Diagnostic, Prognostic, and Therapeutic Value of Non-Coding RNA Expression Profiles in Renal Transplantation
Abstract
1. Introduction
2. Molecular Profiling in Nephrology and Transplantation
3. Non-Coding RNA Profiles as Predictors of Renal Phenotypes
3.1. Post-Transplant AKI
3.2. Allograft Rejection
4. Circulating Non-Coding RNAs in Transplantation
4.1. MicroRNAs
4.2. Long Non-Coding RNAs
4.3. Circular RNAs
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Luyckx, V.A.; Tonelli, M.; Stanifer, J.W. The global burden of kidney disease and the sustainable development goals. Bull World Health Organ. 2018, 96, 414–422D. [Google Scholar] [CrossRef] [PubMed]
- Liyanage, T.; Ninomiya, T.; Jha, V.; Neal, B.; Patrice, H.M.; Okpechi, I.; Zhao, M.H.; Lv, J.; Garg, A.X.; Knight, J.; et al. Worldwide access to treatment for end-stage kidney disease: A systematic review. Lancet 2015, 385, 1975–1982. [Google Scholar] [CrossRef]
- Laupacis, A.; Keown, P.; Pus, N.; Krueger, H.; Ferguson, B.; Wong, C.; Muirhead, N. A study of the quality of life and cost-utility of renal transplantation. Kidney Int. 1996, 50, 235–242. [Google Scholar] [CrossRef]
- Loupy, A.; Lefaucheur, C. Antibody-Mediated Rejection of Solid-Organ Allografts. N. Engl. J. Med. 2018, 379, 1150–1160. [Google Scholar] [CrossRef] [PubMed]
- Kaikkonen, M.U.; Adelman, K. Emerging Roles of Non-Coding RNA Transcription. Trends Biochem. Sci. 2018, 43, 654–667. [Google Scholar] [CrossRef] [PubMed]
- Pennisi, E. Genomics. ENCODE project writes eulogy for junk DNA. Science 2012, 337, 1159–1161. [Google Scholar] [CrossRef] [PubMed]
- Brandenburger, T.; Salgado Somoza, A.; Devaux, Y.; Lorenzen, J.M. Noncoding RNAs in acute kidney injury. Kidney Int. 2018, 94, 870–881. [Google Scholar] [CrossRef] [PubMed]
- Kato, M. Noncoding RNAs as therapeutic targets in early stage diabetic kidney disease. Kidney Res. Clin. Pract. 2018, 37, 197–209. [Google Scholar] [CrossRef]
- Ignarski, M.; Islam, R.; Müller, R.U. Long Non-Coding RNAs in Kidney Disease. Int. J. Mol. Sci. 2019, 20. [Google Scholar] [CrossRef]
- Memczak, S.; Papavasileiou, P.; Peters, O.; Rajewsky, N. Identification and Characterization of Circular RNAs As a New Class of Putative Biomarkers in Human Blood. PLoS ONE 2015, 10, e0141214. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef]
- Lytle, J.R.; Yario, T.A.; Steitz, J.A. Target mRNAs are repressed as efficiently by microRNA-binding sites in the 5’ UTR as in the 3’ UTR. Proc. Natl. Acad. Sci. USA 2007, 104, 9667–9672. [Google Scholar] [CrossRef] [PubMed]
- Cipolla, G.A. A non-canonical landscape of the microRNA system. Front Genet. 2014, 5, 337. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Wang, Y.; Shu, S.; Cai, J.; Tang, C.; Dong, Z. Non-coding RNAs in kidney injury and repair. Am. J. Physiol. Cell Physiol. 2019, 317, C177–C188. [Google Scholar] [CrossRef] [PubMed]
- Jeck, W.R.; Sorrentino, J.A.; Wang, K.; Slevin, M.K.; Burd, C.E.; Liu, J.; Marzluff, W.F.; Sharpless, N.E. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 2013, 19, 141–157. [Google Scholar] [CrossRef] [PubMed]
- Cocquerelle, C.; Mascrez, B.; Hétuin, D.; Bailleul, B. Mis-splicing yields circular RNA molecules. FASEB J. 1993, 7, 155–160. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, K.Y.; Sun, H.S.; Tsai, S.J. Circular RNA—New member of noncoding RNA with novel functions. Exp. Biol. Med. (Maywood) 2017, 242, 1136–1141. [Google Scholar] [CrossRef]
- Li, Z.; Huang, C.; Bao, C.; Chen, L.; Lin, M.; Wang, X.; Zhong, G.; Yu, B.; Hu, W.; Dai, L.; et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 2015, 22, 256–264. [Google Scholar] [CrossRef]
