; McKinley, K.; Pearce, M.M.; Rosenfeld, A.B.; Zilliox, M.J.; Mueller, E.R.; Brubaker, L.; Gai, X.; Wolfe, A.J.; Schreckenberger, P.C. Urine is not sterile: Use of enhanced urine culture techniques to detect resident bacterial flora in the adult female bladder. J. Clin. Microbiol. 2014, 52, 871–876. [Google Scholar] [CrossRef] [PubMed]
- Croxall, G.; Weston, V.; Joseph, S.; Manning, G.; Cheetham, P.; McNally, A. Increased human pathogenic potential of Escherichia coli from polymicrobial urinary tract infections in comparison to isolates from monomicrobial culture samples. J. Med. Microbiol. 2011, 60, 102–109. [Google Scholar] [CrossRef]
- Waller, T.A.; Pantin, S.A.L.; Yenior, A.L.; Pujalte, G.G.A. Urinary Tract Infection Antibiotic Resistance in the United States. Prim. Care 2018, 45, 455–466. [Google Scholar] [CrossRef]
- Palmqvist, E.; Aspevall, O.; Burman, E.; Nordin, G.; Svahn, A.; Forsum, U. Difficulties for primary health care staff in interpreting bacterial findings on a device for simplified urinary culture. Scand. J. Clin. Lab. Invest. 2008, 68, 312–316. [Google Scholar] [CrossRef]
- Lehmann, L.E.; Hauser, S.; Malinka, T.; Klaschik, S.; Weber, S.U.; Schewe, J.-C.; Stüber, F.; Book, M. Rapid qualitative urinary tract infection pathogen identification by SeptiFast real-time PCR. PLoS ONE 2011, 6, e17146. [Google Scholar] [CrossRef]
- Cybulski, Z.; Schmidt, K.; Grabiec, A.; Talaga, Z.; Bociąg, P.; Wojciechowicz, J.; Roszak, A.; Kycler, W. Usability application of multiplex polymerase chain reaction in the diagnosis of microorganisms isolated from urine of patients treated in cancer hospital. Radiol. Oncol. 2013, 47, 296–303. [Google Scholar]
- Schmidt, K.; Stanley, K.K.; Hale, R.; Smith, L.; Wain, J.; O’Grady, J.; Livermore, D.M. Evaluation of multiplex tandem PCR (MT-PCR) assays for the detection of bacterial resistance genes among Enterobacteriaceae in clinical urines. J. Antimicrob. Chemother. 2019, 74, 349–356. [Google Scholar] [CrossRef]
- Rachow, A.; Zumla, A.; Heinrich, N.; Rojas-Ponce, G.; Mtafya, B.; Reither, K.; Ntinginya, E.N.; O’Grady, J.; Huggett, J.; Dheda, K.; et al. Rapid and accurate detection of Mycobacterium tuberculosis in sputum samples by Cepheid Xpert MTB/RIF assay—A clinical validation study. PLoS ONE 2011, 6, e20458. [Google Scholar]
- Wolfe, A.J.; Toh, E.; Shibata, N.; Rong, R.; Kenton, K.S.; Fitzgerald, M.; Mueller, E.R.; Schreckenberger, P.C.; Dong, Q.; Nelson, D.E.; et al. Evidence of uncultivated bacteria in the adult female bladder. J. Clin. Microbiol. 2012, 50, 1376–1383. [Google Scholar] [CrossRef]
- Antunes-Lopes, T.; Vale, L.; Coelho, A.M.; Silva, C.; Rieken, M.; Geavlete, B.; Rashid, T.; Rahnama’I, S.M.; Cornu, J.N.; Marcelissen, T. The Role of Urinary Microbiota in Lower Urinary Tract Dysfunction: A Systematic Review. Eur. Urol. Focus 2020, 6, 361–369. [Google Scholar] [CrossRef]
- Thapaliya, J.; Khadka, P.; Thapa, S.; Gongal, C. Enhanced quantitative urine culture technique, a slight modification, in detecting under-diagnosed pediatric urinary tract infection. BMC Res. Notes 2020, 13, 5. [Google Scholar]
- Jacobs, K.M.; Price, T.K.; Thomas-White, K.; Halverson, T.; Davies, A.; Myers, D.L.; Wolfe, A.J. Cultivable Bacteria in Urine of Women With Interstitial Cystitis: (Not) What We Expected. Female Pelvic Med. Reconstr. Surg. 2020. [Google Scholar] [CrossRef]
- Bresler, L.; Price, T.K.; Hilt, E.E.; Joyce, C.; Fitzgerald, C.M.; Wolfe, A.J. Female lower urinary tract microbiota do not associate with IC/PBS symptoms: A case-controlled study. Int. Urogynecol. J. 2019, 30, 1835–1842. [Google Scholar] [CrossRef]
