Male Breast Cancer: Results of the Application of Multigene Panel Testing to an Italian Cohort of Patients
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Patients’ Selection
2.3. Sample Collection and DNA Extraction
2.4. Next-Generation Sequencing (NGS)
2.5. Bioinformatics Analysis
2.6. Multiplex Ligation-Dependent Probe Amplification (MLPA)
2.7. Variant Classification
3. Results
3.1. Patient Characteristics
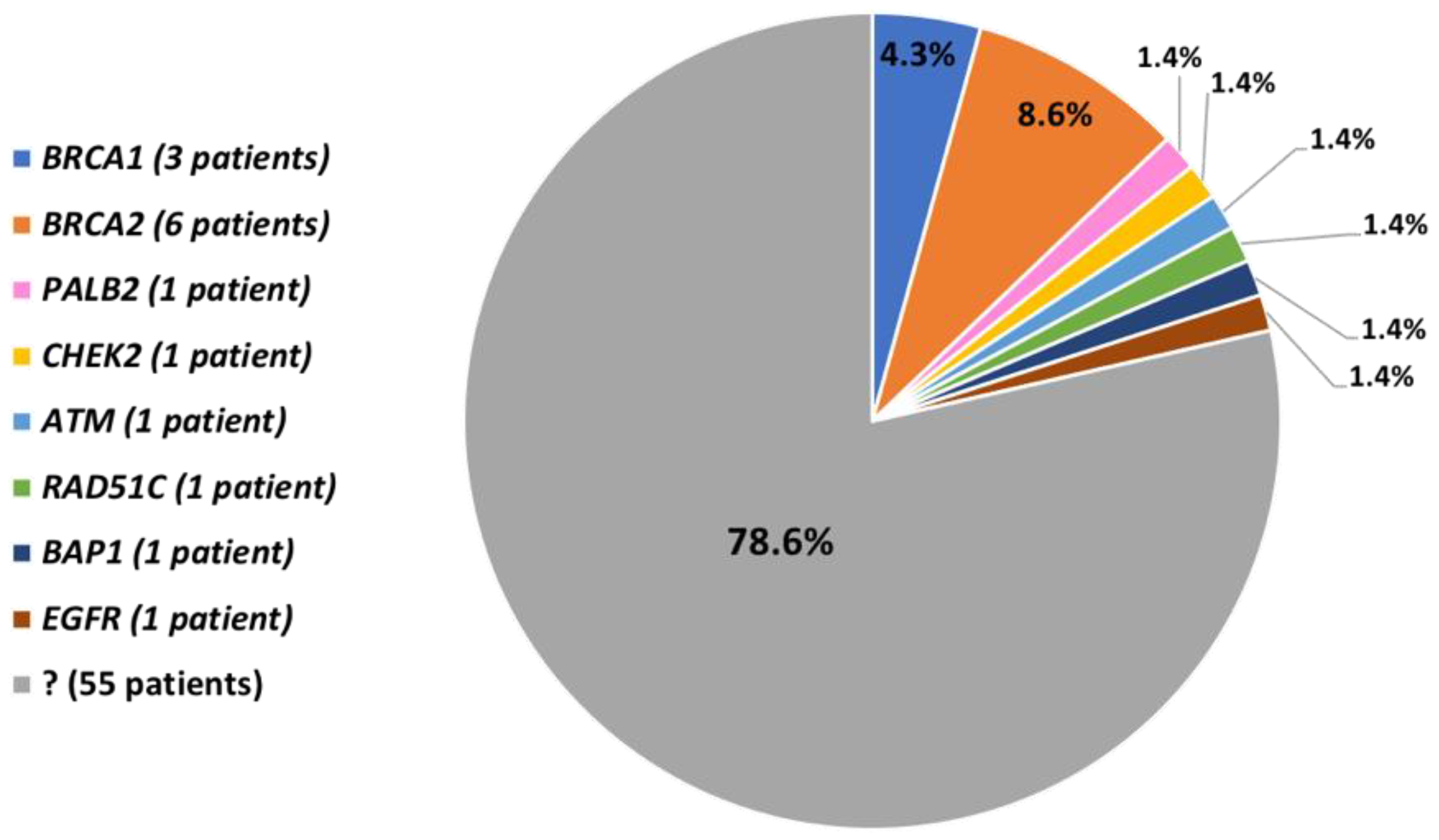
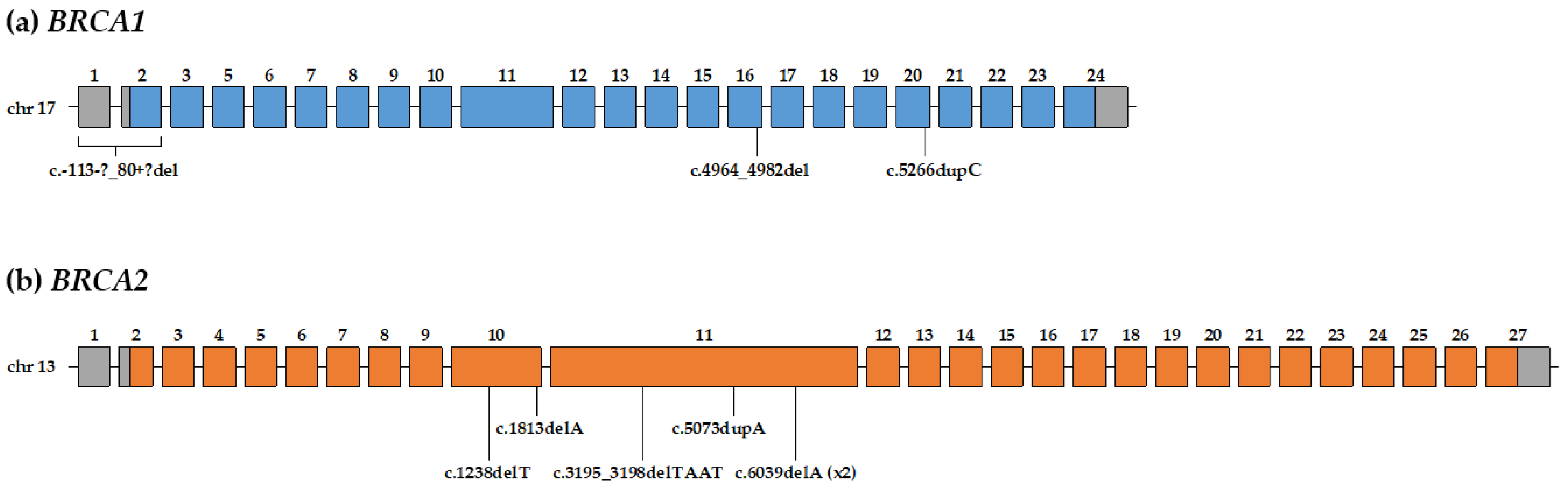
3.2. Pathogenic and Likely-Pathogenic Variants in BRCA1/2 Genes
3.3. Pathogenic and Likely-Pathogenic Variants in other Genes
3.4. Patients without Pathogenic and Likely-Pathogenic Variants
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA. Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Ottini, L. Male breast cancer: A rare disease that might uncover underlying pathways of breast cancer. Nat. Rev. Cancer 2014, 14, 643. [Google Scholar] [CrossRef] [PubMed]
- Gnerlich, J.L.; Deshpande, A.D.; Jeffe, D.B.; Seelam, S.; Kimbuende, E.; Margenthaler, J.A. Poorer survival outcomes for male breast cancer compared with female breast cancer may be attributable to in-stage migration. Ann. Surg. Oncol. 2011, 18, 1837–1844. [Google Scholar] [CrossRef] [PubMed]
- Rizzolo, P.; Silvestri, V.; Tommasi, S.; Pinto, R.; Danza, K.; Falchetti, M.; Gulino, M.; Frati, P.; Ottini, L. Male breast cancer: Genetics, epigenetics, and ethical aspects. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 2013, 24 (Suppl. 8), viii75–viii82. [Google Scholar] [CrossRef] [PubMed]
- Angeli, D.; Salvi, S.; Tedaldi, G. Genetic Predisposition to Breast and Ovarian Cancers: How Many and Which Genes to Test? Int. J. Mol. Sci. 2020, 21, 1128. [Google Scholar] [CrossRef] [PubMed]
- Miki, Y.; Swensen, J.; Shattuck-Eidens, D.; Futreal, P.A.; Harshman, K.; Tavtigian, S.; Liu, Q.; Cochran, C.; Bennett, L.M.; Ding, W. A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 1994, 266, 66–71. [Google Scholar] [CrossRef]
- Wooster, R.; Bignell, G.; Lancaster, J.; Swift, S.; Seal, S.; Mangion, J.; Collins, N.; Gregory, S.; Gumbs, C.; Micklem, G. Identification of the breast cancer susceptibility gene BRCA2. Nature 1995, 378, 789–792. [Google Scholar] [CrossRef]
- Petrucelli, N.; Daly, M.B.; Pal, T. BRCA1- and BRCA2-Associated Hereditary Breast and Ovarian Cancer. In GeneReviews®; University of Washington: Seattle, WA, USA, 1998; (updated 2016). [Google Scholar]
- Tai, Y.C.; Domchek, S.; Parmigiani, G.; Chen, S. Breast cancer risk among male BRCA1 and BRCA2 mutation carriers. J. Natl. Cancer Inst. 2007, 99, 1811–1814. [Google Scholar] [CrossRef]
- Rahman, N. Realizing the promise of cancer predisposition genes. Nature 2014, 505, 302–308. [Google Scholar] [CrossRef]
- Easton, D.F.; Pharoah, P.D.P.; Antoniou, A.C.; Tischkowitz, M.; Tavtigian, S.V.; Nathanson, K.L.; Devilee, P.; Meindl, A.; Couch, F.J.; Southey, M.; et al. Gene-panel sequencing and the prediction of breast-cancer risk. N. Engl. J. Med. 2015, 372, 2243–2257. [Google Scholar] [CrossRef]
- Apostolou, P.; Fostira, F. Hereditary breast cancer: The era of new susceptibility genes. Biomed. Res. Int. 2013, 2013, 747318. [Google Scholar] [CrossRef] [PubMed]
- Kurian, A.W.; Antoniou, A.C.; Domchek, S.M. Refining Breast Cancer Risk Stratification: Additional Genes, Additional Information. Am. Soc. Clin. Oncol. Educ. book. Am. Soc. Clin. Oncol. Annu. Meet. 2016, 35, 44–56. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, F.C.; van Overeem Hansen, T.; Sørensen, C.S. Hereditary breast and ovarian cancer: New genes in confined pathways. Nat. Rev. Cancer 2016, 16, 599–612. [Google Scholar] [CrossRef] [PubMed]
