Considering Strain Variation and Non-Type Strains for Yeast Metabolic Engineering Applications
Abstract
1. Introduction
2. Surveying Innate Metabolic Production and Rewiring Potential across Strains
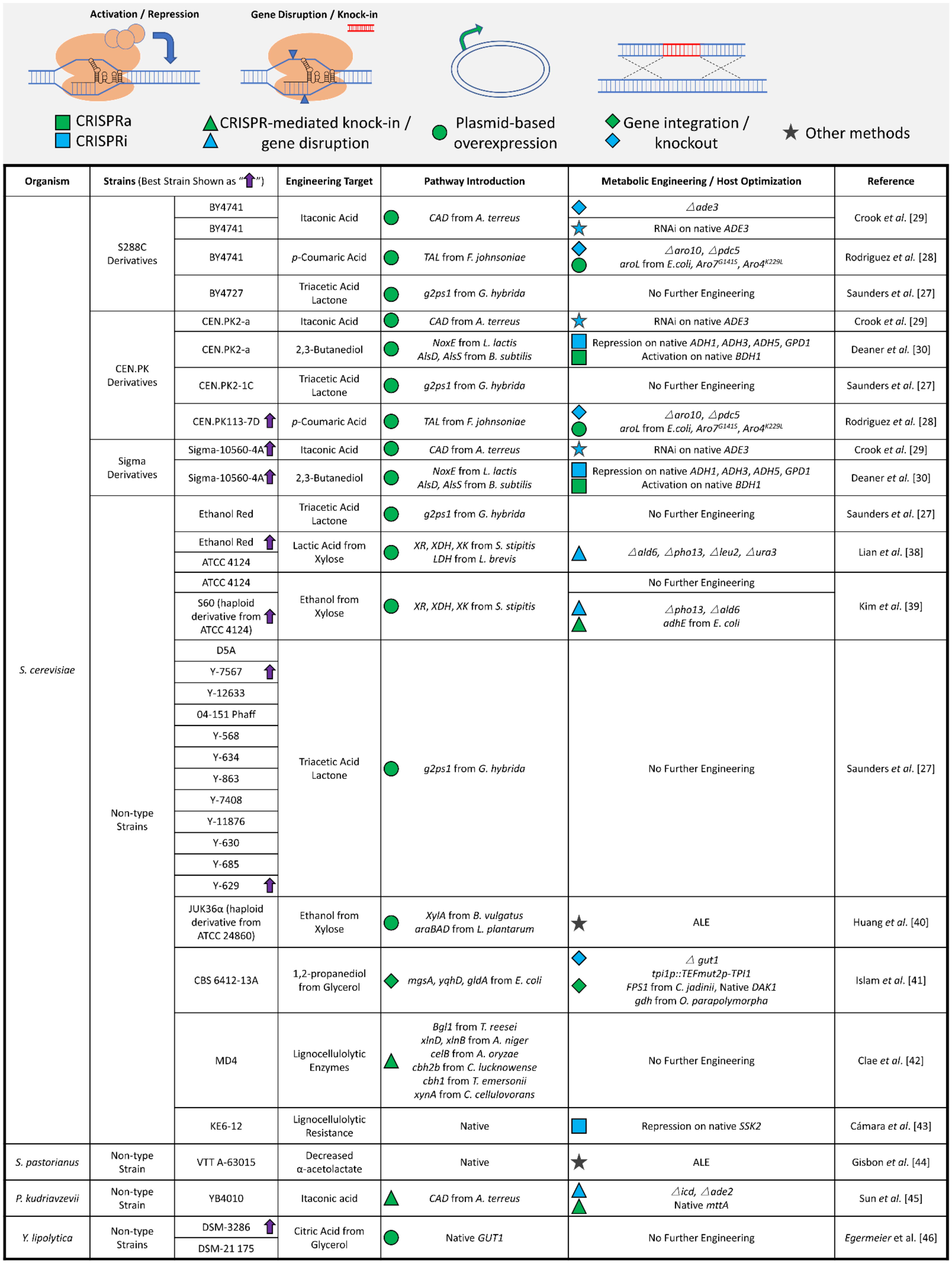
3. Successes in the Metabolic Engineering and Optimization of Non-Type Strains
4. Non-Conventional Yeasts
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ageitos, J.M.; Vallejo, J.A.; Veiga-Crespo, P.; Villa, T.G. Oily Yeasts as Oleaginous Cell Factories. Appl. Microbiol. Biotechnol. 2011, 90, 1219–1227. [Google Scholar] [CrossRef]
- Gustavsson, M.; Lee, S.Y. Prospects of Microbial Cell Factories Developed through Systems Metabolic Engineering. Microb. Biotechnol. 2016, 9, 610–617. [Google Scholar] [CrossRef]
- Liu, Z.; Moradi, H.; Shi, S.; Darvishi, F. Yeasts as Microbial Cell Factories for Sustainable Production of Biofuels. Renew. Sustain. Energy Rev. 2021, 143, 110907. [Google Scholar] [CrossRef]
- Intasian, P.; Prakinee, K.; Phintha, A.; Trisrivirat, D.; Weeranoppanant, N.; Wongnate, T.; Chaiyen, P. Enzymes, in Vivo Biocatalysis, and Metabolic Engineering for Enabling a Circular Economy and Sustainability. Chem. Rev. 2021, 121, 10367–10451. [Google Scholar] [CrossRef]
- Adkins, J.; Pugh, S.; McKenna, R.; Nielsen, D.R. Engineering Microbial Chemical Factories to Produce Renewable “Biomonomers”. Front. Microbiol. 2012, 3, 313. [Google Scholar] [CrossRef]
- Steen, E.J.; Kang, Y.; Bokinsky, G.; Hu, Z.; Schirmer, A.; McClure, A.; Del Cardayre, S.B.; Keasling, J.D. Microbial Production of Fatty-Acid-Derived Fuels and Chemicals from Plant Biomass. Nature 2010, 463, 559–562. [Google Scholar] [CrossRef]
- Lee, J.W.; Trinh, C.T. Towards Renewable Flavors, Fragrances, and Beyond. Curr. Opin. Biotechnol. 2020, 61, 168–180. [Google Scholar] [CrossRef]
- Tippmann, S.; Chen, Y.; Siewers, V.; Nielsen, J. From Flavors and Pharmaceuticals to Advanced Biofuels: Production of Isoprenoids in Saccharomyces cerevisiae. Biotechnol. J. 2013, 8, 1435–1444. [Google Scholar] [CrossRef]
- Romero-Suarez, D.; Keasling, J.D.; Jensen, M.K. Supplying Plant Natural Products by Yeast Cell Factories. Curr. Opin. Green Sustain. Chem. 2022, 33, 100567. [Google Scholar] [CrossRef]
- Liu, L.; Redden, H.; Alper, H.S. Frontiers of Yeast Metabolic Engineering: Diversifying beyond Ethanol and Saccharomyces. Curr. Opin. Biotechnol. 2013, 24, 1023–1030. [Google Scholar] [CrossRef]
- Markham, K.A.; Palmer, C.M.; Chwatko, M.; Wagner, J.M.; Murray, C.; Vazquez, S.; Swaminathan, A.; Chakravarty, I.; Lynd, N.A.; Alper, H.S. Rewiring Yarrowia lipolytica toward Triacetic Acid Lactone for Materials Generation. Proc. Natl. Acad. Sci. USA 2018, 115, 2096–2101. [Google Scholar] [CrossRef]
