Cloning and Molecular Characterization of an Alpha-Glucosidase (MalH) from the Halophilic Archaeon Haloquadratum walsbyi
Abstract
:1. Introduction
2. Materials and Methods
2.1. Microbial Strain
2.2. Construction of the Expression Plasmid
2.3. Protein Expression and Purification
2.4. Enzymatic Assays
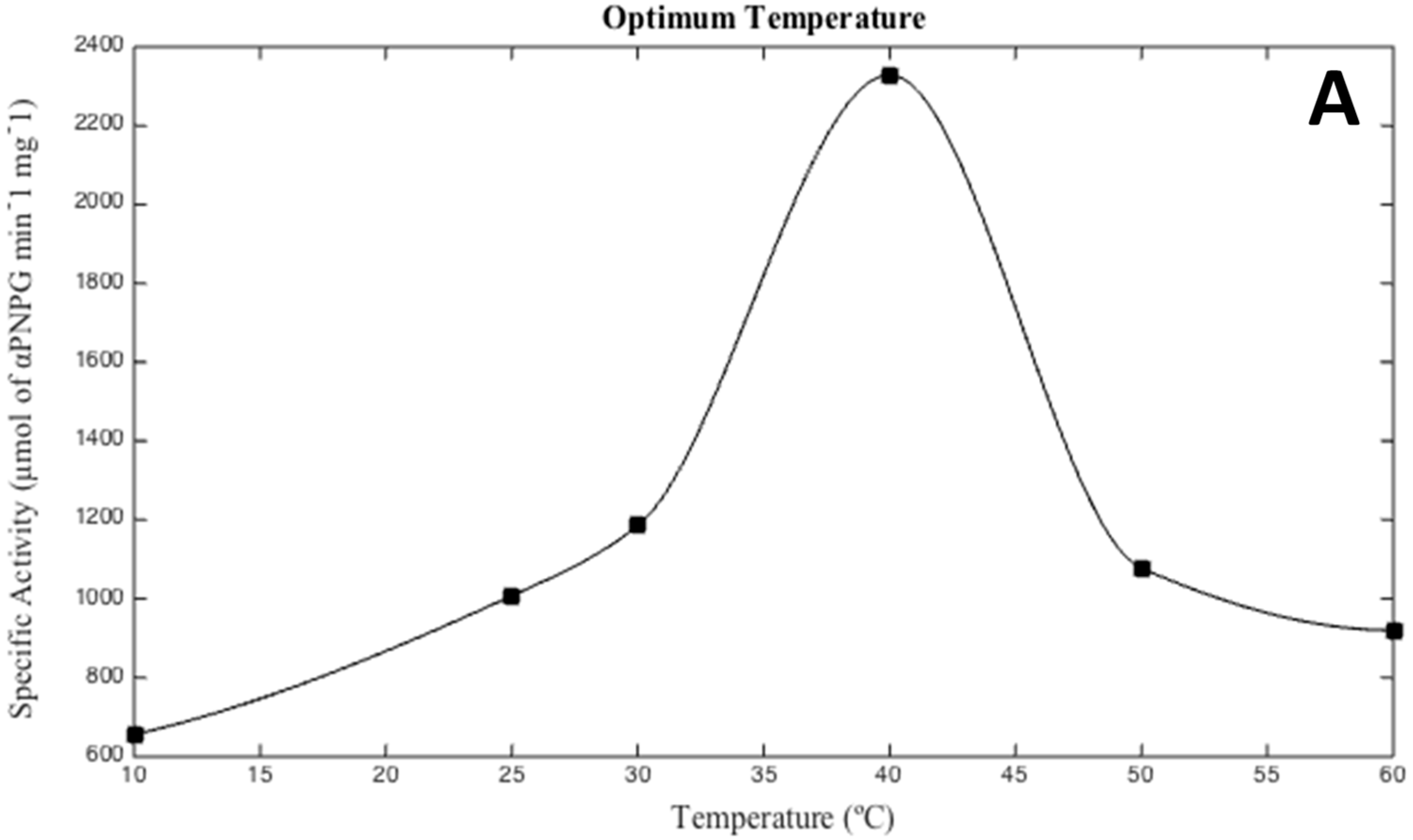
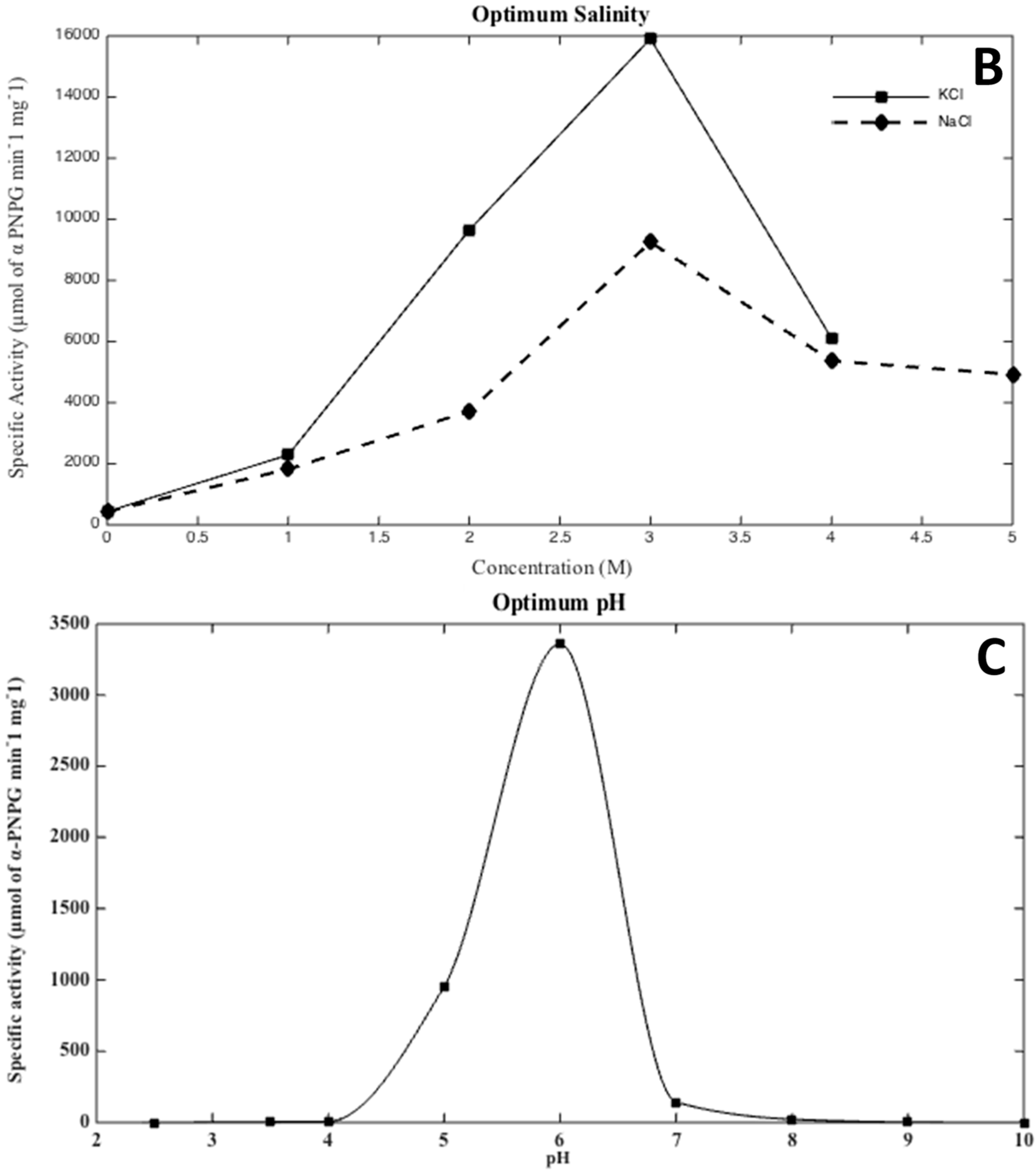
2.5. Effects of Salinity, pH, and Temperature
2.6. Phylogenetic Analyses
2.7. In Silico Functional Characterization of the Hqr. walsbyi Alpha-Glucosidase
3. Results and Discussion
3.1. Identification of a Putative Alpha-Glucosidase Gene in the Hqr. walsbyi C23 Genome
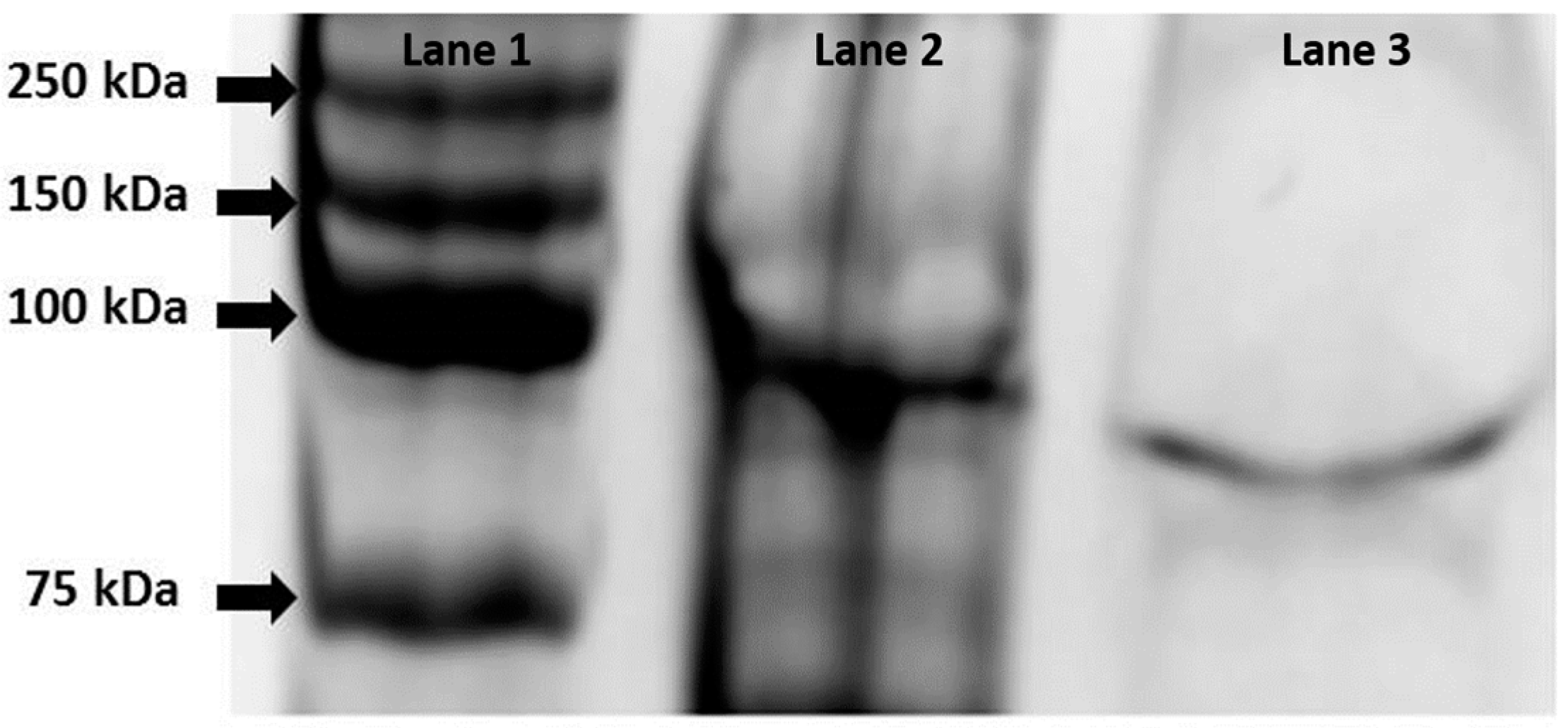
3.2. Biochemical Characterization of the Recombinant Alpha-Glucosidase from Hqr. walsbyi
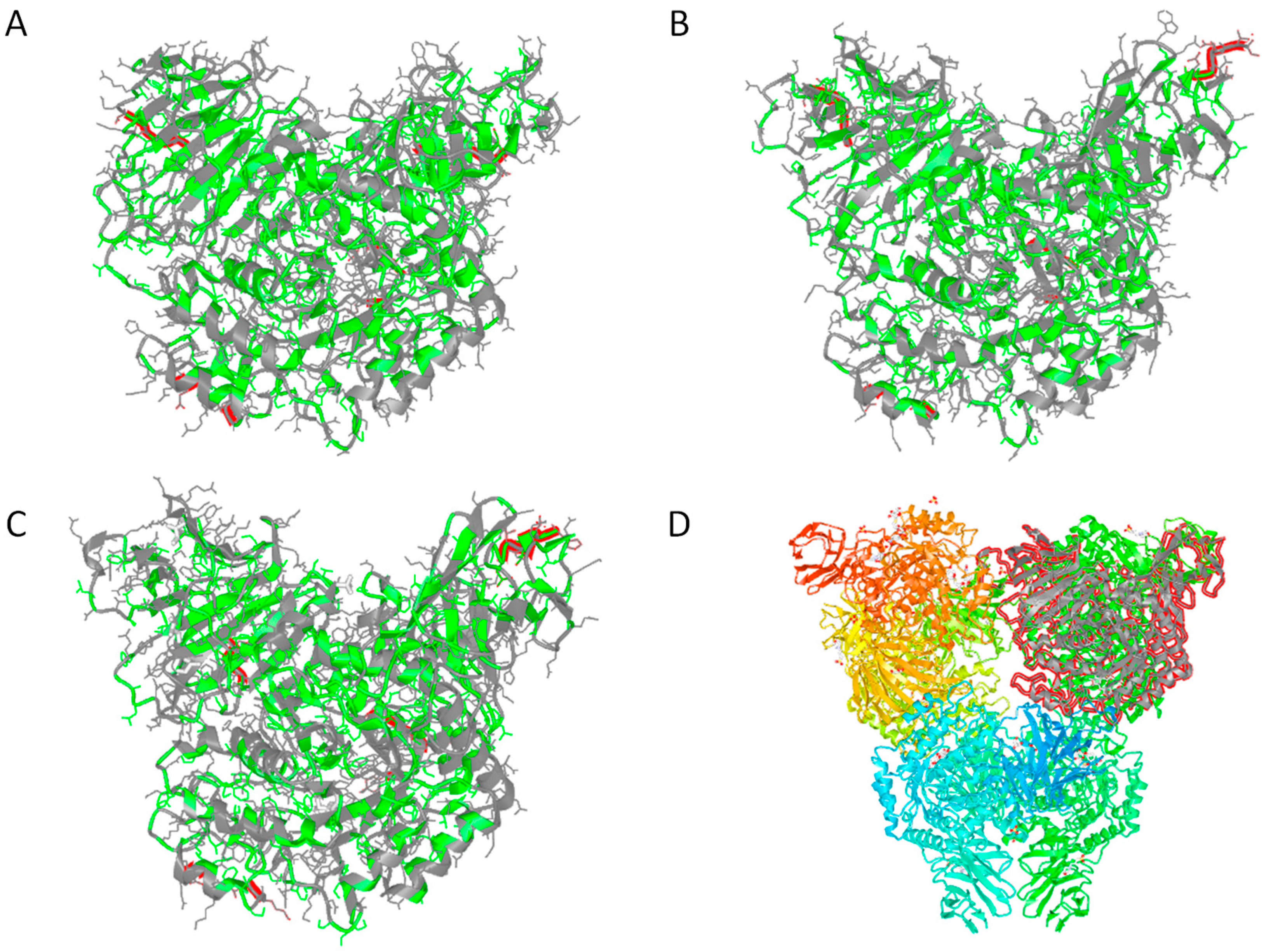
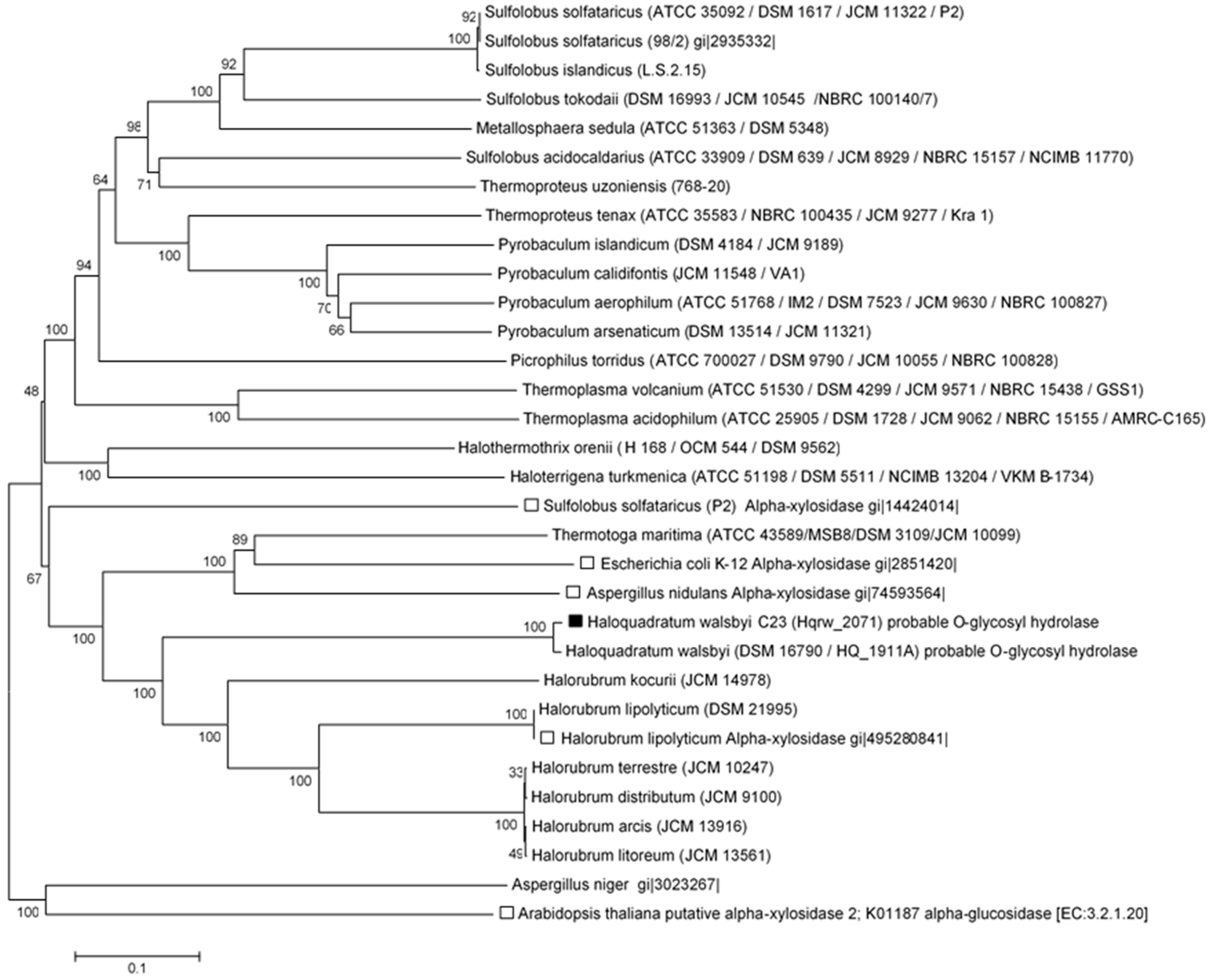
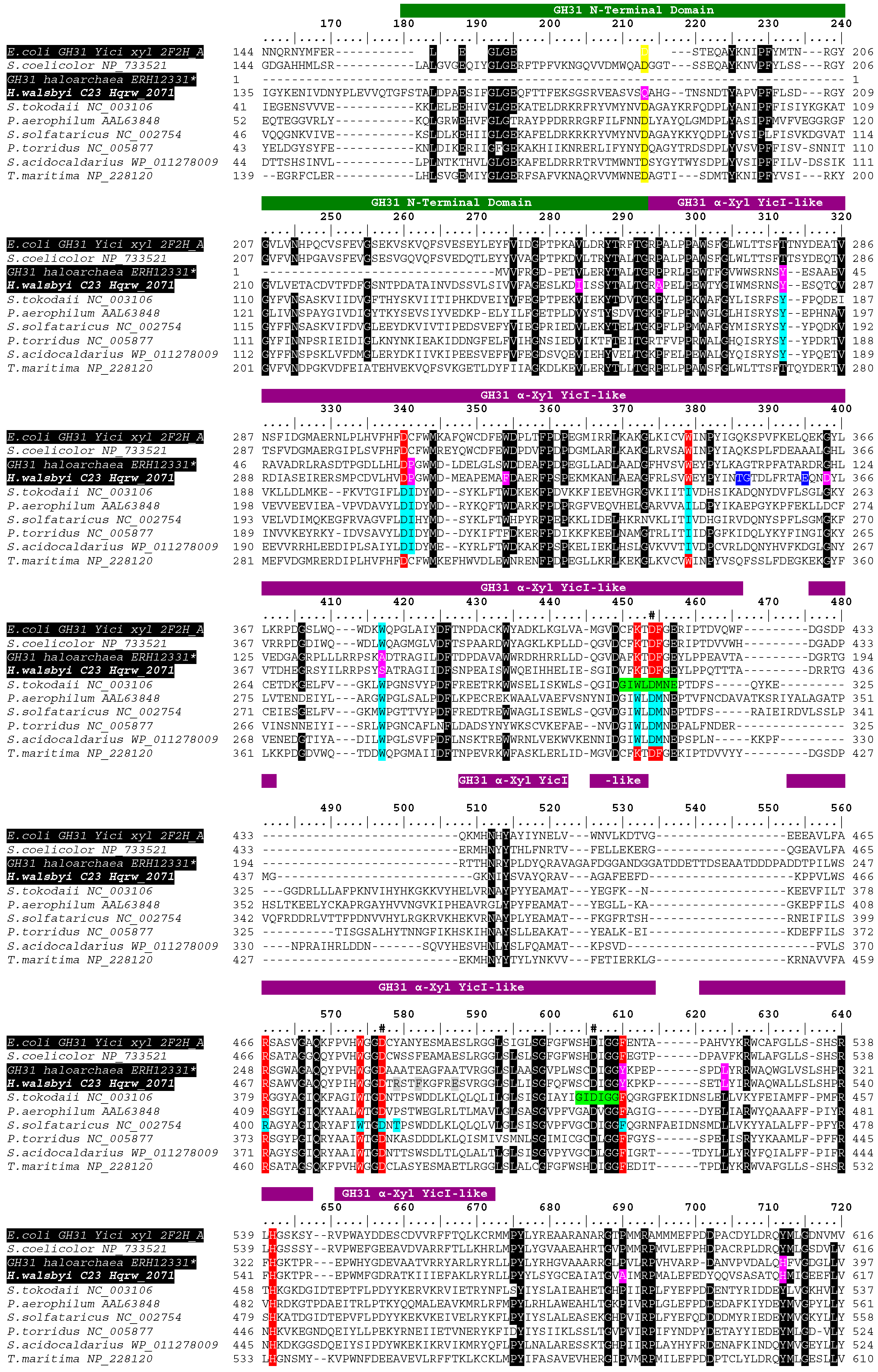
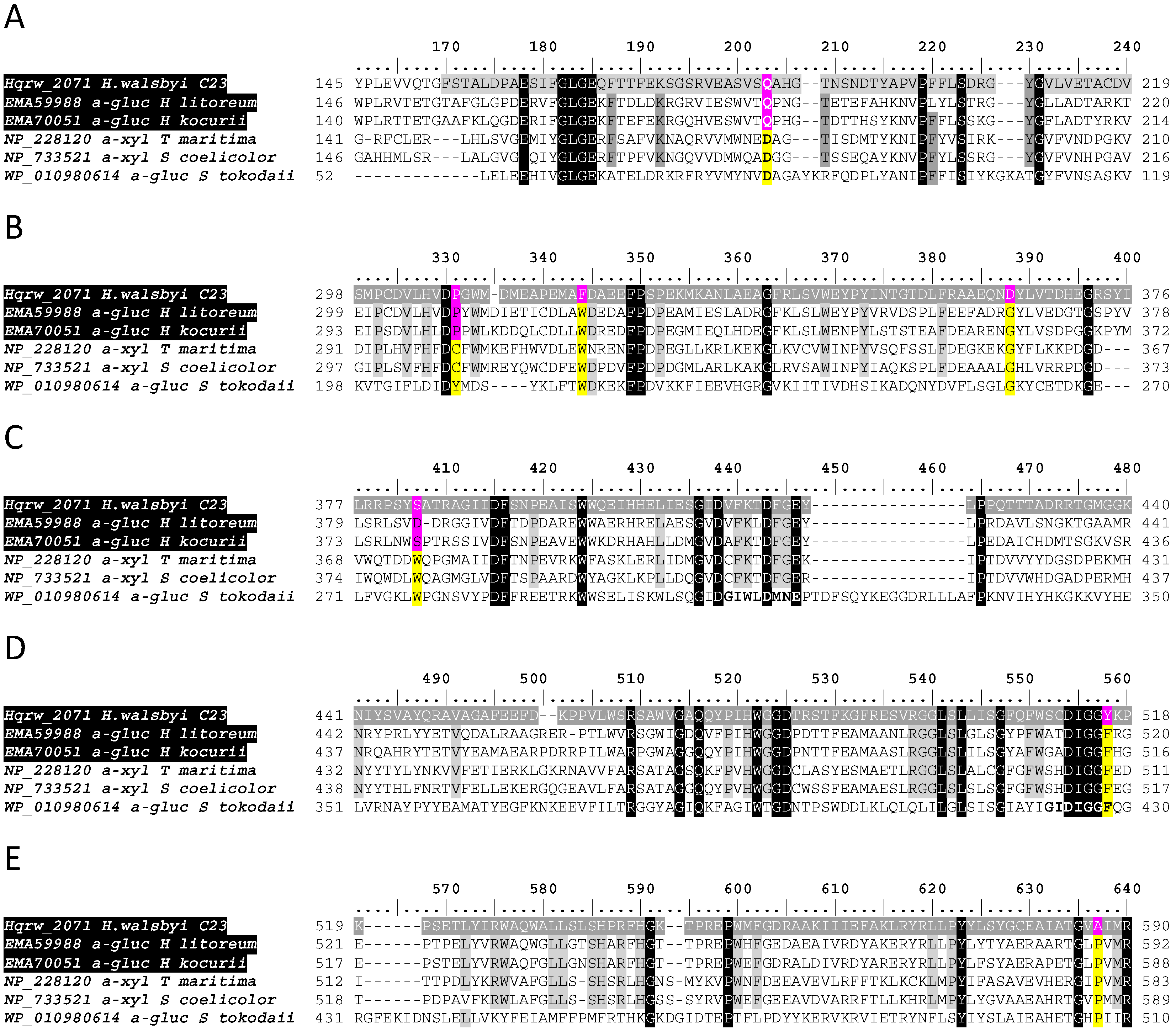
3.3. In Silico Functional Chracterization and Phylogenetic Analysis of the Hqrw_2071 Gene Product
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Burns, D.G.; Janssen, P.H.; Itoh, T.; Kamekura, M.; Li, Z.; Jensen, G.; Rodríguez-Valera, F.; Bolhuis, H.; Dyall-Smith, M.L. Haloquadratum walsbyi gen. nov., sp. nov., the square haloarchaeon of Walsby, isolated from saltern crystallizers in Australia and Spain. Int. J. Syst. Evol. Microbiol. 2007, 57, 387–392. [Google Scholar] [CrossRef] [PubMed]
- Ghai, R.; Pasic, L.; Fernandez, A.B.; Martin-Cuadrado, A.B.; Mizuno, C.M.; McMahon, K.D.; Papke, R.T.; Stepanauskas, R.; Rodriguez-Brito, B.; Rohwer, F.; et al. New abundant microbial groups in aquatic hypersaline environments. Sci. Rep. 2011, 1, 135. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Valera, F.; Ruiz-Berraquero, F.; Ramos-Cormenzana, A. Characteristics of the heterotrophic bacterial populations in hypersaline environments of different salt concentrations. Microb. Ecol. 1981, 7, 235–243. [Google Scholar] [CrossRef] [PubMed]
