Serum Myo-Inositol, Dimethyl Sulfone, and Valine in Combination with Creatinine Allow Accurate Assessment of Renal Insufficiency—A Proof of Concept
Abstract
1. Introduction
2. Materials and Methods
2.1. Cohorts and Samples
2.2. Benchmarking
2.3. mGFR, Serum Creatinine, and Cystatin C Measurements
2.4. NMR Analysis
2.5. Biomarker Quantification
2.5.1. Precision
2.5.2. Linearity
2.5.3. Bias
2.6. GFR Modeling
2.7. Model Selection
3. Results
3.1. Biomarker Quantification
3.2. GFR Estimation
3.3. Subgroup Analysis of GFRNMR According to Sex and Age
3.4. Molecular Phenotyping by Matched Sample Sets
4. Discussion
5. Patents
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Steubl, D.; Inker, L.A. How best to estimate glomerular filtration rate? Novel filtration markers and their application. Curr. Opin. Nephrol. Hypertens. 2018, 27, 398–405. [Google Scholar] [CrossRef] [PubMed]
- Inker, L.A.; Schmid, C.H.; Tighiouart, H.; Eckfeldt, J.H.; Feldman, H.I.; Greene, T.; Kusek, J.W.; Manzi, J.; Van Lente, F.; Zhang, Y.L.; et al. Estimating glomerular filtration rate from serum creatinine and cystatin C. N. Engl. J. Med. 2012, 367, 20–29. [Google Scholar] [CrossRef] [PubMed]
- Chesney, R.W. The future of pediatric nephrology. Pediatric Nephrol. 2005, 20, 867–871. [Google Scholar] [CrossRef] [PubMed]
- Seegmiller, J.C.; Eckfeldt, J.H.; Lieske, J.C. Challenges in Measuring Glomerular Filtration Rate: A Clinical Laboratory Perspective. Adv. Chronic Kidney Dis. 2018, 25, 84–92. [Google Scholar] [CrossRef]
- Guignard, J.-P. Postnatal Development of Glomerular Filtration Rate in Neonates. In Fetal and Neonatal Physiology, 3rd ed.; Elsevier: Amsterdam, The Netherlands, 2003; Volume 2, pp. 1256–1266. [Google Scholar]
- Glassock, R.J.; Warnock, D.G.; Delanaye, P. The global burden of chronic kidney disease: Estimates, variability and pitfalls. Nat. Rev. Nephrol. 2017, 13, 104–114. [Google Scholar] [CrossRef]
- Topf, J.M.; Inker, L.A. Measurement of glomerular filtration rate. In Nephrology Secrets, 4th ed.; Elsevier: Amsterdam, The Netherlands, 2018; pp. 22–29. [Google Scholar]
- Porrini, E.; Ruggenenti, P.; Luis-Lima, S.; Carrara, F.; Jimenez, A.; de Vries, A.P.J.; Torres, A.; Gaspari, F.; Remuzzi, G. Estimated GFR: Time for a critical appraisal. Nat. Rev. Nephrol. 2019, 15, 177–190. [Google Scholar] [CrossRef]
- Hsu, C.Y.; Bansal, N. Measured GFR as “gold standard”—All that glitters is not gold? Clin. J. Am. Soc. Nephrol. 2011, 6, 1813–1814. [Google Scholar] [CrossRef]
- Banas, M.; Neumann, S.; Eiglsperger, J.; Schiffer, E.; Putz, F.J.; Reichelt-Wurm, S.; Kramer, B.K.; Pagel, P.; Banas, B. Identification of a urine metabolite constellation characteristic for kidney allograft rejection. Metabolomics 2018, 14, 116. [Google Scholar] [CrossRef]
- Sekula, P.; Goek, O.N.; Quaye, L.; Barrios, C.; Levey, A.S.; Romisch-Margl, W.; Menni, C.; Yet, I.; Gieger, C.; Inker, L.A.; et al. A Metabolome-Wide Association Study of Kidney Function and Disease in the General Population. J. Am. Soc. Nephrol. 2016, 27, 1175–1188. [Google Scholar] [CrossRef]
