Toward Experimental Evolution with Giant Vesicles
Abstract
1. Introduction
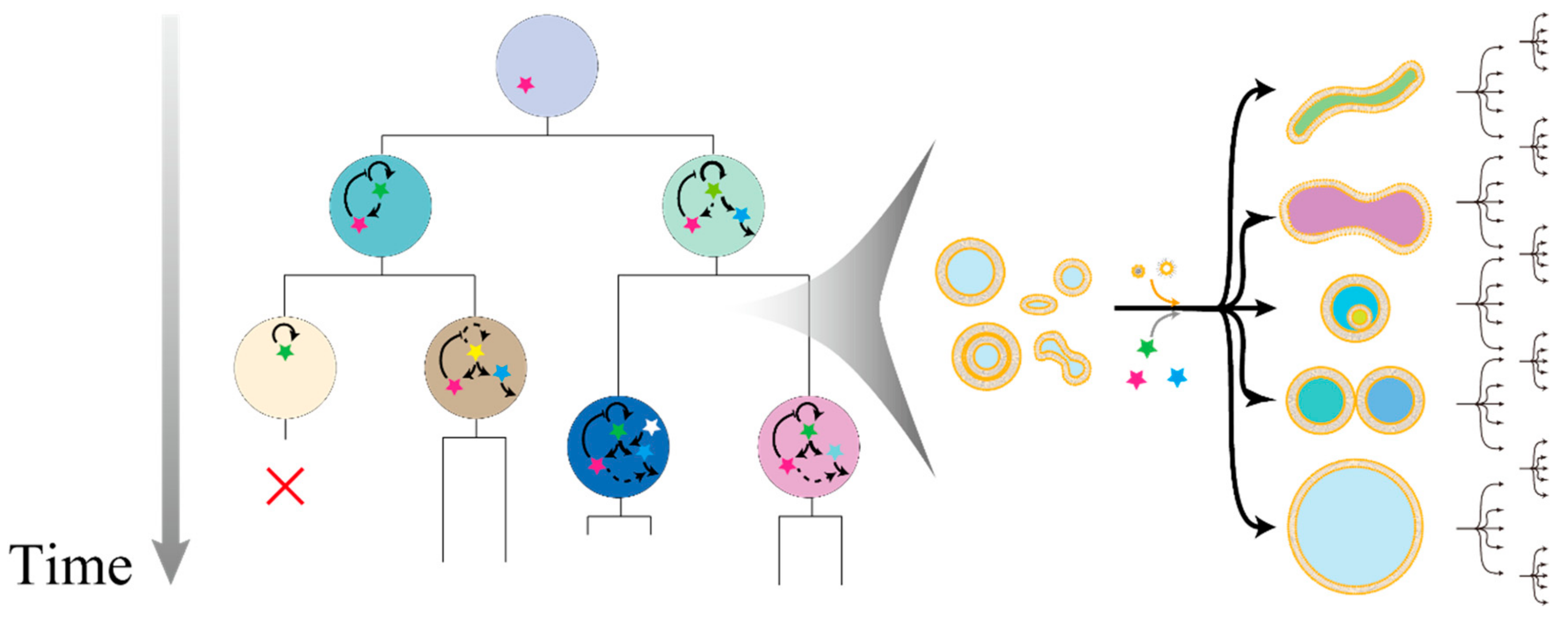
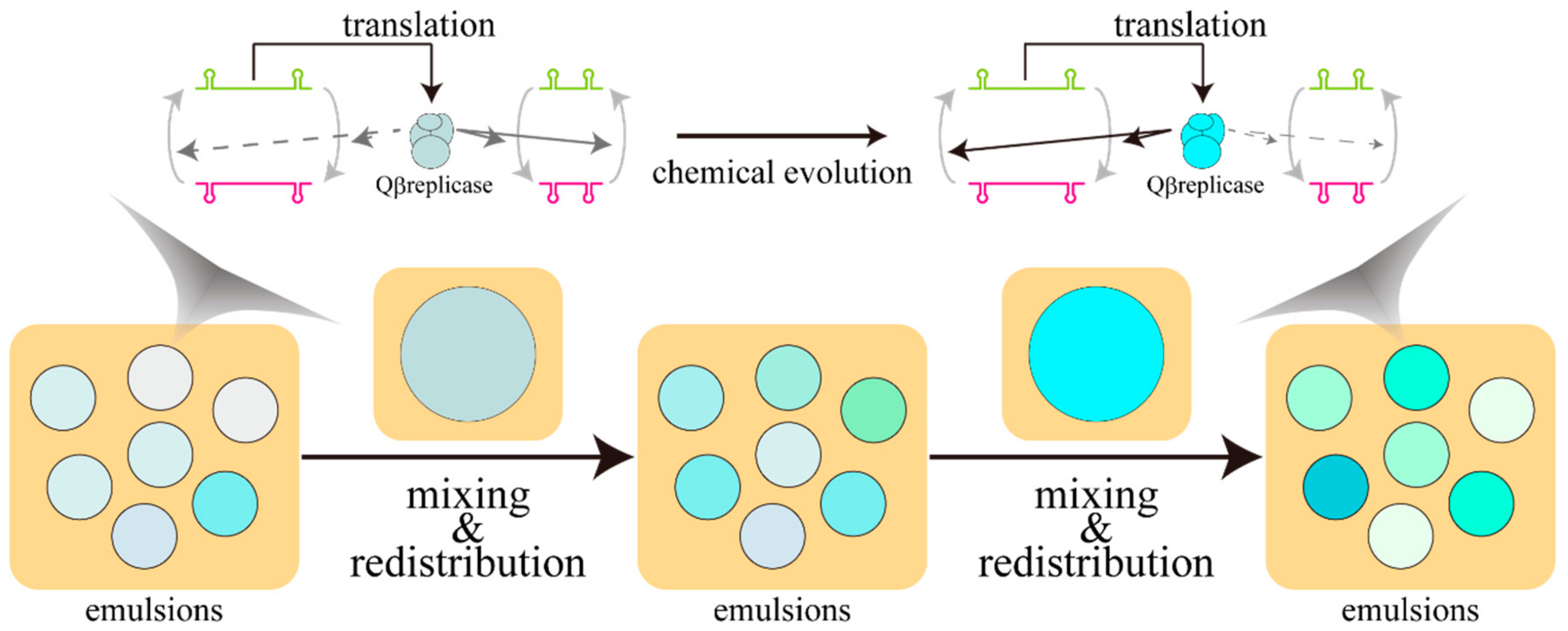
2. Experimental Evolution with Water-in-Oil Emulsions
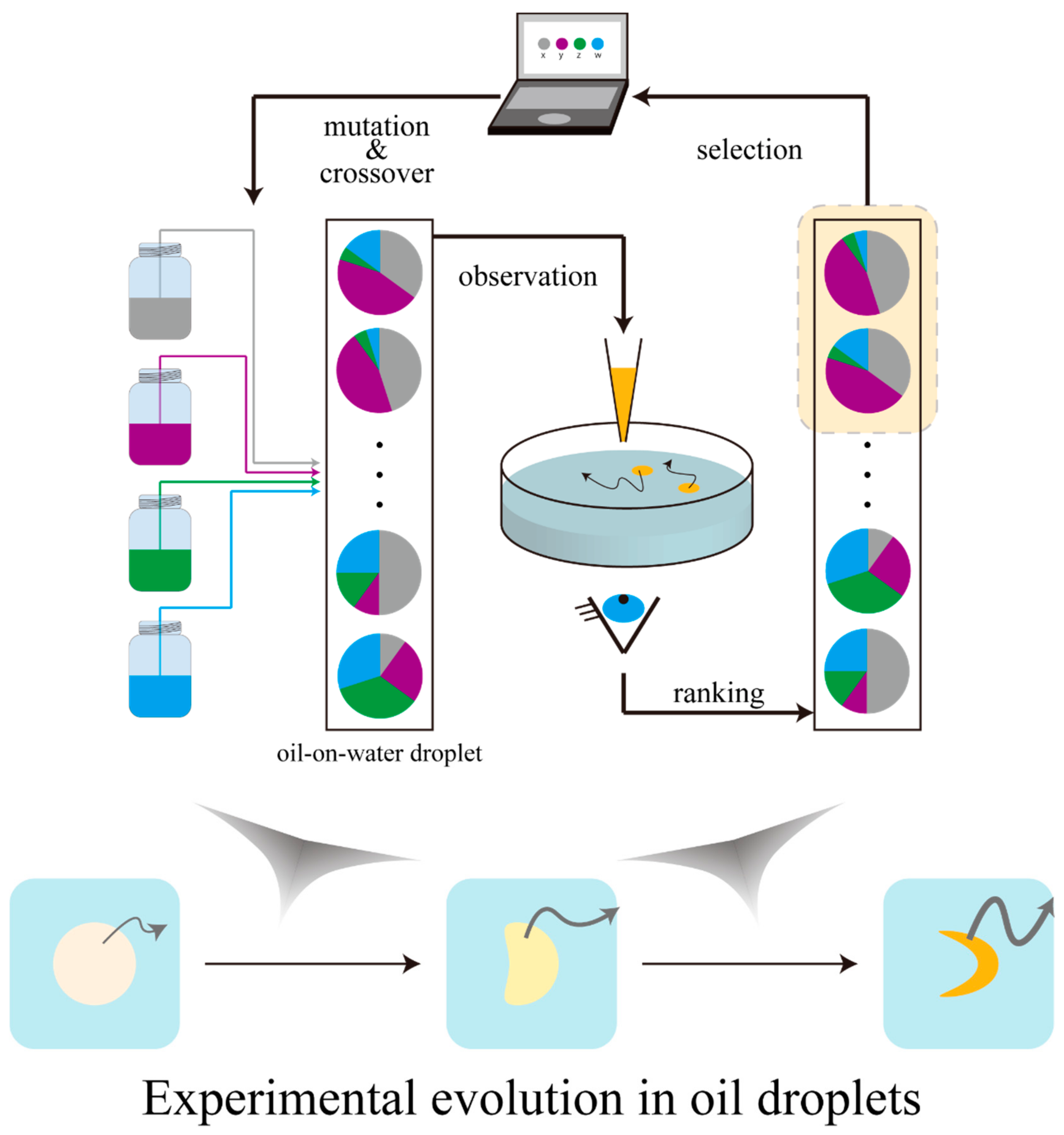
3. Experimental Evolution with Oil-on-Water Droplets
4. Experimental Evolution with Vesicles
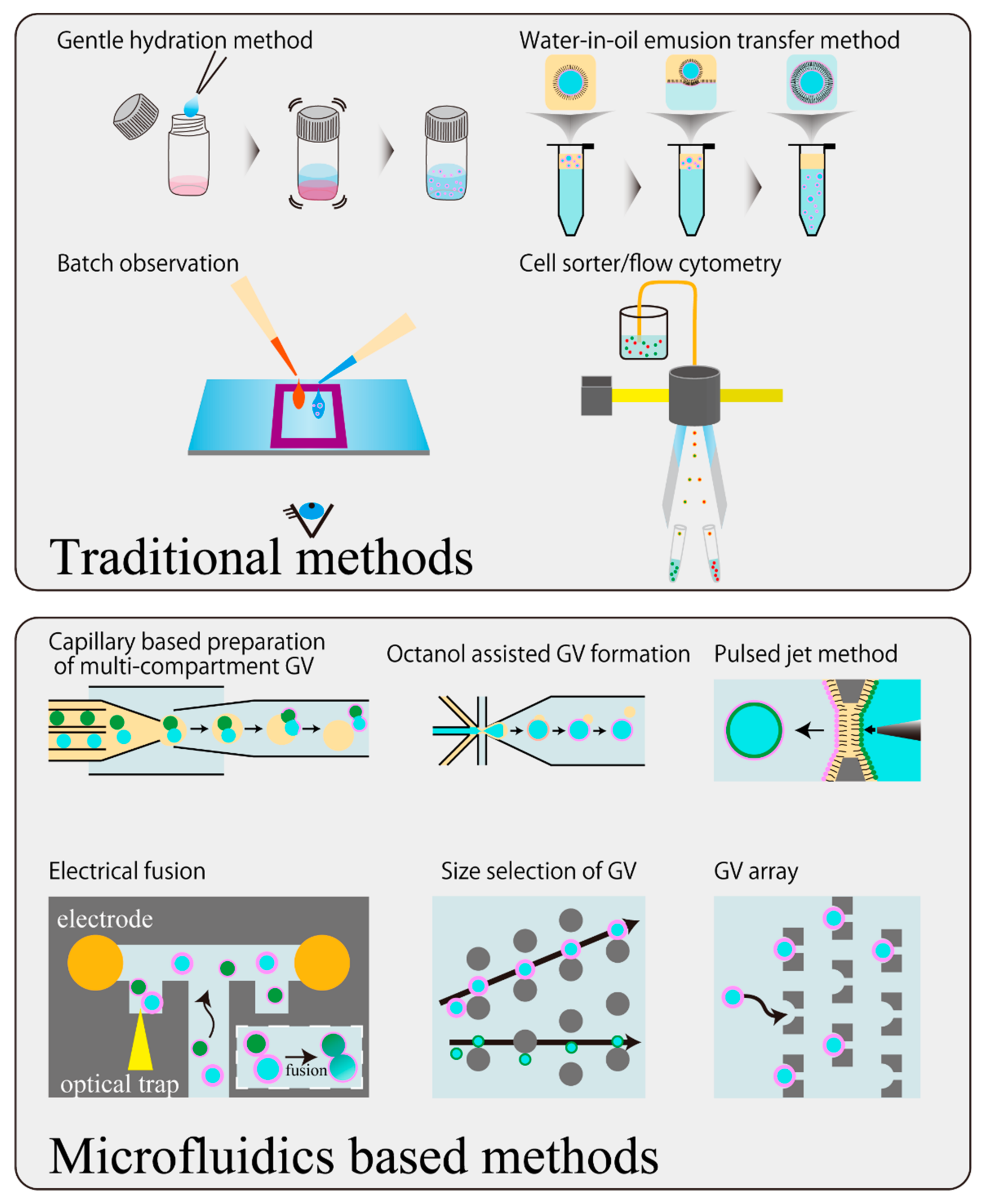
5. Development of Handling Techniques of GVs Using a Microfluidic Device
6. Summary and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hartwell, L.H.; Hopfield, J.J.; Leibler, S.; Murray, A.W. From molecular to modular cell biology. Nature 1999, 402, C47–C52. [Google Scholar] [CrossRef] [PubMed]
- Elowitz, M.B.; Levine, A.J.; Siggia, E.D.; Swain, P.S. Stochastic gene expression in a single cell. Science 2002, 297, 1183–1186. [Google Scholar] [CrossRef] [PubMed]
- Vickers, C.E. The minimal genome comes of age. Nat. Biotechnol. 2016, 34, 623–624. [Google Scholar] [CrossRef] [PubMed]
- Langton, C.G. Artificial Life: Proceedings of an Interdisciplinary Workshop on the Synthesis and Simulation of Living Systems; Addison-Wesley Longman: Boston, MA, USA, 1989. [Google Scholar]
- Blain, J.C.; Szostak, J.W. Progress Toward Synthetic Cells. Annu. Rev. Biochem. 2014, 83, 615–640. [Google Scholar] [CrossRef] [PubMed]
- Eigen, M. Selforganization of matter and evolution of biological macromolecules. Naturwissenschaften 1971, 58, 465–523. [Google Scholar] [CrossRef] [PubMed]
- Eigen, M.; Schuster, P. Hypercycle–principle of natural self-organization. A emergence of hypercycle. Naturwissenschaften 1979, 64, 541–565. [Google Scholar] [CrossRef]
- Bresch, C.; Niesert, U.; Harnasch, D. Hypercycles, parasites and packages. J. Theor. Biol. 1980, 85, 399–405. [Google Scholar] [CrossRef]
