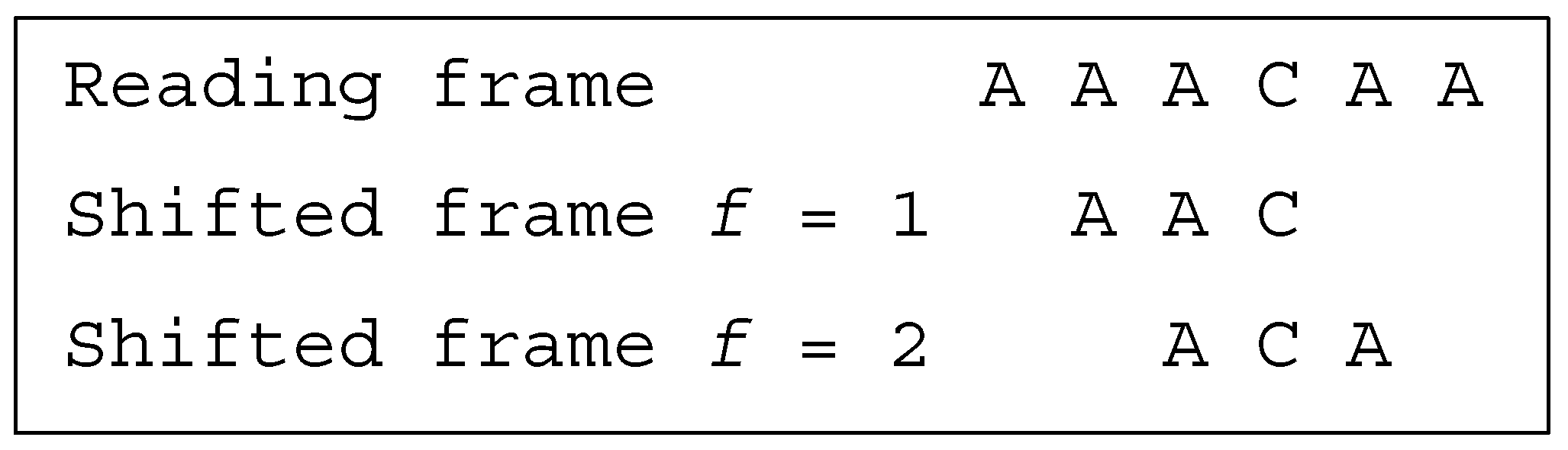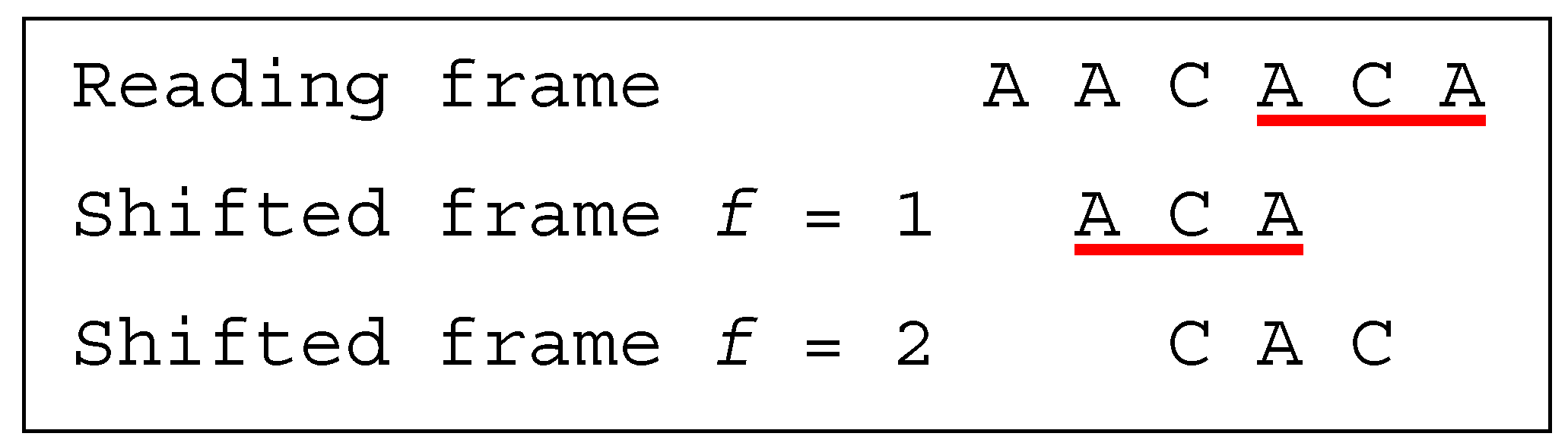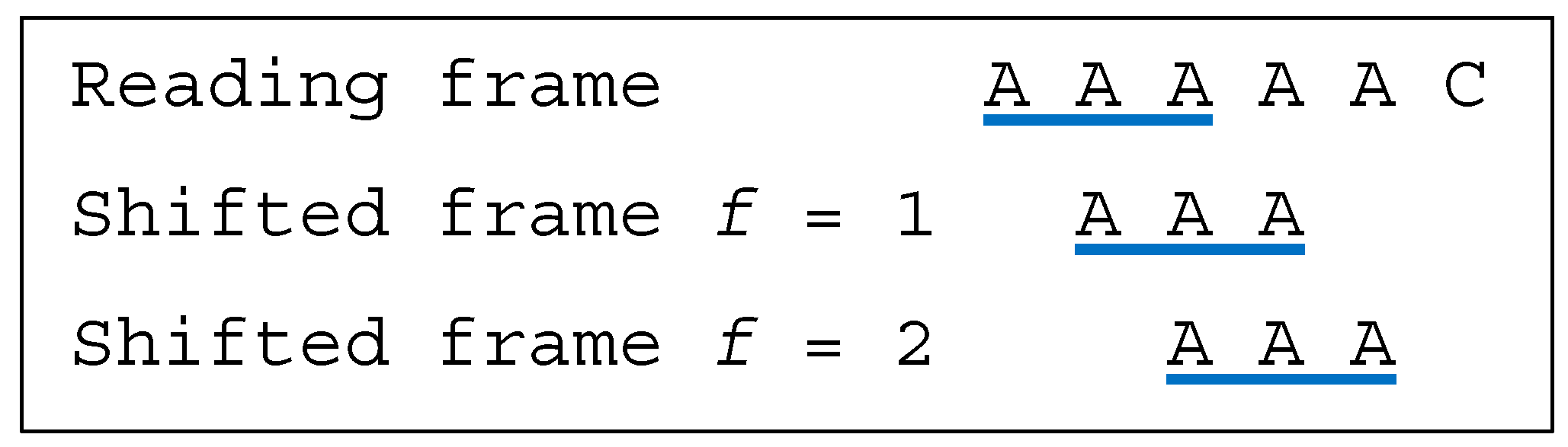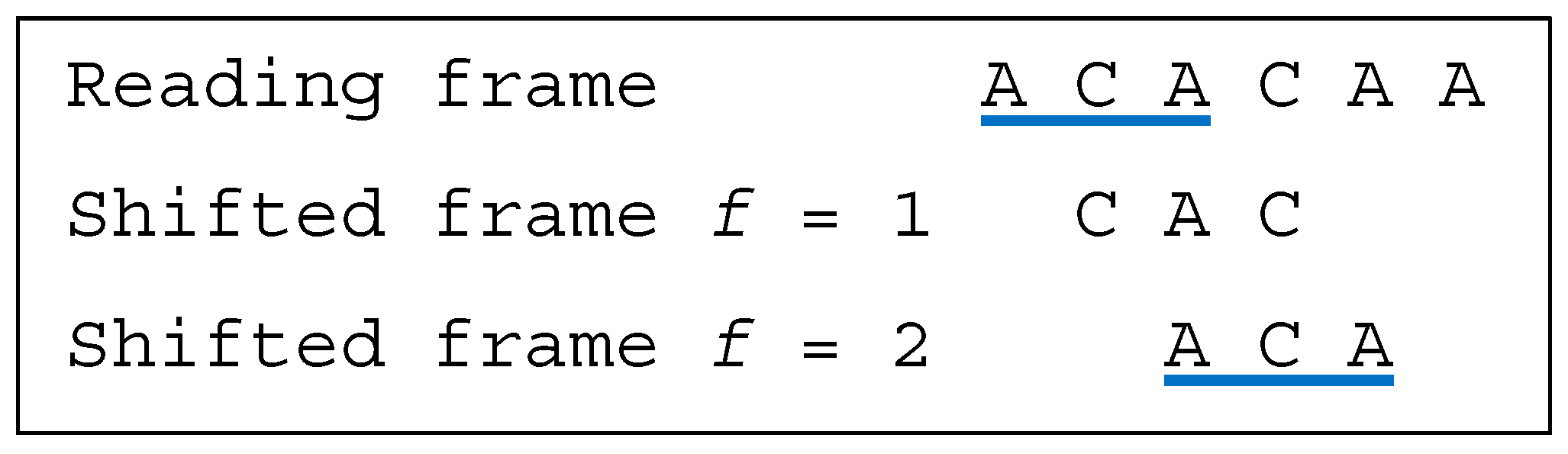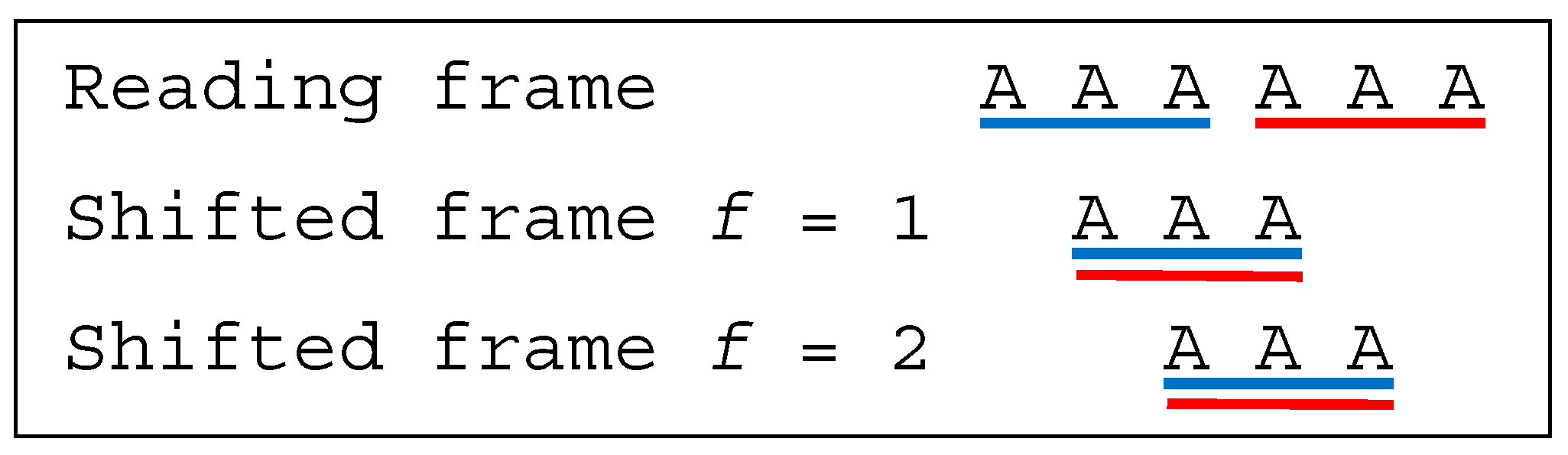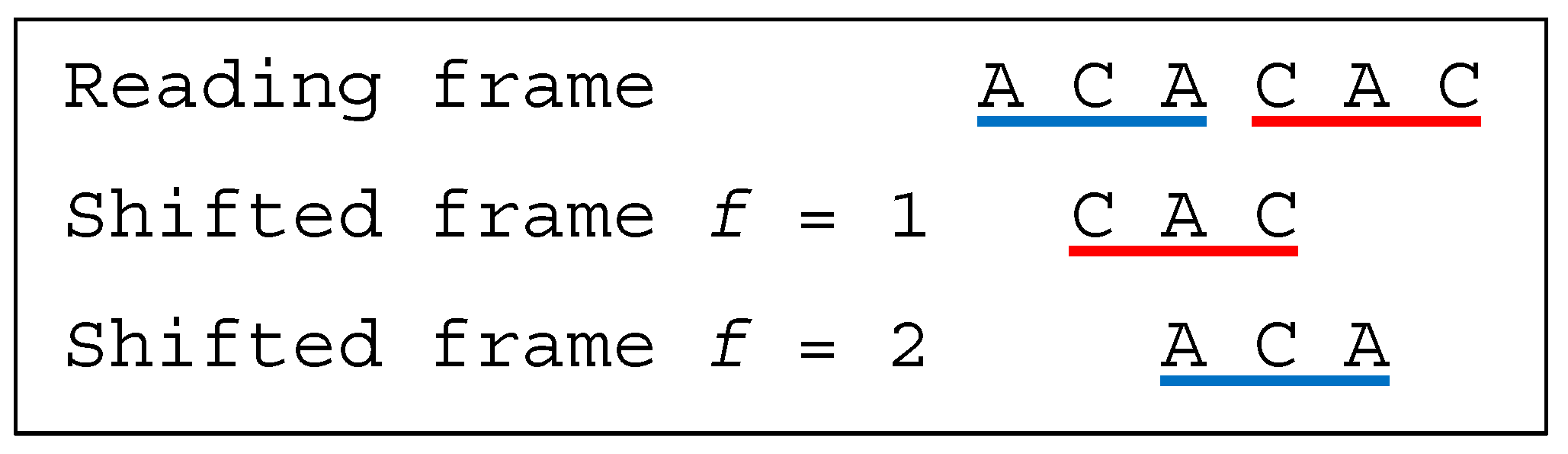1. Introduction
The reading frame coding with trinucleotide sets is a fascinating problem, both theoretical and experimental. Before the discovery of the genetic code, a first code was proposed by Gamow [
1] by considering the “key-and-lock” relation between various amino acids, and the rhomb shaped “holes” formed by various nucleotides in the DNA. The proposed model will later prove to be false. A few years later, a class of trinucleotide codes, called comma-free codes, was proposed by Crick et al. [
2] for explaining how the reading of a sequence of trinucleotides could code amino acids. In particular, how the correct reading frame can be retrieved and maintained. The four nucleotides {
A,
C,
G,
T} as well as the 16 dinucleotides {
AA,…,
TT} are simple codes which are not appropriate for coding 20 amino acids. However, trinucleotides induce a redundancy in their coding. Thus, Crick et al. [
2] conjectured that only 20 trinucleotides among the 64 possible trinucleotides {
AAA,…,
TTT} code for the 20 amino acids. Such a bijective code implies that the coding trinucleotides are found only in one frame—the comma-freeness property. The determination of a set of 20 trinucleotides forming a comma-free code has several necessary conditions:
(i) A periodic trinucleotide from the set {AAA,CCC,GGG,TTT} must be excluded from such a code. Indeed, the concatenation of AAA with itself, for instance, does not allow the (original) reading frame to be retrieved as there are three possible decompositions: …,AAA,AAA,AAA,… (original frame), …A,AAA,AAA,AA… and …AA,AAA,AAA,A…, the commas showing the adopted decomposition.