- Huang, S.; Yang, B.; Chen, B.J.; Bliim, N.; Ueberham, U.; Arendt, T.; Janitz, M. The emerging role of circular RNAs in transcriptome regulation. Genomics 2017, 109, 401–407. [Google Scholar] [CrossRef]
- Haddad, G.; Lorenzen, J.M. Biogenesis and Function of Circular RNAs in Health and in Disease. Front. Pharmacol. 2019, 10, 428. [Google Scholar] [CrossRef]
- Zheng, Q.; Bao, C.; Guo, W.; Li, S.; Chen, J.; Chen, B.; Luo, Y.; Lyu, D.; Li, Y.; Shi, G.; et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat. Commun. 2016, 7, 11215. [Google Scholar] [CrossRef] [PubMed]
- Du, W.W.; Zhang, C.; Yang, W.; Yong, T.; Awan, F.M.; Yang, B.B. Identifying and Characterizing circRNA-Protein Interaction. Theranostics 2017, 7, 4183–4191. [Google Scholar] [CrossRef] [PubMed]
- Luan, J.; Jiao, C.; Kong, W.; Fu, J.; Qu, W.; Chen, Y.; Zhu, X.; Zeng, Y.; Guo, G.; Qi, H.; et al. circHLA-C Plays an Important Role in Lupus Nephritis by Sponging miR-150. Mol. Ther. Nucleic Acids 2018, 10, 245–253. [Google Scholar] [CrossRef] [PubMed]
- Kölling, M.; Seeger, H.; Haddad, G.; Kistler, A.; Nowak, A.; Faulhaber-Walter, R.; Kielstein, J.; Haller, H.; Fliser, D.; Mueller, T.; et al. The Circular RNA ciRs-126 Predicts Survival in Critically Ill Patients With Acute Kidney Injury. Kidney Int. Rep. 2018, 3, 1144–1152. [Google Scholar] [CrossRef]
- Wang, K.C.; Chang, H.Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 2011, 43, 904–914. [Google Scholar] [CrossRef]
- Mongelli, A.; Martelli, F.; Farsetti, A.; Gaetano, C. The Dark That Matters: Long Non-coding RNAs as Master Regulators of Cellular Metabolism in Non-communicable Diseases. Front. Physiol. 2019, 10, 369. [Google Scholar] [CrossRef]
- Lorenzen, J.M.; Thum, T. Long noncoding RNAs in kidney and cardiovascular diseases. Nat. Rev. Nephrol. 2016, 12, 360–373. [Google Scholar] [CrossRef]
- Ge, Y.Z.; Xu, T.; Cao, W.J.; Wu, R.; Yao, W.T.; Zhou, C.C.; Wang, M.; Xu, L.W.; Lu, T.Z.; Zhao, Y.C.; et al. A Molecular Signature of Two Long Non-Coding RNAs in Peripheral Blood Predicts Acute Renal Allograft Rejection. Cell. Physiol. Biochem. 2017, 44, 1213–1223. [Google Scholar] [CrossRef]
- Jiang, X.; Zhang, F. Long noncoding RNA: A new contributor and potential therapeutic target in fibrosis. Epigenomics 2017, 9, 1233–1241. [Google Scholar] [CrossRef]
- Saez-Rodriguez, J.; Rinschen, M.M.; Floege, J.; Kramann, R. Big science and big data in nephrology. Kidney Int. 2019, 95, 1326–1337. [Google Scholar] [CrossRef]
- Kretzler, M.; Cohen, C.D.; Doran, P.; Henger, A.; Madden, S.; Gröne, E.F.; Nelson, P.J.; Schlöndorff, D.; Gröne, H.J. Repuncturing the renal biopsy: Strategies for molecular diagnosis in nephrology. J. Am. Soc. Nephrol. 2002, 13, 1961–1972. [Google Scholar] [CrossRef] [PubMed]
- Yasuda, Y.; Cohen, C.D.; Henger, A.; Kretzler, M. European Renal cDNA Bank (ERCB) Consortium. Gene expression profiling analysis in nephrology: Towards molecular definition of renal disease. Clin. Exp. Nephrol. 2006, 10, 91–98. [Google Scholar] [CrossRef] [PubMed]
- Halloran, P.F.; Merino Lopez, M.; Barreto Pereira, A. Identifying Subphenotypes of Antibody-Mediated Rejection in Kidney Transplants. Am. J. Transplant. 2016, 16, 908–920. [Google Scholar] [CrossRef] [PubMed]
- Halloran, P.F.; Pereira, A.B.; Chang, J.; Matas, A.; Picton, M.; De Freitas, D.; Bromberg, J.; Serón, D.; Sellarés, J.; Einecke, G.; et al. Microarray diagnosis of antibody-mediated rejection in kidney transplant biopsies: An international prospective study (INTERCOM). Am. J. Transplant. 2013, 13, 2865–2874. [Google Scholar] [CrossRef] [PubMed]
- Halloran, P.F.; Famulski, K.S.; Reeve, J. Molecular assessment of disease states in kidney transplant biopsy samples. Nat. Rev. Nephrol. 2016, 12, 534–548. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Yi, Z.; Wei, C.; Keung, K.L.; Sun, Z.; Xi, C.; Woytovich, C.; Farouk, S.; Gallon, L.; Menon, M.C.; et al. Pretransplant transcriptomic signature in peripheral blood predicts early acute rejection. JCI Insight 2019, 4. [Google Scholar] [CrossRef]