- Luck, M.; Bertho, G.; Bateson, M.; Karras, A.; Yartseva, A.; Thervet, E.; Damon, C.; Pallet, N. Rule-Mining for the Early Prediction of Chronic Kidney Disease Based on Metabolomics and Multi-Source Data. PLoS ONE 2016, 11, e0166905. [Google Scholar] [CrossRef]
- Duranton, F.; Lundin, U.; Gayrard, N.; Mischak, H.; Aparicio, M.; Mourad, G.; Daures, J.P.; Weinberger, K.M.; Argiles, A. Plasma and urinary amino acid metabolomic profiling in patients with different levels of kidney function. Clin. J. Am. Soc. Nephrol. 2014, 9, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Mutsaers, H.A.; Engelke, U.F.; Wilmer, M.J.; Wetzels, J.F.; Wevers, R.A.; van den Heuvel, L.P.; Hoenderop, J.G.; Masereeuw, R. Optimized metabolomic approach to identify uremic solutes in plasma of stage 3-4 chronic kidney disease patients. PLoS ONE 2013, 8, e71199. [Google Scholar] [CrossRef] [PubMed]
- Hao, X.; Liu, X.; Wang, W.; Ren, H.; Xie, J.; Shen, P.; Lin, D.; Chen, N. Distinct metabolic profile of primary focal segmental glomerulosclerosis revealed by NMR-based metabolomics. PLoS ONE 2013, 8, e78531. [Google Scholar] [CrossRef] [PubMed]
- Sui, W.; Li, L.; Che, W.; Guimai, Z.; Chen, J.; Li, W.; Dai, Y. A proton nuclear magnetic resonance-based metabonomics study of metabolic profiling in immunoglobulin a nephropathy. Clinics 2012, 67, 363–373. [Google Scholar] [CrossRef]
- Qi, S.; Ouyang, X.; Wang, L.; Peng, W.; Wen, J.; Dai, Y. A pilot metabolic profiling study in serum of patients with chronic kidney disease based on (1) H-NMR-spectroscopy. Clin. Transl. Sci. 2012, 5, 379–385. [Google Scholar] [CrossRef] [PubMed]
- Mao, Y.Y.; Bai, J.Q.; Chen, J.H.; Shou, Z.F.; He, Q.; Wu, J.Y.; Chen, Y.; Cheng, Y.Y. A pilot study of GC/MS-based serum metabolic profiling of acute rejection in renal transplantation. Transpl. Immunol. 2008, 19, 74–80. [Google Scholar] [CrossRef] [PubMed]
- Coresh, J.; Inker, L.A.; Sang, Y.; Chen, J.; Shafi, T.; Post, W.S.; Shlipak, M.G.; Ford, L.; Goodman, K.; Perichon, R.; et al. Metabolomic profiling to improve glomerular filtration rate estimation: A proof-of-concept study. Nephrol. Dial. Transplant. 2019, 34, 825–833. [Google Scholar] [CrossRef] [PubMed]
- Niwa, T.; Yamamoto, N.; Maeda, K.; Yamada, K.; Ohki, T.; Mori, M. Gas chromatographic—Mass spectrometric analysis of polyols in urine and serum of uremic patients. Identification of new deoxyalditols and inositol isomers. J. Chromatogr. 1983, 277, 25–39. [Google Scholar] [CrossRef]
- Inamoto, Y.; Hiraga, Y.; Hanai, T.; Kinosita, T. The development of a sensitive myo-inositol analyser using a liquid chromatograph with a post-label fluorescence detector. Biomed. Chromatogr. 1995, 9, 146–149. [Google Scholar] [CrossRef]
- Michaelis, T.; Videen, J.S.; Linsey, M.S.; Ross, B.D. Dialysis and transplantation affect cerebral abnormalities of end-stage renal disease. J. Magn. Reson. Imaging 1996, 6, 341–347. [Google Scholar] [CrossRef]