- Walsby, A. A square bacterium. Nature 1980, 283, 69–71. [Google Scholar] [CrossRef]
- Bolhuis, H.; Palm, P.; Wende, A.; Falb, M.; Rampp, M.; Rodriguez-Valera, F.; Pfeiffer, F.; Oesterhelt, D. The genome of the square archaeon Haloquadratum walsbyi: Life at the limits of water activity. BMC Genom. 2006, 7, 169. [Google Scholar] [CrossRef] [PubMed]
- Ferrer, M.; Golyshina, O.V.; Plou, F.J.; Timmis, K.N.; Golyshin, P.N. A novel α-glucosidase from the acidophilic archaeon Ferroplasma acidiphilum strain Y with high transglycosylation activity and an unusual catalytic nucleophile. Biochem. J. 2005, 391, 269–276. [Google Scholar] [CrossRef] [PubMed]
- Henrissat, B. Glycosidases families. Biochem. Soc. Trans. 1998, 26, 153–156. [Google Scholar] [CrossRef] [PubMed]
- Amoozegar, M.A.; Siroosi, M.; Atashgahi, S.; Smidt, H.; Ventosa, A. Systematics of haloarchaea and biotechnological potential of their hydrolytic enzymes. Microbiology 2017, 163, 623–645. [Google Scholar] [CrossRef] [PubMed]
- Dalmaso, G.Z.L.; Ferreira, D.; Vermelho, A.B. Marine extremophiles a source of hydrolases for biotechnological applications. Mar. Drugs 2015, 13, 1925–1965. [Google Scholar] [CrossRef] [PubMed]
- Rolfsmeier, M.; Haseltine, C.; Bini, E.; Clark, A.; Blum, P. Molecular characterization of the α-glucosidase gene (malA) from the hyperthermophilic archaeon Sulfolobus solfataricus. J. Bacteriol. 1998, 180, 1287–1295. [Google Scholar] [PubMed]
- Kim, V.T.T.; Ryu, S.I.; Lee, K.J.; Kim, E.J.; Lee, S.B. Cloning and characterization of glycogen-debranching enzyme from hyperthermophilic archaeon Sulfolobus shibatae. J. Microbiol. Biotechnol. 2007, 17, 792–799. [Google Scholar]
- Chang, S.T.; Parker, K.N.; Bauer, M.W.; Kelly, R.M. α-Glucosidase from Pyrococcus furiosus. Methods Enzymol. 2001, 330, 260–269. [Google Scholar] [CrossRef] [PubMed]
