A Novel Ferroptosis-Associated Gene Signature to Predict Prognosis in Patients with Uveal Melanoma
Abstract
1. Introduction
2. Materials and Methods
2.1. Cohorts and Ferroptosis-Related Genes
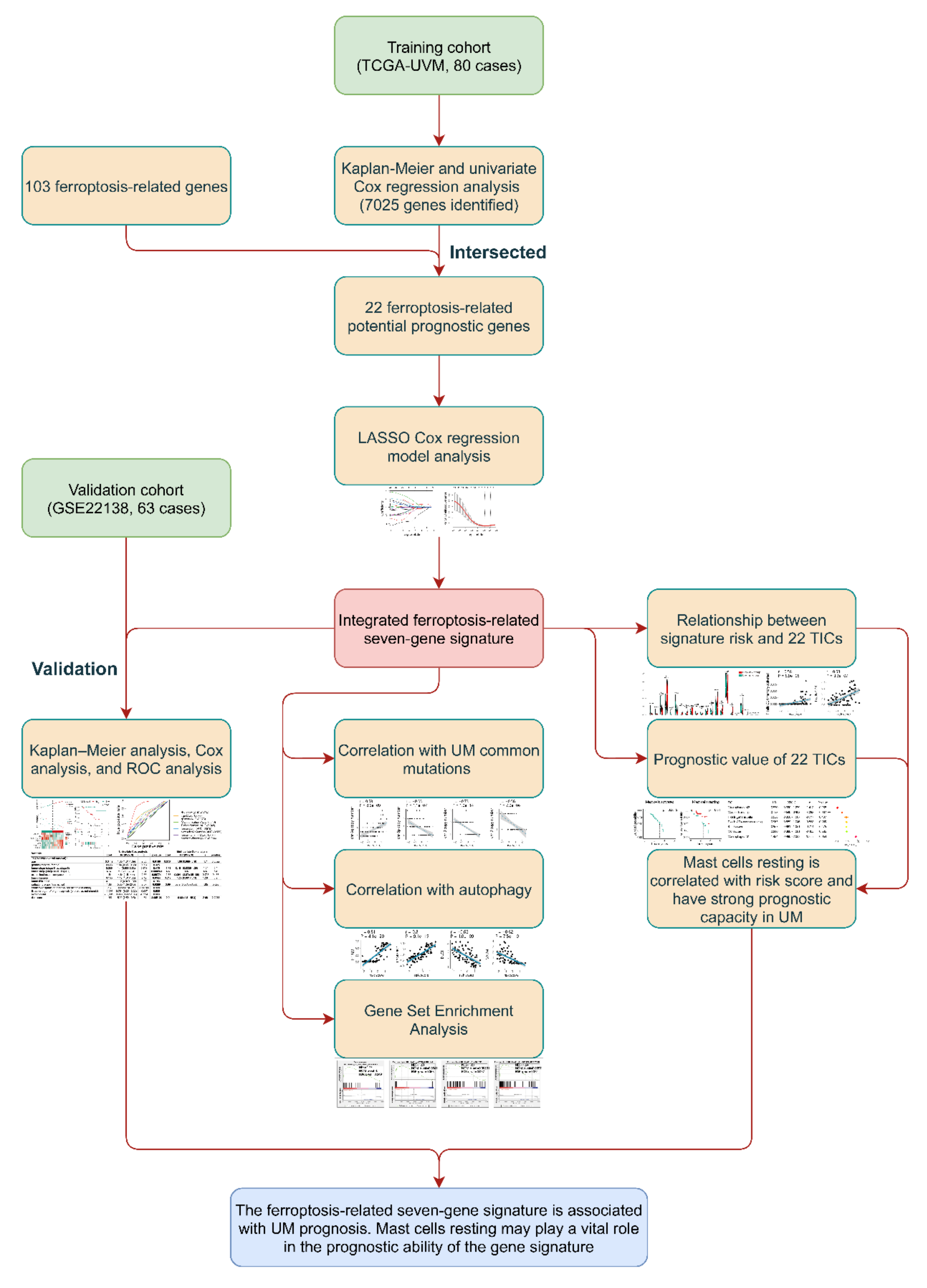
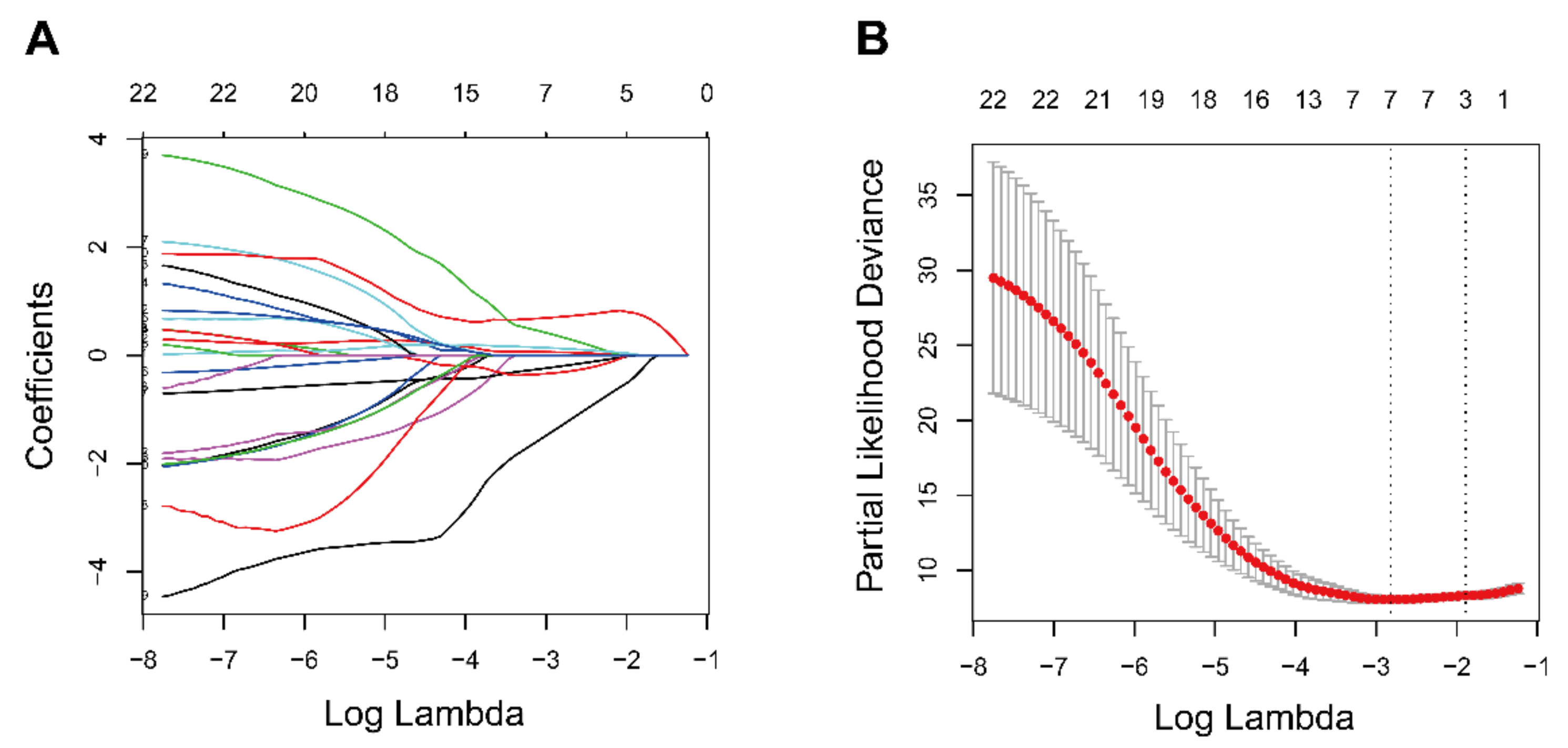
2.2. Identification and Validation of the Prognostic Ferroptosis-Related Gene Signature
2.3. Correlation between Gene Signature and UM Common Mutations
2.4. Relationships between Gene Signature and Autophagy in UM
2.5. Gene Set Enrichment Analysis
2.6. Relationship of Gene Signature and the 22 Tumor-Infiltrating Immune Cells (TICs)
2.7. Statistical Analysis
3. Results
3.1. Characteristics of UMs
3.2. Identification of Prognostic Ferroptosis-Related Gene Signature
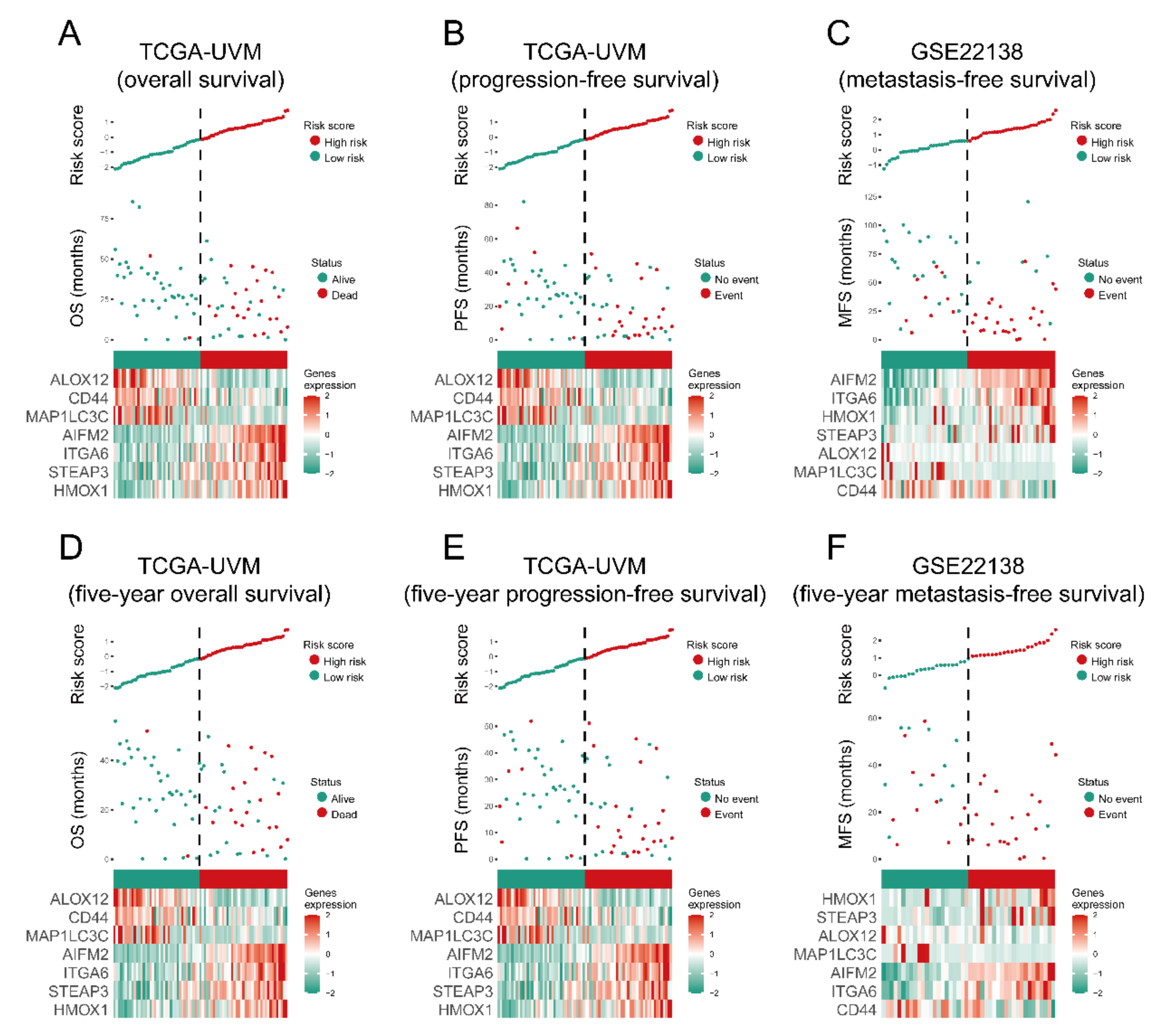
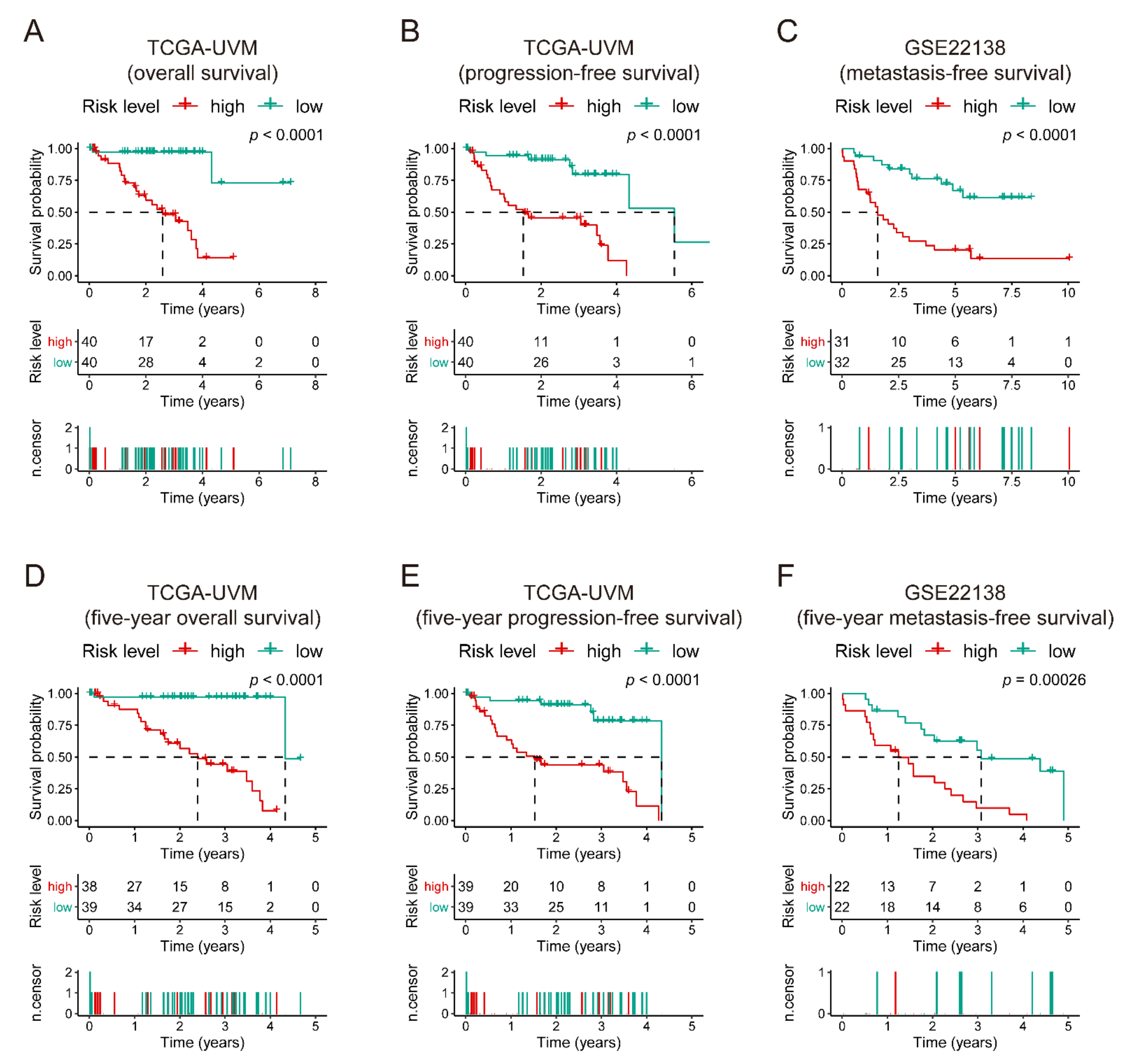
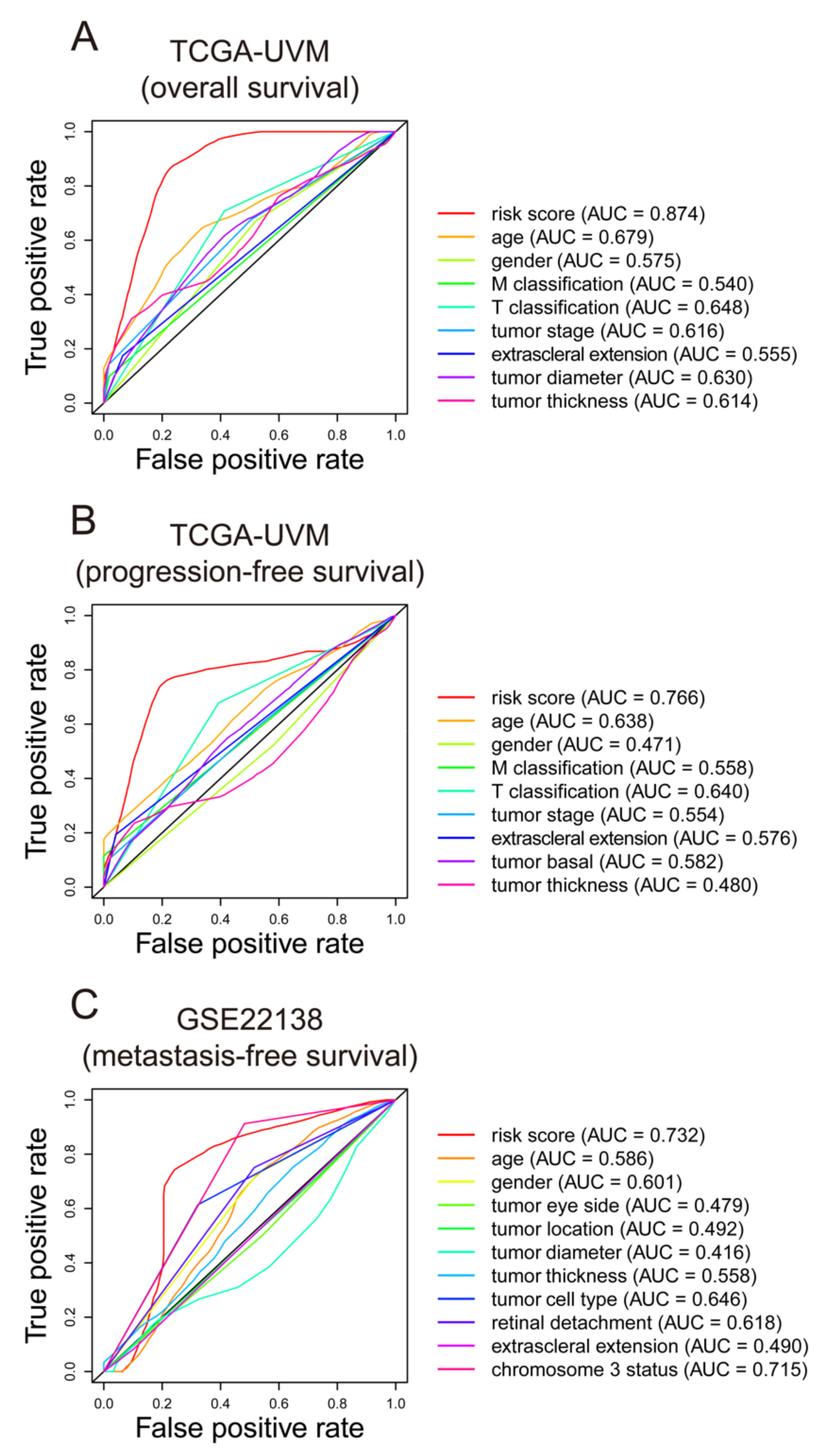
3.3. The Prognostic Capacity of the Seven-Gene Signature
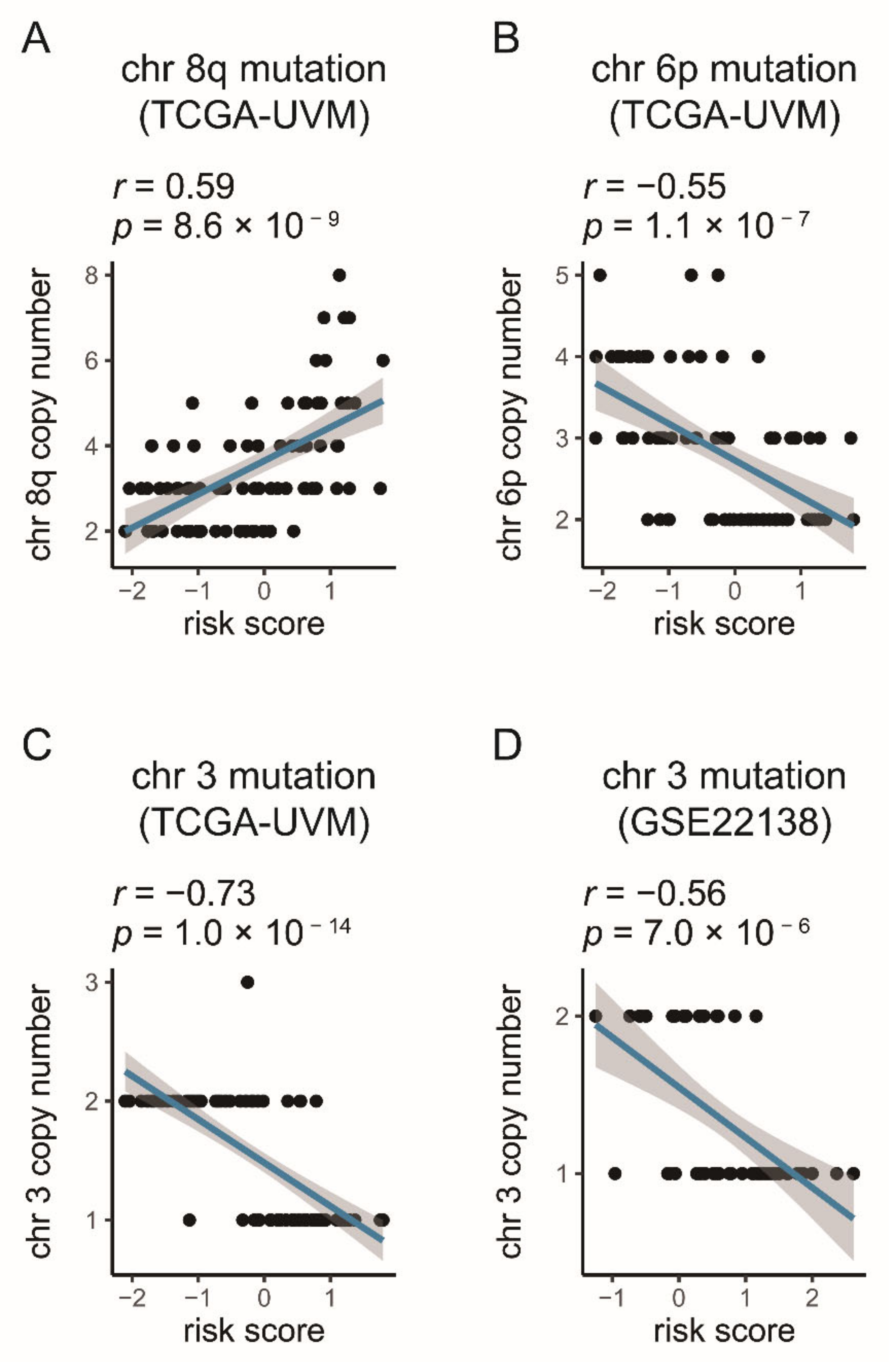
3.4. Identification of the Correlation between Seven-Gene Signature and UM Common Mutations
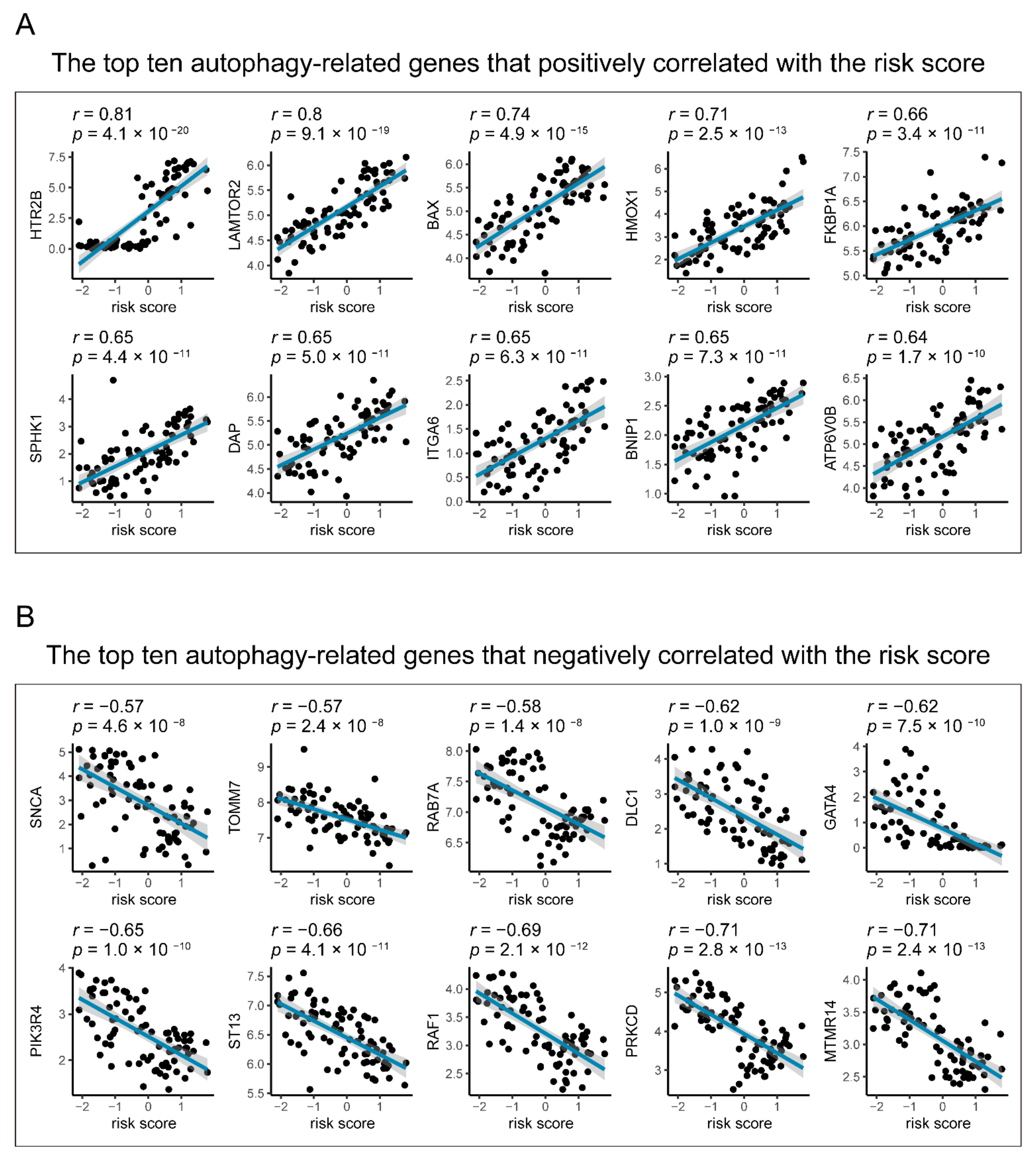
3.5. Identification of the Autophagy Correlation with the Seven-Gene Signature
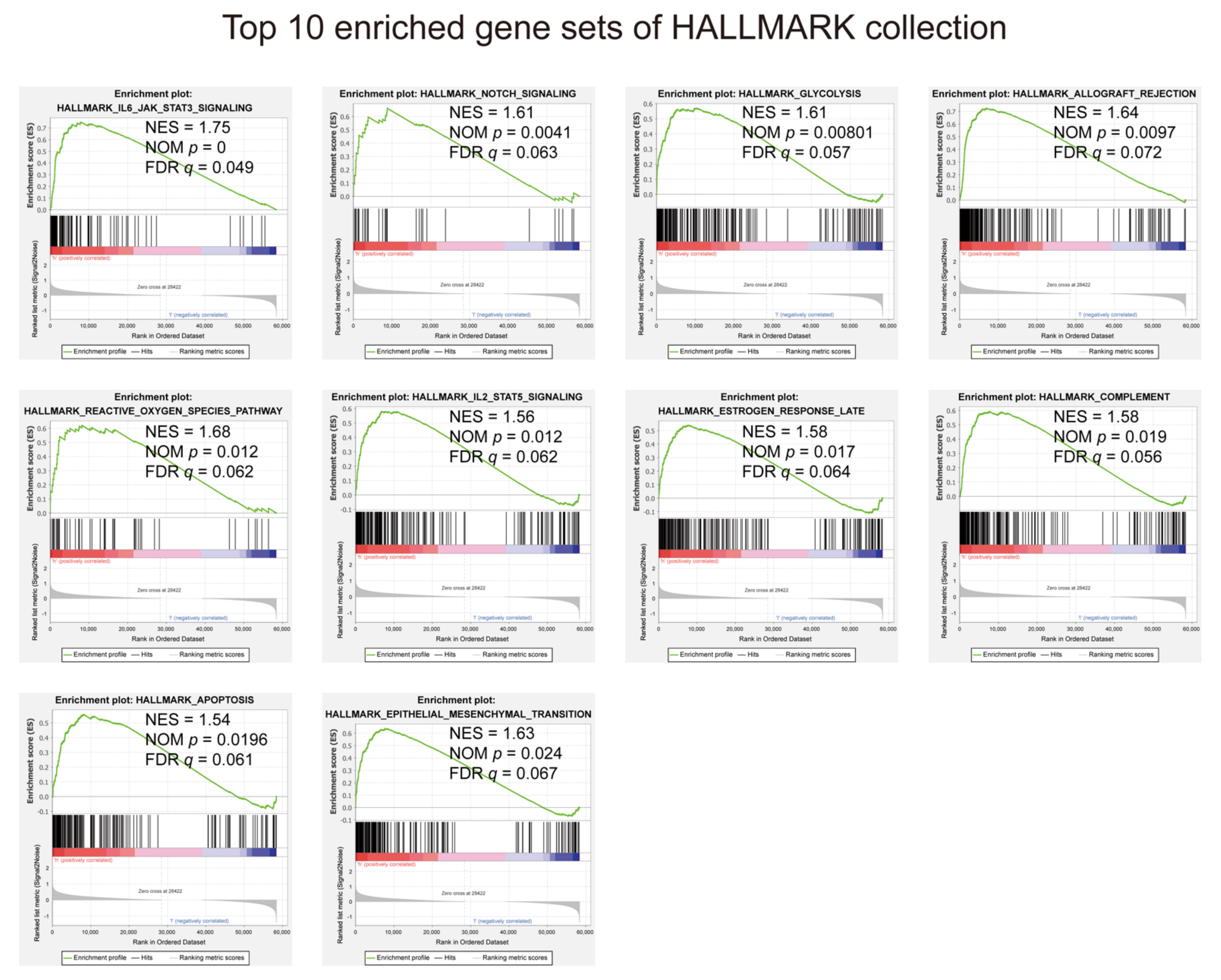
3.6. Gene Set Enrichment Analysis
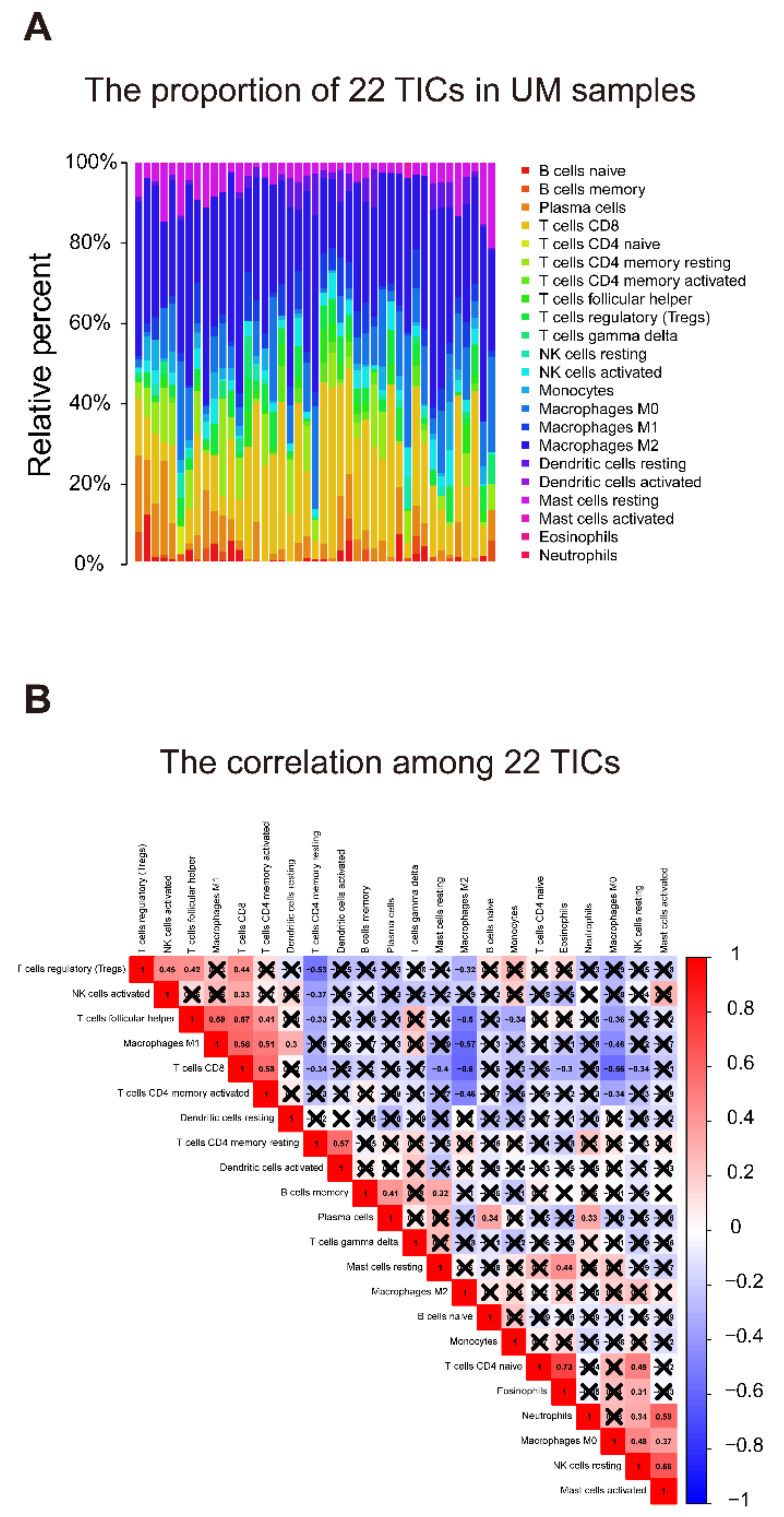
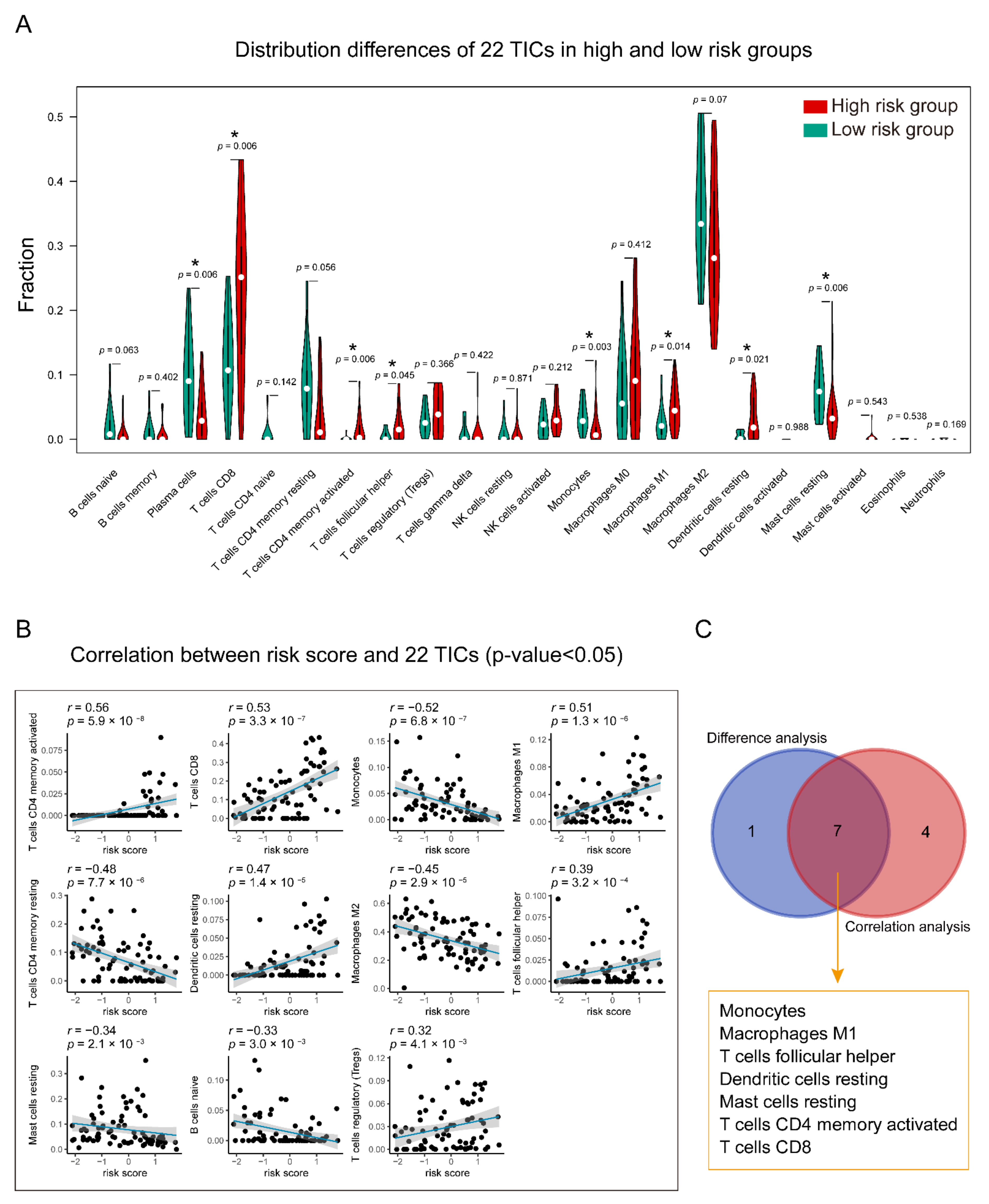
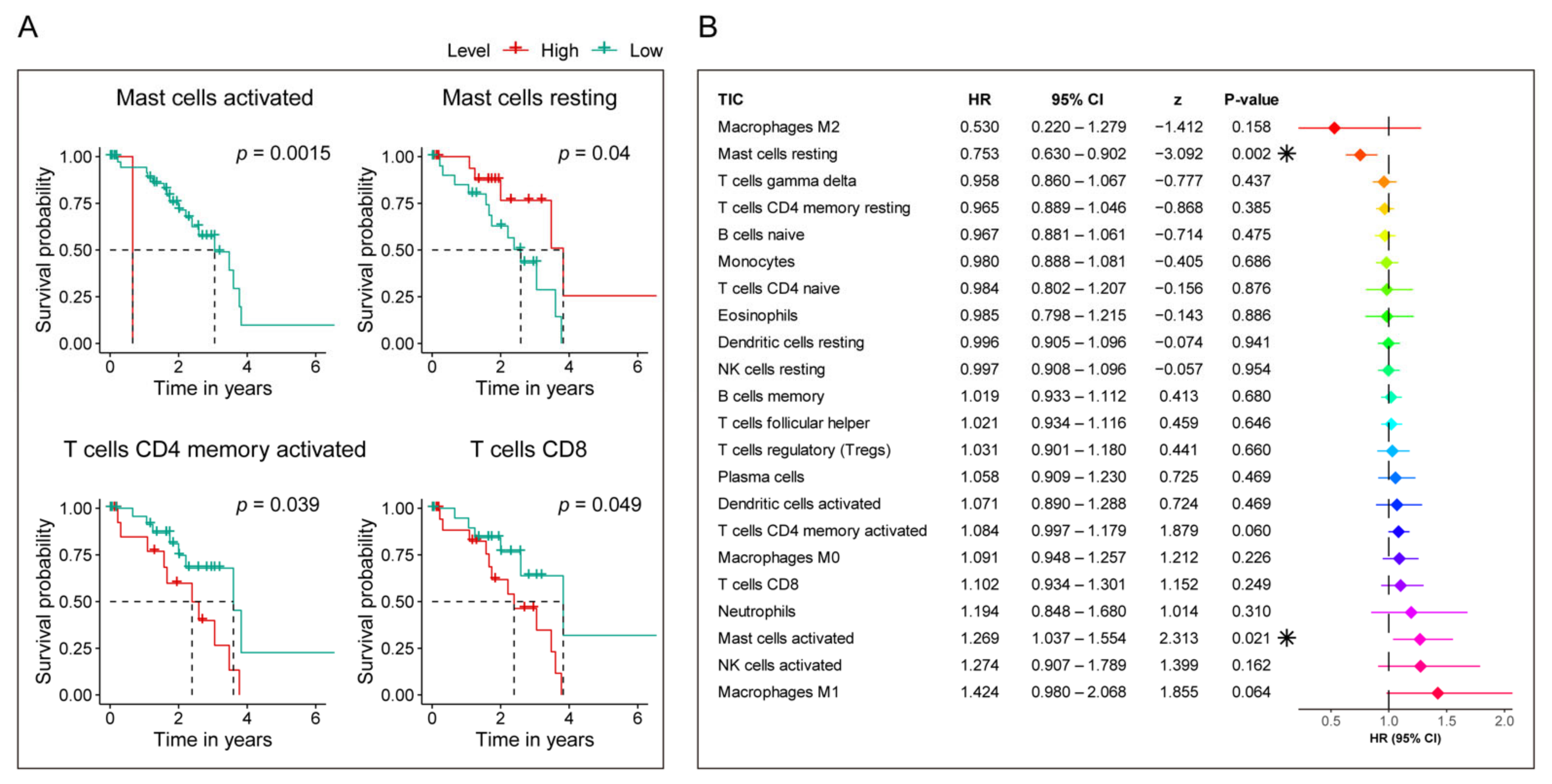
3.7. Identification of the Relationship between the Seven-Gene Signature and 22 TICs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Singh, A.D.; Turell, M.E.; Topham, A.K. Uveal melanoma: Trends in incidence, treatment, and survival. Ophthalmology 2011, 118, 1881–1885. [Google Scholar] [CrossRef]
- McLaughlin, C.C.; Wu, X.C.; Jemal, A.; Martin, H.J.; Roche, L.M.; Chen, V.W. Incidence of noncutaneous melanomas in the U.S. Cancer 2005, 103, 1000–1007. [Google Scholar] [CrossRef]
- Damato, B. Progress in the management of patients with uveal melanoma. The 2012 Ashton Lecture. Eye 2012, 26, 1157–1172. [Google Scholar] [CrossRef] [PubMed]
- Ponti, A.; Denys, A.; Digklia, A.; Schaefer, N.; Hocquelet, A.; Knebel, J.F.; Michielin, O.; Dromain, C.; Duran, R. First-Line Selective Internal Radiation Therapy in Patients with Uveal Melanoma Metastatic to the Liver. J. Nucl. Med. 2020, 61, 350–356. [Google Scholar] [CrossRef] [PubMed]
- Slater, K.; Hoo, P.S.; Buckley, A.M.; Piulats, J.M.; Villanueva, A.; Portela, A.; Kennedy, B.N. Evaluation of oncogenic cysteinyl leukotriene receptor 2 as a therapeutic target for uveal melanoma. Cancer Metastasis Rev. 2018, 37, 335–345. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Manson, D.K.; Marr, B.P.; Carvajal, R.D. Treatment of uveal melanoma: Where are we now? Ther. Adv. Med. Oncol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Cao, F.; Yin, H.L.; Huang, Z.J.; Lin, Z.T.; Mao, N.; Sun, B.; Wang, G. Ferroptosis: Past, present and future. Cell Death Dis. 2020, 11, 88. [Google Scholar] [CrossRef]
- Wu, Y.; Yu, C.; Luo, M.; Cen, C.; Qiu, J.; Zhang, S.; Hu, K. Ferroptosis in Cancer Treatment: Another Way to Rome. Front. Oncol. 2020. [Google Scholar] [CrossRef]
- Bebber, C.M.; Muller, F.; Prieto Clemente, L.; Weber, J.; von Karstedt, S. Ferroptosis in Cancer Cell Biology. Cancers 2020, 12, 164. [Google Scholar] [CrossRef]
- Liang, C.; Zhang, X.; Yang, M.; Dong, X. Recent Progress in Ferroptosis Inducers for Cancer Therapy. Adv. Mater. 2019, 31, e1904197. [Google Scholar] [CrossRef]
- Hassannia, B.; Vandenabeele, P.; Vanden Berghe, T. Targeting Ferroptosis to Iron Out Cancer. Cancer Cell 2019, 35, 830–849. [Google Scholar] [CrossRef] [PubMed]
- Jiang, M.; Qiao, M.; Zhao, C.; Deng, J.; Li, X.; Zhou, C. Targeting ferroptosis for cancer therapy: Exploring novel strategies from its mechanisms and role in cancers. Transl. Lung Cancer Res. 2020, 9, 1569–1584. [Google Scholar] [CrossRef]