- Choi, J.Y.; Yoon, Y.J.; Choi, H.J.; Park, S.H.; Kim, C.D.; Kim, I.S.; Kwon, T.H.; Do, J.Y.; Kim, S.H.; Ryu, D.H.; et al. Dialysis modality-dependent changes in serum metabolites: Accumulation of inosine and hypoxanthine in patients on haemodialysis. Nephrol. Dial. Transplant. 2011, 26, 1304–1313. [Google Scholar] [CrossRef] [PubMed]
- Al-Ani, B.; Fitzpatrick, M.; Al-Nuaimi, H.; Coughlan, A.M.; Hickey, F.B.; Pusey, C.D.; Savage, C.; Benton, C.M.; O’Brien, E.C.; O’Toole, D.; et al. Changes in urinary metabolomic profile during relapsing renal vasculitis. Sci. Rep. 2016, 6, 38074. [Google Scholar] [CrossRef] [PubMed]
- Galle, J. Oxidative stress in chronic renal failure. Nephrol. Dial. Transplant. 2001, 16, 2135–2137. [Google Scholar] [CrossRef] [PubMed]
- Tsuruta, Y.; Ito, Y.; Harada, K.I.; Narita, Y.; Ohbayashi, T.A.; Azekura, H.; Fukagawa, M.; Narita, M.; Maeda, K. Measurements of blood DMSO and DMSO2 in a healthy person and a hemodialysis patient. Clin. Exp. Nephrol. 2001, 5, 158–162. [Google Scholar] [CrossRef]
- Locatelli, F.; Canaud, B.; Eckardt, K.U.; Stenvinkel, P.; Wanner, C.; Zoccali, C. Oxidative stress in end-stage renal disease: An emerging threat to patient outcome. Nephrol. Dial. Transplant. 2003, 18, 1272–1280. [Google Scholar] [CrossRef] [PubMed]
- Vanholder, R.; De Smet, R.; Glorieux, G.; Argiles, A.; Baurmeister, U.; Brunet, P.; Clark, W.; Cohen, G.; De Deyn, P.P.; Deppisch, R.; et al. Review on uremic toxins: Classification, concentration, and interindividual variability. Kidney Int. 2003, 63, 1934–1943. [Google Scholar] [CrossRef] [PubMed]
- Rysz, J.; Kasielski, M.; Apanasiewicz, J.; Krol, M.; Woznicki, A.; Luciak, M.; Nowak, D. Increased hydrogen peroxide in the exhaled breath of uraemic patients unaffected by haemodialysis. Nephrol. Dial. Transplant. 2004, 19, 158–163. [Google Scholar] [CrossRef]
- Juttner, B.; Gehrmann, A.; Breitmeier, D.; Jaeger, K.; Weissig, A.; Bornscheuer, A.; Piepenbrock, S.; Scheinichen, D. Renal transplantation normalized hydrogen peroxide production of neutrophils within the first day. Am. J. Nephrol. 2008, 28, 531–538. [Google Scholar] [CrossRef]
- Al Khodor, S.; Shatat, I.F. Gut microbiome and kidney disease: A bidirectional relationship. Pediatric Nephrol. 2017, 32, 921–931. [Google Scholar] [CrossRef]
- Kraut, J.A.; Kurtz, I. Metabolic acidosis of CKD: Diagnosis, clinical characteristics, and treatment. Am. J. Kidney Dis. 2005, 45, 978–993. [Google Scholar] [CrossRef]
- Kumar, M.A.; Bitla, A.R.; Raju, K.V.; Manohar, S.M.; Kumar, V.S.; Narasimha, S.R. Branched chain amino acid profile in early chronic kidney disease. Saudi J. Kidney Dis. Transplant. 2012, 23, 1202–1207. [Google Scholar] [CrossRef]
- Schaeffner, E.S.; van der Giet, M.; Gaedeke, J.; Tolle, M.; Ebert, N.; Kuhlmann, M.K.; Martus, P. The Berlin initiative study: The methodology of exploring kidney function in the elderly by combining a longitudinal and cross-sectional approach. Eur. J. Epidemiol. 2010, 25, 203–210. [Google Scholar] [CrossRef] [PubMed]