- Xavier, K.B.; Peist, R.; Kossmann, M.; Boos, W.; Santos, H. Maltose metabolism in the hyperthermophilic archaeon Thermococcus litoralis: Purification and characterization of key enzymes. J. Bacteriol. 1999, 181, 3358–3367. [Google Scholar] [PubMed]
- Montalvo-Rodríguez, R.; Ruiz-Acevedo, A.; López-Garriga, J. New Isolates of Extremely halophilic archaeabacteria (Halobacteria) from Puerto Rico and the Caribbean. Caribb. J. Sci. 1997, 33, 98–104. [Google Scholar]
- Giménez, M.I.; Studdert, C.A.; Sánchez, J.J.; De Castro, R.E. Extracellular protease of Natrialba magadii: Purification and biochemical characterization. Extremophiles 2000, 4, 181–188. [Google Scholar] [CrossRef] [PubMed]
- Hutcheon, G.W.; Vasisht, N.; Bolhuis, A. Characterisation of a highly stable α-amylase from the halophilic archaeon Haloarcula hispanica. Extremophiles 2005, 9, 487–495. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Pomares, F.; Bautista, V.; Ferrer, J.; Pire, C.; Marhuenda-Egea, F.C.; Bonete, M.J. α-Amylase activity from the halophilic archaeon Haloferax mediterranei. Extremophiles 2003, 7, 299–306. [Google Scholar] [CrossRef] [PubMed]
- Coronado, M.J.; Vargas, C.; Mellado, E.; Tegos, G.; Drainas, C.; Nieto, J.J.; Ventosa, A. The α-amylase gene amyH of the moderate halophile Halomonas meridiana: Cloning and molecular characterization. Microbiology 2000, 146, 861–868. [Google Scholar] [CrossRef] [PubMed]
- Bonete, M.J.; Pire, C.; LLorca, F.I.; Camacho, M.L. Glucose dehydrogenase from the halophilic archaeon Haloferax mediterranei: Enzyme purification, characterisation and N-terminal sequence. FEBS Lett. 1996, 383, 227–229. [Google Scholar] [CrossRef]
- Cao, Y.; Liao, L.; Xu, X.W.; Oren, A.; Wu, M. Aldehyde dehydrogenase of the haloalkaliphilic archaeon Natronomonas pharaonis and its function in ethanol metabolism. Extremophiles 2008, 12, 849–854. [Google Scholar] [CrossRef] [PubMed]