- Ubellacker, J.M.; Tasdogan, A.; Ramesh, V.; Shen, B.; Mitchell, E.C.; Martin-Sandoval, M.S.; Gu, Z.; McCormick, M.L.; Durham, A.B.; Spitz, D.R.; et al. Lymph protects metastasizing melanoma cells from ferroptosis. Nature 2020, 585, 113–118. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.J.; Hu, H.M.; Li, G.Z.; Zhang, Y.; Wu, F.; Liu, X.; Wang, K.Y.; Zhang, C.B.; Jiang, T. Ferroptosis-Related Gene Signature Predicts Glioma Cell Death and Glioma Patient Progression. Front. Cell Dev. Biol. 2020, 8, 538. [Google Scholar] [CrossRef] [PubMed]
- Liang, J.Y.; Wang, D.S.; Lin, H.C.; Chen, X.X.; Yang, H.; Zheng, Y.; Li, Y.H. A Novel Ferroptosis-related Gene Signature for Overall Survival Prediction in Patients with Hepatocellular Carcinoma. Int. J. Biol. Sci. 2020, 16, 2430–2441. [Google Scholar] [CrossRef]
- Stelzer, G.; Rosen, N.; Plaschkes, I.; Zimmerman, S.; Twik, M.; Fishilevich, S.; Stein, T.I.; Nudel, R.; Lieder, I.; Mazor, Y.; et al. The GeneCards Suite: From Gene Data Mining to Disease Genome Sequence Analyses. Curr. Protoc. Bioinform. 2016, 54, 1–30. [Google Scholar] [CrossRef] [PubMed]
- Friedman, J.; Hastie, T.; Tibshirani, R. Regularization Paths for Generalized Linear Models via Coordinate Descent. J. Stat. Softw. 2010, 33, 1–22. [Google Scholar] [PubMed]
- Goeman, J.J. L1 penalized estimation in the Cox proportional hazards model. Biom. J. 2010, 52, 70–84. [Google Scholar] [PubMed]
- Sauerbrei, W.; Royston, P.; Binder, H. Selection of important variables and determination of functional form for continuous predictors in multivariable model building. Stat. Med. 2007, 26, 5512–5528. [Google Scholar] [CrossRef] [PubMed]
- Tibshirani, R. The lasso method for variable selection in the Cox model. Stat. Med. 1997, 16, 385–395. [Google Scholar] [CrossRef]
- Robertson, A.G.; Shih, J.; Yau, C.; Gibb, E.A.; Oba, J.; Mungall, K.L.; Hess, J.M.; Uzunangelov, V.; Walter, V.; Danilova, L.; et al. Integrative Analysis Identifies Four Molecular and Clinical Subsets in Uveal Melanoma. Cancer Cell 2017, 32, 204–220. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Shen, Y.; Chen, C.; Sui, X.; Yang, J.; Wang, L.; Zhou, J. The crosstalk between autophagy and ferroptosis: What can we learn to target drug resistance in cancer? Cancer Biol. Med. 2019, 16, 630–646. [Google Scholar]
- Abdul Rahim, S.A.; Dirkse, A.; Oudin, A.; Schuster, A.; Bohler, J.; Barthelemy, V.; Muller, A.; Vallar, L.; Janji, B.; Golebiewska, A.; et al. Regulation of hypoxia-induced autophagy in glioblastoma involves ATG9A. Br. J. Cancer. 2017, 117, 813–825. [Google Scholar] [CrossRef] [PubMed]
- Thorsson, V.; Gibbs, D.L.; Brown, S.D.; Wolf, D.; Bortone, D.S.; Ou Yang, T.H.; Porta-Pardo, E.; Gao, G.F.; Plaisier, C.L.; Eddy, J.A.; et al. The Immune Landscape of Cancer. Immunity 2018, 48, 812–830. [Google Scholar] [CrossRef]
- Newman, A.M.; Steen, C.B.; Liu, C.L.; Gentles, A.J.; Chaudhuri, A.A.; Scherer, F.; Khodadoust, M.S.; Esfahani, M.S.; Luca, B.A.; Steiner, D.; et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 2019, 37, 773–782. [Google Scholar] [CrossRef] [PubMed]
- Song, X.; Xie, Y.; Kang, R.; Hou, W.; Sun, X.; Epperly, M.W.; Greenberger, J.S.; Tang, D. FANCD2 protects against bone marrow injury from ferroptosis. Biochem. Biophys. Res. Commun. 2016, 480, 443–449. [Google Scholar] [CrossRef]
- Kang, R.; Zhu, S.; Zeh, H.J.; Klionsky, D.J.; Tang, D. BECN1 is a new driver of ferroptosis. Autophagy 2018, 14, 2173–2175. [Google Scholar] [CrossRef]
- Brown, C.W.; Amante, J.J.; Goel, H.L.; Mercurio, A.M. The alpha6beta4 integrin promotes resistance to ferroptosis. J. Cell Biol. 2017, 216, 4287–4297. [Google Scholar] [CrossRef]
- Chang, L.C.; Chiang, S.K.; Chen, S.E.; Yu, Y.L.; Chou, R.H.; Chang, W.C. Heme oxygenase-1 mediates BAY 11-7085 induced ferroptosis. Cancer Lett. 2018, 416, 124–137. [Google Scholar] [CrossRef]
- Kwon, M.Y.; Park, E.; Lee, S.J.; Chung, S.W. Heme oxygenase-1 accelerates erastin-induced ferroptotic cell death. Oncotarget 2015, 6, 24393–24403. [Google Scholar] [CrossRef]
- Su, L.; Jiang, X.; Yang, C.; Zhang, J.; Chen, B.; Li, Y.; Yao, S.; Xie, Q.; Gomez, H.; Murugan, R.; et al. Pannexin 1 mediates ferroptosis that contributes to renal ischemia/reperfusion injury. J. Biol. Chem. 2019, 294, 19395–19404. [Google Scholar] [CrossRef] [PubMed]