- Ruiz-Mirazo, K.; Peretó, J.; Moreno, A. A universal definition of life: Autonomy and open-ended evolution. Orig. Life Evol. Biospheres 2004, 34, 323–346. [Google Scholar] [CrossRef]
- Luisi, P.L. Chemistry Constraints on the Origin of Life. Isr. J. Chem. 2015, 55, 906–918. [Google Scholar] [CrossRef]
- Gilbert, W. Origin of life—The RNA world. Nature 1986, 319, 618. [Google Scholar] [CrossRef]
- Joyce, G.F. The antiquity of RNA-based evolution. Nature 2002, 418, 214–221. [Google Scholar] [CrossRef] [PubMed]
- Dale, T. Protein and nucleic acid together: A mechanism for the emergence of biological selection. J. Theor. Biol. 2006, 240, 337–342. [Google Scholar] [CrossRef] [PubMed]
- Carny, O.; Gazit, E. A model for the role of short self-assembled peptides in the very early stages of the origin of life. FASEB J. 2005, 19, 1051–1055. [Google Scholar] [CrossRef] [PubMed]
- Segré, D.; Ben-Eli, D.; Deamer, D.W.; Lancet, D. The lipid world. Orig. Life Evol. Biospheres 2001, 31, 119–145. [Google Scholar] [CrossRef]
- Benner, S.A. Defining Life. Astrobiology 2010, 10, 1021–1030. [Google Scholar] [CrossRef] [PubMed]
- Tawfik, D.S.; Griffiths, A.D. Man-made cell-like compartments for molecular evolution. Nat. Biotechnol. 1998, 16, 652–656. [Google Scholar] [CrossRef] [PubMed]
- Griffiths, A.D.; Tawfik, D.S. Miniaturising the laboratory in emulsion droplets. Trends Biotechnol. 2006, 24, 395–402. [Google Scholar] [CrossRef] [PubMed]
- Toyota, T.; Maru, N.; Hanczyc, M.M.; Ikegami, T.; Sugawara, T. Self-Propelled Oil Droplets Consuming “Fuel” Surfactant. J. Am. Chem. Soc. 2009, 131, 5012–5013. [Google Scholar] [CrossRef] [PubMed]
- Banno, T.; Kuroha, R.; Toyota, T. pH-Sensitive Self-Propelled Motion of Oil Droplets in the Presence of Cationic Surfactants Containing Hydrolyzable Ester Linkages. Langmuir 2012, 28, 1190–1195. [Google Scholar] [CrossRef] [PubMed]