- McTaggart, T.L.; Bever, D.; Bassett, S.; Da Silva, N.A. Synthesis of Polyketides from Low Cost Substrates by the Thermotolerant Yeast Kluyveromyces marxianus. Biotechnol. Bioeng. 2019, 116, 1721–1730. [Google Scholar] [CrossRef]
- Worland, A.M.; Czajka, J.J.; Li, Y.; Wang, Y.; Tang, Y.J.; Su, W.W. Biosynthesis of Terpene Compounds Using the Non-Model Yeast Yarrowia lipolytica: Grand Challenges and a Few Perspectives. Curr. Opin. Biotechnol. 2020, 64, 134–140. [Google Scholar] [CrossRef]
- Bhataya, A.; Schmidt-Dannert, C.; Lee, P.C. Metabolic Engineering of Pichia pastoris X-33 for Lycopene Production. Process Biochem. 2009, 44, 1095–1102. [Google Scholar] [CrossRef]
- Hawkins, K.M.; Smolke, C.D. Production of Benzylisoquinoline Alkaloids in Saccharomyces cerevisiae. Nat. Chem. Biol. 2008, 4, 564–573. [Google Scholar] [CrossRef]
- Hori, K.; Okano, S.; Sato, F. Efficient Microbial Production of Stylopine Using a Pichia pastoris Expression System. Sci. Rep. 2016, 6, 22201. [Google Scholar] [CrossRef]
- Park, Y.K.; Nicaud, J.M.; Ledesma-Amaro, R. The Engineering Potential of Rhodosporidium toruloides as a Workhorse for Biotechnological Applications. Trends Biotechnol. 2018, 36, 304–317. [Google Scholar] [CrossRef]
- Blazeck, J.; Hill, A.; Liu, L.; Knight, R.; Miller, J.; Pan, A.; Otoupal, P.; Alper, H.S. Harnessing Yarrowia lipolytica Lipogenesis to Create a Platform for Lipid and Biofuel Production. Nat. Commun. 2014, 5, 3131. [Google Scholar] [CrossRef]
- Konzock, O.; Zaghen, S.; Norbeck, J. Tolerance of Yarrowia lipolytica to Inhibitors Commonly Found in Lignocellulosic Hydrolysates. BMC Microbiol. 2021, 21, 77. [Google Scholar] [CrossRef]
- Ha-Tran, D.M.; Nguyen, T.T.M.; Huang, C.-C. Kluyveromyces marxianus: Current State of Omics Studies, Strain Improvement Strategy and Potential Industrial Implementation. Fermentation 2020, 6, 124. [Google Scholar] [CrossRef]
- Nielsen, J.; Larsson, C.; Van Maris, A.; Pronk, J. Metabolic Engineering of Yeast for Production of Fuels and Chemicals. Curr. Opin. Biotechnol. 2013, 24, 398–404. [Google Scholar] [CrossRef]
- Karim, A.; Gerliani, N.; Aïder, M. Kluyveromyces marxianus: An Emerging Yeast Cell Factory for Applications in Food and Biotechnology. Int. J. Food Microbiol. 2020, 333, 108818. [Google Scholar] [CrossRef]
- Nielsen, J. Yeast Systems Biology: Model Organism and Cell Factory. Biotechnol. J. 2019, 14, 1800421. [Google Scholar] [CrossRef]
- Löbs, A.K.; Schwartz, C.; Wheeldon, I. Genome and Metabolic Engineering in Non-Conventional Yeasts: Current Advances and Applications. Synth. Syst. Biotechnol. 2017, 2, 198–207. [Google Scholar] [CrossRef]