- Tedaldi, G.; Tebaldi, M.; Zampiga, V.; Danesi, R.; Arcangeli, V.; Ravegnani, M.; Cangini, I.; Pirini, F.; Petracci, E.; Rocca, A.; et al. Multiple-gene panel analysis in a case series of 255 women with hereditary breast and ovarian cancer. Oncotarget 2017, 8, 47064–47075. [Google Scholar] [CrossRef] [PubMed]
- Walsh, T.; Lee, M.K.; Casadei, S.; Thornton, A.M.; Stray, S.M.; Pennil, C.; Nord, A.S.; Mandell, J.B.; Swisher, E.M.; King, M.-C. Detection of inherited mutations for breast and ovarian cancer using genomic capture and massively parallel sequencing. Proc. Natl. Acad. Sci. USA 2010, 107, 12629–12633. [Google Scholar] [CrossRef] [PubMed]
- Gracia-Aznarez, F.J.; Fernandez, V.; Pita, G.; Peterlongo, P.; Dominguez, O.; de la Hoya, M.; Duran, M.; Osorio, A.; Moreno, L.; Gonzalez-Neira, A.; et al. Whole exome sequencing suggests much of non-BRCA1/BRCA2 familial breast cancer is due to moderate and low penetrance susceptibility alleles. PLoS ONE 2013, 8, e55681. [Google Scholar] [CrossRef] [PubMed]
- Kurian, A.W.; Hare, E.E.; Mills, M.A.; Kingham, K.E.; McPherson, L.; Whittemore, A.S.; McGuire, V.; Ladabaum, U.; Kobayashi, Y.; Lincoln, S.E.; et al. Clinical evaluation of a multiple-gene sequencing panel for hereditary cancer risk assessment. J. Clin. Oncol. 2014, 32, 2001–2009. [Google Scholar] [CrossRef]
- Desmond, A.; Kurian, A.W.; Gabree, M.; Mills, M.A.; Anderson, M.J.; Kobayashi, Y.; Horick, N.; Yang, S.; Shannon, K.M.; Tung, N.; et al. Clinical Actionability of Multigene Panel Testing for Hereditary Breast and Ovarian Cancer Risk Assessment. JAMA Oncol. 2015, 1, 943–951. [Google Scholar] [CrossRef]
- Kraus, C.; Hoyer, J.; Vasileiou, G.; Wunderle, M.; Lux, M.P.; Fasching, P.A.; Krumbiegel, M.; Uebe, S.; Reuter, M.; Beckmann, M.W.; et al. Gene panel sequencing in familial breast/ovarian cancer patients identifies multiple novel mutations also in genes others than BRCA1/2. Int. J. Cancer 2017, 140, 95–102. [Google Scholar] [CrossRef]
- Yang, X.; Leslie, G.; Doroszuk, A.; Schneider, S.; Allen, J.; Decker, B.; Dunning, A.M.; Redman, J.; Scarth, J.; Plaskocinska, I.; et al. Cancer Risks Associated With Germline PALB2 Pathogenic Variants: An International Study of 524 Families. J. Clin. Oncol. 2019, JCO1901907. [Google Scholar] [CrossRef]
- Silvestri, V.; Rizzolo, P.; Zanna, I.; Falchetti, M.; Masala, G.; Bianchi, S.; Papi, L.; Giannini, G.; Palli, D.; Ottini, L. PALB2 mutations in male breast cancer: A population-based study in Central Italy. Breast Cancer Res. Treat. 2010, 122, 299–301. [Google Scholar] [CrossRef] [PubMed]
- Silvestri, V.; Zelli, V.; Valentini, V.; Rizzolo, P.; Navazio, A.S.; Coppa, A.; Agata, S.; Oliani, C.; Barana, D.; Castrignanò, T.; et al. Whole-exome sequencing and targeted gene sequencing provide insights into the role of PALB2 as a male breast cancer susceptibility gene. Cancer 2017, 123, 210–218. [Google Scholar] [CrossRef] [PubMed]
- Rizzolo, P.; Zelli, V.; Silvestri, V.; Valentini, V.; Zanna, I.; Bianchi, S.; Masala, G.; Spinelli, A.M.; Tibiletti, M.G.; Russo, A.; et al. Insight into genetic susceptibility to male breast cancer by multigene panel testing: Results from a multicenter study in Italy. Int. J. Cancer 2019, 145, 390–400. [Google Scholar] [CrossRef] [PubMed]
- Rizzolo, P.; Silvestri, V.; Bucalo, A.; Zelli, V.; Valentini, V.; Catucci, I.; Zanna, I.; Masala, G.; Bianchi, S.; Spinelli, A.M.; et al. Contribution of MUTYH Variants to Male Breast Cancer Risk: Results From a Multicenter Study in Italy. Front. Oncol. 2018, 8, 583. [Google Scholar] [CrossRef] [PubMed]