(ii) Two non-periodic permuted trinucleotides, i.e., two trinucleotides related by a circular permutation, e.g., ACG and CGA, must also be excluded from such a code. Indeed, the concatenation of ACG with itself, for instance, does not allow the reading frame to be retrieved as there are two possible decompositions: …,ACG,ACG,ACG,… (original frame) and …A,CGA,CGA,CG…
Therefore, by excluding the four periodic trinucleotides and by gathering the 60 remaining trinucleotides in 20 classes of three trinucleotides such that, in each class, the three trinucleotides are deduced from each other by a circular permutation, e.g., ACG, CGA and GAC, we see that a comma-free code can contain only one trinucleotide from each class and thus has at most 20 trinucleotides. This trinucleotide number is identical to the amino acid number, thus leading to a code assigning one trinucleotide per amino acid without ambiguity.
In the beginning 1960’s, the discovery that the trinucleotide
TTT, an excluded trinucleotide in a comma-free code, codes phenylalanine [
3], led to the abandonment of the concepts both of a comma-free code [
2] and a bijective code as the genetic code is degenerate [
4,
5,
6] with a gene translation in one direction [
7].
In 1996, a statistical analysis of occurrence frequencies of the 64 trinucleotides in the three frames of genes of both prokaryotes and eukaryotes showed that the trinucleotides are not uniformly distributed in these three frames [
8]. By excluding the four periodic trinucleotides and by assigning each trinucleotide to a preferential frame (frame of its highest occurrence frequency), three subsets
,
and
of 20 trinucleotides each are found in the frames 0 (reading frame), 1 (frame 0 shifted by one nucleotide in the
direction, i.e., to the right) and 2 (frame 0 shifted by two nucleotides in the
direction) in genes of both prokaryotes and eukaryotes. The same set
of trinucleotides was identified in average in genes (reading frame) of bacteria, archaea, eukaryotes, plasmids and viruses [
9,
10]. It contains the 20 following trinucleotides:
and codes the 12 following amino acids (three and one letter notation):
This set
has a strong mathematical property. Indeed,
is a maximal
self-complementary trinucleotide circular code [
8].
The reading frame coding with trinucleotide codes (sets of words) in general terms, i.e., not particularly the genetic code, is a concept which has been studied in Michel [
11,
12]. We extend it to the motifs (words of codes), a theoretical domain which has been ignored according to our knowledge. Genes (protein coding regions) can be partitioned into two disjoint classes of motifs: the single-frame motifs
with an unambiguous trinucleotide decoding in the two
and
directions, and the multiple-frame motifs
with an ambiguous trinucleotide decoding in at least one direction. A single-frame motif
has no trinucleotide in reading frame (frame 0) that occurs in a shifted frame (frame 1 or 2). In contrast, a multiple-frame motif
has at least one trinucleotide in reading frame that occurs in a shifted frame. Some well-known
motifs are involved in ribosomal frameshifting. The expression of some viral and cellular genes utilizes a -1 programmed ribosomal frameshifting (-1 PRF) [
13,
14]. This -1 PRF sequence is based on three elements: (i) a slippery motif composed of seven nucleotides at which the change in reading frame occurs; (ii) a spacer motif, usually less than 12 nucleotides; and (iii) a down-stream (3′) stimulatory motif, usually a pseudoknot or a stem-loop. In eukaryotes, the slippery motif fits a consensus heptanucleotide
X,XXY,YYZ, where
XXX is any three identical nucleotides,
YYY represents
AAA or
TTT,
Z represents
A,
C or
T, the commas separating the codons in reading frame [
15,
16]. The slippery motifs
and
are multiple-frame
. Indeed, the codon
AAA in reading frame also occurs in the shifted frames 1 and 2 in
, and similarly with the codon
TTT in
. Alternative gene decoding is also possible with +1 programmed ribosomal frameshifting (+1 PRF) which has been particularly observed in
Euplotes [
17]. The identified slippery motif
where
is multiple-frame
. The slippery motifs
AAA,
CCC,
GGG and
TTT may cause frameshifting during transcription, producing RNAs missing specific nucleotides when compared to template DNA [
18,
19]. The slippery motifs are not always multiple-frame while stressing that the spacer and the down-stream stimulatory motifs have been very poorly characterized [
20] and could also be involved in such a multiple-frame definition. From a theoretical point of view, it is important to extend this concept by increasing the length of such multiple-frame slippery motifs and also by considering their different classes. If the multiple-frame motifs may be involved in ribosomal frameshifting, the single-frame motifs
and the framing motifs
(also called circular code motifs; first introduced in Michel [
21,
22]) from the circular codes [
8,
9,
10] (reviews in Michel [
23]; Fimmel and Strüngmann [
24]) may have been important in early life genes for constructing the modern genetic code and the extant genes (see Discussion).
Several classes of motifs are defined: (i) a unidirectional multiple-frame motif has no trinucleotide in reading frame that occurs in a shifted frame after its reading (i.e., its position in the reading frame) but has at least one trinucleotide in reading frame that occurs in a shifted frame before its reading, e.g., the dicodon AACACA is as the trinucleotides AAC and (trivially) ACA do not occur in a shifted frame after their reading and as the trinucleotide ACA occurs in a shifted frame (precisely frame 1) before its reading; (ii) a unidirectional multiple-frame motif , the opposite, has no trinucleotide in reading frame that occurs in a shifted frame before its reading but has at least one trinucleotide in reading frame that occurs in a shifted frame after its reading, e.g., the dicodon ACACAA mirror of AACACA is as the trinucleotides (trivially) ACA and CAA do not occur in a shifted frame before their reading and as the trinucleotide ACA occurs in a shifted frame (precisely frame 2) after its reading; and (iii) a bidirectional multiple-frame motif has at least one trinucleotide in reading frame that occurs in a shifted frame before its reading and has at least one trinucleotide in reading frame that occurs in a shifted frame after its reading (both and ), e.g., the dicodons AAAAAA and ACACAC are . A 5′ unambiguous motif , is either a motif or a motif, e.g., the dicodons AAACAA ( motif) and AACACA ( motif) belong to the class .