- Lu, B.; Chen, X.-B.; Hong, Y.-C.; Zhu, H.; He, Q.-J.; Yang, B.; Ying, M.-D.; Cao, J. Identification of PRDX6 as a regulator of ferroptosis. Acta Pharmacol. Sin. 2019, 40, 1334–1342. [Google Scholar] [CrossRef] [PubMed]
- Adedoyin, O.; Boddu, R.; Traylor, A.M.; Lever, J.M.; Bolisetty, S.; George, J.F.; Agarwal, A. Heme oxygenase-1 mitigates ferroptosis in renal proximal tubule cells. Am. J. Physiol. Ren. Physiol. 2018, 314, F702–F714. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Ou, Z.; Chen, R.; Niu, X.; Chen, D.; Kang, R.; Tang, D. Activation of the p62-Keap1-NRF2 pathway protects against ferroptosis in hepatocellular carcinoma cells. Hepatology 2016, 63, 173–184. [Google Scholar] [CrossRef] [PubMed]
- Eling, N.; Reuter, L.; Hazin, J.; Hamacher-Brady, A.; Brady, N.R. Identification of artesunate as a specific activator of ferroptosis in pancreatic cancer cells. Oncoscience 2015, 2, 517–532. [Google Scholar] [CrossRef] [PubMed]
- Fang, X.; Wang, H.; Han, D.; Xie, E.; Yang, X.; Wei, J.; Gu, S.; Gao, F.; Zhu, N.; Yin, X.; et al. Ferroptosis as a target for protection against cardiomyopathy. Proc. Natl. Acad. Sci. USA 2019, 116, 2672–2680. [Google Scholar] [CrossRef]
- Liu, T.; Jiang, L.; Tavana, O.; Gu, W. The Deubiquitylase OTUB1 Mediates Ferroptosis via Stabilization of SLC7A11. Cancer Res. 2019, 79, 1913–1924. [Google Scholar] [CrossRef]
- Chu, B.; Kon, N.; Chen, D.; Li, T.; Liu, T.; Jiang, L.; Song, S.; Tavana, O.; Gu, W. ALOX12 is required for p53-mediated tumour suppression through a distinct ferroptosis pathway. Nat. Cell Biol. 2019, 21, 579–591. [Google Scholar] [CrossRef]
- Shintoku, R.; Takigawa, Y.; Yamada, K.; Kubota, C.; Yoshimoto, Y.; Takeuchi, T.; Koshiishi, I.; Torii, S. Lipoxygenase-mediated generation of lipid peroxides enhances ferroptosis induced by erastin and RSL3. Cancer Sci. 2017, 108, 2187–2194. [Google Scholar] [CrossRef]
- Yang, W.S.; Kim, K.J.; Gaschler, M.M.; Patel, M.; Shchepinov, M.S.; Stockwell, B.R. Peroxidation of polyunsaturated fatty acids by lipoxygenases drives ferroptosis. Proc. Natl. Acad. Sci. USA 2016, 113, E4966–E4975. [Google Scholar] [CrossRef]
- Shah, R.; Shchepinov, M.S.; Pratt, D.A. Resolving the Role of Lipoxygenases in the Initiation and Execution of Ferroptosis. ACS Cent. Sci. 2018, 4, 387–396. [Google Scholar] [CrossRef] [PubMed]
- Bersuker, K.; Hendricks, J.M.; Li, Z.; Magtanong, L.; Ford, B.; Tang, P.H.; Roberts, M.A.; Tong, B.; Maimone, T.J.; Zoncu, R.; et al. The CoQ oxidoreductase FSP1 acts parallel to GPX4 to inhibit ferroptosis. Nature 2019, 575, 688–692. [Google Scholar] [CrossRef] [PubMed]
- Doll, S.; Freitas, F.P.; Shah, R.; Aldrovandi, M.; Da Silva, M.C.; Ingold, I.; Grocin, A.G.; Da Silva, T.N.X.; Panzilius, E.; Scheel, C.H.; et al. FSP1 is a glutathione-independent ferroptosis suppressor. Nature 2019, 575, 693–698. [Google Scholar] [CrossRef] [PubMed]
- Tao, N.; Li, K.; Liu, J. Molecular Mechanisms of Ferroptosis and Its Role in Pulmonary Disease. Oxid. Med. Cell Longev. 2020, 2020, 9547127. [Google Scholar] [CrossRef] [PubMed]
- Mou, Y.; Wang, J.; Wu, J.; He, D.; Zhang, C.; Duan, C.; Li, B. Ferroptosis, a new form of cell death: Opportunities and challenges in cancer. J. Hematol. Oncol. 2019, 12, 34. [Google Scholar] [CrossRef] [PubMed]
- Ashrafizadeh, M.; Mohammadinejad, R.; Tavakol, S.; Ahmadi, Z.; Roomiani, S.; Katebi, M. Autophagy, anoikis, ferroptosis, necroptosis, and endoplasmic reticulum stress: Potential applications in melanoma therapy. J. Cell Physiol. 2019, 234, 19471–19479. [Google Scholar] [CrossRef]
- Zhang, K.; Wu, L.; Zhang, P.; Luo, M.; Du, J.; Gao, T.; O’Connell, D.; Wang, G.; Wang, H.; Yang, Y. MiR-9 regulates ferroptosis by targeting glutamic-oxaloacetic transaminase GOT1 in melanoma. Mol. Carcinog. 2018, 57, 1566–1576. [Google Scholar] [CrossRef]
- Luo, M.; Wu, L.; Zhang, K.; Wang, H.; Zhang, T.; Gutierrez, L.; O’Connell, D.; Zhang, P.; Li, Y.; Gao, T.; et al. MiR-137 regulates ferroptosis by targeting glutamine transporter SLC1A5 in melanoma. Cell Death Differ. 2018, 25, 1457–1472. [Google Scholar] [CrossRef]