- Silvestri, V.; Rizzolo, P.; Zelli, V.; Valentini, V.; Zanna, I.; Bianchi, S.; Tibiletti, M.G.; Varesco, L.; Russo, A.; Tommasi, S.; et al. A possible role of FANCM mutations in male breast cancer susceptibility: Results from a multicenter study in Italy. Breast 2018, 38, 92–97. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.C.; Steele, L.; Kuan, C.-J.; Greilac, S.; Neuhausen, S.L. Mutations in BRCA2 and PALB2 in male breast cancer cases from the United States. Breast Cancer Res. Treat. 2011, 126, 771–778. [Google Scholar] [CrossRef]
- Fackenthal, J.D.; Marsh, D.J.; Richardson, A.L.; Cummings, S.A.; Eng, C.; Robinson, B.G.; Olopade, O.I. Male breast cancer in Cowden syndrome patients with germline PTEN mutations. J. Med. Genet. 2001, 38, 159–164. [Google Scholar] [CrossRef][Green Version]
- Silvestri, V.; Rizzolo, P.; Falchetti, M.; Zanna, I.; Masala, G.; Bianchi, S.; Palli, D.; Ottini, L. Mutation analysis of BRIP1 in male breast cancer cases: A population-based study in Central Italy. Breast Cancer Res. Treat. 2011, 126, 539–543. [Google Scholar] [CrossRef][Green Version]
- Rizzolo, P.; Silvestri, V.; Valentini, V.; Zelli, V.; Bucalo, A.; Zanna, I.; Bianchi, S.; Tibiletti, M.G.; Russo, A.; Varesco, L.; et al. Evaluation of CYP17A1 and CYP1B1 polymorphisms in male breast cancer risk. Endocr. Connect. 2019, 8, 1224–1229. [Google Scholar] [CrossRef]
- Silvestri, V.; Rizzolo, P.; Falchetti, M.; Zanna, I.; Masala, G.; Palli, D.; Ottini, L. Mutation screening of RAD51C in male breast cancer patients. Breast Cancer Res. 2011, 13, 404. [Google Scholar] [CrossRef]
- Rischio Eredo-familiare di Tumore al Seno. Available online: https://salute.regione.emilia-romagna.it/screening/tumori-femminili/screeningmammografico/rischio-eredo-familiare (accessed on 29 April 2020).
- Collegio Italiano dei Senologi Predisposizione Genetica al Tumore Mammario e Geni BRCA1 e BRCA2. Available online: https://www.senologia.it/wp-content/uploads/2019/10/Carcinoma-eredo-familiare-10.19.pdf (accessed on 29 April 2020).
- Tedaldi, G.; Pirini, F.; Tebaldi, M.; Zampiga, V.; Cangini, I.; Danesi, R.; Arcangeli, V.; Ravegnani, M.; Abou Khouzam, R.; Molinari, C.; et al. Multigene Panel Testing Increases the Number of Loci Associated with Gastric Cancer Predisposition. Cancers 2019, 11, 1340. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Li, M.; Hakonarson, H. ANNOVAR: Functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010, 38, e164. [Google Scholar] [CrossRef]
- Plon, S.E.; Eccles, D.M.; Easton, D.; Foulkes, W.D.; Genuardi, M.; Greenblatt, M.S.; Hogervorst, F.B.L.; Hoogerbrugge, N.; Spurdle, A.B.; Tavtigian, S.V.; et al. Sequence variant classification and reporting: Recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum. Mutat. 2008, 29, 1282–1291. [Google Scholar] [CrossRef] [PubMed]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef]
- BRCA Exchange. Available online: https://brcaexchange.org (accessed on 29 April 2020).
- LOVD-BRCA1. Available online: https://databases.lovd.nl/shared/genes/BRCA1 (accessed on 29 April 2020).
- LOVD-BRCA2. Available online: https://databases.lovd.nl/shared/genes/BRCA2 (accessed on 29 April 2020).
- Breast Cancer Information Core (BIC). Available online: https://research.nhgri.nih.gov/bic/ (accessed on 29 April 2020).
- BRCA Share-BRCA1. Available online: http://www.umd.be/BRCA1 (accessed on 29 April 2020).
- BRCA Share-BRCA2. Available online: http://www.umd.be/BRCA2 (accessed on 29 April 2020).
- dbSNP–NCBI–NIH. Available online: https://www.ncbi.nlm.nih.gov/snp/ (accessed on 29 April 2020).
- ClinVar–NCBI–NIH. Available online: https://www.ncbi.nlm.nih.gov/clinvar/ (accessed on 29 April 2020).
- Varsome. Available online: https://varsome.com (accessed on 29 April 2020).
- Breast Equivalent Terms and Definition. Available online: https://seer.cancer.gov/tools/solidtumor/Breast_STM.pdf (accessed on 29 April 2020).