- Nagai, K.; Sumino, Y.; Kitahata, H.; Yoshikawa, K. Mode selection in the spontaneous motion of an alcohol droplet. Phys. Rev. E 2005, 71, 065301. [Google Scholar] [CrossRef] [PubMed]
- Hanczyc, M.M. Droplets: Unconventional protocell model with life-like dynamics and room to grow. Life 2014, 4, 1038–1049. [Google Scholar] [CrossRef] [PubMed]
- Bangham, A.D.; Horne, R.W. Negative staining of phospholipids and their structural modification by surface-active agents as observed in electron microscope. J. Mol. Biol. 1964, 8, 660–668. [Google Scholar] [CrossRef]
- Stano, P.; Luisi, P.L. Achievements and open questions in the self-reproduction of vesicles and synthetic minimal cells. Chem. Commun. 2010, 46, 3639–3653. [Google Scholar] [CrossRef] [PubMed]
- Wick, R.; Walde, P.; Luisi, P.L. Light-microscopic investigations of the autocatalytic self-reproduction of giant vesicles. J. Am. Chem. Soc. 1995, 117, 1435–1436. [Google Scholar] [CrossRef]
- Zhu, T.F.; Szostak, J.W. Coupled Growth and Division of Model Protocell Membranes. J. Am. Chem. Soc. 2009, 131, 5705–5713. [Google Scholar] [CrossRef] [PubMed]
- Kurihara, K.; Tamura, M.; Shohda, K.; Toyota, T.; Susuki, K.; Sugawara, T. Self-reproduction of supramolecular giant vesicles combined with the amplification of encapsulated DNA. Nat. Chem. 2011, 3, 775–781. [Google Scholar] [CrossRef] [PubMed]
- Hardy, M.D.; Yang, J.; Selimkhanov, J.; Cole, C.M.; Tsimring, L.S.; Devaraj, N.K. Self-reproducing catalyst drives repeated phospholipid synthesis and membrane growth. Proc. Natl. Acad. Sci. USA 2015, 112, 8187–8192. [Google Scholar] [CrossRef] [PubMed]
- Horinouchi, T.; Minamoto, T.; Suzuki, S.; Shimizu, H.; Furusawa, C. Development of an Automated Culture System for Laboratory Evolution. J. Lab. Autom. 2014, 19, 478–482. [Google Scholar] [CrossRef] [PubMed]