- Basit, F.; van Oppen, L.M.; Schockel, L.; Bossenbroek, H.M.; van Emst-de Vries, S.E.; Hermeling, J.C.; Grefte, S.; Kopitz, C.; Heroult, M.; Willems, P.H.; et al. Mitochondrial complex I inhibition triggers a mitophagy-dependent ROS increase leading to necroptosis and ferroptosis in melanoma cells. Cell Death Dis. 2017, 8, e2716. [Google Scholar] [CrossRef]
- Zheng, Z.; Li, Y.; Jin, G.; Huang, T.; Zou, M.; Duan, S. The biological role of arachidonic acid 12-lipoxygenase (ALOX12) in various human diseases. Biomed. Pharmacother. 2020, 129, 110354. [Google Scholar] [CrossRef]
- Gil, J.; Ramsey, D.; Pawlowski, P.; Szmida, E.; Leszczynski, P.; Bebenek, M.; Sasiadek, M.M. The Influence of Tumor Microenvironment on ATG4D Gene Expression in Colorectal Cancer Patients. Med. Oncol. 2018, 35, 159. [Google Scholar] [CrossRef]
- Dai, E.; Zhang, W.; Cong, D.; Kang, R.; Wang, J.; Tang, D. AIFM2 blocks ferroptosis independent of ubiquinol metabolism. Biochem. Biophys. Res. Commun. 2020, 523, 966–971. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.-H.; Yang, Z.-L.; Zhou, E.-X.; Miao, X.-Y.; Zou, Q.; Li, J.-H.; Liang, L.-F.; Zeng, G.-X.; Chen, S.-L. Overexpression of Thy1 and ITGA6 is associated with invasion, metastasis and poor prognosis in human gallbladder carcinoma. Oncol. Lett. 2016, 12, 5136–5144. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Brooks, D.L.P.; Schwab, L.P.; Krutilina, R.; Parke, D.N.; Sethuraman, A.; Hoogewijs, D.; Schörg, A.; Gotwald, L.; Fan, M.; Wenger, R.H.; et al. ITGA6 is directly regulated by hypoxia-inducible factors and enriches for cancer stem cell activity and invasion in metastatic breast cancer models. Mol. Cancer 2016, 15, 26. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.-Y.; Nam, M.; Son, H.Y.; Hyun, K.; Jang, S.Y.; Kim, J.W.; Kim, M.W.; Jung, Y.; Jang, E.; Yoon, S.-J.; et al. Polyunsaturated fatty acid biosynthesis pathway determines ferroptosis sensitivity in gastric cancer. Proc. Natl. Acad. Sci. USA 2020, 117, 32433–32442. [Google Scholar] [CrossRef] [PubMed]
- Hangauer, M.J.; Viswanathan, V.S.; Ryan, M.J.; Bole, D.; Eaton, J.K.; Matov, A.; Galeas, J.; Dhruv, H.D.; Berens, M.E.; Schreiber, S.L.; et al. Drug-tolerant persister cancer cells are vulnerable to GPX4 inhibition. Nature 2017, 551, 247–250. [Google Scholar] [CrossRef] [PubMed]
- Viswanathan, V.S.; Ryan, M.J.; Dhruv, H.D.; Gill, S.; Eichhoff, O.M.; Seashore-Ludlow, B.; Kaffenberger, S.D.; Eaton, J.K.; Shimada, K.; Aguirre, A.J.; et al. Dependency of a therapy-resistant state of cancer cells on a lipid peroxidase pathway. Nature 2017, 547, 453–457. [Google Scholar] [CrossRef]
- Ahrens, T.; Sleeman, J.P.; Schempp, C.M.; Howells, N.; Hofmann, M.; Ponta, H.; Herrlich, P.; Simon, J.C. Soluble CD44 inhibits melanoma tumor growth by blocking cell surface CD44 binding to hyaluronic acid. Oncogene 2001, 20, 3399–3408. [Google Scholar] [CrossRef]
- Louderbough, J.M.; Schroeder, J.A. Understanding the dual nature of CD44 in breast cancer progression. Mol. Cancer Res. 2011, 9, 1573–1586. [Google Scholar] [CrossRef]
- Gomes, I.M.; Maia, C.J.; Santos, C.R. STEAP proteins: From structure to applications in cancer therapy. Mol. Cancer Res. 2012, 10, 573–587. [Google Scholar] [CrossRef]
- Hassannia, B.; Wiernicki, B.; Ingold, I.; Qu, F.; Van Herck, S.; Tyurina, Y.Y.; Bayır, H.; Abhari, B.A.; Angeli, J.P.F.; Choi, S.M.; et al. Nano-targeted induction of dual ferroptotic mechanisms eradicates high-risk neuroblastoma. J. Clin. Investig. 2018, 128, 3341–3355. [Google Scholar] [CrossRef] [PubMed]
- Scholes, A.G.; Damato, B.E.; Nunn, J.; Hiscott, P.; Grierson, I.; Field, J.K. Monosomy 3 in uveal melanoma: Correlation with clinical and histologic predictors of survival. Invest. Ophthalmol. Vis. Sci. 2003, 44, 1008–1011. [Google Scholar] [CrossRef] [PubMed]
- Sisley, K.; Rennie, I.G.; Parsons, M.A.; Jacques, R.; Hammond, D.W.; Bell, S.M.; Potter, A.M.; Rees, R.C. Abnormalities of chromosomes 3 and 8 in posterior uveal melanoma correlate with prognosis. Genes Chromosomes Cancer 1997, 19, 22–28. [Google Scholar] [CrossRef]