We will only investigate here the distribution of the single-frame motifs associated with an unambiguous trinucleotide decoding in the two and directions, and the 5′ unambiguous motifs associated with an unambiguous trinucleotide decoding in the direction only, i.e., a less restrictive class of motifs. The distributions of and motifs will be analysed without and with constraints. The constraints studied are: (i) with initiation and stop codons; (ii) without periodic codons ; (iii) with antiparallel complementarity; and (iv) with parallel complementarity.
We will also investigate the particular case of motifs made up of two codons, i.e., the dicodons. The definitions of
and
dicodons will thus identify two new classes of dipeptides, the
and
dipeptides. The
dipeptides are coded by dicodons with an unambiguous trinucleotide decoding, in contrast to the
dipeptides which are coded by dicodons with an ambiguous trinucleotide decoding. The concept of
and
dipeptides might be of predictive value to studies of prebiotic metabolites [
25]. Peptide evolution on the primitive earth is an active and exciting field of research with cyclic dipeptides [
26] and selective formation of
SerHis dipeptide via phosphorus activation [
27,
28].
3. Results
3.1. Single-Frame Motifs
I first investigated the probability
(Equation (3)) of single-frame
-motifs
(Definition 9). The probability
is equal to 1 (1-motifs,
Section 2.5). The probability
is equal to 94.9% (2-motifs,
Section 2.6). The probability
for
is given in
Table 4. The probability
for
is obtained by computer simulation (
Section 2.7).
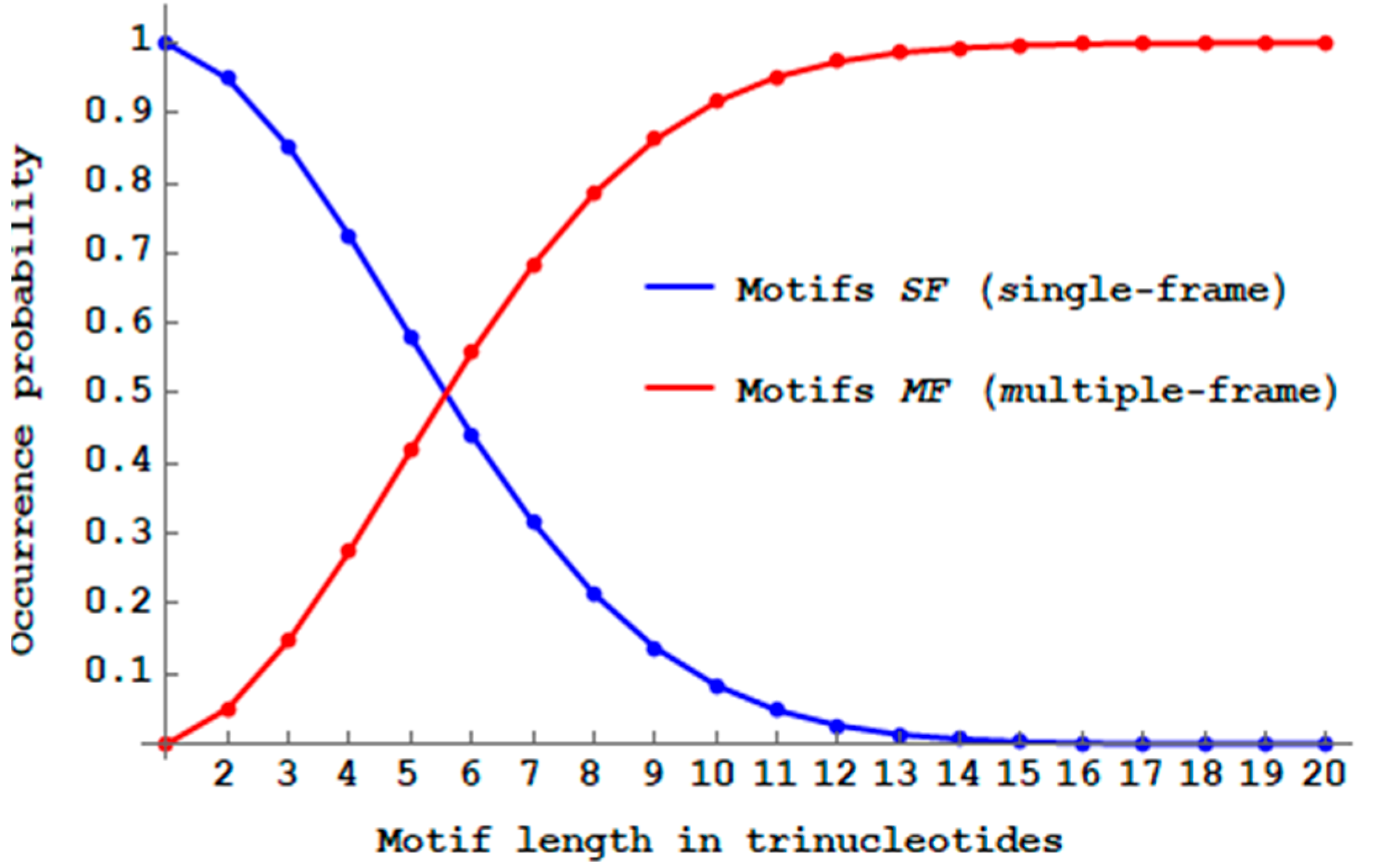
While the proportion of multiple-frame
-motifs
(Definition 10) is minimal (
for dicodons,
Section 2.6),
Figure 8 shows that their propagation will drastically reduce the proportion of
-motifs when the trinucleotide length
increases. There are almost no more
motifs with a length of 14 trinucleotides (
) and the number of
motifs becomes already higher than the number of
motifs with a length of six trinucleotides (
Figure 8).
Thus, only short genes, i.e., with up to five trinucleotides, have a higher proportion of single-frame motifs compared to the multiple-frame motifs. Thus, primitive translation, without the extant complex ribosome, could only generate short peptides without frameshift errors.
3.2. 5′ Unambiguous Motifs
I then compared the probability
(Equation (3)) of single-frame
-motifs
(Definition 9) and the probability
(Equation (4)) of 5′ unambiguous
-motifs
(Definition 14).