- Miller, O.J.; Bernath, K.; Agresti, J.J.; Amitai, G.; Kelly, B.T.; Mastrobattista, E.; Taly, V.; Magdassi, S.; Tawfik, D.S.; Griffiths, A.D. Directed evolution by in vitro compartmentalization. Nat. Methods 2006, 3, 561–570. [Google Scholar] [CrossRef] [PubMed]
- Ichihashi, N.; Usui, K.; Kazuta, Y.; Sunami, T.; Matsuura, T.; Yomo, T. Darwinian evolution in a translation-coupled RNA replication system within a cell-like compartment. Nat. Commun. 2013, 4, 2494. [Google Scholar] [CrossRef] [PubMed]
- Holland, J.H. Genetic algorithms. Sci. Am. 1992, 267, 66–72. [Google Scholar] [CrossRef]
- Matsumura, S.; Kun, A.; Ryckelynck, M.; Coldren, F.; Szilágyi, A.; Jossinet, F.; Rick, C.; Nghe, P.; Szathmáry, E.; Griffiths, A.D. Transient compartmentalization of RNA replicators prevents extinction due to parasites. Science 2016, 354, 1293–1296. [Google Scholar] [CrossRef] [PubMed]
- Baret, J.C.; Miller, O.J.; Taly, V.; Ryckelynck, M.; El-Harrak, A.; Frenz, L.; Rick, C.; Samuels, M.L.; Hutchison, J.B.; Agresti, J.J.; et al. Fluorescence-activated droplet sorting (FADS): Efficient microfluidic cell sorting based on enzymatic activity. Lab Chip 2009, 9, 1850–1858. [Google Scholar] [CrossRef] [PubMed]
- Hanczyc, M.M.; Toyota, T.; Ikegami, T.; Packard, N.; Sugawara, T. Fatty acid chemistry at the oil-water interface: Self-propelled oil droplets. J. Am. Chem. Soc. 2007, 129, 9386–9391. [Google Scholar] [CrossRef] [PubMed]
- Banno, T.; Asami, A.; Ueno, N.; Kitahata, H.; Koyano, Y.; Asakura, K.; Toyota, T. Deformable Self-Propelled Micro-Object Comprising Underwater Oil Droplets. Sci. Rep. 2016, 6, 31292. [Google Scholar] [CrossRef] [PubMed]
- Čejková, J.; Novák, M.; Štěpánek, F.; Hanczyc, M.M. Dynamics of Chemotactic Droplets in Salt Concentration Gradients. Langmuir 2014, 30, 11937–11944. [Google Scholar] [CrossRef] [PubMed]
- Scriven, L.E.; Sternling, C.V. Marangoni effects. Nature 1960, 187, 186–188. [Google Scholar] [CrossRef]