- Versluis, M.; de Lange, M.J.; van Pelt, S.I.; Ruivenkamp, C.A.; Kroes, W.G.; Cao, J.; Jager, M.J.; Luyten, G.P.; van der Velden, P.A. Digital PCR validates 8q dosage as prognostic tool in uveal melanoma. PLoS ONE 2015, 10, e0116371. [Google Scholar] [CrossRef] [PubMed]
- Caines, R.; Eleuteri, A.; Kalirai, H.; Fisher, A.C.; Heimann, H.; Damato, B.E.; Coupland, S.E.; Taktak, A.F. Cluster analysis of multiplex ligation-dependent probe amplification data in choroidal melanoma. Mol. Vis. 2015, 21, 1–11. [Google Scholar]
- Cassoux, N.; Rodrigues, M.J.; Plancher, C.; Asselain, B.; Levy-Gabriel, C.; Rouic, L.L.-L.; Piperno-Neumann, S.; Dendale, R.; Sastre, X.; Desjardins, L.; et al. Genome-wide profiling is a clinically relevant and affordable prognostic test in posterior uveal melanoma. Br. J. Ophthalmol. 2014, 98, 769–774. [Google Scholar] [CrossRef]
- Damato, B.; Dopierala, J.A.; Coupland, S.E. Genotypic profiling of 452 choroidal melanomas with multiplex ligation-dependent probe amplification. Clin. Cancer Res. 2010, 16, 6083–6092. [Google Scholar] [CrossRef]
- Onken, M.D.; Worley, L.A.; Harbour, J.W. A metastasis modifier locus on human chromosome 8p in uveal melanoma identified by integrative genomic analysis. Clin. Cancer Res. 2008, 14, 3737–3745. [Google Scholar] [CrossRef]
- White, V.A.; Chambers, J.D.; Courtright, P.D.; Chang, W.Y.; Horsman, D.E. Correlation of cytogenetic abnormalities with the outcome of patients with uveal melanoma. Cancer 1998, 83, 354–359. [Google Scholar] [CrossRef]
- Mizushima, N.; Komatsu, M. Autophagy: Renovation of cells and tissues. Cell 2011, 147, 728–741. [Google Scholar] [CrossRef]
- Kang, R.; Tang, D. Autophagy and Ferroptosis—What’s the Connection? Curr. Pathobiol. Rep. 2017, 5, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Ndoye, A.; Weeraratna, A.T. Autophagy—An emerging target for melanoma therapy. F1000Research 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Giatromanolaki, A.N.; Charitoudis, G.; Bechrakis, E.N.; Kozobolis, V.P.; Koukourakis, I.M.; Foerster, M.H.; Sivridis, E.L. Autophagy patterns and prognosis in uveal melanomas. Mod. Pathol. 2011, 24, 1036–1045. [Google Scholar] [CrossRef] [PubMed]
- Johnson, D.E.; O’Keefe, R.A.; Grandis, J.R. Targeting the IL-6/JAK/STAT3 signalling axis in cancer. Nat. Rev. Clin. Oncol. 2018, 15, 234–248. [Google Scholar] [CrossRef]
- Terai, M.; Eto, M.; Young, G.D.; Berd, D.; Mastrangelo, M.J.; Tamura, Y.; Harigaya, K.; Sato, T. Interleukin 6 mediates production of interleukin 10 in metastatic melanoma. Cancer Immunol. Immunother. 2012, 61, 145–155. [Google Scholar] [CrossRef]
- Balint, K.; Xiao, M.; Pinnix, C.C.; Soma, A.; Veres, I.; Juhasz, I.; Brown, E.J.; Capobianco, A.J.; Herlyn, M.; Liu, Z.J. Activation of Notch1 signaling is required for beta-catenin-mediated human primary melanoma progression. J. Clin. Invest. 2005, 115, 3166–3176. [Google Scholar] [CrossRef]
- Bedogni, B. Notch signaling in melanoma: Interacting pathways and stromal influences that enhance Notch targeting. Pigment. Cell Melanoma Res. 2014, 27, 162–168. [Google Scholar] [CrossRef]
- Liu, Z.J.; Xiao, M.; Balint, K.; Smalley, K.S.; Brafford, P.; Qiu, R.; Pinnix, C.C.; Li, X.; Herlyn, M. Notch1 signaling promotes primary melanoma progression by activating mitogen-activated protein kinase/phosphatidylinositol 3-kinase-Akt pathways and up-regulating N-cadherin expression. Cancer Res. 2006, 66, 4182–4190. [Google Scholar]
- Liberti, M.V.; Locasale, J.W. The Warburg Effect: How Does it Benefit Cancer Cells? Trends Biochem. Sci. 2016, 41, 211–218. [Google Scholar] [CrossRef]
- Ganapathy-Kanniappan, S.; Geschwind, J.F. Tumor glycolysis as a target for cancer therapy: Progress and prospects. Mol. Cancer 2013, 12, 152. [Google Scholar] [CrossRef]
- Kaesler, S.; Wölbing, F.; Kempf, W.E.; Skabytska, Y.; Köberle, M.; Volz, T.; Sinnberg, T.; Amaral, T.; Möckel, S.; Yazdi, A.; et al. Targeting tumor-resident mast cells for effective anti-melanoma immune responses. JCI Insight 2019, 4, e125057. [Google Scholar] [CrossRef] [PubMed]