- Holmes, M.L.; Scopes, R.K.; Moritz, R.L.; Simpson, R.J.; Englert, C.; Pfeifer, F.; Dyall-Smith, M.L. Purification and analysis of an extremely halophilic β-galactosidase from Haloferax alicantei. Biochim. Biophys. Acta Protein Struct. Mol. Enzymol. 1997, 1337, 276–286. [Google Scholar] [CrossRef]
- Holmes, M.L.; Dyall-Smith, M.L. Sequence and expression of a halobacterial beta-galactosidase gene. Mol. Microbiol. 2000, 36, 114–122. [Google Scholar] [CrossRef] [PubMed]
- Nuttall, S.; Bath, C.; Pfeiffer, M.; Santos, F.; Eichler, J.; Mcalpine, T. Protocols for haloarchaeal genetics. In The Halohandbook, version 7.2; Dyall-Smith, M., Ed.; Max Planck Institute: Martinsried, Germany, 2009; pp. 1–144. [Google Scholar]
- Okuyama, M. Function and structure studies of GH family 31 and 97 alpha-glycosidases. Biosci. Biotechnol. Biochem. 2011, 75, 2269–2277. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J.; et al. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, 222–230. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, A.; Chang, H.Y.; Daugherty, L.; Fraser, M.; Hunter, S.; Lopez, R.; McAnulla, C.; McMenamin, C.; Nuka, G.; Pesseat, S.; et al. The InterPro protein families database: The classification resource after 15 years. Nucleic Acids Res. 2015, 43, D213–D221. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Saitou, N.; Nei, M. The neighbour-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [PubMed]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Marchler-Bauer, A.; Bo, Y.; Han, L.; He, J.; Lanczycki, C.J.; Lu, S.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; et al. CDD/SPARCLE: Functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 2017, 45, D200–D203. [Google Scholar] [CrossRef] [PubMed]
- Papadopoulos, J.S.; Agarwala, R. COBALT: Constraint-based alignment tool for multiple protein sequences. Bioinformatics 2007, 23, 1073–1079. [Google Scholar] [CrossRef] [PubMed]