- Galichon, P.; Xu-Dubois, Y.C.; Buob, D.; Tinel, C.; Anglicheau, D.; Benbouzid, S.; Dahan, K.; Ouali, N.; Hertig, A.; Brocheriou, I.; et al. Urinary transcriptomics reveals patterns associated with subclinical injury of the renal allograft. Biomark. Med. 2018, 12, 427–438. [Google Scholar] [CrossRef]
- Reeve, J.; Böhmig, G.A.; Eskandary, F.; Einecke, G.; Lefaucheur, C.; Loupy, A.; Halloran, P.F.; MMDx-Kidney study group. Assessing rejection-related disease in kidney transplant biopsies based on archetypal analysis of molecular phenotypes. JCI Insight 2017, 2. [Google Scholar] [CrossRef]
- Chen, W.; Peng, W.; Huang, J.; Yu, X.; Tan, K.; Chen, Y.; Lin, X.; Chen, D.; Dai, Y. Microarray analysis of long non-coding RNA expression in human acute rejection biopsy samples following renal transplantation. Mol. Med. Rep. 2014, 10, 2210–2216. [Google Scholar] [CrossRef]
- Wilflingseder, J.; Regele, H.; Perco, P.; Kainz, A.; Soleiman, A.; Mühlbacher, F.; Mayer, B.; Oberbauer, R. miRNA profiling discriminates types of rejection and injury in human renal allografts. Transplantation 2013, 95, 835–841. [Google Scholar] [CrossRef]
- Wilflingseder, J.; Sunzenauer, J.; Toronyi, E.; Heinzel, A.; Kainz, A.; Mayer, B.; Perco, P.; Telkes, G.; Langer, R.M.; Oberbauer, R. Molecular pathogenesis of post-transplant acute kidney injury: Assessment of whole-genome mRNA and miRNA profiles. PLoS ONE 2014, 9, e104164. [Google Scholar] [CrossRef] [PubMed]
- Milhoransa, P.; Montanari, C.C.; Montenegro, R.; Manfro, R.C. Micro RNA 146a-5p expression in Kidney transplant recipients with delayed graft function. J. Bras Nefrol 2019, 41, 242–251. [Google Scholar] [CrossRef] [PubMed]
- Roest, H.P.; Ooms, L.S.S.; Gillis, A.J.M.; IJzermans, J.N.M.; Looijenga, L.H.J.; Dorssers, L.C.J.; Dor, F.J.M.F.; van der Laan, L.J.W. Cell-free MicroRNA miR-505-3p in Graft Preservation Fluid Is an Independent Predictor of Delayed Graft Function After Kidney Transplantation. Transplantation 2019, 103, 329–335. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Dos-Santos, V.; Ramos-Muñoz, E.; García-Bermejo, M.L.; Ruiz-Hernández, M.; Rodríguez-Serrano, E.M.; Saiz-González, A.; Martínez-Perez, A.; Burgos-Revilla, F.J. MicroRNAs in Kidney Machine Perfusion Fluid as Novel Biomarkers for Graft Function. Normalization Methods for miRNAs Profile Analysis. Transplant. Proc. 2019, 51, 307–310. [Google Scholar] [CrossRef]
- Wang, J.; Li, X.; Wu, X.; Wang, Z.; Zhang, C.; Cao, G.; Yan, T. Expression Profiling of Exosomal miRNAs Derived from the Peripheral Blood of Kidney Recipients with DGF Using High-Throughput Sequencing. BioMed Res. Int. 2019, 2019, 1759697. [Google Scholar] [CrossRef] [PubMed]
- Sui, W.; Dai, Y.; Huang, Y.; Lan, H.; Yan, Q.; Huang, H. Microarray analysis of MicroRNA expression in acute rejection after renal transplantation. Transpl. Immunol. 2008, 19, 81–85. [Google Scholar] [CrossRef]
- Anglicheau, D.; Sharma, V.K.; Ding, R.; Hummel, A.; Snopkowski, C.; Dadhania, D.; Seshan, S.V.; Suthanthiran, M. MicroRNA expression profiles predictive of human renal allograft status. Proc. Natl. Acad. Sci. USA 2009, 106, 5330–5335. [Google Scholar] [CrossRef]
- Soltaninejad, E.; Nicknam, M.H.; Nafar, M.; Ahmadpoor, P.; Pourrezagholi, F.; Sharbafi, M.H.; Hosseinzadeh, M.; Foroughi, F.; Yekaninejad, M.S.; Bahrami, T.; et al. Differential expression of microRNAs in renal transplant patients with acute T-cell mediated rejection. Transpl. Immunol. 2015, 33, 1–6. [Google Scholar] [CrossRef]