- Levin, A.S.; Bilous, R.W.; Coresh, J. Chapter 1: Definition and classification of CKD. Kidney Int. Suppl. 2013, 3, 19–62. [Google Scholar] [CrossRef]
- Levey, A.S.; Stevens, L.A.; Schmid, C.H.; Zhang, Y.L.; Castro, A.F., III; Feldman, H.I.; Kusek, J.W.; Eggers, P.; Van Lente, F.; Greene, T.; et al. A new equation to estimate glomerular filtration rate. Ann. Intern. Med. 2009, 150, 604–612. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, G.J.; Munoz, A.; Schneider, M.F.; Mak, R.H.; Kaskel, F.; Warady, B.A.; Furth, S.L. New equations to estimate GFR in children with CKD. J. Am. Soc. Nephrol. 2009, 20, 629–637. [Google Scholar] [CrossRef]
- Pottel, H.; Bjork, J.; Courbebaisse, M.; Couzi, L.; Ebert, N.; Eriksen, B.O.; Dalton, R.N.; Dubourg, L.; Gaillard, F.; Garrouste, C.; et al. Development and Validation of a Modified Full Age Spectrum Creatinine-Based Equation to Estimate Glomerular Filtration Rate: A Cross-sectional Analysis of Pooled Data. Ann. Intern. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, G.J.; Schneider, M.F.; Maier, P.S.; Moxey-Mims, M.; Dharnidharka, V.R.; Warady, B.A.; Furth, S.L.; Munoz, A. Improved equations estimating GFR in children with chronic kidney disease using an immunonephelometric determination of cystatin C. Kidney Int. 2012, 82, 445–453. [Google Scholar] [CrossRef]
- da Silva Selistre, L.; Rech, D.L.; de Souza, V.; Iwaz, J.; Lemoine, S.; Dubourg, L. Diagnostic Performance of Creatinine-Based Equations for Estimating Glomerular Filtration Rate in Adults 65 Years and Older. JAMA Intern. Med. 2019, 179, 796–804. [Google Scholar] [CrossRef]
- Ebert, N.; Loesment, A.; Martus, P.; Jakob, O.; Gaedeke, J.; Kuhlmann, M.; Bartel, J.; Schuchardt, M.; Tolle, M.; Huang, T.; et al. Iohexol plasma clearance measurement in older adults with chronic kidney disease-sampling time matters. Nephrol. Dial. Transplant. 2015, 30, 1307–1314. [Google Scholar] [CrossRef]
- Grewal, G.S.; Blake, G.M. Reference data for 51Cr-EDTA measurements of the glomerular filtration rate derived from live kidney donors. Nucl. Med. Commun. 2005, 26, 61–65. [Google Scholar] [CrossRef]
- Soveri, I.; Berg, U.B.; Bjork, J.; Elinder, C.G.; Grubb, A.; Mejare, I.; Sterner, G.; Back, S.E.; Group, S.G.R. Measuring GFR: A systematic review. Am. J. Kidney Dis. 2014, 64, 411–424. [Google Scholar] [CrossRef] [PubMed]
- Thienpont, L.M.; Van Landuyt, K.G.; Stockl, D.; De Leenheer, A.P. Candidate reference method for determining serum creatinine by isocratic HPLC: Validation with isotope dilution gas chromatography-mass spectrometry and application for accuracy assessment of routine test kits. Clin. Chem. 1995, 41, 995–1003. [Google Scholar] [CrossRef] [PubMed]
- Aguilar, J.A.; Nilsson, M.; Bodenhausen, G.; Morris, G.A. Spin echo NMR spectra without J modulation. Chem. Commun. 2012, 48, 811–813. [Google Scholar] [CrossRef]
- R Core Team. R Foundation for Statistical Computing R: A Language and Environment for Statistical Computing. (v3.5.1). 2017. Available online: https://R-project.org (accessed on 17 May 2020).
