Abnormal Circulating Maternal miRNA Expression Is Associated with a Low (<4%) Cell-Free DNA Fetal Fraction
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Samples Selection
2.3. Total RNA Extraction from Plasma
2.4. NanoString nCounter Human v3 miRNA Assay
2.5. Data Analysis
3. Results
3.1. Demographic Data
3.2. Evaluation of Placenta-Specific miRNAs with Changes of Expression
3.3. Evaluation of miRNAs with Significant Changes of Expression of at Least ±1.5-Fold
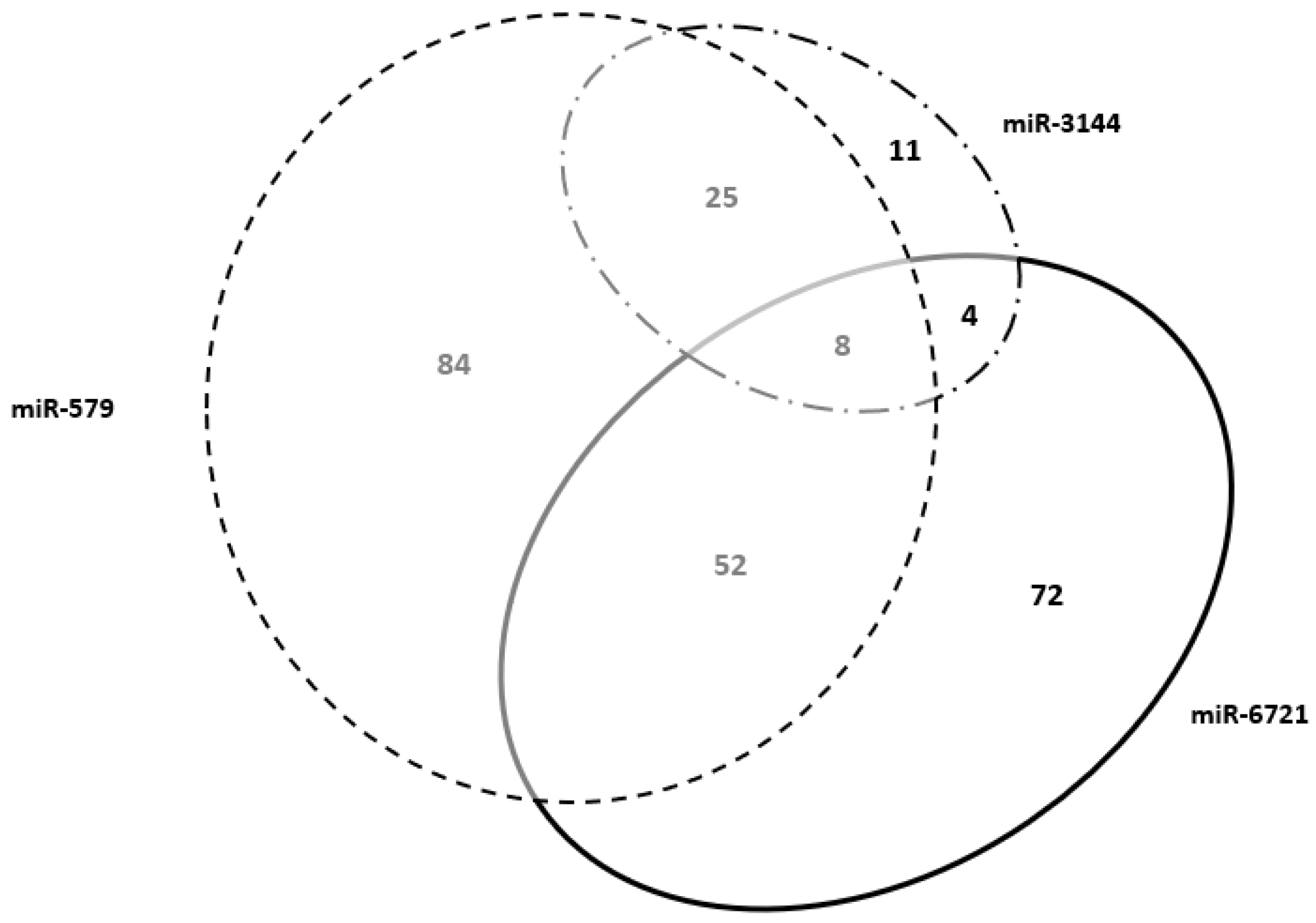
3.4. Predicted Biological Processes for the Four miRNA Target Genes Selected
3.5. Selected Maternal miRNAs and Placenta-Specific Genes
3.6. Placental Proteins Encoded by miRNAs Target Genes
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Qiao, L.; Mao, J.; Liu, M.; Liu, Y.; Song, X.; Tang, H.; Zhang, Q.; Li, H.; Lu, Y.; Liang, Y.; et al. Experimental factors are associated with fetal fraction in size selection noninvasive prenatal testing. Am. J. Transl. Res. 2019, 11, 6370–6381. [Google Scholar] [PubMed]
- Barrett, A.N.; Xiong, L.; Tan, T.Z.; Advani, H.V.; Hua, R.; Laureano-Asibal, C.; Soong, R.; Biswas, A.; Nagarajan, N.; Choolani, M. Measurement of fetal fraction in cell-free DNA from maternal plasma using a panel of insertion/deletion polymorphisms. PLoS ONE 2017, 12, e0186771. [Google Scholar] [CrossRef] [Green Version]
- Rafi, I.; Hill, M.; Hayward, J.; Chitty, L.S. Non-invasive prenatal testing: Use of cell-free fetal DNA in Down syndrome screening. Br. J. Gen. Pr. 2017, 67, 298–299. [Google Scholar] [CrossRef] [Green Version]
- La Verde, M.; De Falco, L.; Torella, A.; Savarese, G.; Savarese, P.; Ruggiero, R.; Conte, A.; Fico, V.; Torella, M.; Fico, A. Performance of cell-free DNA sequencing-based non-invasive prenatal testing: Experience on 36,456 singleton and multiple pregnancies. BMC Med. Genom. 2021, 14, 1–11. [Google Scholar] [CrossRef]
- Taylor-Phillips, S.; Freeman, K.; Geppert, J.; Agbebiyi, A.; Uthman, O.A.; Madan, J.; Clarke, A.; Quenby, S.; Clarke, A. Accuracy of non-invasive prenatal testing using cell-free DNA for detection of Down, Edwards and Patau syndromes: A systematic review and meta-analysis. BMJ Open 2016, 6, e010002. [Google Scholar] [CrossRef] [Green Version]
- Hartwig, T.S.; Ambye, L.; Werge, L.; Weiergang, M.K.; Nørgaard, P.; Sørensen, S.; Jørgensen, F.S. Non-Invasive Prenatal Testing (NIPT) in pregnancies with trisomy 21, 18 and 13 performed in a public setting—Factors of importance for correct interpretation of results. Eur. J. Obstet. Gynecol. Reprod. Biol. 2018, 226, 35–39. [Google Scholar] [CrossRef]
- Burns, W.; Koelper, N.; Barberio, A.; DeAgostino-Kelly, M.; Mennuti, M.; Sammel, M.D.; Dugoff, L. The association between anticoagulation therapy, maternal characteristics, and a failed cfDNA test due to a low fetal fraction. Prenat. Diagn. 2017, 37, 1125–1129. [Google Scholar] [CrossRef] [PubMed]