- Swensen, J.; Hoffman, M.; Skolnick, M.H.; Neuhausen, S.L. Identification of a 14 kb deletion involving the promoter region of BRCA1 in a breast cancer family. Hum. Mol. Genet. 1997, 6, 1513–1517. [Google Scholar] [CrossRef]
- Engert, S.; Wappenschmidt, B.; Betz, B.; Kast, K.; Kutsche, M.; Hellebrand, H.; Goecke, T.O.; Kiechle, M.; Niederacher, D.; Schmutzler, R.K.; et al. MLPA screening in the BRCA1 gene from 1,506 German hereditary breast cancer cases: Novel deletions, frequent involvement of exon 17, and occurrence in single early-onset cases. Hum. Mutat. 2008, 29, 948–958. [Google Scholar] [CrossRef]
- Yassaee, V.R.; Emamalizadeh, B.; Omrani, M.D. Screening for genomic rearrangements at BRCA1 locus in Iranian women with breast cancer using multiplex ligation-dependent probe amplification. J. Genet. 2013, 92, 131–134. [Google Scholar] [CrossRef]
- Puget, N.; Stoppa-Lyonnet, D.; Sinilnikova, O.M.; Pagès, S.; Lynch, H.T.; Lenoir, G.M.; Mazoyer, S. Screening for germ-line rearrangements and regulatory mutations in BRCA1 led to the identification of four new deletions. Cancer Res. 1999, 59, 455–461. [Google Scholar]
- Stegel, V.; Krajc, M.; Zgajnar, J.; Teugels, E.; De Grève, J.; Hočevar, M.; Novaković, S. The occurrence of germline BRCA1 and BRCA2 sequence alterations in Slovenian population. BMC Med. Genet. 2011, 12, 9. [Google Scholar] [CrossRef] [PubMed]
- Caux-Moncoutier, V.; Castéra, L.; Tirapo, C.; Michaux, D.; Rémon, M.-A.; Laugé, A.; Rouleau, E.; De Pauw, A.; Buecher, B.; Gauthier-Villars, M.; et al. EMMA, a cost- and time-effective diagnostic method for simultaneous detection of point mutations and large-scale genomic rearrangements: Application to BRCA1 and BRCA2 in 1,525 patients. Hum. Mutat. 2011, 32, 325–334. [Google Scholar] [CrossRef] [PubMed]
- Iyevleva, A.G.; Suspitsin, E.N.; Kroeze, K.; Gorodnova, T.V.; Sokolenko, A.P.; Buslov, K.G.; Voskresenskiy, D.A.; Togo, A.V.; Kovalenko, S.P.; van der Stoep, N.; et al. Non-founder BRCA1 mutations in Russian breast cancer patients. Cancer Lett. 2010, 298, 258–263. [Google Scholar] [CrossRef] [PubMed]
- Arnold, A.G.; Otegbeye, E.; Fleischut, M.H.; Glogowski, E.A.; Siegel, B.; Boyar, S.R.; Salo-Mullen, E.; Amoroso, K.; Sheehan, M.; Berliner, J.L.; et al. Assessment of individuals with BRCA1 and BRCA2 large rearrangements in high-risk breast and ovarian cancer families. Breast Cancer Res. Treat. 2014, 145, 625–634. [Google Scholar] [CrossRef] [PubMed]
- Mangone, L.; Ferrari, F.; Mancuso, P.; Carrozzi, G.; Michiara, M.; Falcini, F.; Piffer, S.; Filiberti, R.A.; Caldarella, A.; Vitale, F.; et al. Epidemiology and biological characteristics of male breast cancer in Italy. Breast Cancer 2020. [Google Scholar] [CrossRef] [PubMed]
- Antoniou, A.; Pharoah, P.D.P.; Narod, S.; Risch, H.A.; Eyfjord, J.E.; Hopper, J.L.; Loman, N.; Olsson, H.; Johannsson, O.; Borg, A.; et al. Average risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case Series unselected for family history: A combined analysis of 22 studies. Am. J. Hum. Genet. 2003, 72, 1117–1130. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Parmigiani, G. Meta-analysis of BRCA1 and BRCA2 penetrance. J. Clin. Oncol. 2007, 25, 1329–1333. [Google Scholar] [CrossRef]
- Mavaddat, N.; Peock, S.; Frost, D.; Ellis, S.; Platte, R.; Fineberg, E.; Evans, D.G.; Izatt, L.; Eeles, R.A.; Adlard, J.; et al. Cancer risks for BRCA1 and BRCA2 mutation carriers: Results from prospective analysis of EMBRACE. J. Natl. Cancer Inst. 2013, 105, 812–822. [Google Scholar] [CrossRef]
- Ford, D.; Easton, D.F.; Bishop, D.T.; Narod, S.A.; Goldgar, D.E. Risks of cancer in BRCA1-mutation carriers. Breast Cancer Linkage Consortium. Lancet 1994, 343, 692–695. [Google Scholar] [CrossRef]
- Leongamornlert, D.; Mahmud, N.; Tymrakiewicz, M.; Saunders, E.; Dadaev, T.; Castro, E.; Goh, C.; Govindasami, K.; Guy, M.; O’Brien, L.; et al. Germline BRCA1 mutations increase prostate cancer risk. Br. J. Cancer 2012, 106, 1697–1701. [Google Scholar] [CrossRef]
- Thompson, D.; Easton, D.F. Breast Cancer Linkage Consortium Cancer Incidence in BRCA1 mutation carriers. J. Natl. Cancer Inst. 2002, 94, 1358–1365. [Google Scholar] [CrossRef] [PubMed]
- Ghiorzo, P.; Pensotti, V.; Fornarini, G.; Sciallero, S.; Battistuzzi, L.; Belli, F.; Bonelli, L.; Borgonovo, G.; Bruno, W.; Gozza, A.; et al. Contribution of germline mutations in the BRCA and PALB2 genes to pancreatic cancer in Italy. Fam. Cancer 2012, 11, 41–47. [Google Scholar] [CrossRef] [PubMed]
- Kote-Jarai, Z.; Leongamornlert, D.; Saunders, E.; Tymrakiewicz, M.; Castro, E.; Mahmud, N.; Guy, M.; Edwards, S.; O’Brien, L.; Sawyer, E.; et al. BRCA2 is a moderate penetrance gene contributing to young-onset prostate cancer: Implications for genetic testing in prostate cancer patients. Br. J. Cancer 2011, 105, 1230–1234. [Google Scholar] [CrossRef] [PubMed]
- Iqbal, J.; Ragone, A.; Lubinski, J.; Lynch, H.T.; Moller, P.; Ghadirian, P.; Foulkes, W.D.; Armel, S.; Eisen, A.; Neuhausen, S.L.; et al. The incidence of pancreatic cancer in BRCA1 and BRCA2 mutation carriers. Br. J. Cancer 2012, 107, 2005–2009. [Google Scholar] [CrossRef] [PubMed]
- Moran, A.; O’Hara, C.; Khan, S.; Shack, L.; Woodward, E.; Maher, E.R.; Lalloo, F.; Evans, D.G.R. Risk of cancer other than breast or ovarian in individuals with BRCA1 and BRCA2 mutations. Fam. Cancer 2012, 11, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Breast Cancer Linkage Consortium. Cancer risks in BRCA2 mutation carriers. J. Natl. Cancer Inst. 1999, 91, 1310–1316. [Google Scholar] [CrossRef]
- National Comprehensive Cancer Network (NCCN). Guidelines: Genetic/Familial High-Risk Assessment: Breast, Ovarian and Pancreatic. Version 1.2020. Available online: https://www.nccn.org/professionals/physician_gls/pdf/genetics_bop.pdf (accessed on 29 April 2020).