- Mackensen-Haen, S.; Bader, R.; Grund, K.E.; Bohle, A. Correlations between renal cortical interstitial fibrosis, atrophy of the proximal tubules and impairment of the glomerular filtration rate. Clin. Nephrol. 1981, 15, 167–171. [Google Scholar] [PubMed]
- Snauwaert, E.; Van Biesen, W.; Raes, A.; Holvoet, E.; Glorieux, G.; Van Hoeck, K.; Van Dyck, M.; Godefroid, N.; Vanholder, R.; Roels, S.; et al. Accumulation of uraemic toxins is reflected only partially by estimated GFR in paediatric patients with chronic kidney disease. Pediatric Nephrol. 2018, 33, 315–323. [Google Scholar] [CrossRef]
- Chen, Y.; Zelnick, L.R.; Wang, K.; Hoofnagle, A.N.; Becker, J.O.; Hsu, C.Y.; Feldman, H.I.; Mehta, R.C.; Lash, J.P.; Waikar, S.S.; et al. Kidney Clearance of Secretory Solutes Is Associated with Progression of CKD: The CRIC Study. J. Am. Soc. Nephrol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Freed, T.A.; Coresh, J.; Inker, L.A.; Toal, D.R.; Perichon, R.; Chen, J.; Goodman, K.D.; Zhang, Q.; Conner, J.K.; Hauser, D.M.; et al. Validation of a Metabolite Panel for a More Accurate Estimation of Glomerular Filtration Rate Using Quantitative LC-MS/MS. Clin. Chem. 2019, 65, 406–418. [Google Scholar] [CrossRef] [PubMed]
- Mendu, M.L.; Lundquist, A.; Aizer, A.A.; Leaf, D.E.; Robinson, E.; Steele, D.J.; Waikar, S.S. The usefulness of diagnostic testing in the initial evaluation of chronic kidney disease. JAMA Intern. Med. 2015, 175, 853–856. [Google Scholar] [CrossRef]