- Rebello, S.; Abraham, A.; Madhavan, A.; Sindhu, R.; Binod, P.; Karthika Bahuleyan, A.; Aneesh, E.M.; Pandey, A. Non-Conventional Yeast Cell Factories for Sustainable Bioprocesses. FEMS Microbiol. Lett. 2018, 365, fny222. [Google Scholar] [CrossRef]
- Sun, L.; Alper, H.S. Non-Conventional Hosts for the Production of Fuels and Chemicals. Curr. Opin. Chem. Biol. 2020, 59, 15–22. [Google Scholar] [CrossRef]
- Saunders, L.P.; Bowman, M.J.; Mertens, J.A.; Da Silva, N.A.; Hector, R.E. Triacetic Acid Lactone Production in Industrial Saccharomyces Yeast Strains. J. Ind. Microbiol. Biotechnol. 2015, 42, 711–721. [Google Scholar] [CrossRef]
- Rodriguez, A.; Chen, Y.; Khoomrung, S.; Özdemir, E.; Borodina, I.; Nielsen, J. Comparison of the Metabolic Response to Over-Production of p-Coumaric Acid in Two Yeast Strains. Metab. Eng. 2017, 44, 265–272. [Google Scholar] [CrossRef]
- Crook, N.C.; Schmitz, A.C.; Alper, H.S. Optimization of a Yeast RNA Interference System for Controlling Gene Expression and Enabling Rapid Metabolic Engineering. ACS Synth. Biol. 2014, 3, 307–313. [Google Scholar] [CrossRef]
- Deaner, M.; Holzman, A.; Alper, H.S. Modular Ligation Extension of Guide RNA Operons (LEGO) for Multiplexed DCas9 Regulation of Metabolic Pathways in Saccharomyces cerevisiae. Biotechnol. J. 2018, 13, 1700582. [Google Scholar] [CrossRef]
- Jakočiunas, T.; Jensen, M.K.; Keasling, J.D. CRISPR/Cas9 Advances Engineering of Microbial Cell Factories. Metab. Eng. 2016, 34, 44–59. [Google Scholar] [CrossRef]
- Bao, Z.; HamediRad, M.; Xue, P.; Xiao, H.; Tasan, I.; Chao, R.; Liang, J.; Zhao, H. Genome-Scale Engineering of Saccharomyces cerevisiae with Single-Nucleotide Precision. Nat. Biotechnol. 2018, 36, 505–508. [Google Scholar] [CrossRef]
- Schwartz, C.; Shabbir-Hussain, M.; Frogue, K.; Blenner, M.; Wheeldon, I. Standardized Markerless Gene Integration for Pathway Engineering in Yarrowia lipolytica. ACS Synth. Biol. 2017, 6, 402–409. [Google Scholar] [CrossRef]
- Cai, P.; Gao, J.; Zhou, Y. CRISPR-Mediated Genome Editing in Non-Conventional Yeasts for Biotechnological Applications. Microb. Cell Factories 2019, 18, 63. [Google Scholar] [CrossRef]
- Shi, T.Q.; Huang, H.; Kerkhoven, E.J.; Ji, X.J. Advancing Metabolic Engineering of Yarrowia lipolytica Using the CRISPR/Cas System. Appl. Microbiol. Biotechnol. 2018, 102, 9541–9548. [Google Scholar] [CrossRef]
- Nishida, K.; Kondo, A. CRISPR-Derived Genome Editing Technologies for Metabolic Engineering. Metab. Eng. 2021, 63, 141–147. [Google Scholar] [CrossRef]
- Raschmanová, H.; Weninger, A.; Glieder, A.; Kovar, K.; Vogl, T. Implementing CRISPR-Cas Technologies in Conventional and Non-Conventional Yeasts: Current State and Future Prospects. Biotechnol. Adv. 2018, 36, 641–665. [Google Scholar] [CrossRef]