- Tatusova, T.; Dicuccio, M.; Badretdin, A.; Chetvernin, V.; Nawrocki, E.P.; Zaslavsky, L.; Lomsadze, A.; Pruitt, K.D.; Borodovsky, M.; Ostell, J. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016, 44, 6614–6624. [Google Scholar] [CrossRef] [PubMed]
- Bouffard, G.G.; Rudd, K.E.; Adhya, S.L. Dependence of lactose metabolism upon mutarotase encoded in the gal operon in Escherichia coli. J. Mol. Biol. 1994, 244, 269–278. [Google Scholar] [CrossRef] [PubMed]
- Jeon, H.; Lee, H.; Byun, D.; Choi, H.; Shim, J.H. Molecular cloning, characterization, and application of a novel thermostable α-glucosidase from the hyperthermophilic archaeon Pyrobaculum aerophilum strain IM2. Food Sci. Biotechnol. 2015, 24, 175–182. [Google Scholar] [CrossRef]
- Rolfsmeier, M.; Blum, P. Purification and characterization of a maltase from the extremely thermophilic crenarchaeote Sulfolobus solfataricus. J. Bacteriol. 1995, 177, 482–485. [Google Scholar] [CrossRef] [PubMed]
- Park, J.E.; Park, S.H.; Woo, J.Y.; Hwang, H.S.; Cha, J.; Lee, H. Enzymatic properties of a thermostable-glucosidase from acidothermophilic crenarchaeon Sulfolobus tokodaii strain 7. J. Microbiol. Biotechnol. 2013, 23, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Angelov, A.; Putyrski, M.; Liebl, W. Molecular and biochemical characterization of α-glucosidase and α-mannosidase and their clustered genes from the thermoacidophilic archaeon Picrophilus torridus. J. Bacteriol. 2006, 188, 7123–7131. [Google Scholar] [CrossRef] [PubMed]
- Oren, A. Microbial life at high salt concentrations: Phylogenetic and metabolic diversity. Saline Syst. 2008, 4, 2. [Google Scholar] [CrossRef] [PubMed]