- Morano, D.; Rossi, S.; Lapucci, C.; Pittalis, M.C.; Farina, A. Cell-Free DNA (cfDNA) Fetal Fraction in Early- and Late-Onset Fetal Growth Restriction. Mol. Diagn. Ther. 2018, 22, 613–619. [Google Scholar] [CrossRef]
- Rolnik, D.L.; O’Gorman, N.; Fiolna, M.; van den Boom, D.; Nicolaides, K.H.; Poon, L.C. Maternal plasma cell-free DNA in the pre-diction of pre-eclampsia. Ultrasound Obstet Gynecol. 2015, 45, 106–111. [Google Scholar] [CrossRef] [PubMed]
- Koide, K.; Sekizawa, A.; Iwasaki, M.; Matsuoka, R.; Honma, S.; Farina, A.; Saito, H.; Okai, T. Fragmentation of cell-free fetal DNA in plasma and urine of pregnant women. Prenat. Diagn. 2005, 25, 604–607. [Google Scholar] [CrossRef] [PubMed]
- Nagy, B. Cell-free nucleic acids in prenatal diagnosis and pregnancy-associated diseases. EJIFCC 2019, 30, 215–223. [Google Scholar] [PubMed]
- Tsochandaridis, M.; Nasca, L.; Toga, C.; Levy-Mozziconacci, A. Circulating MicroRNAs as Clinical Biomarkers in the Predictions of Pregnancy Complications. BioMed Res. Int. 2015, 2015, 1–8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Curti, A.; Lapucci, C.; Berto, S.; Prandstraller, D.; Perolo, A.; Rizzo, N.; Farina, A. Maternal plasma mRNA species in fetal heart defects: A potential for molecular screening. Prenat. Diagn. 2016, 36, 738–743. [Google Scholar] [CrossRef] [PubMed]
- Contro, E.; Stefani, L.; Berto, S.; Lapucci, C.; Arcelli, D.; Prandstraller, D. Circulating mRNA in Maternal Plasma at the Second Trimester of Pregnancy: A Possible Screening Tool for Cardiac Conotruncal and Left Ventricular Outflow Tract Abnormalities. Mol. Diagn. Ther. 2017, 21, 653–661. [Google Scholar] [CrossRef]
- Morano, D.; Berto, S.; Lapucci, C.; Walczer Baldinazzo, L.; Prandstraller, D.; Farina, A. Levels of Circulating mRNA for the Tenas-cin-X (TNXB) Gene in Maternal Plasma at the Second Trimester in Pregnancies with Isolated Congenital Ventricular Septal Defects. Mol. Diagn. Ther. 2018, 22, 235–240. [Google Scholar] [CrossRef]
- Lytle, J.R.; Yario, T.A.; Steitz, J.A. Target mRNAs are repressed as efficiently by microRNA-binding sites in the 5′ UTR as in the 3′ UTR. Proc. Natl. Acad. Sci. USA 2007, 104, 9667–9672. [Google Scholar] [CrossRef] [Green Version]
- Catalanotto, C.; Cogoni, C.; Zardo, G. MicroRNA in Control of Gene Expression: An Overview of Nuclear Functions. Int. J. Mol. Sci. 2016, 17, 1712. [Google Scholar] [CrossRef] [Green Version]
- Liu, B.; Li, J.; Cairns, M.J. Identifying miRNAs, targets and functions. Brief Bioinform. 2014, 15, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Mouillet, J.-F.; Ouyang, Y.; Coyne, C.B.; Sadovsky, Y. MicroRNAs in placental health and disease. Am. J. Obstet. Gynecol. 2015, 213, S163–S172. [Google Scholar] [CrossRef] [Green Version]