- Dudeck, A.; Köberle, M.; Goldmann, O.; Meyer, N.; Dudeck, J.; Lemmens, S.; Rohde, M.; Roldán, N.G.; Dietze-Schwonberg, K.; Orinska, Z.; et al. Mast cells as protectors of health. J. Allergy Clin. Immunol. 2019, 144, S4–S18. [Google Scholar] [CrossRef] [PubMed]
| Characteristics | Training Cohort (TCGA-UVM, 80 Cases) | Validation Cohort (GSE22138, 63 Cases) | |
|---|---|---|---|
| age | |||
| <65 | 45(56.25%) | 36(57.14%) | |
| ≥65 | 35(43.75%) | 27(42.86%) | |
| gender | |||
| female | 35(43.75%) | 24(38.1%) | |
| male | 45(56.25%) | 39(61.9%) | |
| T classification | |||
| T1 | 0 | NA | |
| T2 | 4(5%) | NA | |
| T3 | 36(45%) | NA | |
| T4 | 38(47.5%) | NA | |
| unknown | 2(2.5%) | NA | |
| M classification | |||
| M0 | 73(91.25%) | 28(44.44%) | |
| M1 | 3(3.75%) | 35(55.56%) | |
| unknown | 4(5%) | 0 | |
| tumor stage | |||
| stage I | 0 | NA | |
| stage II | 36(45%) | NA | |
| stage III | 40(50%) | NA | |
| stage IV | 4(5%) | NA | |
| extrascleral extension | |||
| yes | 7(8.75%) | 5(7.94%) | |
| no | 68(85%) | 48(76.19%) | |
| unknown | 5(6.25%) | 10(15.87%) | |
| tumor diameter, mm | |||
| <20 | 60(75%) | 44(69.84%) | |
| ≥20 | 19(23.75%) | 9(14.29%) | |
| unknown | 1(1.25%) | 10(15.87%) | |
| tumor thickness, mm | |||
| <10 | 29(36.25%) | 10(15.87%) | |
| ≥10 | 51(63.75%) | 53(84.13%) | |
| tumor eye side | |||
| left | NA | 33(52.38%) | |
| right | NA | 30(47.62%) | |
| tumor location | |||
| all over the eye | NA | 1(1.59%) | |
| anterior to equator | NA | 3(4.76%) | |
| on equator | NA | 42(66.67%) | |
| posterior and on equator | NA | 3(4.76%) | |
| posterior to equator | NA | 9(14.29%) | |
| unknown | NA | 5(7.94%) | |
| tumor cell type | |||
| epithelioid | NA | 21(33.33%) | |
| mixed | NA | 23(36.51%) | |
| unknown | NA | 19(30.16%) | |
| eye color | |||
| blue | 9(11.25%) | NA | |
| brown | 15(18.75%) | NA | |
| green | 6(7.5%) | NA | |
| unknown | 50(62.5%) | NA | |
| person neoplasm cancer status | |||
| with tumor | 9(11.25%) | NA | |
| tumor free | 56(70%) | NA | |
| unknown | 15(18.75%) | NA | |
| radiation therapy | |||
| yes | 3(3.75%) | NA | |
| no | 63(78.75%) | NA | |
| unknown | 14(17.5%) | NA | |
| ethnicity | |||
| hispanic or latino | 1(1.25%) | NA | |
| not hispanic or latino | 52(65%) | NA | |
| unknown | 27(33.75%) | NA | |
| tissue or organ of origin diagnosis | |||
| choroid | 67(83.75%) | NA | |
| ciliary body | 5(6.25%) | NA | |
| overlapping lesion of eye and adnexa | 8(10%) | NA | |
| retinal detachment | |||
| yes | NA | 36(57.14%) | |
| no | NA | 22(34.92%) | |
| unknown | NA | 5(7.94%) | |
| mitotic count | |||
| <20 | 42(52.5%) | NA | |
| ≥20 | 11(13.75%) | NA | |
| unknown | 27(33.75%) | NA | |
| chromosome 3 status | |||
| disomy | NA | 18(28.57%) | |
| monosomy | NA | 37(58.73%) | |
| unknown | NA | 8(12.7%) | |
| Gene Symbol | Description | Category | Genomic Location | Kaplan–Meier Analysis (p-Value) | Univariate Cox Regression Analysis | |||
|---|---|---|---|---|---|---|---|---|
| HR | HR_95L | HR_95H | p-Value | |||||
| VDAC1 | Voltage Dependent Anion Channel 1 | Protein Coding | chr5 | 7.12 × 10−6 | 5.291343594 | 1.781142996 | 15.71929771 | 2.71 × 10−3 |
| STEAP3 | STEAP3 Metalloreductase | Protein Coding | chr2 | 3.19 × 10−3 | 4.206162616 | 2.092921898 | 8.453160136 | 5.49 × 10−5 |
| SLC39A8 | Solute Carrier Family 39 Member 8 | Protein Coding | chr4 | 3.77 × 10−2 | 3.869852511 | 1.326333156 | 11.29109861 | 1.33 × 10−2 |
| SLC11A2 | Solute Carrier Family 11 Member 2 | Protein Coding | chr12 | 4.76 × 10−3 | 3.094601274 | 1.570769273 | 6.096730571 | 1.09 × 10−3 |
| PEBP1 | Phosphatidylethanolamine Binding Protein 1 | Protein Coding | chr12 | 4.86 × 10−2 | 0.234529606 | 0.075635069 | 0.727230595 | 1.20 × 10−2 |
| MAPK1 | Mitogen−Activated Protein Kinase 1 | Protein Coding | chr22 | 9.03 × 10−3 | 2.895607401 | 1.163501743 | 7.206299663 | 2.23 × 10−2 |
| MAP1LC3C | Microtubule Associated Protein 1 Light Chain 3 Gamma | Protein Coding | chr1 | 1.31 × 10−2 | 0.459748503 | 0.271733125 | 0.777853955 | 3.78 × 10−3 |