- Lian, J.; Bao, Z.; Hu, S.; Zhao, H. Engineered CRISPR/Cas9 System for Multiplex Genome Engineering of Polyploid Industrial Yeast Strains. Biotechnol. Bioeng. 2018, 115, 1630–1635. [Google Scholar] [CrossRef]
- Kim, S.R.; Skerker, J.M.; Kong, I.I.; Kim, H.; Maurer, M.J.; Zhang, G.C.; Peng, D.; Wei, N.; Arkin, A.P.; Jin, Y.S. Metabolic Engineering of a Haploid Strain Derived from a Triploid Industrial Yeast for Producing Cellulosic Ethanol. Metab. Eng. 2017, 40, 176–185. [Google Scholar] [CrossRef]
- Huang, S.; Liu, T.; Peng, B.; Geng, A. Enhanced Ethanol Production from Industrial Lignocellulose Hydrolysates by a Hydrolysate-Cofermenting Saccharomyces cerevisiae Strain. Bioprocess Biosyst. Eng. 2019, 42, 883–896. [Google Scholar] [CrossRef]
- Ul Islam, Z.; Klein, M.; Aßkamp, M.R.; Ødum, A.S.R.; Nevoigt, E. A Modular Metabolic Engineering Approach for the Production of 1,2-Propanediol from Glycerol by Saccharomyces cerevisiae. Metab. Eng. 2017, 44, 223–235. [Google Scholar] [CrossRef][Green Version]
- Claes, A.; Deparis, Q.; Foulquié-Moreno, M.R.; Thevelein, J.M. Simultaneous Secretion of Seven Lignocellulolytic Enzymes by an Industrial Second-Generation Yeast Strain Enables Efficient Ethanol Production from Multiple Polymeric Substrates. Metab. Eng. 2020, 59, 131–141. [Google Scholar] [CrossRef]
- Cámara, E.; Lenitz, I.; Nygård, Y. A CRISPR Activation and Interference Toolkit for Industrial Saccharomyces cerevisiae Strain KE6-12. Sci. Rep. 2020, 10, 14605. [Google Scholar] [CrossRef]
- Gibson, B.; Vidgren, V.; Peddinti, G.; Krogerus, K. Diacetyl Control during Brewery Fermentation via Adaptive Laboratory Engineering of the Lager Yeast Saccharomyces pastorianus. J. Ind. Microbiol. Biotechnol. 2018, 45, 1103–1112. [Google Scholar] [CrossRef]
- Sun, W.; Vila-Santa, A.; Liu, N.; Prozorov, T.; Xie, D.; Faria, N.T.; Ferreira, F.C.; Mira, N.P.; Shao, Z. Metabolic Engineering of an Acid-Tolerant Yeast Strain Pichia kudriavzevii for Itaconic Acid Production. Metab. Eng. Commun. 2020, 10, e00124. [Google Scholar] [CrossRef]
- Egermeier, M.; Sauer, M.; Marx, H. Golden Gate-Based Metabolic Engineering Strategy for Wild-Type Strains of Yarrowia lipolytica. FEMS Microbiol. Lett. 2019, 366, fnz022. [Google Scholar] [CrossRef]
- Winzeler, E.A.; Shoemaker, D.D.; Astromoff, A.; Liang, H.; Anderson, K.; Andre, B.; Bangham, R.; Benito, R.; Boeke, J.D.; Bussey, H.; et al. Functional Characterization of the S. cerevisiae Genome by Gene Deletion and Parallel Analysis. Science 1999, 285, 901–906. [Google Scholar] [CrossRef]
- Lu, H.; Li, F.; Sánchez, B.J.; Zhu, Z.; Li, G.; Domenzain, I.; Marcišauskas, S.; Anton, P.M.; Lappa, D.; Lieven, C.; et al. A Consensus S. cerevisiae Metabolic Model Yeast8 and Its Ecosystem for Comprehensively Probing Cellular Metabolism. Nat. Commun. 2019, 10, 3586. [Google Scholar] [CrossRef]