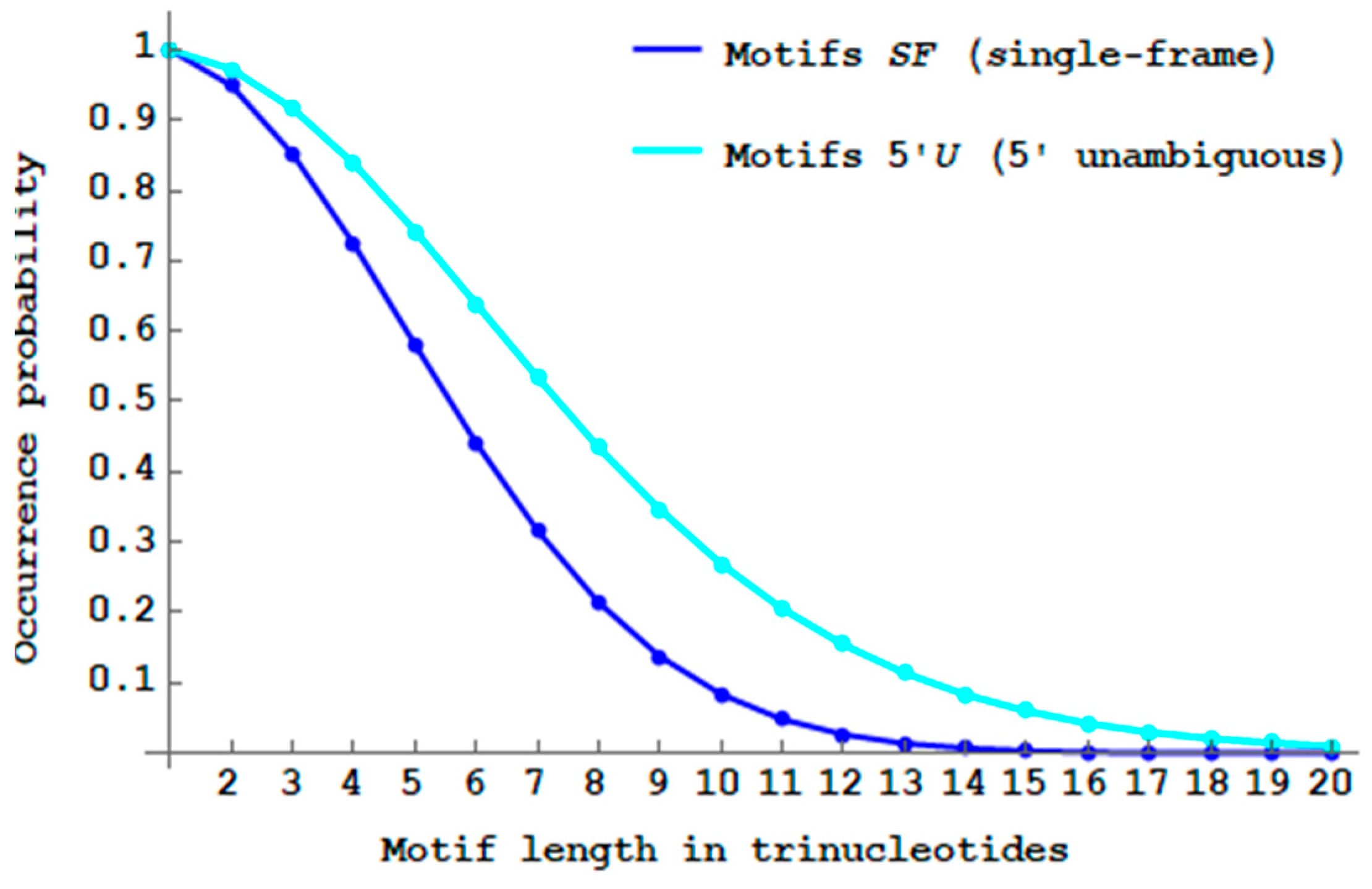
Figure 9 shows the decreasing probability
of
-motifs when the trinucleotide length
increases. As expected (see Remark 1), its decrease is slower than that of
-motifs. There are almost no more
motifs with a length of 20 trinucleotides (
). Thus with the
motifs, there is a length increase of
trinucleotides in the trinucleotide decoding. The maximum probability difference
is 22.0% at length
trinucleotides.
The 5′ unambiguous -motifs, a less restrictive class of motifs with an unambiguous trinucleotide decoding in the direction only, can generate a slightly longer peptides without frameshift error compared to the single-frame motifs.
I now evaluate the single-frame motifs and the 5′ unambiguous motifs with constraints.
3.3. Single-Frame and 5′ Unambiguous Motifs with Initiation and Stop Codons
The single-frame
-motifs
and the 5′ unambiguous motifs
are investigated with an initiation codon
ATG and a stop codon
. The case
does not exist. For
, there are only three dicodons:
ATGTAA,
ATGTAG and
ATGTGA which are all obviously
. Thus, the probabilities of
and
-motifs are obviously
.
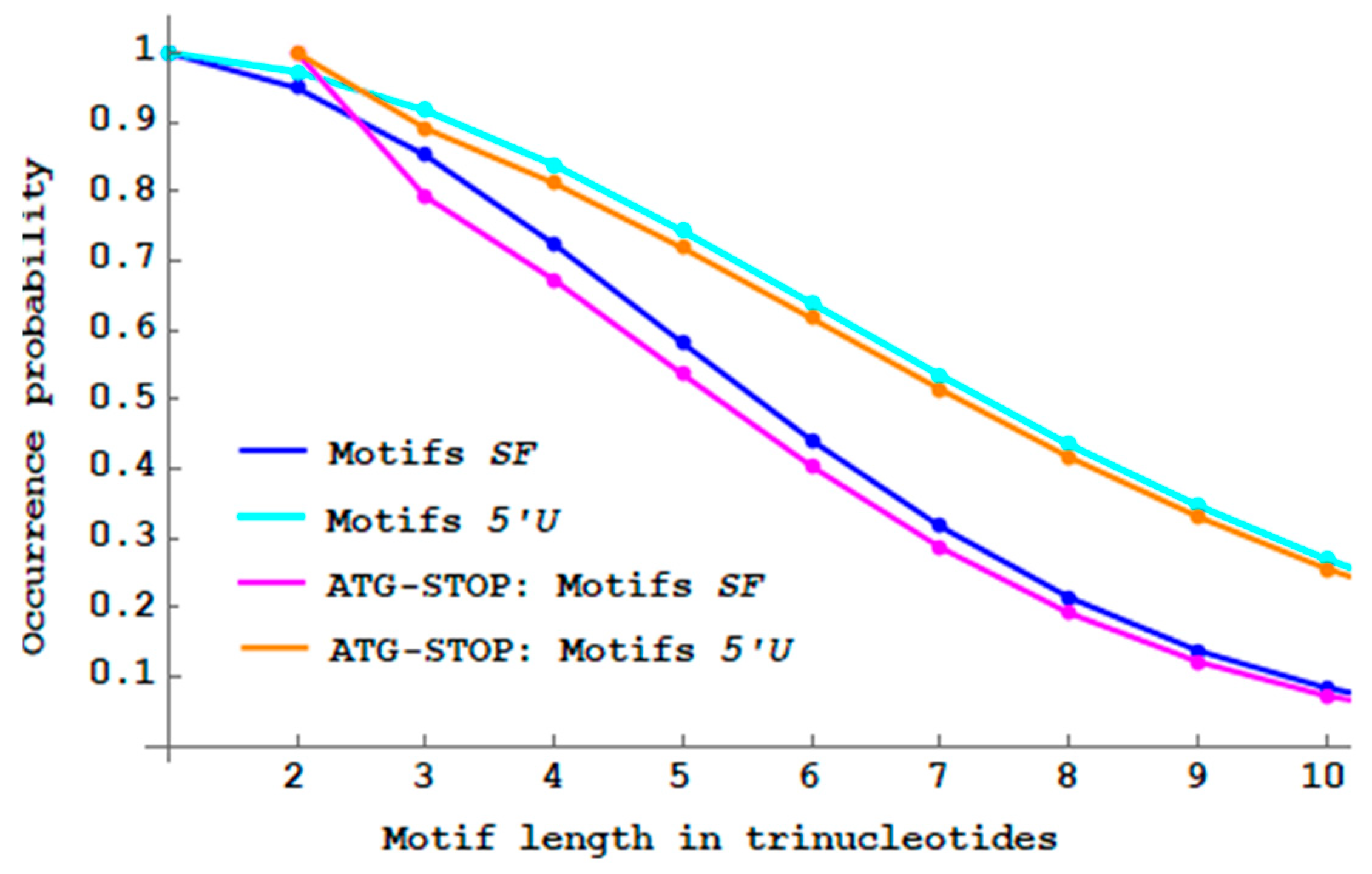
Figure 10 shows that the proportions of
and
motifs with initiation and stop codons are lower than their respective non-constrained motifs.