- Sui, W.; Lin, H.; Peng, W.; Huang, Y.; Chen, J.; Zhang, Y.; Dai, Y. Molecular dysfunctions in acute rejection after renal transplantation revealed by integrated analysis of transcription factor, microRNA and long noncoding RNA. Genomics 2013, 102, 310–322. [Google Scholar] [CrossRef]
- Oghumu, S.; Bracewell, A.; Nori, U.; Maclean, K.H.; Balada-Lasat, J.M.; Brodsky, S.; Pelletier, R.; Henry, M.; Satoskar, A.R.; Nadasdy, T.; et al. Acute pyelonephritis in renal allografts: A new role for microRNAs? Transplantation 2014, 97, 559–568. [Google Scholar] [CrossRef]
- Scian, M.J.; Maluf, D.G.; David, K.G.; Archer, K.J.; Suh, J.L.; Wolen, A.R.; Mba, M.U.; Massey, H.D.; King, A.L.; Gehr, T.; et al. MicroRNA profiles in allograft tissues and paired urines associate with chronic allograft dysfunction with IF/TA. Am. J. Transplant. 2011, 11, 2110–2122. [Google Scholar] [CrossRef] [PubMed]
- Heinemann, F.M.; Jindra, P.T.; Bockmeyer, C.L.; Zeuschner, P.; Wittig, J.; Höflich, H.; Eßer, M.; Abbas, M.; Dieplinger, G.; Stolle, K.; et al. Glomerulocapillary miRNA response to HLA-class I antibody in vitro and in vivo. Sci. Rep. 2017, 7, 14554. [Google Scholar] [CrossRef] [PubMed]
- Tao, J.; Yang, X.; Han, Z.; Lu, P.; Wang, J.; Liu, X.; Wu, B.; Wang, Z.; Huang, Z.; Lu, Q.; et al. Serum MicroRNA-99a Helps Detect Acute Rejection in Renal Transplantation. Transplant. Proc. 2015, 47, 1683–1687. [Google Scholar] [CrossRef] [PubMed]
- Ledeganck, K.J.; Gielis, E.M.; Abramowicz, D.; Stenvinkel, P.; Shiels, P.G.; Van Craenenbroeck, A.H. MicroRNAs in AKI and Kidney Transplantation. Clin. J. Am. Soc. Nephrol. 2019, 14, 454–468. [Google Scholar] [CrossRef]
- Ben-Dov, I.Z.; Muthukumar, T.; Morozov, P.; Mueller, F.B.; Tuschl, T.; Suthanthiran, M. MicroRNA sequence profiles of human kidney allografts with or without tubulointerstitial fibrosis. Transplantation 2012, 94, 1086–1094. [Google Scholar] [CrossRef]
- Zununi Vahed, S.; Omidi, Y.; Ardalan, M.; Samadi, N. Dysregulation of urinary miR-21 and miR-200b associated with interstitial fibrosis and tubular atrophy (IFTA) in renal transplant recipients. Clin. Biochem. 2017, 50, 32–39. [Google Scholar] [CrossRef]
- Janszky, N.; Süsal, C. Circulating and urinary microRNAs as possible biomarkers in kidney transplantation. Transplant. Rev. (Orlando) 2018, 32, 110–118. [Google Scholar] [CrossRef]
- Ulbing, M.; Kirsch, A.H.; Leber, B.; Lemesch, S.; Münzker, J.; Schweighofer, N.; Hofer, D.; Trummer, O.; Rosenkranz, A.R.; Müller, H.; et al. MicroRNAs 223-3p and 93-5p in patients with chronic kidney disease before and after renal transplantation. Bone 2017, 95, 115–123. [Google Scholar] [CrossRef]
- Lorenzen, J.M.; Volkmann, I.; Fiedler, J.; Schmidt, M.; Scheffner, I.; Haller, H.; Gwinner, W.; Thum, T. Urinary miR-210 as a mediator of acute T-cell mediated rejection in renal allograft recipients. Am. J. Transplant. 2011, 11, 2221–2227. [Google Scholar] [CrossRef]
- Qiu, J.; Chen, Y.; Huang, G.; Zhang, Z.; Chen, L.; Na, N. Transforming growth factor-β activated long non-coding RNA ATB plays an important role in acute rejection of renal allografts and may impacts the postoperative pharmaceutical immunosuppression therapy. Nephrology (Carlton) 2017, 22, 796–803. [Google Scholar] [CrossRef]
- Zou, Y.; Zhang, W.; Zhou, H.H.; Liu, R. Analysis of long noncoding RNAs for acute rejection and graft outcome in kidney transplant biopsies. Biomark. Med. 2019, 13, 185–195. [Google Scholar] [CrossRef]
- Xu, J.; Hu, J.; Xu, H.; Zhou, H.; Liu, Z.; Zhou, Y.; Liu, R.; Zhang, W. Long Non-coding RNA Expression Profiling in Biopsy to Identify Renal Allograft at Risk of Chronic Damage and Future Graft Loss. Appl. Biochem. Biotechnol. 2019. [Google Scholar] [CrossRef]