- Davies, D.F.; Shock, N.W. The variability of measurement of insulin and diodrast tests of kidney function. J. Clin. Investig. 1950, 29, 491–495. [Google Scholar] [CrossRef] [PubMed]
- Markley, J.L.; Bruschweiler, R.; Edison, A.S.; Eghbalnia, H.R.; Powers, R.; Raftery, D.; Wishart, D.S. The future of NMR-based metabolomics. Curr. Opin. Biotechnol. 2017, 43, 34–40. [Google Scholar] [CrossRef]
- Weir, M.R. Improving the estimating equation for GFR—A clinical perspective. N. Engl. J. Med. 2012, 367, 75–76. [Google Scholar] [CrossRef] [PubMed]
| Training Set | Test Set | |
|---|---|---|
| N | 95 | 189 |
| Age (years, range) | 4–76 | 3–88 |
| Age (years, mean ± SD) | 34 ± 23 a | 51 ± 24 a |
| Sex (% male) | 53 | 60 |
| mGFR | ||
| range | 5–147 | 8–178 |
| mean ± SD | 75 ± 35 | 71 ± 31 |
| iohexol | 54 | 101 |
| inulin | 22 | 60 |
| 51Cr-EDTA | 19 | 28 |
| CKD stage | ||
| 1 | 34 | 49 |
| 2 | 27 | 70 |
| 3 | 22 | 57 |
| 4 | 9 | 11 |
| 5 | 3 | 2 |
| Storage # time (years, range) | 0.4–13.4 | 0.4–13.3 |
| Storage # time (mean ± SD) | 2.5 ± 3.6 | 4.2 ± 3.9 |
| Precision | Linearity | Trueness | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Adult Low | Pediatric Low | Adult Normal | |||||||
| Mean (µmol/L) | CV (%) | Mean (µmol/L) | CV (%) | Mean (µmol/L) | CV (%) | Low (µmol/L) | High (µmol/L) | Pearson Correlation | |
| creatinine | 189.1 | 6.4 | 108.3 | 6.2 | 107.9 | 7.0 | 21 | 928 | 0.993 |