| LINC00472 | Long Intergenic Non-Protein Coding RNA 472 | RNA Gene | chr6 | 6.36 × 10−3 | 0.043258919 | 0.003322917 | 0.563160037 | 1.65 × 10−2 |
| ITGA6 | Integrin Subunit Alpha 6 | Protein Coding | chr2 | 1.13 × 10−3 | 4.613594536 | 2.148024539 | 9.909223176 | 8.85 × 10−5 |
| HSPA5 | Heat Shock Protein Family A (Hsp70) Member 5 | Protein Coding | chr9 | 8.91 × 10−3 | 2.25326069 | 1.228917946 | 4.131426149 | 8.63 × 10−3 |
| HMOX1 | Heme Oxygenase 1 | Protein Coding | chr22 | 2.19 × 10−3 | 2.334473768 | 1.59857086 | 3.409149952 | 1.14 × 10−5 |
| GSS | Glutathione Synthetase | Protein Coding | chr20 | 3.31 × 10−3 | 3.851728269 | 1.85357871 | 8.003874117 | 3.02 × 10−4 |
| FTH1 | Ferritin Heavy Chain 1 | Protein Coding | chr11 | 4.64 × 10− 3 | 4.040699109 | 1.10619952 | 14.75976891 | 3.46 × 10−2 |
| CD44 | CD44 Molecule (Indian Blood Group) | Protein Coding | chr11 | 6.66 × 10− 3 | 0.304760194 | 0.142947413 | 0.649740862 | 2.10 × 10−3 |
| CASP8 | Caspase 8 | Protein Coding | chr2 | 3.91 × 10−2 | 2.604965341 | 1.209052632 | 5.612530215 | 1.45 × 10−2 |
| BAP1 | BRCA1 Associated Protein 1 | Protein Coding | chr3 | 1.40 × 10−6 | 0.561778701 | 0.412394493 | 0.765275275 | 2.56 × 10−4 |
| AURKA | Aurora Kinase A | Protein Coding | chr20 | 2.54 × 10−2 | 3.390492663 | 1.565174215 | 7.344511806 | 1.96 × 10−3 |
| ANO6 | Anoctamin 6 | Protein Coding | chr12 | 2.56 × 10−2 | 2.263254914 | 1.292723147 | 3.962428319 | 4.26 × 10−3 |
| ALOX12 | Arachidonate 12-Lipoxygenase, 12S Type | Protein Coding | chr17 | 1.48 × 10−3 | 0.022909689 | 0.002539428 | 0.20668188 | 7.66 × 10−4 |
| AIFM2/FSP1 | Apoptosis Inducing Factor Mitochondria Associated 2 | Protein Coding | chr10 | 6.03 × 10−6 | 6.104896507 | 2.780109603 | 13.40586045 | 6.55 × 10−6 |
| ACSL6 | Acyl-CoA Synthetase Long Chain Family Member 6 | Protein Coding | chr5 | 4.56 × 10−4 | 2.283441779 | 1.099533069 | 4.742109633 | 2.68 × 10−2 |
| ACSL1 | Acyl-CoA Synthetase Long Chain Family Member 1 | Protein Coding | chr4 | 6.64 × 10−3 | 1.873686292 | 1.236619189 | 2.838950221 | 3.06 × 10−3 |
| Gene Symbol | Description | Role | Risk Coefficient |
|---|---|---|---|
| STEAP3 | STEAP3 Metalloreductase | Marker [26] | 0.055060532 |
| MAP1LC3C | Microtubule Associated Protein 1 Light Chain 3 Gamma | Driver [27] | −0.202884346 |
| ITGA6 | Integrin Subunit Alpha 6 | Suppressor [28] | 0.34461317 |
| HMOX1 | Heme Oxygenase 1 | Driver [29,30,31,32], Suppressor [33,34], Marker [35,36] | 0.125266141 |
| CD44 | CD44 Molecule (Indian Blood Group) | Suppressor [37] | −0.316011897 |
| ALOX12 | Arachidonate 12-Lipoxygenase, 12S Type | Driver [38,39,40], Marker [41] | −1.311120914 |
| AIFM2/FSP1 | Apoptosis Inducing Factor Mitochondria Associated 2 | Suppressor [42,43] | 0.710789029 |
| Variable | Univariate Cox Analysis | Multivariate Cox Analysis | ||||||
|---|---|---|---|---|---|---|---|---|
| Coef | HR (95% CI) | z | p-Value | Coef | HR (95% CI) | z | p-Value | |
| TCGA-UVM (overall survival) * | ||||||||
| age | 0.0447 | 1.05 (1.01–1.09) | 2.35 | 0.0186 | 0.101 | 1.11 (0.976–1.25) | 1.57 | 0.115 |
| gender (male vs. female) | 0.433 | 1.54 (0.651–3.65) | 0.984 | 0.325 | ||||
| tumor stage (stage III vs. stage II) | 0.336 | 1.4 (0.556–3.52) | 0.713 | 0.476 | −3.06 | 0.047 (0.00199–1.11) | −1.9 | 0.0579 |
| tumor stage (stage IV vs. stage II) | 4.37 | 79.3 (7.55–834) | 3.64 | 0.000269 | NA | NA | NA | NA |
| extrascleral extension (yes vs. no) | 1.54 | 4.64 (1.5–14.4) | 2.66 | 0.00774 | −4.25 | 0.0142 (3.98× 10−13–5.1× 10 8) | −0.343 | 0.732 |
| tumor diameter | 0.155 | 1.17 (1.01–1.35) | 2.12 | 0.0344 | 0.723 | 2.06 (1.11–3.83) | 2.29 | 0.0221 |
| tumor thickness | 0.111 | 1.12 (0.949–1.32) | 1.33 | 0.183 | ||||
| radiation therapy (yes vs. no) | 1.68 | 5.35 (1.09–26.3) | 2.07 | 0.0389 | 7.79 | 2410 (4.34 × 10−8–1.33 × 10 14) | 0.617 | 0.537 |
| ethnicity (hispanic or latino vs. not hispanic or latino) | −16 | 1.09 × 10−7 (0-Inf) | −0.00205 | 0.998 | ||||
| tissue or organ of origin diagnosis (choroid vs. not choroid) | −0.286 | 0.751 (0.254–2.22) | −0.517 | 0.605 | ||||