- Bajic, P.; Van Kuiken, M.E.; Burge, B.K.; Kirshenbaum, E.J.; Joyce, C.J.; Wolfe, A.J.; Branch, J.D.; Bresler, L.; Farooq, A.V. Male Bladder Microbiome Relates to Lower Urinary Tract Symptoms. Eur. Urol. Focus 2020, 6, 376–382. [Google Scholar] [CrossRef]
- Pearce, M.M.; Zilliox, M.J.; Rosenfeld, A.B.; Thomas-White, K.J.; Richter, H.E.; Nager, C.W.; Visco, A.G.; Nygaard, I.E.; Barber, M.D.; Schaffer, J.; et al. The female urinary microbiome in urgency urinary incontinence. Am. J. Obstet. Gynecol. 2015, 213, 347.e1–347.e11. [Google Scholar] [CrossRef]
- Thomas-White, K.; Brady, M.; Wolfe, A.J.; Mueller, E.R. The bladder is not sterile: History and current discoveries on the urinary microbiome. Curr. Bladder Dysfunct. Rep. 2016, 11, 18–24. [Google Scholar] [CrossRef]
- Dornbier, R.A.; Bajic, P.; Van Kuiken, M.; Jardaneh, A.; Lin, H.; Gao, X.; Knudsen, B.; Dong, Q.; Wolfe, A.J.; Schwaderer, A.L. The microbiome of calcium-based urinary stones. Urolithiasis 2020, 48, 191–199. [Google Scholar] [CrossRef]
- Gasiorek, M.; Hsieh, M.H.; Forster, C.S. Utility of DNA Next-Generation Sequencing and Expanded Quantitative Urine Culture in Diagnosis and Management of Chronic or Persistent Lower Urinary Tract Symptoms. J. Clin. Microbiol. 2019, 58. [Google Scholar] [CrossRef]
- Dubnau, D.; Smith, I.; Morell, P.; Marmur, J. Gene conservation in Bacillus species. I. Conserved genetic and nucleic acid base sequence homologies. Proc. Natl. Acad. Sci. USA 1965, 54, 491–498. [Google Scholar] [CrossRef]
- Hasman, H.; Saputra, D.; Sicheritz-Ponten, T.; Lund, O.; Svendsen, C.A.; Frimodt-Møller, N.; Aarestrup, F.M. Rapid whole-genome sequencing for detection and characterization of microorganisms directly from clinical samples. J. Clin. Microbiol. 2014, 52, 139–146. [Google Scholar] [CrossRef]
- McDonald, M.; Kameh, D.; Johnson, M.E.; Johansen, T.E.B.; Albala, D.; Mouraviev, V. A Head-to-Head Comparative Phase II Study of Standard Urine Culture and Sensitivity Versus DNA Next-generation Sequencing Testing for Urinary Tract Infections. Rev. Urol. 2017, 19, 213–220. [Google Scholar]
- Ishihara, T.; Watanabe, N.; Inoue, S.; Aoki, H.; Tsuji, T.; Yamamoto, B.; Yanagi, H.; Oki, M.; Kryukov, K.; Nakagawa, S.; et al. Usefulness of next-generation DNA sequencing for the diagnosis of urinary tract infection. Drug Discov. Ther. 2020, 14, 42–49. [Google Scholar] [CrossRef]
- Lewis, D.A.; Brown, R.; Williams, J.; White, P.; Jacobson, S.K.; Marchesi, J.R.; Drake, M.J. The human urinary microbiome; bacterial DNA in voided urine of asymptomatic adults. Front. Cell Infect. Microbiol. 2013, 3, 41. [Google Scholar] [CrossRef] [PubMed]
- Dixon, M.; Stefil, M.; McDonald, M.; Bjerklund-Johansen, T.E.; Naber, K.; Wagenlehner, F.; Mouraviev, V. Metagenomics in diagnosis and improved targeted treatment of UTI. World J. Urol. 2020, 38, 35–43. [Google Scholar] [CrossRef]
- Liss, M.R.K.; Wheeler, A.; Arellanes, A.; Tseng, T. MP77-02 Antibiotic selection using next generation sequencing versus urine culture for stewardship prior to surgery using and interprofessional model. J. Urol. 2019, 201, e1131–e1132. [Google Scholar] [CrossRef]
- Siddiqui, H.; Nederbragt, A.J.; Lagesen, K.; Jeansson, S.L.; Jakobsen, K.S. Assessing diversity of the female urine microbiota by high throughput sequencing of 16S rDNA amplicons. BMC Microbiol. 2011, 11, 244. [Google Scholar] [CrossRef]
- Siddiqui, H.; Lagesen, K.; Nederbragt, A.J.; Jeansson, S.L.; Jakobsen, K.S. Alterations of microbiota in urine from women with interstitial cystitis. BMC Microbiol. 2012, 12, 205. [Google Scholar] [CrossRef] [PubMed]
- Abernethy, M.G.; Rosenfeld, A.; White, J.R.; Mueller, M.G.; Lewicky-Gaupp, C.; Kenton, K. Urinary Microbiome and Cytokine Levels in Women With Interstitial Cystitis. Obstet. Gynecol. 2017, 129, 500–506. [Google Scholar] [CrossRef] [PubMed]