- Podell, S.; Ugalde, J.A.; Narasingarao, P.; Banfield, J.F.; Heidelberg, K.B.; Allen, E.E. Assembly-Driven Community Genomics of a Hypersaline Microbial Ecosystem. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Becker, E.A.; Seitzer, P.M.; Tritt, A.; Larsen, D.; Krusor, M.; Yao, A.I.; Wu, D.; Madern, D.; Eisen, J.A.; Darling, A.E.; et al. Phylogenetically Driven Sequencing of Extremely Halophilic Archaea Reveals Strategies for Static and Dynamic Osmo-response. PLoS Genet. 2014, 10. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.W.; Lovering, A.L.; Chen, H.; Kantner, T.; McIntosh, L.P.; Strynadka, N.C.J.; Withers, S.G. Expanding the thioglycoligase strategy to the synthesis of α-linked thioglycosides allows structural investigation of the parent enzyme/substrate complex. J. Am. Chem. Soc. 2006, 128, 2202–2203. [Google Scholar] [CrossRef] [PubMed]
- McCarter, J.D.; Stephen Withers, G. Mechanisms of enzymatic glycoside hydrolysis. Curr. Opin. Struct. Biol. 1994, 4, 885–892. [Google Scholar] [CrossRef]
- Chaillou, S.; Lokman, B.C.; Leer, R.J.; Posthuma, C.; Postma, P.W.; Pouwels, P.H. Cloning, sequence analysis, and characterization of the genes involved in isoprimeverose metabolism in Lactobacillus pentosus. J. Bacteriol. 1998, 180, 2312–2320. [Google Scholar] [PubMed]
- Tan, K.; Tesar, C.; Wilton, R.; Keigher, L.; Babnigg, G.; Joachimiak, A. Novel α-glucosidase from human gut microbiome: Substrate specificities and their switch. FASEB J. 2010, 24, 3939–3949. [Google Scholar] [CrossRef] [PubMed]
- Tagami, T.; Yamashita, K.; Okuyama, M.; Mori, H.; Yao, M.; Kimura, A. Molecular Basis for the Recognition of Long-chain Substrates by Plant α-Glucosidases. J. Biol. Chem. 2013, 288, 19296–19303. [Google Scholar] [CrossRef] [PubMed]
- De Lourdes Moreno, M.; Pérez, D.; García, M.T.; Mellado, E. Halophilic bacteria as a source of novel hydrolytic enzymes. Life 2013, 3, 38–51. [Google Scholar] [CrossRef] [PubMed]