| dimethyl sulfone | 12.6 | 13.4 | 12.2 | 19.8 | 8.5 | 20.7 | 4 | 90 | 0.983 |
| myo-inositol | 110.5 | 11.1 | 78.5 | 11.2 | 68.5 | 14.2 | 39 | 441 | 0.991 |
| valine | 418.0 | 1.5 | 310.6 | 1.9 | 437.8 | 3.8 | 27 | 1250 | 0.998 |
| GFRNMR | 43.5 | 9.5 | 68.3 | 6.0 | 82.4 | 4.2 | n.a. | n.a. | 0.84 |
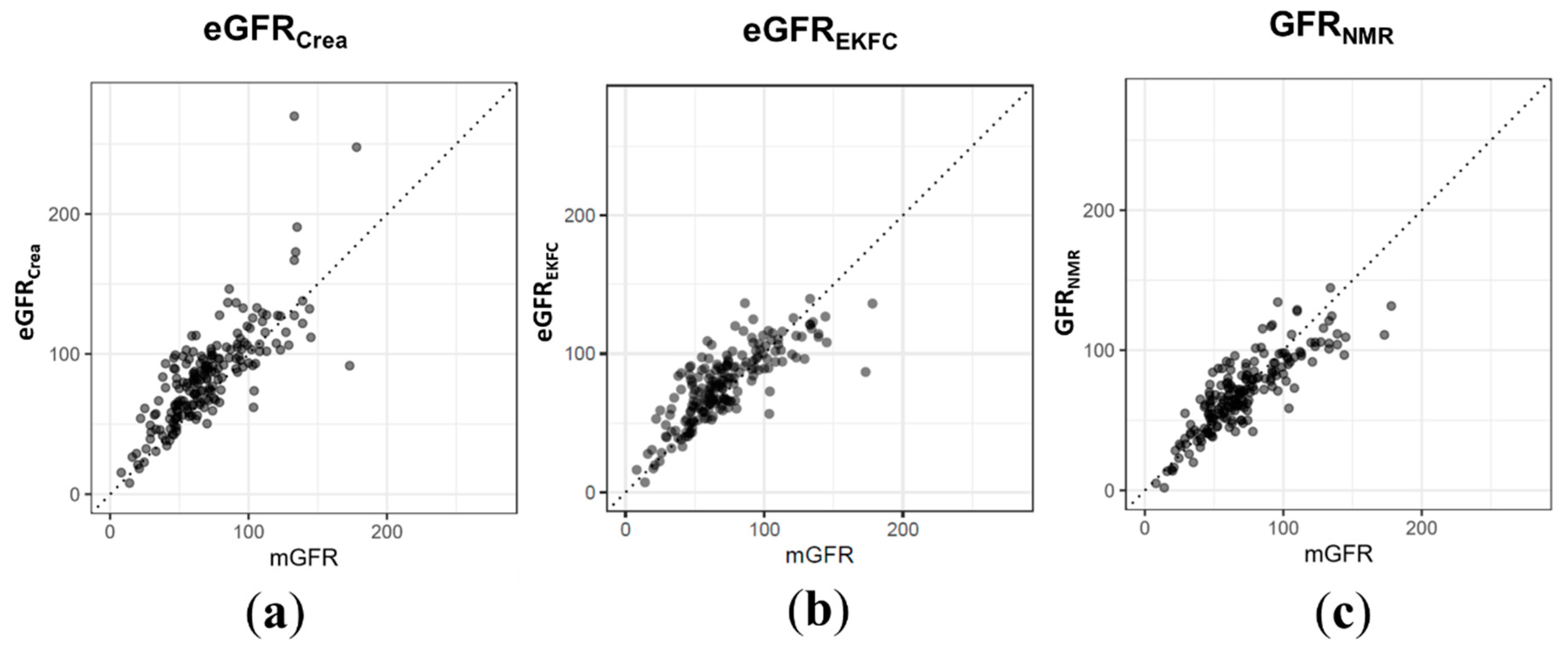
| eGFRCrea | eGFREKFC | GFRNMR | |
|---|---|---|---|
| RMSE | 25.30 | 19.61 | 16.51 |
| Pearson correlation | 0.79 (0.73–0.84) | 0.80 (0.74–0.84) | 0.84 (0.80–0.88) |
| P10 | 0.27 (0.20–0.35) | 0.37 (0.29–0.45) | 0.35 (0.27–0.43) |
| P15 | 0.40 (0.32–0.49) a | 0.50 (0.41–0.58) | 0.52 (0.44–0.60) a |
| P30 | 0.64 (0.56–0.72) b | 0.74 (0.66–0.81). | 0.81 (0.74–0.87) b |
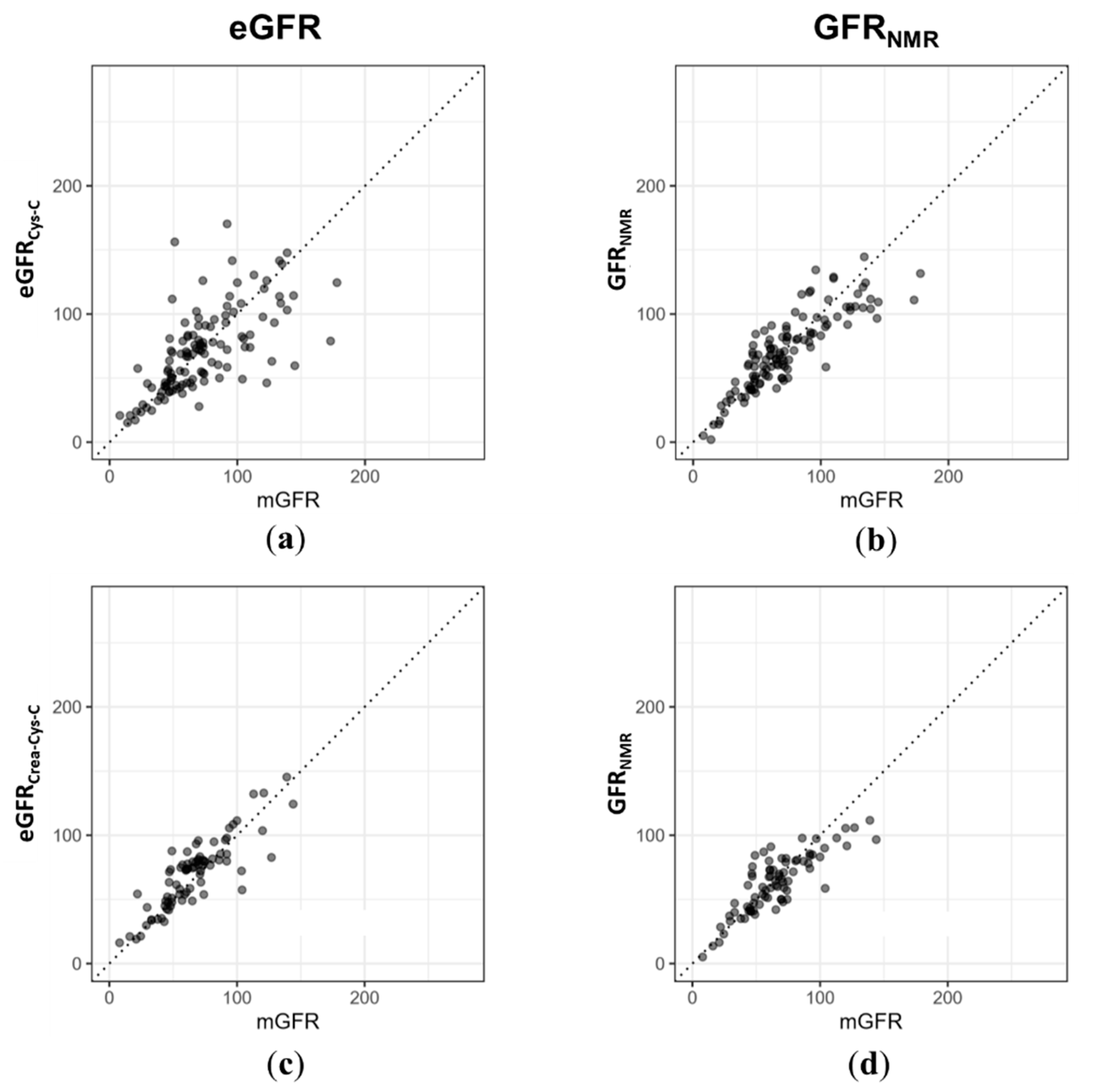
| (a) | eGFRCys-C * | GFRNMR | (b) | eGFRCrea-Cys-C ** | GFRNMR |
|---|---|---|---|---|---|
| n | 118 | 118 | n | 79 | 79 |
| RMSE | 27.59 | 17.60 | RMSE | 14.87 | 15.32 |
| Pearson correlation | 0.66 (0.54–0.75) | 0.86 (0.80–0.90) | Pearson correlation | 0.86 (0.79–0.91) | 0.83 (0.75–0.89) |
| P10 | 0.28 (0.19–0.38) | 0.31 (0.22–0.42) | P10 | 0.42 (0.29–0.55) | 0.37 (0.25–0.50) |
| P15 | 0.40 (0.30–0.51) | 0.52 (0.41–0.62) | P15 | 0.57 (0.44–0.69) | 0.58 (0.45–0.71) |
| P30 | 0.72 (0.62–0.81) | 0.81 (0.72–0.89) | P30 | 0.81 (0.69–0.90) | 0.81 (0.69–0.90) |
| Df | Sum Sq | Mean Sq | f Value | p Value | |
|---|---|---|---|---|---|
| GFRNMR | 1 | 126,206 | 126,206 | 464 | <0.0001 |
| Age | 1 | 518 | 518 | 1.90 | 0.17 |