- National Comprehensive Cancer Network (NCCN). Guidelines: Prostate Cancer Early Detection. Version 2.2019. Available online: https://www.nccn.org/professionals/physician_gls/pdf/prostate_detection.pdf (accessed on 29 April 2020).
- Pritzlaff, M.; Summerour, P.; McFarland, R.; Li, S.; Reineke, P.; Dolinsky, J.S.; Goldgar, D.E.; Shimelis, H.; Couch, F.J.; Chao, E.C.; et al. Male breast cancer in a multi-gene panel testing cohort: Insights and unexpected results. Breast Cancer Res. Treat. 2017, 161, 575–586. [Google Scholar] [CrossRef]
- Wolpert, N.; Warner, E.; Seminsky, M.F.; Futreal, A.; Narod, S.A. Prevalence of BRCA1 and BRCA2 mutations in male breast cancer patients in Canada. Clin. Breast Cancer 2000, 1, 57–63. [Google Scholar] [CrossRef]
- Silvestri, V.; Barrowdale, D.; Mulligan, A.M.; Neuhausen, S.L.; Fox, S.; Karlan, B.Y.; Mitchell, G.; James, P.; Thull, D.L.; Zorn, K.K.; et al. Male breast cancer in BRCA1 and BRCA2 mutation carriers: Pathology data from the Consortium of Investigators of Modifiers of BRCA1/2. Breast Cancer Res. 2016, 18, 15. [Google Scholar] [CrossRef]
- Antoniou, A.C.; Casadei, S.; Heikkinen, T.; Barrowdale, D.; Pylkäs, K.; Roberts, J.; Lee, A.; Subramanian, D.; De Leeneer, K.; Fostira, F.; et al. Breast-cancer risk in families with mutations in PALB2. N. Engl. J. Med. 2014, 371, 497–506. [Google Scholar] [CrossRef]
- Antoniou, A.C.; Foulkes, W.D.; Tischkowitz, M. Breast-cancer risk in families with mutations in PALB2. N. Engl. J. Med. 2014, 371, 1651–1652. [Google Scholar] [CrossRef] [PubMed]
- Casadei, S.; Norquist, B.M.; Walsh, T.; Stray, S.; Mandell, J.B.; Lee, M.K.; Stamatoyannopoulos, J.A.; King, M.-C. Contribution of inherited mutations in the BRCA2-interacting protein PALB2 to familial breast cancer. Cancer Res. 2011, 71, 2222–2229. [Google Scholar] [CrossRef] [PubMed]
- Adank, M.A.; Jonker, M.A.; Kluijt, I.; van Mil, S.E.; Oldenburg, R.A.; Mooi, W.J.; Hogervorst, F.B.L.; van den Ouweland, A.M.W.; Gille, J.J.P.; Schmidt, M.K.; et al. CHEK2*1100delC homozygosity is associated with a high breast cancer risk in women. J. Med. Genet. 2011, 48, 860–863. [Google Scholar] [CrossRef] [PubMed]
- Walsh, T.; Casadei, S.; Coats, K.H.; Swisher, E.; Stray, S.M.; Higgins, J.; Roach, K.C.; Mandell, J.; Lee, M.K.; Ciernikova, S.; et al. Spectrum of mutations in BRCA1, BRCA2, CHEK2, and TP53 in families at high risk of breast cancer. JAMA 2006, 295, 1379–1388. [Google Scholar] [CrossRef] [PubMed]
- Desrichard, A.; Bidet, Y.; Uhrhammer, N.; Bignon, Y.-J. CHEK2 contribution to hereditary breast cancer in non-BRCA families. Breast Cancer Res. 2011, 13, R119. [Google Scholar] [CrossRef] [PubMed]
- Tedaldi, G.; Danesi, R.; Zampiga, V.; Tebaldi, M.; Bedei, L.; Zoli, W.; Amadori, D.; Falcini, F.; Calistri, D. First evidence of a large CHEK2 duplication involved in cancer predisposition in an Italian family with hereditary breast cancer. BMC Cancer 2014, 14, 478. [Google Scholar] [CrossRef][Green Version]
- Meijers-Heijboer, H.; van den Ouweland, A.; Klijn, J.; Wasielewski, M.; de Snoo, A.; Oldenburg, R.; Hollestelle, A.; Houben, M.; Crepin, E.; van Veghel-Plandsoen, M.; et al. Low-penetrance susceptibility to breast cancer due to CHEK2(*)1100delC in noncarriers of BRCA1 or BRCA2 mutations. Nat. Genet. 2002, 31, 55–59. [Google Scholar]
- Weischer, M.; Bojesen, S.E.; Ellervik, C.; Tybjaerg-Hansen, A.; Nordestgaard, B.G. CHEK2*1100delC genotyping for clinical assessment of breast cancer risk: Meta-analyses of 26,000 patient cases and 27,000 controls. J. Clin. Oncol. 2008, 26, 542–548. [Google Scholar] [CrossRef]
- Cybulski, C.; Wokołorczyk, D.; Jakubowska, A.; Huzarski, T.; Byrski, T.; Gronwald, J.; Masojć, B.; Deebniak, T.; Górski, B.; Blecharz, P.; et al. Risk of breast cancer in women with a CHEK2 mutation with and without a family history of breast cancer. J. Clin. Oncol. 2011, 29, 3747–3752. [Google Scholar] [CrossRef]
- Cybulski, C.; Górski, B.; Huzarski, T.; Masojć, B.; Mierzejewski, M.; Debniak, T.; Teodorczyk, U.; Byrski, T.; Gronwald, J.; Matyjasik, J.; et al. CHEK2 is a multiorgan cancer susceptibility gene. Am. J. Hum. Genet. 2004, 75, 1131–1135. [Google Scholar] [CrossRef]
- Dong, X.; Wang, L.; Taniguchi, K.; Wang, X.; Cunningham, J.M.; McDonnell, S.K.; Qian, C.; Marks, A.F.; Slager, S.L.; Peterson, B.J.; et al. Mutations in CHEK2 associated with prostate cancer risk. Am. J. Hum. Genet. 2003, 72, 270–280. [Google Scholar] [CrossRef] [PubMed]
- Cybulski, C.; Huzarski, T.; Górski, B.; Masojć, B.; Mierzejewski, M.; Debniak, T.; Gliniewicz, B.; Matyjasik, J.; Złowocka, E.; Kurzawski, G.; et al. A novel founder CHEK2 mutation is associated with increased prostate cancer risk. Cancer Res. 2004, 64, 2677–2679. [Google Scholar] [CrossRef] [PubMed]
- Cybulski, C.; Wokołorczyk, D.; Huzarski, T.; Byrski, T.; Gronwald, J.; Górski, B.; Debniak, T.; Masojć, B.; Jakubowska, A.; Gliniewicz, B.; et al. A large germline deletion in the Chek2 kinase gene is associated with an increased risk of prostate cancer. J. Med. Genet. 2006, 43, 863–866. [Google Scholar] [CrossRef] [PubMed]