- Cai, M.; Kolluru, G.K.; Ahmed, A. Small Molecule, Big Prospects: MicroRNA in Pregnancy and Its Complications. J. Pregnancy 2017, 2017, 6972732. [Google Scholar] [CrossRef] [Green Version]
- Higashijima, A.; Miura, K.; Mishima, H.; Kinoshita, A.; Jo, O.; Abe, S.; Hasegawa, Y.; Miura, S.; Yamasaki, K.; Yoshida, A.; et al. Characterization of placenta-specific microRNAs in fetal growth restriction pregnancy. Prenat. Diagn. 2013, 33, 214–222. [Google Scholar] [CrossRef]
- Hromadnikova, I.; Dvorakova, L.; Kotlabova, K.; Krofta, L. The Prediction of Gestational Hypertension, Preeclampsia and Fe-tal Growth Restriction via the First Trimester Screening of Plasma Exosomal C19MC microRNAs. Int. J. Mol. Sci. 2019, 20, 2972. [Google Scholar] [CrossRef]
- Donker, R.B.; Mouillet, J.F.; Chu, T.; Hubel, C.A.; Stolz, D.B.; Morelli, A.E.; Sadovsky, Y. The expression profile of C19MC microRNAs in pri-mary human trophoblast cells and exosomes. Mol. Hum. Reprod. 2012, 18, 417–424. [Google Scholar] [CrossRef] [Green Version]
- Yuan, Z.; Sun, X.; Jiang, D.; Ding, Y.; Lu, Z.; Gong, L.; Liu, H.; Xie, J. Origin and evolution of a placental-specific microRNA family in the human genome. BMC Evol. Biol. 2010, 10, 346. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dieckmann, K.-P.; Spiekermann, M.; Balks, T.; Flor, I.; Löning, T.; Bullerdiek, J.; Belge, G. MicroRNAs miR-371-3 in serum as diagnostic tools in the management of testicular germ cell tumours. Br. J. Cancer 2012, 107, 1754–1760. [Google Scholar] [CrossRef] [Green Version]
- Concepcion, C.P.; Bonetti, C.; Ventura, A. The miR-17-92 family of microRNA clusters in development and disease. Cancer J. 2012, 18, 262–267. [Google Scholar] [CrossRef] [PubMed]
- Friedrich, M.; Pracht, K.; Mashreghi, M.F.; Jäck, H.M.; Radbruch, A.; Seliger, B. The role of the miR-148/-152 family in physiology and disease. Eur. J. Immunol. 2017, 47, 2026–2638. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chang, W.-L.; Liu, Y.-W.; Dang, Y.-L.; Jiang, X.-X.; Xu, H.; Huang, X.; Wang, Y.-L.; Wang, H.; Zhu, C.; Xue, L.-Q.; et al. PLAC8, a new marker for human interstitial extravillous trophoblast cells, promotes their invasion and migration. Development 2018, 145. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Woods, L.; Perez-Garcia, V.; Hemberger, M. Regulation of Placental Development and Its Impact on Fetal Growth-New In-sights From Mouse Models. Front. Endocrinol. 2018, 9, 570. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Windsperger, K.; Dekan, S.; Pils, S.; Golletz, C.; Kunihs, V.; Fiala, C.; Kristiansen, G.; Knöfler, M.; Pollheimer, J. Extravillous trophoblast invasion of venous as well as lymphatic vessels is altered in idiopathic, recurrent, spontaneous abortions. Hum. Reprod. 2017, 32, 1208–1217. [Google Scholar] [CrossRef] [Green Version]