- Lorenzen, J.M.; Schauerte, C.; Kölling, M.; Hübner, A.; Knapp, M.; Haller, H.; Thum, T. Long Noncoding RNAs in Urine Are Detectable and May Enable Early Detection of Acute T Cell-Mediated Rejection of Renal Allografts. Clin. Chem. 2015, 61, 1505–1514. [Google Scholar] [CrossRef]
- Nagarajah, S.; Xia, S.; Rasmussen, M.; Tepel, M. Endogenous intronic antisense long non-coding RNA, MGAT3-AS1, and kidney transplantation. Sci. Rep. 2019, 9, 14743. [Google Scholar] [CrossRef]
- Kölling, M.; Haddad, G.; Wegmann, U.; Kistler, A.; Bosakova, A.; Seeger, H.; Hübel, K.; Haller, H.; Mueller, T.; Wüthrich, R.P.; et al. Circular RNAs in Urine of Kidney Transplant Patients with Acute T Cell-Mediated Allograft Rejection. Clin. Chem. 2019, 65, 1287–1294. [Google Scholar] [CrossRef]
- Yarlagadda, S.G.; Coca, S.G.; Formica, R.N.; Poggio, E.D.; Parikh, C.R. Association between delayed graft function and allograft and patient survival: A systematic review and meta-analysis. Nephrol. Dial. Transplant. 2009, 24, 1039–1047. [Google Scholar] [CrossRef]
- Hall, I.E.; Schröppel, B.; Doshi, M.D.; Ficek, J.; Weng, F.L.; Hasz, R.D.; Thiessen-Philbrook, H.; Reese, P.P.; Parikh, C.R. Associations of deceased donor kidney injury with kidney discard and function after transplantation. Am. J. Transplant. 2015, 15, 1623–1631. [Google Scholar] [CrossRef]
- Menke, J.; Sollinger, D.; Schamberger, B.; Heemann, U.; Lutz, J. The effect of ischemia/reperfusion on the kidney graft. Curr. Opin. Organ Transplant. 2014, 19, 395–400. [Google Scholar] [CrossRef]
- Wilflingseder, J.; Jelencsics, K.; Bergmeister, H.; Sunzenauer, J.; Regele, H.; Eskandary, F.; Reindl-Schwaighofer, R.; Kainz, A.; Oberbauer, R. miR-182-5p Inhibition Ameliorates Ischemic Acute Kidney Injury. Am. J. Pathol. 2017, 187, 70–79. [Google Scholar] [CrossRef]
- Ichii, O.; Horino, T. MicroRNAs associated with the development of kidney diseases in humans and animals. J. Toxicol. Pathol. 2018, 31, 23–34. [Google Scholar] [CrossRef]
- Zhang, W.; Shao, M.; He, X.; Wang, B.; Li, Y.; Guo, X. Overexpression of microRNA-146 protects against oxygen-glucose deprivation/recovery-induced cardiomyocyte apoptosis by inhibiting the NF-κB/TNF-α signaling pathway. Mol. Med. Rep. 2018, 17, 1913–1918. [Google Scholar] [CrossRef] [PubMed]
- Aguado-Fraile, E.; Ramos, E.; Conde, E.; Rodríguez, M.; Martín-Gómez, L.; Lietor, A.; Candela, Á.; Ponte, B.; Liaño, F.; García-Bermejo, M.L. A Pilot Study Identifying a Set of microRNAs As Precise Diagnostic Biomarkers of Acute Kidney Injury. PLoS ONE 2015, 10, e0127175. [Google Scholar] [CrossRef]
- Wang, X.; Ha, T.; Liu, L.; Zou, J.; Zhang, X.; Kalbfleisch, J.; Gao, X.; Williams, D.; Li, C. Increased expression of microRNA-146a decreases myocardial ischaemia/reperfusion injury. Cardiovasc. Res. 2013, 97, 432–442. [Google Scholar] [CrossRef] [PubMed]
- Gómez-Dos-Santos, V.; Garcia-Bermejo, L.; Ramos, E.; Diez-Nicolas, V.; Alvarez-Rodriguez, S.; Hevia, V.; Burgos-Revilla, F.J. MicroRNAs in kidney hypothermic machine perfusion fluid as novel biomarkers for graft function: Have changes in normalization guidelines support previous results? Eur. Urol. Suppl. 2018, 17, e768. [Google Scholar] [CrossRef]
- Bhome, R.; Del Vecchio, F.; Lee, G.H.; Bullock, M.D.; Primrose, J.N.; Sayan, A.E.; Mirnezami, A.H. Exosomal microRNAs (exomiRs): Small molecules with a big role in cancer. Cancer Lett. 2018, 420, 228–235. [Google Scholar] [CrossRef]
- Roufosse, C.; Simmonds, N.; Clahsen-van Groningen, M.; Haas, M.; Henriksen, K.J.; Horsfield, C.; Loupy, A.; Mengel, M.; Perkowska-Ptasińska, A.; Rabant, M.; et al. A 2018 Reference Guide to the Banff Classification of Renal Allograft Pathology. Transplantation 2018, 102, 1795–1814. [Google Scholar] [CrossRef] [PubMed]