- Segré, D.; Lancet, D.; Kedem, O.; Pilpel, Y. Graded autocatalysis replication domain (GARD): Kinetic analysis of self-replication in mutually catalytic sets. Orig. Life Evol. Biosph. 1998, 28, 501–514. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, J.M.; Hinkley, T.; Taylor, J.W.; Yanev, K.; Cronin, L. Evolution of oil droplets in a chemorobotic platform. Nat. Commun. 2014, 5, 5571. [Google Scholar] [CrossRef] [PubMed]
- Parrilla-Gutierrez, J.M.; Tsuda, S.; Grizou, J.; Taylor, J.; Henson, A.; Cronin, L. Adaptive artificial evolution of droplet protocells in a 3D-printed fluidic chemorobotic platform with configurable environments. Nat. Commun. 2017, 8, 1144. [Google Scholar] [CrossRef] [PubMed]
- Points, L.J.; Taylor, J.W.; Grizou, J.; Donkers, K.; Cronin, L. Artificial intelligence exploration of unstable protocells leads to predictable properties and discovery of collective behavior. Proc. Natl. Acad. Sci. USA 2018, 115, 885–890. [Google Scholar] [CrossRef] [PubMed]
- Fujii, S.; Matsuura, T.; Sunami, T.; Kazuta, Y.; Yomo, T. In vitro evolution of alpha-hemolysin using a liposome display. Proc. Natl. Acad. Sci. USA 2013, 110, 16796–16801. [Google Scholar] [CrossRef] [PubMed]
- Fujii, S.; Matsuura, T.; Sunami, T.; Nishikawa, T.; Kazuta, Y.; Yomo, T. Liposome display for in vitro selection and evolution of membrane proteins. Nat. Protoc. 2014, 9, 1578–1591. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, S.; Terai, T.; Yoshikawa, Y.; Ohkawa, R.; Ebihara, M.; Hayashi, M.; Taniguchi, K.; Nemoto, N. In vitro selection of random peptides against artificial lipid bilayers: A potential tool to immobilize molecules on membranes. Chem. Commun. 2017, 53, 3458–3461. [Google Scholar] [CrossRef] [PubMed]
- Noireaux, V.; Libchaber, A. A vesicle bioreactor as a step toward an artificial cell assembly. Proc. Natl. Acad. Sci. USA 2004, 101, 17669–17674. [Google Scholar] [CrossRef] [PubMed]