| Sex | 1 | 400 | 400 | 1.47 | 0.23 |
| Residuals | 185 | 50,298 | 272 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ehrich, J.; Dubourg, L.; Hansson, S.; Pape, L.; Steinle, T.; Fruth, J.; Höckner, S.; Schiffer, E. Serum Myo-Inositol, Dimethyl Sulfone, and Valine in Combination with Creatinine Allow Accurate Assessment of Renal Insufficiency—A Proof of Concept. Diagnostics 2021, 11, 234. https://doi.org/10.3390/diagnostics11020234
Ehrich J, Dubourg L, Hansson S, Pape L, Steinle T, Fruth J, Höckner S, Schiffer E. Serum Myo-Inositol, Dimethyl Sulfone, and Valine in Combination with Creatinine Allow Accurate Assessment of Renal Insufficiency—A Proof of Concept. Diagnostics. 2021; 11(2):234. https://doi.org/10.3390/diagnostics11020234
Chicago/Turabian StyleEhrich, Jochen, Laurence Dubourg, Sverker Hansson, Lars Pape, Tobias Steinle, Jana Fruth, Sebastian Höckner, and Eric Schiffer. 2021. "Serum Myo-Inositol, Dimethyl Sulfone, and Valine in Combination with Creatinine Allow Accurate Assessment of Renal Insufficiency—A Proof of Concept" Diagnostics 11, no. 2: 234. https://doi.org/10.3390/diagnostics11020234
APA StyleEhrich, J., Dubourg, L., Hansson, S., Pape, L., Steinle, T., Fruth, J., Höckner, S., & Schiffer, E. (2021). Serum Myo-Inositol, Dimethyl Sulfone, and Valine in Combination with Creatinine Allow Accurate Assessment of Renal Insufficiency—A Proof of Concept. Diagnostics, 11(2), 234. https://doi.org/10.3390/diagnostics11020234