- Siebolts, U.; Varnholt, H.; Drebber, U.; Dienes, H.P.; Wickenhauser, C.; Odenthal, M. Tissues from routine pathology archives are suitable for microRNA analyses by quantitative PCR. J. Clin. Pathol. 2009, 62, 84–88. [Google Scholar] [CrossRef]
- Landgraf, P.; Rusu, M.; Sheridan, R.; Sewer, A.; Iovino, N.; Aravin, A.; Pfeffer, S.; Rice, A.; Kamphorst, A.O.; Landthaler, M.; et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 2007, 129, 1401–1414. [Google Scholar] [CrossRef]
- Alivernini, S.; Gremese, E.; McSharry, C.; Tolusso, B.; Ferraccioli, G.; McInnes, I.B.; Kurowska-Stolarska, M. MicroRNA-155-at the Critical Interface of Innate and Adaptive Immunity in Arthritis. Front Immunol. 2017, 8, 1932. [Google Scholar] [CrossRef]
- Berrien-Elliott, M.M.; Sun, Y.; Neal, C.; Ireland, A.; Trissal, M.C.; Sullivan, R.P.; Wagner, J.A.; Leong, J.W.; Wong, P.; Mah-Som, A.Y.; et al. MicroRNA-142 Is Critical for the Homeostasis and Function of Type 1 Innate Lymphoid Cells. Immunity 2019, 51, 479–490.e6. [Google Scholar] [CrossRef]
- Yuan, X.; Berg, N.; Lee, J.W.; Le, T.T.; Neudecker, V.; Jing, N.; Eltzschig, H. MicroRNA miR-223 as regulator of innate immunity. J. Leukoc Biol. 2018, 104, 515–524. [Google Scholar] [CrossRef]
- Goral, J.; Shenoy, S.; Mohanakumar, T.; Clancy, J. Antibodies to 70 kD and 90 kD heat shock proteins are associated with graft-versus-host disease in peripheral blood stem cell transplant recipients. Clin. Exp. Immunol. 2002, 127, 553–559. [Google Scholar] [CrossRef]
- Takahashi, K.; Kawai, T.; Kumar, H.; Sato, S.; Yonehara, S.; Akira, S. Roles of caspase-8 and caspase-10 in innate immune responses to double-stranded RNA. J. Immunol. 2006, 176, 4520–4524. [Google Scholar] [CrossRef]
- Tam, W. Identification and characterization of human BIC, a gene on chromosome 21 that encodes a noncoding RNA. Gene 2001, 274, 157–167. [Google Scholar] [CrossRef]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef]
- Badal, S.S.; Danesh, F.R. MicroRNAs and their applications in kidney diseases. Pediatr. Nephrol. 2015, 30, 727–740. [Google Scholar] [CrossRef][Green Version]
- Lin, H.; Yang, L.; Tian, F.; Nie, S.; Zhou, H.; Liu, J.; Chen, W. Up-regulated LncRNA-ATB regulates the growth and metastasis of cholangiocarcinoma via miR-200c signals. Onco Targets Ther. 2019, 12, 7561–7571. [Google Scholar] [CrossRef]
- Li, R.H.; Chen, M.; Liu, J.; Shao, C.C.; Guo, C.P.; Wei, X.L.; Li, Y.C.; Huang, W.H.; Zhang, G. Long noncoding RNA ATB promotes the epithelial-mesenchymal transition by upregulating the miR-200c/Twist1 axe and predicts poor prognosis in breast cancer. Cell Death Dis. 2018, 9, 1171. [Google Scholar] [CrossRef]
- Liu, Y.; Li, Y.; Xu, Q.; Yao, W.; Wu, Q.; Yuan, J.; Yan, W.; Xu, T.; Ji, X.; Ni, C. Long non-coding RNA-ATB promotes EMT during silica-induced pulmonary fibrosis by competitively binding miR-200c. Biochim. Biophys. Acta Mol. Basis Dis. 2018, 1864, 420–431. [Google Scholar] [CrossRef]
- Jin, Y.; Tymen, S.D.; Chen, D.; Fang, Z.J.; Zhao, Y.; Dragas, D.; Dai, Y.; Marucha, P.T.; Zhou, X. MicroRNA-99 family targets AKT/mTOR signaling pathway in dermal wound healing. PLoS ONE 2013, 8, e64434. [Google Scholar] [CrossRef]
- Truini, A.; Coco, S.; Nadal, E.; Genova, C.; Mora, M.; Dal Bello, M.G.; Vanni, I.; Alama, A.; Rijavec, E.; Biello, F.; et al. Downregulation of miR-99a/let-7c/miR-125b miRNA cluster predicts clinical outcome in patients with unresected malignant pleural mesothelioma. Oncotarget 2017, 8, 68627–68640. [Google Scholar] [CrossRef]
- Bao, M.H.; Li, J.M.; Luo, H.Q.; Tang, L.; Lv, Q.L.; Li, G.Y.; Zhou, H.H. NF-κB-Regulated miR-99a Modulates Endothelial Cell Inflammation. Mediat. Inflamm. 2016, 2016, 5308170. [Google Scholar] [CrossRef]