- Meijers-Heijboer, H.; Wijnen, J.; Vasen, H.; Wasielewski, M.; Wagner, A.; Hollestelle, A.; Elstrodt, F.; van den Bos, R.; de Snoo, A.; Fat, G.T.A.; et al. The CHEK2 1100delC mutation identifies families with a hereditary breast and colorectal cancer phenotype. Am. J. Hum. Genet. 2003, 72, 1308–1314. [Google Scholar] [CrossRef] [PubMed]
- Teodorczyk, U.; Cybulski, C.; Wokołorczyk, D.; Jakubowska, A.; Starzyńska, T.; Lawniczak, M.; Domagała, P.; Ferenc, K.; Marlicz, K.; Banaszkiewicz, Z.; et al. The risk of gastric cancer in carriers of CHEK2 mutations. Fam. Cancer 2013, 12, 473–478. [Google Scholar] [CrossRef] [PubMed]
- CHEK2 Breast Cancer Case-Control Consortium. CHEK2*1100delC and susceptibility to breast cancer: A collaborative analysis involving 10,860 breast cancer cases and 9,065 controls from 10 studies. Am. J. Hum. Genet. 2004, 74, 1175–1182. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, J.L.; Teraoka, S.N.; John, E.M.; Andrulis, I.L.; Knight, J.A.; Lapinski, R.; Olson, E.R.; Wolitzer, A.L.; Seminara, D.; Whittemore, A.S.; et al. The CHEK2*1100delC allelic variant and risk of breast cancer: Screening results from the Breast Cancer Family Registry. Cancer Epidemiol. Biomark. Prev. 2006, 15, 348–352. [Google Scholar] [CrossRef][Green Version]
- Weischer, M.; Bojesen, S.E.; Tybjaerg-Hansen, A.; Axelsson, C.K.; Nordestgaard, B.G. Increased risk of breast cancer associated with CHEK2*1100delC. J. Clin. Oncol. 2007, 25, 57–63. [Google Scholar] [CrossRef]
- Thompson, D.; Duedal, S.; Kirner, J.; McGuffog, L.; Last, J.; Reiman, A.; Byrd, P.; Taylor, M.; Easton, D.F. Cancer risks and mortality in heterozygous ATM mutation carriers. J. Natl. Cancer Inst. 2005, 97, 813–822. [Google Scholar] [CrossRef]
- Renwick, A.; Thompson, D.; Seal, S.; Kelly, P.; Chagtai, T.; Ahmed, M.; North, B.; Jayatilake, H.; Barfoot, R.; Spanova, K.; et al. ATM mutations that cause ataxia-telangiectasia are breast cancer susceptibility alleles. Nat. Genet. 2006, 38, 873–875. [Google Scholar] [CrossRef]
- Goldgar, D.E.; Healey, S.; Dowty, J.G.; Da Silva, L.; Chen, X.; Spurdle, A.B.; Terry, M.B.; Daly, M.J.; Buys, S.M.; Southey, M.C.; et al. Rare variants in the ATM gene and risk of breast cancer. Breast Cancer Res. 2011, 13, R73. [Google Scholar] [CrossRef] [PubMed]
- Marabelli, M.; Cheng, S.-C.; Parmigiani, G. Penetrance of ATM Gene Mutations in Breast Cancer: A Meta-Analysis of Different Measures of Risk. Genet. Epidemiol. 2016, 40, 425–431. [Google Scholar] [CrossRef] [PubMed]
- Van Os, N.J.H.; Roeleveld, N.; Weemaes, C.M.R.; Jongmans, M.C.J.; Janssens, G.O.; Taylor, A.M.R.; Hoogerbrugge, N.; Willemsen, M.A.A.P. Health risks for ataxia-telangiectasia mutated heterozygotes: A systematic review, meta-analysis and evidence-based guideline. Clin. Genet. 2016, 90, 105–117. [Google Scholar] [CrossRef] [PubMed]
- Tavtigian, S.V.; Oefner, P.J.; Babikyan, D.; Hartmann, A.; Healey, S.; Le Calvez-Kelm, F.; Lesueur, F.; Byrnes, G.B.; Chuang, S.-C.; Forey, N.; et al. Rare, evolutionarily unlikely missense substitutions in ATM confer increased risk of breast cancer. Am. J. Hum. Genet. 2009, 85, 427–446. [Google Scholar] [CrossRef] [PubMed]
- Fostira, F.; Saloustros, E.; Apostolou, P.; Vagena, A.; Kalfakakou, D.; Mauri, D.; Tryfonopoulos, D.; Georgoulias, V.; Yannoukakos, D.; Fountzilas, G.; et al. Germline deleterious mutations in genes other than BRCA2 are infrequent in male breast cancer. Breast Cancer Res. Treat. 2018, 169, 105–113. [Google Scholar] [CrossRef]
- Tung, N.; Battelli, C.; Allen, B.; Kaldate, R.; Bhatnagar, S.; Bowles, K.; Timms, K.; Garber, J.E.; Herold, C.; Ellisen, L.; et al. Frequency of mutations in individuals with breast cancer referred for BRCA1 and BRCA2 testing using next-generation sequencing with a 25-gene panel. Cancer 2015, 121, 25–33. [Google Scholar] [CrossRef]
- Loveday, C.; Turnbull, C.; Ruark, E.; Xicola, R.M.M.; Ramsay, E.; Hughes, D.; Warren-Perry, M.; Snape, K.; Breast Cancer Susceptibility Collaboration (UK); Eccles, D.; et al. Germline RAD51C mutations confer susceptibility to ovarian cancer. Nat. Genet. 2012, 44, 475–476. [Google Scholar] [CrossRef]
- Song, H.; Dicks, E.; Ramus, S.J.; Tyrer, J.P.; Intermaggio, M.P.; Hayward, J.; Edlund, C.K.; Conti, D.; Harrington, P.; Fraser, L.; et al. Contribution of Germline Mutations in the RAD51B, RAD51C, and RAD51D Genes to Ovarian Cancer in the Population. J. Clin. Oncol. 2015, 33, 2901–2907. [Google Scholar] [CrossRef]
- Akbari, M.R.; Tonin, P.; Foulkes, W.D.; Ghadirian, P.; Tischkowitz, M.; Narod, S.A. RAD51C germline mutations in breast and ovarian cancer patients. Breast Cancer Res. 2010, 12, 404. [Google Scholar] [CrossRef]