- Karstens, L.; Asquith, M.; Davin, S.; Stauffer, P.; Fair, D.; Gregory, W.T.; Rosenbaum, J.T.; McWeeney, S.K.; Nardos, R. Does the Urinary Microbiome Play a Role in Urgency Urinary Incontinence and Its Severity? Front. Cell Infect. Microbiol. 2016, 6, 78. [Google Scholar] [CrossRef] [PubMed]
- E Fouts, D.; Pieper, R.; Szpakowski, S.; Pohl, H.; Knoblach, S.; Suh, M.-J.; Huang, S.-T.; Ljungberg, I.; Sprague, B.M.; Lucas, S.K.; et al. Integrated next-generation sequencing of 16S rDNA and metaproteomics differentiate the healthy urine microbiome from asymptomatic bacteriuria in neuropathic bladder associated with spinal cord injury. J. Transl. Med. 2012, 10, 174. [Google Scholar] [CrossRef]
- Mouraviev, V.; McDonald, M. An implementation of next generation sequencing for prevention and diagnosis of urinary tract infection in urology. Can. J. Urol. 2018, 25, 9349–9356. [Google Scholar] [PubMed]
| SUC | EQUC | PCR | NGS | |
|---|---|---|---|---|
| Diagnostic Tool | Microbiology; developed in 1880’s, designed for acute infections | Advanced Microbiology; developed in 2014, designed for acute infections | Molecular Method; developed in 1970’s | Advanced Molecular Method; developed in 2004 |
| Methodology | Detects species by growing bacteria or fungi on petri or agar plates | Detects atypical and subthreshold species by growing bacteria or fungi on petri or agar plates under modified conditions | Matches extracted microbial DNA to limited PCR panels | Matches extracted microbial DNA to large curated species libraries |
| Ability to Detect Dominant Species | Can identify dominant species with high sensitivity for common uropathogens | Identification of multiple isolates may represent contamination (i.e., requires catheter-collected urine for accuracy) | Cannot reliably discern uropathogen from normal uromicrobiome | Identifies all species in a sample and lists by dominance |
| Antibiotic Sensitivities | Reliable identification of antibiotic sensitivity profiles | Reliable identification of antibiotic sensitivity profiles | Inconsistent inclusion of resistance genes in PCR panels | Dependent on public genomic reference libraries for resistance genes results |
| Specimen Management | Sensitive to time and temperature | Sensitive to time and temperature | Not easily affected by time or temperature | Not easily affected by time or temperature |
| Turn-Around Time | 1–2 days for bacteria; 20+ days for fungi | 1–2 days for bacteria; 20+ days for fungi | 1 day | 3–5 days |
| Costs | −$34 −$71 (fungal) −$114 (acid fast) -Variable (anaerobe) | −$67 | −$15–30 | −$200 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, R.; Deebel, N.; Casals, R.; Dutta, R.; Mirzazadeh, M. A New Gold Rush: A Review of Current and Developing Diagnostic Tools for Urinary Tract Infections. Diagnostics 2021, 11, 479. https://doi.org/10.3390/diagnostics11030479
Xu R, Deebel N, Casals R, Dutta R, Mirzazadeh M. A New Gold Rush: A Review of Current and Developing Diagnostic Tools for Urinary Tract Infections. Diagnostics. 2021; 11(3):479. https://doi.org/10.3390/diagnostics11030479
Chicago/Turabian StyleXu, Raymond, Nicholas Deebel, Randy Casals, Rahul Dutta, and Majid Mirzazadeh. 2021. "A New Gold Rush: A Review of Current and Developing Diagnostic Tools for Urinary Tract Infections" Diagnostics 11, no. 3: 479. https://doi.org/10.3390/diagnostics11030479
APA StyleXu, R., Deebel, N., Casals, R., Dutta, R., & Mirzazadeh, M. (2021). A New Gold Rush: A Review of Current and Developing Diagnostic Tools for Urinary Tract Infections. Diagnostics, 11(3), 479. https://doi.org/10.3390/diagnostics11030479