Genes with initiation and stop codons do not increase translation fidelity compared to non-constrained genes (according to this approach).
3.4. Single-Frame and 5′ Unambiguous Motifs without Periodic Codons
The single-frame motifs
and the 5′ unambiguous motifs
are now studied without periodic codons
. As expected,
Figure 11 shows that the proportions of
and
motifs without periodic codons are higher than their respective non-constrained motifs.
Genes without periodic codons slightly increase frame translation fidelity compared to non-constrained genes (according to this approach).
3.5. Single-Frame and 5′ Unambiguous Motifs with Antiparallel Complementarity
The single-frame
-motifs
and the 5′ unambiguous
-motifs
are now investigated with the following antiparallel complementary sequence:
where the trinucleotide antiparallel complementarity map
applied to a trinucleotide
is recalled in Definition 1. As an example, if
then the antiparallel complementary sequence studied is
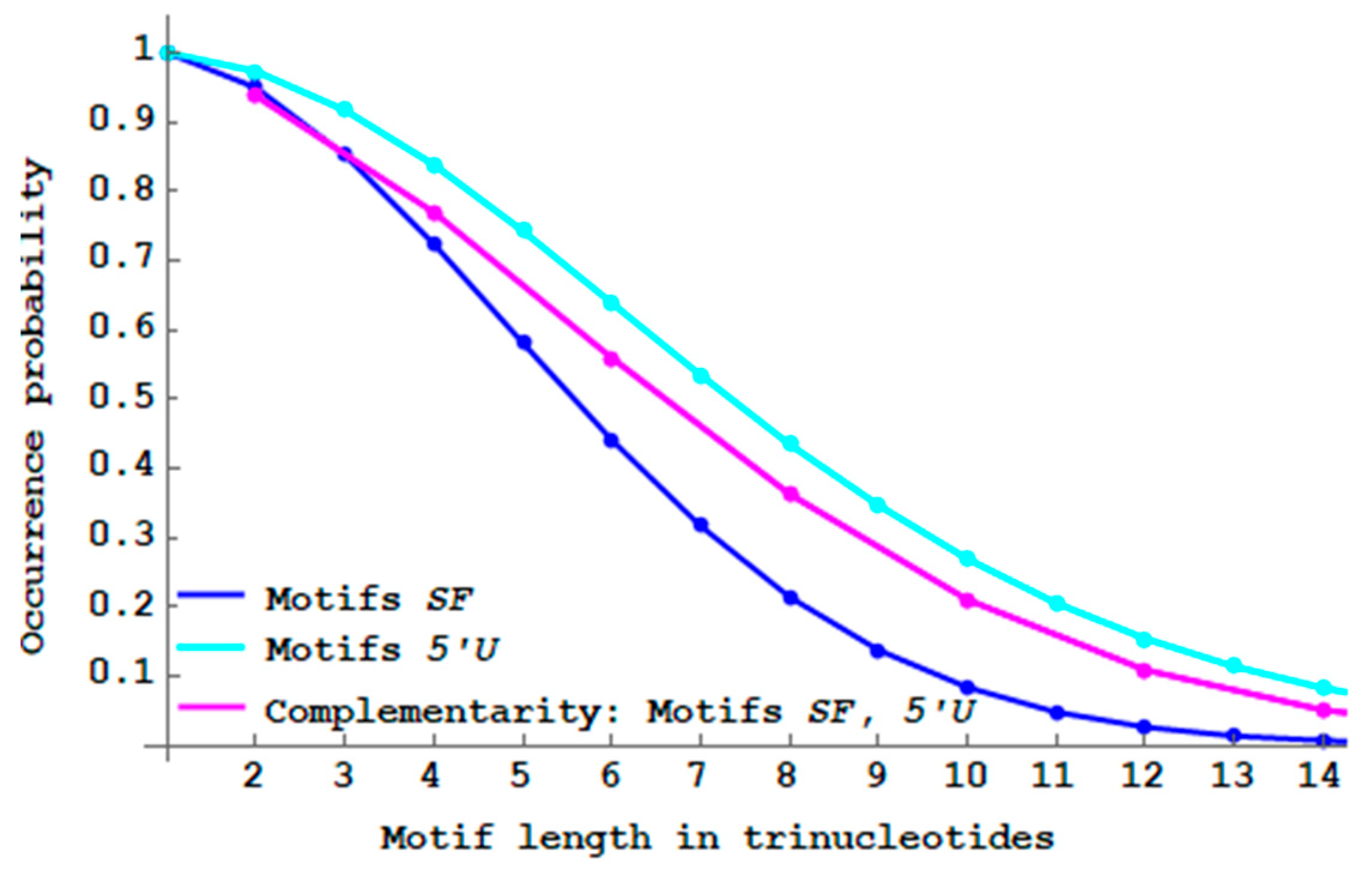
. Note that the trinucleotide length of such motifs is even. Classical antiparallel complementary structures are the DNA double helix and the RNA stem. Interesting results are observed. As expected, the two probability curves
of
motifs and
of
motifs with antiparallel complementarity are identical (
Figure 12). The proof is based on the following property: if
with
(
motif) then
with
(
motif) and
. Furthermore, antiparallel complementarity increases the proportion of
motifs but decreases the proportion of
motifs, compared to their respective non-constrained motifs.
The “antiparallel complementary” genes have a higher proportion of single-frame motifs compared to the non-complementary genes. Thus, primitive translation associated with a DNA property could generate a greater number of peptides without frameshift errors.