- Li, N.; Huang, L.; Li, Y.; Chen, X.; Yang, Y.; Hou, Y.; Qiao, C. Lin28B/miR-92b Promote the Proliferation, Migration, and Invasion in the Pathogenesis of Preeclampsia via the DKK1/Wnt/β-Catenin Pathway. Reprod. Sci. 2020, 27, 815–822. [Google Scholar] [CrossRef]
- Peng, Y.; Croce, C.M. The role of microRNAs in human cancer. Signal. Transduct. Target Ther. 2016, 1, 15004. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fattore, L.; Mancini, R.; Acunzo, M.; Romano, G.; Laganà, A.; Pisanu, M.E.; Malpicci, D.; Madonna, G.; Mallardo, D.; Capone, M.; et al. miR-579-3p controls melanoma progression and resistance to target therapy. Proc. Natl. Acad. Sci. USA 2016, 113, E5005–E5013. [Google Scholar] [CrossRef] [Green Version]
- Jin, Y.; Zhou, X.; Yao, X.; Zhang, Z.; Cui, M.; Lin, Y. MicroRNA-612 inhibits cervical cancer progression by targeting NOB1. J. Cell. Mol. Med. 2020, 24, 3149–3156. [Google Scholar] [CrossRef] [PubMed]
- Kang, X.; Kong, F.; Wu, S.; Liu, Q.; Yang, C.; Wu, X. MicroRNA-612 suppresses the malignant development of non-small-cell lung cancer by directly targeting bromodomain-containing protein 4. Onco Targets Ther. 2019, 12, 4167–4179. [Google Scholar] [CrossRef] [Green Version]
- Tseng, A.M.; Mahnke, A.H.; Wells, A.B.; Salem, N.A.; Allan, A.M.; Roberts, V.H.; Newman, N.; Walter, N.A.; Kroenke, C.D.; Grant, K.A.; et al. Maternal circulating miRNAs that predict infant FASD outcomes influence placental maturation. Life Sci. Alliance 2019, 2, e201800252. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Balaraman, S.; Schafer, J.J.; Tseng, A.M.; Wertelecki, W.; Yevtushok, L.; Zymak-Zakutnya, N.; Chambers, C.D.; Miranda, R.C. Plasma miRNA Profiles in Pregnant Women Predict Infant Outcomes following Prenatal Alcohol Exposure. PLoS ONE 2016, 11, e0165081. [Google Scholar] [CrossRef] [PubMed]
- Torrente, Y.; Bella, P.; Tripodi, L.; Villa, C.; Farini, A. Role of Insulin-Like Growth Factor Receptor 2 across Muscle Homeosta-sis: Implications for Treating Muscular Dystrophy. Cells 2020, 9, 441. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harris, L.; Crocker, I.P.; Baker, P.; Aplin, J.; Westwood, M. IGF2 Actions on Trophoblast in Human Placenta Are Regulated by the Insulin-Like Growth Factor 2 Receptor, Which Can Function as Both a Signaling and Clearance Receptor1. Biol. Reprod. 2011, 84, 440–446. [Google Scholar] [CrossRef] [Green Version]
- Lightowlers, R.N.; Chrzanowska-Lightowlers, Z.M.A. Human pentatricopeptide proteins: Only a few and what do they do? RNA Biol. 2013, 10, 1433–1438. [Google Scholar] [CrossRef] [Green Version]