- Gánti, T. On the early evolutionary origin of biological periodicity. Cell Biol. Int. 2002, 26, 729–735. [Google Scholar] [CrossRef] [PubMed]
- Paterson, D.J.; Reboud, J.; Wilson, R.; Tassieri, M.; Cooper, J.M. Integrating microfluidic generation, handling and analysis of biomimetic giant unilamellar vesicles. Lab Chip 2014, 14, 1806–1810. [Google Scholar] [CrossRef] [PubMed]
- Jang, H.; Hu, P.C.; Jung, S.; Kim, W.Y.; Kim, S.M.; Malmstadt, N.; Jeon, T.J. Automated formation of multicomponent-encapuslating vesosomes using continuous flow microcentrifugation. Biotechnol. J. 2013, 8, 1341–1346. [Google Scholar] [CrossRef] [PubMed]
- Deng, N.N.; Yelleswarapu, M.; Huck, W.T.S. Monodisperse Uni- and Multicompartment Liposomes. J. Am. Chem. Soc. 2016, 138, 7584–7591. [Google Scholar] [CrossRef] [PubMed]
- Deng, N.N.; Yelleswarapu, M.; Zheng, L.; Huck, W.T. Microfluidic Assembly of Monodisperse Vesosomes as Artificial Cell Models. J. Am. Chem. Soc. 2017, 139, 587–590. [Google Scholar] [CrossRef] [PubMed]
- Deshpande, S.; Dekker, C. On-chip microfluidic production of cell-sized liposomes. Nat. Protoc. 2018, 13, 856–874. [Google Scholar] [CrossRef] [PubMed]
- Deshpande, S.; Caspi, Y.; Meijering, A.E.C.; Dekker, C. Octanol-assisted liposome assembly on chip. Nat. Commun. 2016, 7, 10447. [Google Scholar] [CrossRef] [PubMed]
- Abate, A.R.; Hung, T.; Mary, P.; Agresti, J.J.; Weitz, D.A. High-throughput injection with microfluidics using picoinjectors. Proc. Natl. Acad. Sci. USA 2010, 107, 19163–19166. [Google Scholar] [CrossRef] [PubMed]
- Weiss, M.; Frohnmayer, J.P.; Benk, L.T.; Haller, B.; Janiesch, J.W.; Heitkamp, T.; Börsch, M.; Lira, R.B.; Dimova, R.; Lipowsky, R.; et al. Sequential bottom-up assembly of mechanically stabilized synthetic cells by microfluidics. Nat. Mater. 2018, 17, 89–96. [Google Scholar] [CrossRef] [PubMed]