- Meindl, A.; Hellebrand, H.; Wiek, C.; Erven, V.; Wappenschmidt, B.; Niederacher, D.; Freund, M.; Lichtner, P.; Hartmann, L.; Schaal, H.; et al. Germline mutations in breast and ovarian cancer pedigrees establish RAD51C as a human cancer susceptibility gene. Nat. Genet. 2010, 42, 410–414. [Google Scholar] [CrossRef]
- Jensen, D.E.; Proctor, M.; Marquis, S.T.; Gardner, H.P.; Ha, S.I.; Chodosh, L.A.; Ishov, A.M.; Tommerup, N.; Vissing, H.; Sekido, Y.; et al. BAP1: A novel ubiquitin hydrolase which binds to the BRCA1 RING finger and enhances BRCA1-mediated cell growth suppression. Oncogene 1998, 16, 1097–1112. [Google Scholar] [CrossRef] [PubMed]
- Pilarski, R.; Rai, K.; Cebulla, C.; Abdel-Rahman, M. BAP1 Tumor Predisposition Syndrome. In GeneReviews®; University of Washington: Seattle, WA, USA, 2016; (updated 2020). [Google Scholar]
- Testa, J.R.; Cheung, M.; Pei, J.; Below, J.E.; Tan, Y.; Sementino, E.; Cox, N.J.; Dogan, A.U.; Pass, H.I.; Trusa, S.; et al. Germline BAP1 mutations predispose to malignant mesothelioma. Nat. Genet. 2011, 43, 1022–1025. [Google Scholar] [CrossRef] [PubMed]
- Njauw, C.-N.J.; Kim, I.; Piris, A.; Gabree, M.; Taylor, M.; Lane, A.M.; DeAngelis, M.M.; Gragoudas, E.; Duncan, L.M.; Tsao, H. Germline BAP1 inactivation is preferentially associated with metastatic ocular melanoma and cutaneous-ocular melanoma families. PLoS ONE 2012, 7, e35295. [Google Scholar] [CrossRef] [PubMed]
- Popova, T.; Hebert, L.; Jacquemin, V.; Gad, S.; Caux-Moncoutier, V.; Dubois-d’Enghien, C.; Richaudeau, B.; Renaudin, X.; Sellers, J.; Nicolas, A.; et al. Germline BAP1 mutations predispose to renal cell carcinomas. Am. J. Hum. Genet. 2013, 92, 974–980. [Google Scholar] [CrossRef] [PubMed]
- Pilarski, R.; Cebulla, C.M.; Massengill, J.B.; Rai, K.; Rich, T.; Strong, L.; McGillivray, B.; Asrat, M.-J.; Davidorf, F.H.; Abdel-Rahman, M.H. Expanding the clinical phenotype of hereditary BAP1 cancer predisposition syndrome, reporting three new cases. Genes. Chromosomes Cancer 2014, 53, 177–182. [Google Scholar] [CrossRef]
- Coupier, I.; Cousin, P.-Y.; Hughes, D.; Legoix-Né, P.; Trehin, A.; Sinilnikova, O.M.; Stoppa-Lyonnet, D. BAP1 and breast cancer risk. Fam. Cancer 2005, 4, 273–277. [Google Scholar] [CrossRef]
- Wood, L.D.; Parsons, D.W.; Jones, S.; Lin, J.; Sjöblom, T.; Leary, R.J.; Shen, D.; Boca, S.M.; Barber, T.; Ptak, J.; et al. The genomic landscapes of human breast and colorectal cancers. Science 2007, 318, 1108–1113. [Google Scholar] [CrossRef]
- Lynch, T.J.; Bell, D.W.; Sordella, R.; Gurubhagavatula, S.; Okimoto, R.A.; Brannigan, B.W.; Harris, P.L.; Haserlat, S.M.; Supko, J.G.; Haluska, F.G.; et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N. Engl. J. Med. 2004, 350, 2129–2139. [Google Scholar] [CrossRef]
- Bell, D.W.; Gore, I.; Okimoto, R.A.; Godin-Heymann, N.; Sordella, R.; Mulloy, R.; Sharma, S.V.; Brannigan, B.W.; Mohapatra, G.; Settleman, J.; et al. Inherited susceptibility to lung cancer may be associated with the T790M drug resistance mutation in EGFR. Nat. Genet. 2005, 37, 1315–1316. [Google Scholar] [CrossRef]
- Van der Leest, C.; Wagner, A.; Pedrosa, R.M.; Aerts, J.G.; Dinjens, W.N.M.; Dubbink, H.J. Novel EGFR V834L Germline Mutation Associated With Familial Lung Adenocarcinoma. JCO Precis. Oncol. 2018, 2, 1–5. [Google Scholar] [CrossRef]
- Ohtsuka, K.; Ohnishi, H.; Fujiwara, M.; Morii, T.; Matsushima, S.; Ogura, W.; Yamasaki, S.; Kishino, T.; Tanaka, R.; Watanabe, T. Predisposition to Lung Adenocarcinoma in a Family Harboring the Germline EGFR V843I Mutation. JCO Precis. Oncol. 2019, 3, 1–4. [Google Scholar] [CrossRef]
- Oxnard, G.R.; Nguyen, K.-S.H.; Costa, D.B. Germline mutations in driver oncogenes and inherited lung cancer risk independent of smoking history. J. Natl. Cancer Inst. 2014, 106, djt361. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ikeda, K.; Nomori, H.; Mori, T.; Sasaki, J.; Kobayashi, T. Novel germline mutation: EGFR V843I in patient with multiple lung adenocarcinomas and family members with lung cancer. Ann. Thorac. Surg. 2008, 85, 1430–1432. [Google Scholar] [CrossRef] [PubMed]
- Ohtsuka, K.; Ohnishi, H.; Kurai, D.; Matsushima, S.; Morishita, Y.; Shinonaga, M.; Goto, H.; Watanabe, T. Familial lung adenocarcinoma caused by the EGFR V843I germ-line mutation. J. Clin. Oncol. 2011, 29, e191–e192. [Google Scholar] [CrossRef]
- Demierre, N.; Zoete, V.; Michielin, O.; Stauffer, E.; Zimmermann, D.R.; Betticher, D.C.; Peters, S. A dramatic lung cancer course in a patient with a rare EGFR germline mutation exon 21 V843I: Is EGFR TKI resistance predictable? Lung Cancer 2013, 80, 81–84. [Google Scholar] [CrossRef]
- Campbell, P.; Morton, P.E.; Takeichi, T.; Salam, A.; Roberts, N.; Proudfoot, L.E.; Mellerio, J.E.; Aminu, K.; Wellington, C.; Patil, S.N.; et al. Epithelial inflammation resulting from an inherited loss-of-function mutation in EGFR. J. Investig. Dermatol. 2014, 134, 2570–2578. [Google Scholar] [CrossRef]