- Mei, L.L.; Qiu, Y.T.; Huang, M.B.; Wang, W.J.; Bai, J.; Shi, Z.Z. MiR-99a suppresses proliferation, migration and invasion of esophageal squamous cell carcinoma cells through inhibiting the IGF1R signaling pathway. Cancer Biomark. 2017, 20, 527–537. [Google Scholar] [CrossRef]
- Chau, B.N.; Xin, C.; Hartner, J.; Ren, S.; Castano, A.P.; Linn, G.; Li, J.; Tran, P.T.; Kaimal, V.; Huang, X.; et al. MicroRNA-21 promotes fibrosis of the kidney by silencing metabolic pathways. Sci. Transl. Med. 2012, 4, 121ra18. [Google Scholar] [CrossRef]
- Chen, C.; Lu, C.; Qian, Y.; Li, H.; Tan, Y.; Cai, L.; Weng, H. Urinary miR-21 as a potential biomarker of hypertensive kidney injury and fibrosis. Sci. Rep. 2017, 7, 17737. [Google Scholar] [CrossRef]
- Loboda, A.; Sobczak, M.; Jozkowicz, A.; Dulak, J. TGF-β1/Smads and miR-21 in Renal Fibrosis and Inflammation. Mediat. Inflamm. 2016, 2016, 8319283. [Google Scholar] [CrossRef]
- Kurowska-Stolarska, M.; Alivernini, S.; Ballantine, L.E.; Asquith, D.L.; Millar, N.L.; Gilchrist, D.S.; Reilly, J.; Ierna, M.; Fraser, A.R.; Stolarski, B.; et al. MicroRNA-155 as a proinflammatory regulator in clinical and experimental arthritis. Proc. Natl. Acad. Sci. USA 2011, 108, 11193–11198. [Google Scholar] [CrossRef] [PubMed]
- Omrane, I.; Benammar-Elgaaied, A. The immune microenvironment of the colorectal tumor: Involvement of immunity genes and microRNAs belonging to the TH17 pathway. Biochim. Biophys. Acta 2015, 1856, 28–38. [Google Scholar] [CrossRef]
- Millán, O.; Budde, K.; Sommerer, C.; Aliart, I.; Rissling, O.; Bardaji, B.; Matz, M.; Zeier, M.; Silva, I.; Guirado, L.; et al. Urinary miR-155-5p and CXCL10 as prognostic and predictive biomarkers of rejection, graft outcome and treatment response in kidney transplantation. Br. J. Clin. Pharmacol. 2017, 83, 2636–2650. [Google Scholar] [CrossRef]
- Gao, J.; Gu, J.; Pan, X.; Gan, X.; Ju, Z.; Zhang, S.; Xia, Y.; Lu, L.; Wang, X. Blockade of miR-142-3p promotes anti-apoptotic and suppressive function by inducing KDM6A-mediated H3K27me3 demethylation in induced regulatory T cells. Cell Death Dis. 2019, 10, 332. [Google Scholar] [CrossRef]
- Danger, R.; Paul, C.; Giral, M.; Lavault, A.; Foucher, Y.; Degauque, N.; Pallier, A.; Durand, M.; Castagnet, S.; Duong Van Huyen, J.P.; et al. Expression of miR-142-5p in peripheral blood mononuclear cells from renal transplant patients with chronic antibody-mediated rejection. PLoS ONE 2013, 8, e60702. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.L.; Li, D.; He, Y.L.; Zhou, Y.Z.; Gong, S.H.; Wu, L.Y.; Zhao, Y.Q.; Huang, X.; Zhao, T.; Xu, L.; et al. miR-210 protects renal cell against hypoxia-induced apoptosis by targeting HIF-1 alpha. Mol. Med. 2017, 23, 258–271. [Google Scholar] [CrossRef] [PubMed]
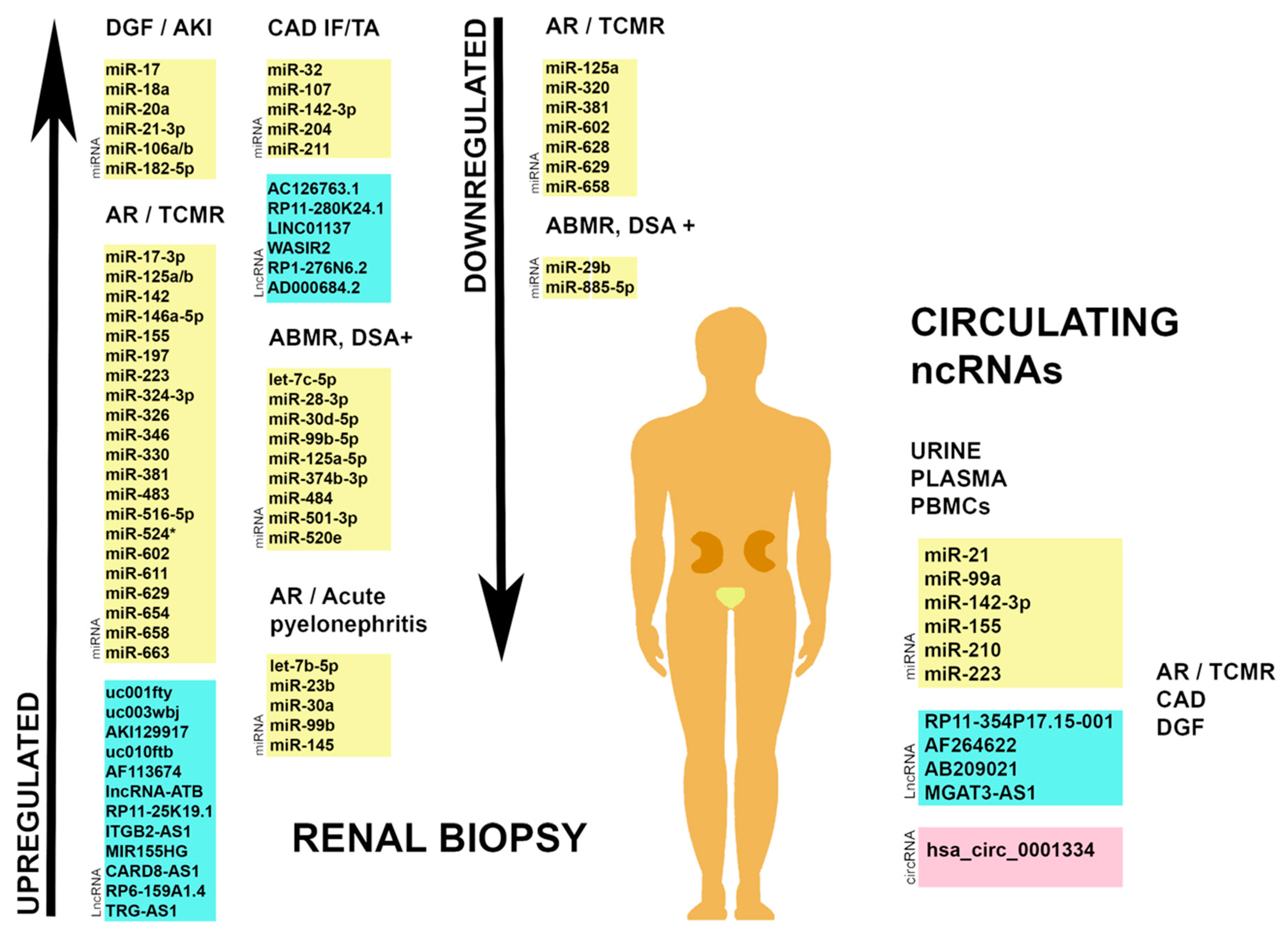
| Authors | Non-Coding RNAs | Expression | Localization | Disease State |
|---|---|---|---|---|
| miRNAs | ||||
| Wilflingseder, J. et al. [40,41] | miR-182-5p, miR-21-3p, miR-106a/b, miR-20a, miR-18a, miR-17 | Upregulated | Biopsy | DGF, AKI |
| Milhoransa, P. et al. [42] | miR-146a-5p | Upregulated | Biopsy | AR |
| Roest, H.P. et al. [43] | miR-505-3p | Upregulated | Preservation fluid | DGF |
| Gómez-Dos-Santos, V. et al. [44] | miR-486, miR-18a, miR-20a, miR-363-3p, miR-144-3p, miR-454-3p, miR-223-3p, miR-142-5p, miR-502-3p, miR-144-3p, miR-144-5p | Upregulated | Preservation solution | DGF |
| Wang, J. et al. [45] | miR-33a-5p, miR-151a-5p, miR-98-5p | Upregulated | Exosomes from peripheral blood | DGF |
| Sui, W. et al. [46] | miR-324-3p, miR-611, miR-654, miR-330, miR-524 *, miR-17-3p, miR-483, miR-663, miR-516-5p, miR-326, miR-197, miR-346 | Upregulated | Biopsy | AR |
| miR-658, miR-125a, miR-320, miR-381, miR-628, miR-602, miR-629 | Downregulated | |||