- Gotanda, M.; Kamiya, K.; Osaki, T.; Fujii, S.; Misawa, N.; Miki, N.; Takeuchi, S. Sequential generation of asymmetric lipid vesicles using a pulsed-jetting method in rotational wells. Sens. Actuators B-Chem. 2018, 261, 392–397. [Google Scholar] [CrossRef]
- Shiomi, H.; Tsuda, S.; Suzuki, H.; Yomo, T. Liposome-Based Liquid Handling Platform Featuring Addition, Mixing, and Aliquoting of Femtoliter Volumes. PLoS ONE 2014, 9, e101820. [Google Scholar] [CrossRef] [PubMed]
- Robinson, T.; Verboket, P.E.; Eyer, K.; Dittrich, P.S. Controllable electrofusion of lipid vesicles: Initiation and analysis of reactions within biomimetic containers. Lab Chip 2014, 14, 2852–2859. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.Y.; Hu, N.; Yeh, L.H.; Zheng, X.; Yang, J.; Joo, S.W.; Qian, S. Electroformation and electrofusion of giant vesicles in a microfluidic device. Colloids Surf. B 2013, 110, 81–87. [Google Scholar] [CrossRef] [PubMed]
- Deshpande, S.; Spoelstra, W.K.; van Doom, M.; Kerssemakers, J.; Dekker, C. Mechanical Division of Cell-Sized Liposomes. ACS Nano 2018, 12, 2560–2568. [Google Scholar] [CrossRef] [PubMed]
- Nuss, H.; Chevallard, C.; Guenoun, P.; Malloggi, F. Microfluidic trap-and-release system for Lab Chip-based studies on giant vesicles. Lab Chip 2012, 12, 5257–5261. [Google Scholar] [CrossRef] [PubMed]
- Robinson, T.; Kuhn, P.; Eyer, K.; Dittrich, P.S. Microfluidic trapping of giant unilamellar vesicles to study transport through a membrane pore. Biomicrofluidics 2013, 7, 044105. [Google Scholar] [CrossRef] [PubMed]
- Kazayama, Y.; Teshima, T.; Osaki, T.; Takeuchi, S.; Toyota, T. Integrated Microfluidic System for Size-Based Selection and Trapping of Giant Vesicles Anal. Chem. 2016, 88, 1111–1116. [Google Scholar] [CrossRef]