- Ganetzky, R.; Finn, E.; Bagchi, A.; Zollo, O.; Conlin, L.; Deardorff, M.; Harr, M.; Simpson, M.A.; McGrath, J.A.; Zackai, E.; et al. EGFR mutations cause a lethal syndrome of epithelial dysfunction with progeroid features. Mol. Genet. Genom. Med. 2015, 3, 452–458. [Google Scholar] [CrossRef]
- Downward, J.; Parker, P.; Waterfield, M.D. Autophosphorylation sites on the epidermal growth factor receptor. Nature 1984, 311, 483–485. [Google Scholar] [CrossRef]
| Genes | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| AIP | ALK | APC | ATM | BAP1 | BLM | BMPR1A | BRCA1 | BRCA2 | BRIP1 |
| BUB1B | CDC73 | CDH1 | CDK4 | CDKN1C | CDKN2A | CEBPA | CEP57 | CHEK2 | CYLD |
| DDB2 | DICER1 | DIS3L2 | EGFR | EPCAM | ERCC2 | ERCC3 | ERCC4 | ERCC5 | EXT1 |
| EXT2 | EZH2 | FANCA | FANCB | FANCC | FANCD2 | FANCE | FANCF | FANCG | FANCI |
| FANCL | FANCM | FH | FLCN | GATA2 | GPC3 | HNF1A | HRAS | KIT | MAX |
| MEN1 | MET | MLH1 | MSH2 | MSH6 | MUTYH | NBN | NF1 | NF2 | NSD1 |
| PALB2 | PHOX2B | PMS1 | PMS2 | PRF1 | PRKAR1A | PTCH1 | PTEN | RAD51C | RAD51D |
| RB1 | RECQL4 | RET | RHBDF2 | RUNX1 | SBDS | SDHAF2 | SDHB | SDHC | SDHD |
| SLX4 | SMAD4 | SMARCB1 | STK11 | SUFU | TMEM127 | TP53 | TSC1 | TSC2 | VHL |
| WRN | WT1 | XPA | XPC | ||||||
| Patient ID | Cancer | Age at Onset | Gene | Chr | cDNA (Transcript) | Protein | Variant Type | IARC Class [38] | dbSNP [46] | ClinVar [47] |
|---|---|---|---|---|---|---|---|---|---|---|
| A142 | IDC | 55y | BRCA1 | 17q21.31 | c.-113-?_80+?del (NM_007294) | p.? | large deletion | 5 | – | – |
| A774 | IDC | 69y | BRCA1 | 17q21.31 | c.4964_4982del (NM_007294) | p.Ser1655Tyrfs*16 | frameshift deletion | 5 | rs80359876 | pathogenic |
| TR140 | IDC | 57y | BRCA1 | 17q21.31 | c.5266dupC (NM_007294) | p.Gln1756Profs*74 | frameshift duplication | 5 | rs80357906 | pathogenic |
| A379 | IDC | 58y | BRCA2 | 13q13.1 | c.1238delT (NM_000059) | p.Leu413Hisfs*17 | frameshift deletion | 5 | rs80359271 | pathogenic |
| A581 | IDC | 77y | BRCA2 | 13q13.1 | c.1813delA (NM_000059) | p.Ile605Tyrfs*9 | frameshift deletion | 5 | rs80359306 | pathogenic |
| T096 | DCIS | 68y | BRCA2 | 13q13.1 | c.3195_3198delTAAT (NM_000059) | p.Asn1066Leufs*10 | frameshift deletion | 5 | rs80359375 | pathogenic |
| B156 | IDC | 64y | BRCA2 | 13q13.1 | c.5073dupA (NM_000059) | p.Trp1692Metfs*3 | frameshift duplication | 5 | rs80359479 | pathogenic |
| A933 | IDC | 59y | BRCA2 | 13q13.1 | c.6039delA (NM_000059) | p.Val2014Tyrfs*26 | frameshift deletion | 5 | rs876660637 | pathogenic |
| A98 | IDC | 56y | BRCA2 | 13q13.1 | c.6039delA (NM_000059) | p.Val2014Tyrfs*26 | frameshift deletion | 5 | rs876660637 | pathogenic |
| Patient ID | Cancer | Age at Onset | Gene | Chr | cDNA | Protein | Variant Type | IARC Class [38] | dbSNP [46] | ClinVar [47] |
|---|---|---|---|---|---|---|---|---|---|---|
| A841 | IDC | 38y | ATM | 11q22.3 | c.8319_8323dupTGTCC (NM_000051) | p.Pro2775Leufs*33 | frameshift duplication | 5 | rs1555135596 | pathogenic |
| A625 | IDC | 65y | BAP1 | 3p21.1 | c.1110_1116delCATGCAG (NM_004656) | p.Met371Argfs*57 | frameshift deletion | 4 | – | – |
| A512 | IDC | 36y | CHEK2 | 22q12.1 | c.1100delC (NM_007194) | p.Thr367Metfs*15 | frameshift deletion | 5 | rs555607708 | pathogenic |
| A225 | DCIS | 62y | EGFR | 7p11.2 | c.3538_3541delGAAG (NM_005228) | p.Glu1180Profs*18 | frameshift deletion | 4 | rs781064539 | – |
| B887 | IDC | 75y | PALB2 | 16p12.2 | c.73A>T (NM_024675) | p.Lys25* | nonsense variant | 5 | rs1248579792 | pathogenic |
| A334 | IDC | 59y | RAD51C | 17q22 | c.181_182delCT (NM_058216) | p.Leu61Alafs*11 | frameshift deletion | 5 | rs786203945 | pathogenic |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tedaldi, G.; Tebaldi, M.; Zampiga, V.; Cangini, I.; Pirini, F.; Ferracci, E.; Danesi, R.; Arcangeli, V.; Ravegnani, M.; Martinelli, G.; et al. Male Breast Cancer: Results of the Application of Multigene Panel Testing to an Italian Cohort of Patients. Diagnostics 2020, 10, 269. https://doi.org/10.3390/diagnostics10050269
Tedaldi G, Tebaldi M, Zampiga V, Cangini I, Pirini F, Ferracci E, Danesi R, Arcangeli V, Ravegnani M, Martinelli G, et al. Male Breast Cancer: Results of the Application of Multigene Panel Testing to an Italian Cohort of Patients. Diagnostics. 2020; 10(5):269. https://doi.org/10.3390/diagnostics10050269
Chicago/Turabian StyleTedaldi, Gianluca, Michela Tebaldi, Valentina Zampiga, Ilaria Cangini, Francesca Pirini, Elisa Ferracci, Rita Danesi, Valentina Arcangeli, Mila Ravegnani, Giovanni Martinelli, and et al. 2020. "Male Breast Cancer: Results of the Application of Multigene Panel Testing to an Italian Cohort of Patients" Diagnostics 10, no. 5: 269. https://doi.org/10.3390/diagnostics10050269
APA StyleTedaldi, G., Tebaldi, M., Zampiga, V., Cangini, I., Pirini, F., Ferracci, E., Danesi, R., Arcangeli, V., Ravegnani, M., Martinelli, G., Falcini, F., Ulivi, P., & Calistri, D. (2020). Male Breast Cancer: Results of the Application of Multigene Panel Testing to an Italian Cohort of Patients. Diagnostics, 10(5), 269. https://doi.org/10.3390/diagnostics10050269