| mitotic count | −0.0119 | 0.988 (0.931–1.05) | −0.394 | 0.693 | ||||
| chromosome 3 copy number | −1.86 | 0.156 (0.0574–0.422) | −3.65 | 0.00026 | 2.76 | 15.9 (0.894–281) | 1.88 | 0.0597 |
| chromosome 6p copy number | −1.06 | 0.348 (0.176–0.687) | −3.04 | 0.00237 | −0.874 | 0.417 (0.0559–3.11) | −0.852 | 0.394 |
| chromosome 8q copy number | 0.516 | 1.67 (1.27–2.2) | 3.68 | 0.000235 | −0.88 | 0.415 (0.126–1.37) | −1.44 | 0.149 |
| risk score | 1.65 | 5.22 (2.59–10.5) | 4.61 | 3.99 × 10−6 | 4.23 | 68.6 (3.36–1400) | 2.75 | 0.00598 |
| TCGA-UVM (progression-free survival) # | ||||||||
| age | 0.0271 | 1.03 (0.996–1.06) | 1.7 | 0.0886 | ||||
| gender (male vs. female) | −0.139 | 0.87 (0.422–1.8) | −0.376 | 0.707 | ||||
| tumor stage (stage III vs. stage II) | 0.381 | 1.46 (0.665–3.22) | 0.946 | 0.344 | −0.959 | 0.383 (0.094–1.56) | −1.34 | 0.181 |
| tumor stage (stage IV vs. stage II) | 3.31 | 27.4 (5.06–149) | 3.84 | 0.000124 | 2.87 | 17.6 (0.677–455) | 1.73 | 0.0845 |
| extrascleral extension (yes vs. no) | 1.45 | 4.26 (1.57–11.5) | 2.85 | 0.0044 | −0.682 | 0.506 (0.00917–27.9) | −0.333 | 0.739 |
| tumor diameter | 0.113 | 1.12 (0.999–1.25) | 1.94 | 0.0527 | ||||
| tumor thickness | 0.00859 | 1.01 (0.875–1.16) | 0.119 | 0.905 | ||||
| radiation therapy (yes vs. no) | 0.0846 | 1.09 (0.142–8.33) | 0.0814 | 0.935 | ||||
| ethnicity (hispanic or latino vs. not hispanic or latino) | −17 | 3.98 × 10−8 (0-Inf) | −0.00217 | 0.998 | ||||
| tissue or organ of origin diagnosis (choroid vs. not choroid) | 0.0872 | 1.09 (0.377–3.16) | 0.161 | 0.872 | ||||
| mitotic count | 0.0545 | 1.06 (1.01–1.1) | 2.64 | 0.00829 | 0.0575 | 1.06 (0.976–1.15) | 1.37 | 0.17 |
| chromosome 3 copy number | −1.86 | 0.156 (0.0631–0.386) | −4.02 | 5.88 × 10−5 | −1.3 | 0.272 (0.0293–2.52) | −1.15 | 0.252 |
| chromosome 6p copy number | −0.628 | 0.534 (0.321–0.888) | −2.42 | 0.0155 | 0.709 | 2.03 (0.611–6.75) | 1.16 | 0.247 |
| chromosome 8q copy number | 0.521 | 1.68 (1.31–2.16) | 4.08 | 4.56 × 10−5 | 0.146 | 1.16 (0.546–2.45) | 0.381 | 0.703 |
| risk score | 0.933 | 2.54 (1.65–3.91) | 4.25 | 2.13 × 10−5 | 0.703 | 2.02 (0.486–8.39) | 0.967 | 0.0334 |
| GSE22138 (metastasis-free survival) & | ||||||||
| age | 0.0213 | 1.02 (0.995–1.05) | 1.59 | 0.113 | ||||
| gender (male vs. female) | 0.353 | 1.42 (0.714–2.84) | 1 | 0.316 | ||||
| tumor eye side (left vs. right) | −0.193 | 0.824 (0.424–1.6) | −0.57 | 0.569 | ||||
| tumor location (on equator vs. others) | −0.325 | 0.723 (0.357–1.46) | −0.903 | 0.366 | ||||
| tumor diameter | −0.0165 | 0.984 (0.893–1.08) | −0.336 | 0.737 | ||||
| tumor thickness | 0.116 | 1.12 (0.951–1.33) | 1.37 | 0.171 | ||||
| tumor cell type (epithelioid vs. mixed) | 0.753 | 2.12 (0.954–4.72) | 1.85 | 0.0649 | ||||
| retinal detachment (yes vs. no) | 1.06 | 2.87 (1.24–6.68) | 2.45 | 0.0142 | 0.857 | 2.36 (0.981–5.66) | 1.92 | 0.0553 |
| extrascleral extension (yes vs. no) | 0.563 | 1.76 (0.668–4.62) | 1.14 | 0.253 | ||||
| chromosome 3 status (monosomy vs. disomy) | 1.67 | 5.29 (1.82–15.3) | 3.07 | 0.00217 | 1.24 | 3.45 (1.04–11.5) | 2.02 | 0.0435 |
| risk score | 0.646 | 1.91 (1.31–2.78) | 3.37 | 0.000745 | 0.523 | 1.69 (1.03–2.77) | 2.07 | 0.0388 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Luo, H.; Ma, C. A Novel Ferroptosis-Associated Gene Signature to Predict Prognosis in Patients with Uveal Melanoma. Diagnostics 2021, 11, 219. https://doi.org/10.3390/diagnostics11020219
Luo H, Ma C. A Novel Ferroptosis-Associated Gene Signature to Predict Prognosis in Patients with Uveal Melanoma. Diagnostics. 2021; 11(2):219. https://doi.org/10.3390/diagnostics11020219
Chicago/Turabian StyleLuo, Huan, and Chao Ma. 2021. "A Novel Ferroptosis-Associated Gene Signature to Predict Prognosis in Patients with Uveal Melanoma" Diagnostics 11, no. 2: 219. https://doi.org/10.3390/diagnostics11020219
APA StyleLuo, H., & Ma, C. (2021). A Novel Ferroptosis-Associated Gene Signature to Predict Prognosis in Patients with Uveal Melanoma. Diagnostics, 11(2), 219. https://doi.org/10.3390/diagnostics11020219