- Adamala, K.P.; Martin-Alarcon, D.A.; Guthrie-Honea, K.R.; Boyden, E.S. Engineering genetic circuit interactions within and between synthetic minimal cells. Nat. Chem. 2017, 9, 431–439. [Google Scholar] [CrossRef] [PubMed]
- Guttenberg, N.; Virgo, N.; Chandru, K.; Scharf, C.; Mamajanov, I. Bulk measurements of messy chemistries are needed for a theory of the origins of life. Philos. Trans. R. Soc. A 2017, 375, 20160347. [Google Scholar] [CrossRef] [PubMed]
- Mamajanov, I.; Cody, G.D. Protoenzymes: The case of hyperbranched polyesters. Philos. Trans. R. Soc. A 2017, 375, 20160357. [Google Scholar] [CrossRef] [PubMed]
- Nozoe, T.; Kussell, E.; Wakamoto, Y. Inferring fitness landscapes and selection on phenotypic states from single-cell genealogical data. PLoS Genet. 2017, 13, e1006653. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, K.; Furusawa, C. An evolutionary relationship between genetic variation and phenotypic fluctuation. J. Theor. Biol. 2006, 240, 78–86. [Google Scholar] [CrossRef] [PubMed]
- Mocciaro, A.; Roth, T.L.; Bennet, H.M.; Soumillon, M.; Shah, A.; Hiatt, J.; Chapman, K.; Marson, A.; Lavieu, G. Light-activated cell identification and sorting (LACIS) for selection of edited clones on a nanofluidic device. Commun. Biol. 2018, 1, 41. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sugiyama, H.; Toyota, T. Toward Experimental Evolution with Giant Vesicles. Life 2018, 8, 53. https://doi.org/10.3390/life8040053
Sugiyama H, Toyota T. Toward Experimental Evolution with Giant Vesicles. Life. 2018; 8(4):53. https://doi.org/10.3390/life8040053
Chicago/Turabian StyleSugiyama, Hironori, and Taro Toyota. 2018. "Toward Experimental Evolution with Giant Vesicles" Life 8, no. 4: 53. https://doi.org/10.3390/life8040053
APA StyleSugiyama, H., & Toyota, T. (2018). Toward Experimental Evolution with Giant Vesicles. Life, 8(4), 53. https://doi.org/10.3390/life8040053