- Naik, R.; Galande, S. SATB family chromatin organizers as master regulators of tumor progression. Oncogene 2018, 38, 1989–2004. [Google Scholar] [CrossRef] [PubMed]
- Asanoma, K.; Kubota, K.; Chakraborty, D.; Renaud, S.J.; Wake, N.; Fukushima, K.; Soares, M.J.; Rumi, M.K. SATB homeobox proteins regulate troph-oblast stem cell renewal and differentiation. J. Biol. Chem. 2012, 287, 2257–2268. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gerson, K.D.; Truong, S.; Haviland, M.J.; O’Brien, B.M.; Hacker, M.R.; Spiel, M.H. Low fetal fraction of cell-free DNA predicts placen-tal dysfunction and hypertensive disease in pregnancy. Pregnancy Hypertens. 2019, 16, 148–153. [Google Scholar] [CrossRef] [PubMed]

| Variable | Group A cfDNAff < 4% (n = 6) | Group B cfDNAff > 4% (n = 6) | p-Value * |
|---|---|---|---|
| GA (days) | 82.0 (72–99) | 84.5 (71–91) | >0.99 |
| BMI | 22.48 (20.43–27.72) | 20.03 (17.26–25.82) | 0.065 |
| Maternal age (yrs) | 33.7 (29–42) | 33.6 (26–41) | >0.99 |
| Female (%) | 50 | 33.3 | 0.500 |
| cfDNAff | 0.5 (0–1) | 15.5 (15–17) | - |
| Neonatal weight (gr.) | 2955 (2100–3140) | 3080 (2860–3680) | 0.121 |
| FGR (%) | 33.3 | 0 | <0.001 |
| miRNAs | Localization |
|---|---|
| C19MC14 [23] | chromosome 19 encoding 46 intronic miRNAs |
| C14MC11 [19] | on chromosome 14 encoding 39 miRNAs |
| placenta-specific 1302 miRNA family [24] | derived from the MER53 transposon element |
| as miR-371/miR-373 [25] | chromosome 19 encoding 7 miRNAs |
| miR-17-92 11. [26] | chromosome 13 encoding for 6 miRNAs |
| miRNA 148-152 family [27] | chromosomes 7, 12 and 17 encoding 3 miRNAs |
| Placenta-Related miRNAs |
|---|
| hsa-let-7a-5p |
| hsa-miR-210-3p |
| hsa-miR-369-3p |
| hsa-miR-371a-5p |
| hsa-miR-512-5p |
| hsa-miR-517a-3p |
| hsa-miR-519a-3p |
| hsa-miR-93-5p |
| hsa-miR-96-5p |
| hsa-miR-206 |
| miRNAs | Placenta-Specific Target Genes | Gene Ontology |
|---|---|---|
| miR-579 | ADAMTSL4, ADM, AFF1, AFF2, AGTR1, AKR1B15, ARL14EPL, ATP6V1C2, BCAR3, C18orf54, C2orf83, CCDC102B, CLEC1A, COBLL1, COL11A1, CRYBG1, CYP19A1, DAB2, DKK1, EDNRB, EPB42, EPYC, ERVV-1, ESRRG, F13A1, F5, FAM162B, FAM46A, FBN1, FBN2, GDF6, GJB7, GM2A, GPR1, HPGDS, HSD17B2, HTR1F, HTR2B, IDO2, IGFBP1, IGSF3, IL1RAP, IL1RL1, JAM2, KATNBL1, LUM, LYPD6, MAGEA10, MB21D2, MEST, MFAP5, MUC15, NLRP10, NNAT, NRK, OLIG3, OLR1, PAPPA, PEG3, PHLDA2, PHLDB2, PKIB, PSG1, PSG2, PSG4, PSG8, RASA1, SEMA6D, SIGLEC6, SLC25A35, SPP1, STS, TBX4, TBX5, TFAP2A, TFRC, TMEM2, TWIST1, TXK, VGLL1, VGLL3, WNT2, XKRX, ZNF468 | cellular process (GO:0009987) |
| metabolic process (GO:0008152) | ||
| biological regulation (GO:0065007) | ||
| localization (GO:0051179) | ||
| multicellular organismal process (GO:0032501) | ||
| response to stimulus (GO:0050896) | ||
| biological adhesion (GO:0022610) | ||
| developmental process (GO:0032502) | ||