- Cota, J.; Corrêa, T.L.R.; Damásio, A.R.L.; Diogo, J.A.; Hoffmam, Z.B.; Garcia, W.; Oliveira, L.C.; Prade, R.A.; Squina, F.M. Comparative analysis of three hyperthermophilic GH1 and GH3 family members with industrial potential. New Biotechnol. 2015, 32, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Thoden, J.B.; Holden, H.M. High Resolution X-ray Structure of Galactose Mutarotase from Lactococcus lactis. J. Biol. Chem. 2002, 277, 20854–20861. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.M.I.; Kim, Y.R.; Kim, J.K.; Jeong, G.T.; Ha, J.C.; Kong, I.S. Characterization of salt-tolerant β-glucosidase with increased thermostability under high salinity conditions from Bacillus sp. SJ-10 isolated from jeotgal, a traditional Korean fermented seafood. Bioprocess Biosyst. Eng. 2015, 38, 1335–1346. [Google Scholar] [CrossRef] [PubMed]
- Lovering, A.L.; Seo, S.; Kim, Y.; Withers, S.G.; Strynadka, N.C.J. Mechanistic and Structural Analysis of a Family 31 α-Glycosidase. J. Biol. Chem. 2005, 280, 2105–2115. [Google Scholar] [CrossRef] [PubMed]
- Sinnott, M.L. Catalytic Mechanisms of Enzymic Glycosyl Transfer. Chem. Rev. 1990, 90, 1171–1202. [Google Scholar] [CrossRef]

| Archaea Species | Conserved Domains | Percent Identity (%) | |
|---|---|---|---|
| Galactose Mutarotase-Like 2 | Glycosyl Hydrolases Family 31 | ||
| Haloquadratum walsbyi | 161–222 | 243–670 | This study |
| Halorubrum kocurii | 156–217 | 238–668 | 43 |
| Halorubrum terrestre | 162–223 | 244–672 | 41 |
| Halorubrum arcis | 162–223 | 244–672 | 41 |
| Halorubrum litoreum | 162–233 | 244–672 | 41 |
| Halorubrum distributum | 162–233 | 244–672 | 41 |
| Halorubrum lipolyticum | 160–221 | 242–670 | 40 |
| Halothermothrix orenii | 144–211 | 232–672 | 29 |
| Thermoplasma volcanium | 177–244 | 265–697 | 27 |
| Thermoproteus uzoniensis | 67–136 | 157–602 | 27 |
| Pyrobaculum aerophilum * | 67–133 | 152–612 | 27 |
| Haloterrigena turkmenica | 126–193 | 226–696 | 26 |
| Sulfolobus islandicus | 61–127 | 148–608 | 26 |