3.6. Single-Frame Motifs and 5′ Unambiguous with Parallel Complementarity
The single-frame
-motifs
and the 5′ unambiguous
-motifs
are now analysed with the following parallel complementary sequence:
where the trinucleotide parallel complementarity map
applied to a trinucleotide
is recalled in Definition 1. As an example, if
then the parallel complementary sequence studied is
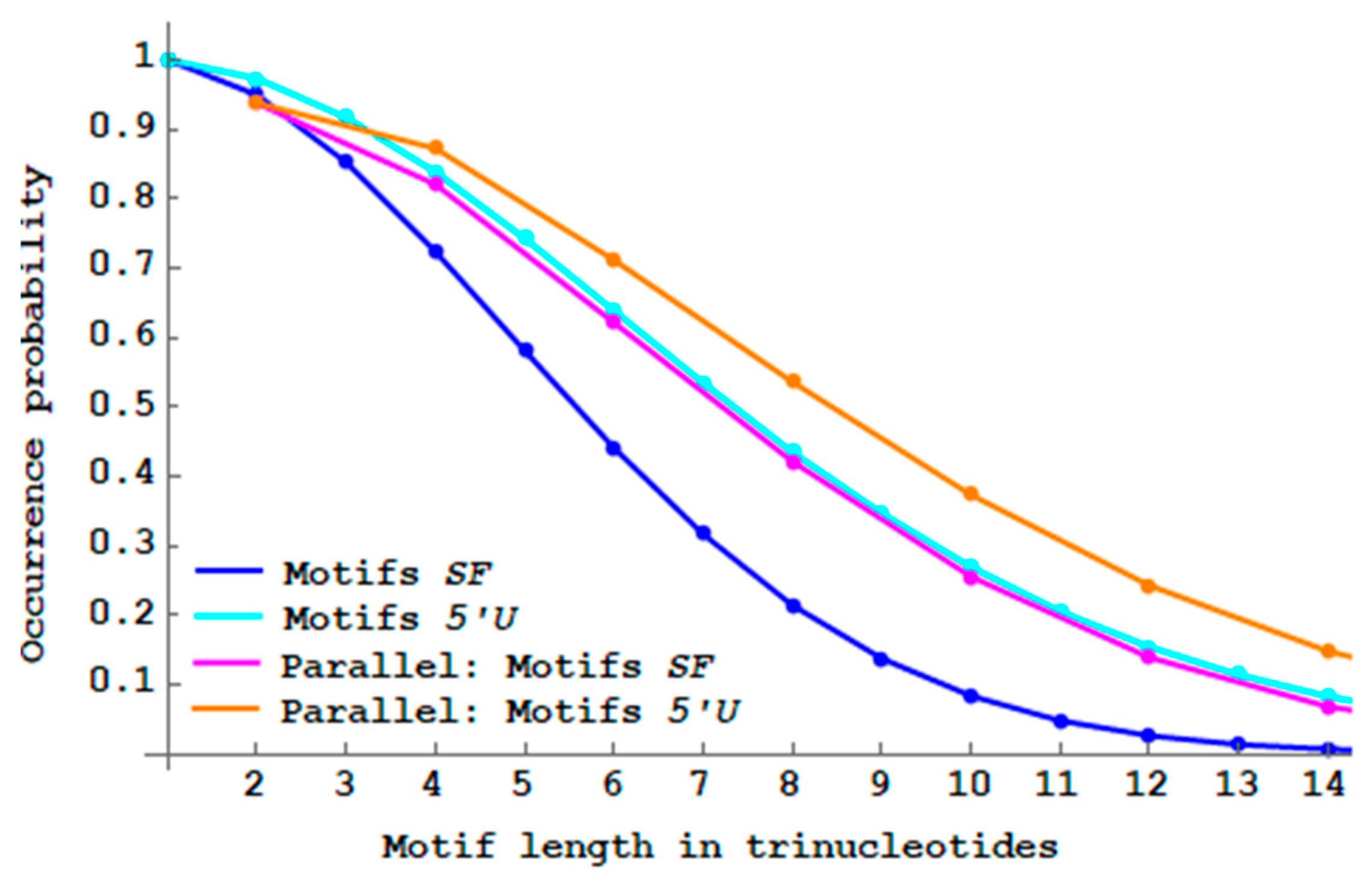
. Note that the trinucleotide length of such motifs is also even. Interesting results are also observed. The two probability curves
of
motifs with parallel complementarity and
of
motifs without constraints are superposable (
Figure 13). Parallel complementarity increases the proportions of both
motifs and
motifs compared to their respective non-constrained motifs.
“Parallel complementary” genes have a slightly higher proportion of single-frame motifs compared to the “antiparallel complementary” genes (compare the magenta curves in
Figure 12 and
Figure 13). The biological meaning is not yet explained.
3.7. Framing Motifs
There are framing motifs which are single-frame or multiple-frame .
Proposition 1. A framing motifcan be single-frame.
Proof. Take the following motif . The motif can be generated by the code . By Theorem 1, it is easy to verify that the graph is acyclic, and thus is circular. Furthermore, the set of trinucleotides in reading frame is , the set of trinucleotides in the shifted frame 1 is and the set of trinucleotides in the shifted frame 2 is . We have and . Thus, the motif is both framing and single-frame .
Proposition 2. A framing motifcan be multiple-frame.
Proof. Take the following motif . The motif can be generated by the code . By Theorem 1, it is easy to verify that the graph is acyclic, and thus is circular. Furthermore, we have the trinucleotide sets , and leading to and . Thus, the motif is both framing and multiple-frame , precisely unidirectional multiple-frame .
There are single-frame motifs or multiple-frame motifs which are not framing .
Proposition 3. A single-frame motifcan be non-framing.
Proof. Take the following motif . The motif can be generated by the code . We have the trinucleotide sets , and leading to and . However, as contains the periodic trinucleotide AAA, is not circular. Thus, the motif is single-frame but not framing .
Proposition 4. A multiple-frame motifcan be non-framing.
Proof. Take the following motif . The motif can be generated by the code . We have the trinucleotide sets , and leading to and . However, as contains the two permuted trinucleotides ACT and TAC, is not circular. Thus, the motif is multiple-frame , precisely unidirectional multiple-frame , but not framing .
Genes which are both framing and single-frame retrieve the reading frame and code for a unique peptide as the shifted frames would lead to a different peptide product.
3.8. A New Class of Theoretical Parameters Relating the Circular Codes and Their Circular Code Motifs
The idea is to define a new class of parameters in order to measure the intensity of a motif of a circular code to retrieve the reading frame. Thus, we have to associate information from the circular code theory with information from words (motifs).
In the circular code theory, the most important and the simplest parameter is the length
of a longest path (maximal arrow-length of a path) in the associated graph
of a circular code
(see Definition 5). Note that the longest path
has a finite length as the graph
is acyclic (Theorem 1). The longest path
can classify the circular codes, from the strong comma-free codes with
and the comma-free codes with
up to the general circular codes with a maximal longest path
when
(i.e., for the trinucleotide circular codes) [
29]. It is also related to the reading frame number
of
, i.e., the number of nucleotides to retrieve the reading frame. This reading frame number
can also be used to classify the circular codes, from the strong comma-free codes with
nucleotides and the comma-free codes with
nucleotides up to the general circular codes with a maximal number
nucleotides when
[
30]. However, this parameter
needs to know the structure of the longest path
which is one of the four cases:
,
,
and
where the nucleotide
and the dinucleotide
for any
(see Definition 5). In summary, for the circular codes
, the longest path
belongs to the interval
and the reading frame number
belongs to the interval
nucleotides. The definition of the reading frame number
can still be generalized to arbitrary sequences, i.e., not entirely consisting of trinucleotides from
[
30]. For these two reasons, i.e., the knowledge of the structure of
and the generalized definition of
, the parameter
, mentioned here to take date, will not be considered here.
A motif of a code, circular or not, can be characterized by its length , given here in trinucleotides for convenience, for measuring its expansion; and its cardinality of the set (see Notation 2) of trinucleotides (in reading frame ) of for measuring its variety (complexity). In the case of a motif of a trinucleotide circular code , .
It is important to stress the following condition: with a trinucleotide circular code . The case is associated with a trinucleotide circular code constructed from the motif .