| immune system process (GO:0002376) | ||
| cellular component organization or biogenesis (GO:0071840) | ||
| reproduction (GO:0000003) | ||
| cell proliferation (GO:0008283) | ||
| rhythmic process (GO:0048511) | ||
| biological phase (GO:0044848) | ||
| nitrogen utilization (GO:0019740) | ||
| miR-3144 | C2orf72, ELOVL2, HAPLN1, MORN3, P2RY1, SKP2, SLC30A2, TFPI2, TRAPPC3L, TRIM10, TRIM64B | cellular process (GO:0009987) |
| metabolic process (GO:0008152) | ||
| biological regulation (GO:0065007) | ||
| localization (GO:0051179) | ||
| multicellular organismal process (GO:0032501) | ||
| response to stimulus (GO:0050896) | ||
| developmental process (GO:0032502) | ||
| biological adhesion (GO:0022610) | ||
| locomotion (GO:0040011) | ||
| pigmentation (GO:0043473) | ||
| miR-6721 | AADACL3, ALPP, AOC1, APLNR, ARID3A, ASCL2, ATG9B, BIRC7, C1QTNF6, CAPN6, CD248, CDH5, CLDN19, CLDN9, COL4A2, COX4I2, CXorf67, CYP11A1, DACT2, DLX3, DUSP9, ERVFRD-1, ERVV-2, FAM129B, FSTL3, FURIN, GABRE, GDPD5, GJA5, GPR78, GRHL2, HES2, INSL4, ISM2, KLF14, LAMC3, LARGE2, LEP, LGR5, MAFK, MFSD2B, N4BP3, NOTUM, NOX5, OC90, PGF, PODNL1, PROCR, SDC1, SEMA7A, SH2D5, SLC13A4, SLC22A11, SLC43A2, SLC4A1, SLC7A4, SP6, ST3GAL4, STRA6, SYNPO, SYTL5, TIMP2, TMEM139, TNS4, TREML2, TRIM58, TRPV6, UNC13D, WNT1, WNT3A, WNT7A, ZFAT | cellular process (GO:0009987) |
| metabolic process (GO:0008152) | ||
| biological regulation (GO:0065007) | ||
| localization (GO:0051179) | ||
| multicellular organismal process (GO:0032501) | ||
| response to stimulus (GO:0050896) | ||
| developmental process (GO:0032502) | ||
| immune system process (GO:0002376) | ||
| biological adhesion (GO:0022610) | ||
| cellular component organization or biogenesis (GO:0071840) | ||
| reproduction (GO:0000003) | ||
| cell proliferation (GO:0008283) | ||
| rhythmic process (GO:0048511) | ||
| biological phase (GO:0044848) | ||
| signaling (GO:0023052) | ||
| pigmentation (GO:0043473) | ||
| multi-organism process (GO:0051704) | ||
| Genes shared between miR-579 and miR-6721 | ACKR2, ADAM12, ADGRG6, APLN, ART4, CADM3, CCSAP, CD59, CLDN6, CREB3L2, CYTH3, DLK1, EGFL7, EXPH5, FHDC1, GADD45G, GCM1, GDF15, GRAMD2A, GSE1, HPGD, IGDCC3, IGF2, IGF2BP1, IL2RB, INHBA, ITIH5, KMO, LRRC15, LVRN, MBNL3, MORC4, NCMAP, PAEP, PDPN, PLAU, PRG2, QSOX1, RAI14, RGPD1, RSPO2, RXFP1, SCIN, SERPINE1, SLC2A1, SLC38A9, SLC6A2, TGM2, THSD7A, TIMP3, UCK2, ZFP42 | cellular process (GO:0009987) |
| metabolic process (GO:0008152) | ||
| biological regulation (GO:0065007) | ||
| localization (GO:0051179) | ||
| response to stimulus (GO:0050896) | ||
| developmental process (GO:0032502) | ||
| immune system process (GO:0002376) | ||
| biological adhesion (GO:0022610) | ||
| cell proliferation (GO:0008283) | ||