| Anglicheau, D. et al. [47] | miR-142-5p, miR-155, miR-223 | Upregulated | Biopsy | AR |
| Soltaninejad, E. et al. [48] | miR-142-5p, miR-142-3p, miR-155, miR-223 | Upregulated | Biopsy | TCMR |
| Sui, W. et al. [49] | miR-483, miR-381, miR-602, miR-629, miR-658, miR-524 *, miR-125a/b, miR-324-3p, miR-663, miR-326, miR-346 | Varies | Biopsy | AR |
| Oghumu, S. et al. [50] | miR-99b, miR-23b, let-7b-5p, miR-30a, miR-145 | Varies | Biopsy | AR, acute pyelonephritis |
| Scian, M.J. et al. [51] | miR-142-3p, miR-204, miR-107, miR-211, miR-32 | Varies | Biopsy, urine | CAD IF/TA |
| Heinemann, F.M. et al. [52] | let-7c-5p, miR-28-3p, miR-30d-5p, miR-99b-5p, miR-125a-5p, miR-195-5p, miR-374b-3p, miR-484, miR-501-3p, miR-520e | Upregulated | Biopsy | ABMR DSA+ |
| miR-29b-3p, miR-885-5p | Downregulated | |||
| Anglicheau, D. et al. [47] Tao, J. et al. [53] Ledeganck, K.J. et al. [54] | miR-99a | Varies | Plasma, PBMCs, urine | AR, CAD IF/TA, TCMR |
| Ben-Dov, I.Z. et al. [55] Zununi Vahed, S. et al. [56] | miR-21 | Upregulated | Biopsy, urine, PBMCs | CAD |
| Anglicheau, D. et al. [47] Soltaninejad, E. et al. [48] Zununi Vahed, S. et al. [56] | miR-155 | Upregulated | PBMCs, urine | AR, CAD, Responsive to immunossuppresive therapy |
| Soltaninejad, E. et al. [48] Janszky, N. et al. [57] | miR-142-3p | Upregulated | PBMCs, plasma, urine | CAD IF/TA |
| Soltaninejad, E. et al. [48] Ulbing, M. et al. [58] | miR-223 | Varies | PBMCs | CAD TCMR |
| Lorenzen, J.M. et al. [59] | miR-210 | Downregulated | Urine | AR |
| lncRNAs | ||||
| Sui, W. et al. [49] | NR_026695, NR_024080, uc010kwo, NR_023318, NR_026576, NR_002909, NR_026550, NR_003024, NR_027303, NR_003130, NR_024418, uc003wcs, uc003syy, uc002zic, uc010gqe, uc003bgk, uc003akf, NR_024400, NR_024332, uc001pyd, uc010lqx, NR_024611, uc002zpx, NR_001562, NR_002941, uc010akv, uc002nyb, NR_003573, NR_002791, uc003zfx, uc003dwf, and uc003tsq | Varies | Biopsy | AR |
| Chen, W. et al. [39] | uc001fty, uc003wbj, AKI129917, uc010ftb, AF113674 | Upregulated | Biopsy | AR |
| Qiu, J. et al. [60] | lncRNA-ATB | Upregulated | Biopsy | AR, Cyclosporine-induced nephrotoxicity |
| Zou, Y. et al. [61] | RP11-25K19.1, ITGB2- AS1, MIR155HG, CARD8-AS1, RP6-159A1.4, TRG-AS1 | Upregulated | Biopsy ** | AR |
| Xu, J. et al. [62] | AC126763.1, RP11-280K24.1, LINC01137, WASIR2, RP1-276N6.2, AD000684.2 | Upregulated | Biopsy ** | CAD |
| Lorenzen, J.M. et al. [63] | RP11-354P17.15-001 | Upregulated | Urine | TCMR |
| Ge, Y.Z. et al. [28] | AF264622, AB209021 | Upregulated | Blood | AR |
| Nagarajah, S. et al. [64] | MGAT3-AS1 | Downregulated | PBMCs | DGF |
| circRNAs | ||||
| Kölling, M. et al. [65] | hsa_circ_0001334, has_circ_0071475 | Upregulated | Urine | TCMR |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Franco-Acevedo, A.; Melo, Z.; Echavarria, R. Diagnostic, Prognostic, and Therapeutic Value of Non-Coding RNA Expression Profiles in Renal Transplantation. Diagnostics 2020, 10, 60. https://doi.org/10.3390/diagnostics10020060
Franco-Acevedo A, Melo Z, Echavarria R. Diagnostic, Prognostic, and Therapeutic Value of Non-Coding RNA Expression Profiles in Renal Transplantation. Diagnostics. 2020; 10(2):60. https://doi.org/10.3390/diagnostics10020060
Chicago/Turabian StyleFranco-Acevedo, Adriana, Zesergio Melo, and Raquel Echavarria. 2020. "Diagnostic, Prognostic, and Therapeutic Value of Non-Coding RNA Expression Profiles in Renal Transplantation" Diagnostics 10, no. 2: 60. https://doi.org/10.3390/diagnostics10020060
APA StyleFranco-Acevedo, A., Melo, Z., & Echavarria, R. (2020). Diagnostic, Prognostic, and Therapeutic Value of Non-Coding RNA Expression Profiles in Renal Transplantation. Diagnostics, 10(2), 60. https://doi.org/10.3390/diagnostics10020060