- Rodrigues, C.I.S.; Wahl, A.; Gombert, A.K. Aerobic Growth Physiology of Saccharomyces cerevisiae on Sucrose Is Strain-Dependent. FEMS Yeast Res. 2021, 21, foab021. [Google Scholar] [CrossRef] [PubMed]
- Kvitek, D.J.; Will, J.L.; Gasch, A.P. Variations in Stress Sensitivity and Genomic Expression in Diverse S. cerevisiae Isolates. PLoS Genet. 2008, 4, e1000223. [Google Scholar] [CrossRef] [PubMed]
- Rywińska, A.; Rymowicz, W.; Zarowska, B.; Skrzypiński, A. Comparison of Citric Acid Production from Glycerol and Glucose by Different Strains of Yarrowia lipolytica. World J. Microbiol. Biotechnol. 2010, 26, 1217–1224. [Google Scholar] [CrossRef] [PubMed]
- Lane, M.M.; Burke, N.; Karreman, R.; Wolfe, K.H.; O’Byrne, C.P.; Morrissey, J.P. Physiological and Metabolic Diversity in the Yeast Kluyveromyces marxianus. Antonie Van Leeuwenhoek Int. J. Gen. Mol. Microbiol. 2011, 100, 507–519. [Google Scholar] [CrossRef] [PubMed]
- Drinnenberg, I.; Weinberg, D.; Xie, K.; Mower, J.; Wolfe, K.; Fink, G.; Bartel, D. RNAi in Budding Yeast. Science 2009, 326, 544–550. [Google Scholar] [CrossRef] [PubMed]
- Kuenz, A.; Gallenmüller, Y.; Willke, T.; Vorlop, K.D. Microbial Production of Itaconic Acid: Developing a Stable Platform for High Product Concentrations. Appl. Microbiol. Biotechnol. 2012, 96, 1209–1216. [Google Scholar] [CrossRef]
- Blazeck, J.; Miller, J.; Pan, A.; Gengler, J.; Holden, C.; Jamoussi, M.; Alper, H.S. Metabolic Engineering of Saccharomyces cerevisiae for Itaconic Acid Production. Appl. Microbiol. Biotechnol. 2014, 98, 8155–8164. [Google Scholar] [CrossRef]
- Alper, H.; Stephanopoulos, G. Engineering for Biofuels: Exploiting Innate Microbial Capacity or Importing Biosynthetic Potential? Nat. Rev. Microbiol. 2009, 7, 715–723. [Google Scholar] [CrossRef]
- Steensels, J.; Snoek, T.; Meersman, E.; Nicolino, M.P.; Voordeckers, K.; Verstrepen, K.J. Improving Industrial Yeast Strains: Exploiting Natural and Artificial Diversity. FEMS Microbiol. Rev. 2014, 38, 947–995. [Google Scholar] [CrossRef]
- Takita, Y.; Takahara, M.; Nogami, S.; Anraku, Y.; Ohya, Y. Applications of the Long and Accurate Polymerase Chain Reaction Method in Yeast Molecular Biology: Direct Sequencing of the Amplified DNA and Its Introduction into Yeast. Yeast 1997, 13, 763–768. [Google Scholar] [CrossRef]
- Papathanasiou, S.; Markoulaki, S.; Blaine, L.J.; Leibowitz, M.L.; Zhang, C.Z.; Jaenisch, R.; Pellman, D. Whole Chromosome Loss and Genomic Instability in Mouse Embryos after CRISPR-Cas9 Genome Editing. Nat. Commun. 2021, 12, 5855. [Google Scholar] [CrossRef]
- Miller, K.K.; Alper, H.S. Yarrowia lipolytica: More than an Oleaginous Workhorse. Appl. Microbiol. Biotechnol. 2019, 103, 9251–9262. [Google Scholar] [CrossRef]