| Genes shared between miR-579 and miR-3144 | ADAMTS20, ADAMTS5, ATP10D, CHSY1, CYSLTR2, DEPDC1B, FN1, GULP1, HGF, HMGB3, LGALS13, LIN28B, LIPG, LNPEP, MAN1A2, NECTIN3, NFE2L3, NIPAL1, PLAC8, SH3TC2, SPTLC3, TDRP, TFPI, TLR3, TUSC3 | cellular process (GO:0009987) |
| metabolic process (GO:0008152) | ||
| biological regulation (GO:0065007) | ||
| localization (GO:0051179) | ||
| multicellular organismal process (GO:0032501) | ||
| response to stimulus (GO:0050896) | ||
| developmental process (GO:0032502) | ||
| biological adhesion (GO:0022610) | ||
| immune system process (GO:0002376) | ||
| reproduction (GO:0000003) | ||
| cellular component organization or biogenesis (GO:0071840) | ||
| Genes shared between miR-3144 and miR-6721 | ERVMER34-1, FAM89A, MFAP2, SPIRE2 | cellular process (GO:0009987) |
| metabolic process (GO:0008152) | ||
| biological regulation (GO:0065007) | ||
| localization (GO:0051179) | ||
| multicellular organismal process (GO:0032501) | ||
| response to stimulus (GO:0050896) | ||
| developmental process (GO:0032502) | ||
| biological adhesion (GO:0022610) | ||
| Genes shared Among miR-579, miR-3144 and miR-6721 | ARHGAP42, FOXO4, GUCY1A2, IGF2R, PHACTR2, PTCD2, SATB2, VAV3 | cellular process (GO:0009987) |
| metabolic process (GO:0008152) | ||
| biological regulation (GO:0065007) | ||
| localization (GO:0051179) | ||
| multicellular organismal process (GO:0032501) | ||
| response to stimulus (GO:0050896) | ||
| developmental process (GO:0032502) | ||
| biological adhesion (GO:0022610) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Santoro, G.; Lapucci, C.; Giannoccaro, M.; Caporilli, S.; Rusin, M.; Seidenari, A.; Ferrari, M.; Farina, A. Abnormal Circulating Maternal miRNA Expression Is Associated with a Low (<4%) Cell-Free DNA Fetal Fraction. Diagnostics 2021, 11, 2108. https://doi.org/10.3390/diagnostics11112108
Santoro G, Lapucci C, Giannoccaro M, Caporilli S, Rusin M, Seidenari A, Ferrari M, Farina A. Abnormal Circulating Maternal miRNA Expression Is Associated with a Low (<4%) Cell-Free DNA Fetal Fraction. Diagnostics. 2021; 11(11):2108. https://doi.org/10.3390/diagnostics11112108
Chicago/Turabian StyleSantoro, Graziano, Cristina Lapucci, Marco Giannoccaro, Simona Caporilli, Martina Rusin, Anna Seidenari, Maurizio Ferrari, and Antonio Farina. 2021. "Abnormal Circulating Maternal miRNA Expression Is Associated with a Low (<4%) Cell-Free DNA Fetal Fraction" Diagnostics 11, no. 11: 2108. https://doi.org/10.3390/diagnostics11112108
APA StyleSantoro, G., Lapucci, C., Giannoccaro, M., Caporilli, S., Rusin, M., Seidenari, A., Ferrari, M., & Farina, A. (2021). Abnormal Circulating Maternal miRNA Expression Is Associated with a Low (<4%) Cell-Free DNA Fetal Fraction. Diagnostics, 11(11), 2108. https://doi.org/10.3390/diagnostics11112108