A simple parameter measuring the expansion intensity
of reading frame retrieval of a circular code motif
can be defined as follows:
where
,
, is the trinucleotide length of the motif
and
,
, is the length of a longest path in the associated graph
of a trinucleotide circular code
. Note that
and if
then
.
A second parameter measuring both the expansion and variety intensity
of a circular code motif
can also be defined as follows:
where
is defined in Equation (5) and
,
, is the cardinality of the set
(Notation 2) of trinucleotides (in reading frame
) of
. Note that
and if
then
. Thus, for the circular code motifs
of a given trinucleotide length
, the intensity
of reading frame retrieval increases according to their cardinality
.
For a sequence
containing several circular code motifs
, the formulas (5) and (6) can be expressed as follows:
with the hypothesis that
is identical for the motifs
, a realistic case when the motifs
are obtained from a same studied trinucleotide circular code
, and thus:
Note also that the formulas and can also be normalized in order to weight the different lengths of sequences .
3.9. Dipeptides
The series of multi-frame motifs
starts with the dicodons. We will now focus on the
dipeptides which are two consecutive amino acids coded by the
dicodons. The 16 bidirectional multiple-frame dicodons
(
Table 1) code 16
dipeptides according to the universal genetic code (
Table 5). They include the four obvious
dipeptides
GlyGly (
GGGGGG),
LysLys (
AAAAAA),
PhePhe (
TTTTTT) and
ProPro (
CCCCCC). 15 amino acids out of 20 are involved in these 16
dipeptides (
Table 6):
Ala,
Arg,
Cys,
Glu,
Gly,
His,
Ile,
Leu,
Lys,
Phe,
Pro,
Ser,
Thr,
Tyr and
Val (except
Asn,
Asp,
Gln,
Met and
Trp), each amino acid occurring once in a position of a
dipeptide, except
Arg occurring twice in a position of a
dipeptide: ArgAla, ArgGlu, AlaArg and GluArg.
The 96 unidirectional multiple-frame dicodons
(
Table 2) code 83
dipeptides and four pairs (stop codon, amino acid):
TAGArg,
TAGGly,
TGAGlu and
TerLys where
Ter can be the two stop codons
TAA and
TGA (
Table 7). All the 20 amino acids are involved in the 83
dipeptides (
Table 8). All the 20 amino acids occur in the first position of
dipeptides. Five amino acids
Asn,
Asp,
Gln, Met and
Trp do not occur in their second position which are the five amino acids not involved in the
dipeptides. In the 83
dipeptides,
Pro and
Gly are involved 20 and 19 times, respectively, while
Met and
Trp only twice and once, respectively.
The 96 unidirectional multiple-frame dicodons
(
Table 3) code 40
dipeptides and three pairs (amino acid, stop codon):
IleTer where
Ter can be the two stop codons
TAA and
TAG,
PheTer where
Ter can be the three stop codons
TAA,
TAG and
TGA, and
ValTGA (
Table 9). All the 20 amino acids are involved in the 40
dipeptides (
Table 10). Five amino acids are
Asn,
Asp,
Gln, Met and
Trp do not occur in the first position of
dipeptides which are the five amino acids not involved in the
dipeptides. All the 20 amino acids occur in their second position. In the 40
dipeptides, two amino acids
Lys and
Phe are involved eight times while
Asn only once.
The
dipeptides among 400, i.e., 28.5%, are coded by
dicodons (
,
,
) among 4096, i.e., 5.1% (
Table 11). As a consequence, 286
dipeptides, i.e., 71.5%, are coded by 3888 single-frame dicodons
, i.e., 94.9%. There is also a strong asymmetry between the number of
dipeptides coded by one direction or other direction: 83
dipeptides (
Table 7) versus 40
dipeptides (
Table 9). This asymmetry may be related to the gene translation in the
direction, the
dicodons having an unambiguous trinucleotide decoding in the
direction.
Five dipeptides
GlyAla,
GlyVal,
PheSer,
ProLeu and
ProArg are the most strongly coded, each by five
dicodons (
Table 12), e.g.,
GlyAla is coded by one
dicodon
GGCGCG (
Table 7), and four
dicodons
GGGGCA, GGGGCC, GGGGCG and
GGGGCT (
Table 9). The
and
dipeptides could have particular spatial structures and biological functions in extant and primitive proteins which remain to be identified.
4. Discussion
For the first time to our knowledge, new definitions of motifs in genes are presented. The single-frame motifs (unambiguous trinucleotide decoding in the two and directions) and the multiple-frame motifs (ambiguous trinucleotide decoding in at least one direction) form a partition of genes. Several classes of motifs are defined and analysed: (i) unidirectional multiple-frame motifs (ambiguous trinucleotide decoding in the direction only); (ii) unidirectional multiple-frame motifs (ambiguous trinucleotide decoding in the direction only); and (iii) bidirectional multiple-frame motifs (ambiguous trinucleotide decoding in the two and directions). The distribution of the single-frame motifs and the 5′ unambiguous motifs (unambiguous trinucleotide decoding in the direction only) are studied without and with constraints.
The proportion of motifs drastically decreases with their trinucleotide length. The motifs become absent () when their length trinucleotides and the number of motifs becomes already higher than the number of motifs when their length trinucleotides. As expected, the proportion of motifs decreases more slowly than that of motifs. The motifs become absent () when their length trinucleotides. Thus with the motifs, there is a length increase of trinucleotides in the trinucleotide decoding.
The proportions of and motifs with initiation and stop codons are lower than their respective non-constrained motifs. In contrasts, their proportions in motifs without periodic codons are higher than their respective non-constrained motifs. The proportions of and motifs with antiparallel complementarity are identical. Antiparallel complementarity increases the proportion of motifs but decreases the proportion of motifs, compared to their respective non-constrained motifs. The proportions of motifs with parallel complementarity and motifs without constraints follow a similar distribution. Finally, parallel complementarity increases the proportions of both motifs and motifs compared to their respective non-constrained motifs. Taken together, these results suggest that the complementarity property involved in the antiparallel (DNA double helix, RNA stem) and parallel sequences could also be fundamental for coding genes with unambiguous trinucleotide decoding, strictly in the two and directions ( motifs) or conserved in the direction but relaxed-lost in the direction ( motifs).
The single-frame motifs
with a property of trinucleotide decoding and the framing motifs
with a property of reading frame decoding could have operated in the primitive soup for constructing the modern genetic code and the extant genes [
31]. They could have been involved in the stage without anticodon-amino acid interactions to form peptides from prebiotically amino acids [
32]. They could also have been related in the Implicated Site Nucleotides (ISN) of RNA interacting with the amino acids at the primitive step of life (review in [
33]). According to a great number of biological experiments, the ISN structure contains nucleotides in fixed and variable positions, as well as an important trinucleotide for interacting with the amino acid (see e.g., the recent review in [
34]). However, the general structure of the aptamers binding amino acids, in particular its nucleotide length, its amino acid binding loop and its nucleotide position, is still an open problem. Similar arguments could hold for the ribonucleopeptides which could be implicated in a primitive T box riboswitch functioning as an aminoacyl-tRNA synthetase and a peptidyl-transferase ribozyme [
35]. The single-frame motifs
and the framing motifs
with their properties to decode the trinucleotides and the reading frame could have been necessary for the evolutionary construction of the modern genetic code.