| Sulfolobus solfataricus * | 161–222 | 243–670 | 26 |
| Thermoproteus tenax | 69–135 | 155–616 | 26 |
| Pyrobaculum arsenaticum | 68–134 | 154–613 | 26 |
| Pyrobaculum calidifontis | 67–133 | 153–610 | 26 |
| Thermoplasma acidophilum | 147–214 | 234–667 | 25 |
| Sulfolobus acidocaldarius | 59–124 | 145–574 | 25 |
| Sulfolobus tokodaii * | 56–122 | 143–586 | 25 |
| Metallosphaera sedula | 57–123 | 144–598 | 25 |
| Pyrobaculum islandicum | 67–133 | 153–612 | 25 |
| Picrophilus torridus * | 58–123 | 144–572 | 24 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cuebas-Irizarry, M.F.; Irizarry-Caro, R.A.; López-Morales, C.; Badillo-Rivera, K.M.; Rodríguez-Minguela, C.M.; Montalvo-Rodríguez, R. Cloning and Molecular Characterization of an Alpha-Glucosidase (MalH) from the Halophilic Archaeon Haloquadratum walsbyi. Life 2017, 7, 46. https://doi.org/10.3390/life7040046
Cuebas-Irizarry MF, Irizarry-Caro RA, López-Morales C, Badillo-Rivera KM, Rodríguez-Minguela CM, Montalvo-Rodríguez R. Cloning and Molecular Characterization of an Alpha-Glucosidase (MalH) from the Halophilic Archaeon Haloquadratum walsbyi. Life. 2017; 7(4):46. https://doi.org/10.3390/life7040046
Chicago/Turabian StyleCuebas-Irizarry, Mara F., Ricardo A. Irizarry-Caro, Carol López-Morales, Keyla M. Badillo-Rivera, Carlos M. Rodríguez-Minguela, and Rafael Montalvo-Rodríguez. 2017. "Cloning and Molecular Characterization of an Alpha-Glucosidase (MalH) from the Halophilic Archaeon Haloquadratum walsbyi" Life 7, no. 4: 46. https://doi.org/10.3390/life7040046
APA StyleCuebas-Irizarry, M. F., Irizarry-Caro, R. A., López-Morales, C., Badillo-Rivera, K. M., Rodríguez-Minguela, C. M., & Montalvo-Rodríguez, R. (2017). Cloning and Molecular Characterization of an Alpha-Glucosidase (MalH) from the Halophilic Archaeon Haloquadratum walsbyi. Life, 7(4), 46. https://doi.org/10.3390/life7040046