- Prielhofer, R.; Barrero, J.J.; Steuer, S.; Gassler, T.; Zahrl, R.; Baumann, K.; Sauer, M.; Mattanovich, D.; Gasser, B.; Marx, H. GoldenPiCS: A Golden Gate-Derived Modular Cloning System for Applied Synthetic Biology in the Yeast Pichia pastoris. BMC Syst. Biol. 2017, 11, 123. [Google Scholar] [CrossRef]
- Wong, L.; Engel, J.; Jin, E.; Holdridge, B.; Xu, P. YaliBricks, a Versatile Genetic Toolkit for Streamlined and Rapid Pathway Engineering in Yarrowia lipolytica. Metab. Eng. Commun. 2017, 5, 68–77. [Google Scholar] [CrossRef]
- Holkenbrink, C.; Dam, M.I.; Kildegaard, K.R.; Beder, J.; Dahlin, J.; Doménech Belda, D.; Borodina, I. EasyCloneYALI: CRISPR/Cas9-Based Synthetic Toolbox for Engineering of the Yeast Yarrowia lipolytica. Biotechnol. J. 2018, 13, 1700543. [Google Scholar] [CrossRef]
- Lian, J.; Hamedirad, M.; Hu, S.; Zhao, H. Combinatorial Metabolic Engineering Using an Orthogonal Tri-Functional CRISPR System. Nat. Commun. 2017, 8, 1688. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, J.; Wang, Z.; Zhang, Y.; Shi, S.; Nielsen, J.; Liu, Z. A gRNA-tRNA Array for CRISPR-Cas9 Based Rapid Multiplexed Genome Editing in Saccharomyces cerevisiae. Nat. Commun. 2019, 10, 1053. [Google Scholar] [CrossRef]
- Nambu-Nishida, Y.; Nishida, K.; Hasunuma, T.; Kondo, A. Development of a Comprehensive Set of Tools for Genome Engineering in a Cold- And Thermo-Tolerant Kluyveromyces marxianus Yeast Strain. Sci. Rep. 2017, 7, 8993. [Google Scholar] [CrossRef]
- Cai, P.; Duan, X.; Wu, X.; Gao, L.; Ye, M.; Zhou, Y.J. Recombination Machinery Engineering Facilitates Metabolic Engineering of the Industrial Yeast Pichia pastoris. Nucleic Acids Res. 2021, 49, 7791–7805. [Google Scholar] [CrossRef]
- Katayama, T.; Maruyama, J. CRISPR/Cpf1-Mediated Mutagenesis and Gene Deletion in Industrial Filamentous Fungi Aspergillus oryzae and Aspergillus sojae. J. Biosci. Bioeng. 2022, 133, 353–361. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yi, X.; Alper, H.S. Considering Strain Variation and Non-Type Strains for Yeast Metabolic Engineering Applications. Life 2022, 12, 510. https://doi.org/10.3390/life12040510
Yi X, Alper HS. Considering Strain Variation and Non-Type Strains for Yeast Metabolic Engineering Applications. Life. 2022; 12(4):510. https://doi.org/10.3390/life12040510
Chicago/Turabian StyleYi, Xiunan, and Hal S. Alper. 2022. "Considering Strain Variation and Non-Type Strains for Yeast Metabolic Engineering Applications" Life 12, no. 4: 510. https://doi.org/10.3390/life12040510
APA StyleYi, X., & Alper, H. S. (2022). Considering Strain Variation and Non-Type Strains for Yeast Metabolic Engineering Applications. Life, 12(4), 510. https://doi.org/10.3390/life12040510





