Small Molecules of Natural Origin as Potential Anti-HIV Agents: A Computational Approach
Abstract
1. Introduction
2. Materials and Methods
2.1. Shape-Based Virtual Screening
2.2. ADMETox Profile
2.3. Antiviral Activity Prediction
2.4. Molecular Docking
2.5. MM-GBSA Free Energies
3. Results and Discussion
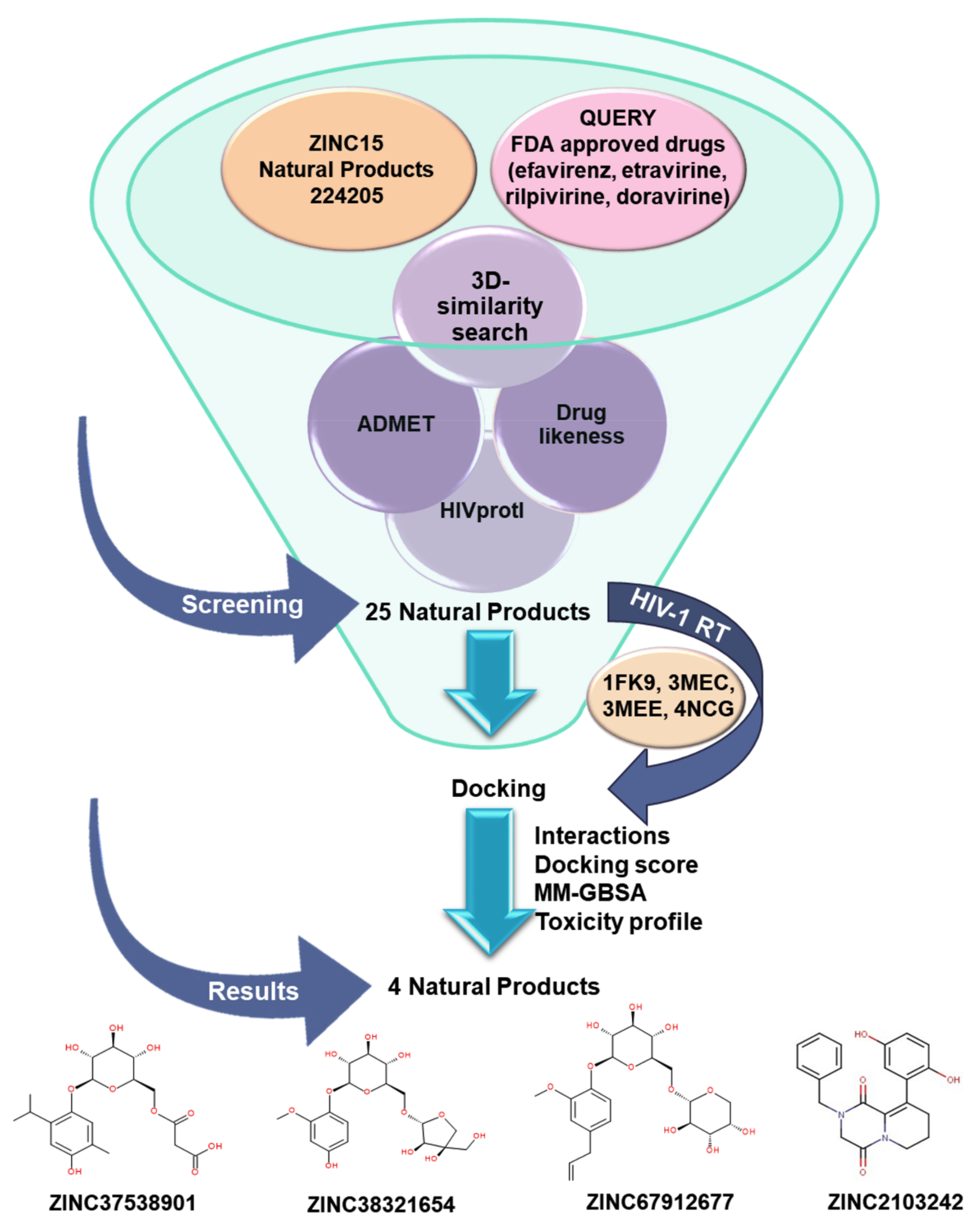
3.1. Workflow
3.2. ZINC15 NPs Subset Analysis
3.3. Shape-Based Virtual Screening Analysis
3.4. ADMETox Analysis
3.5. Antiviral Activity Prediction Analysis
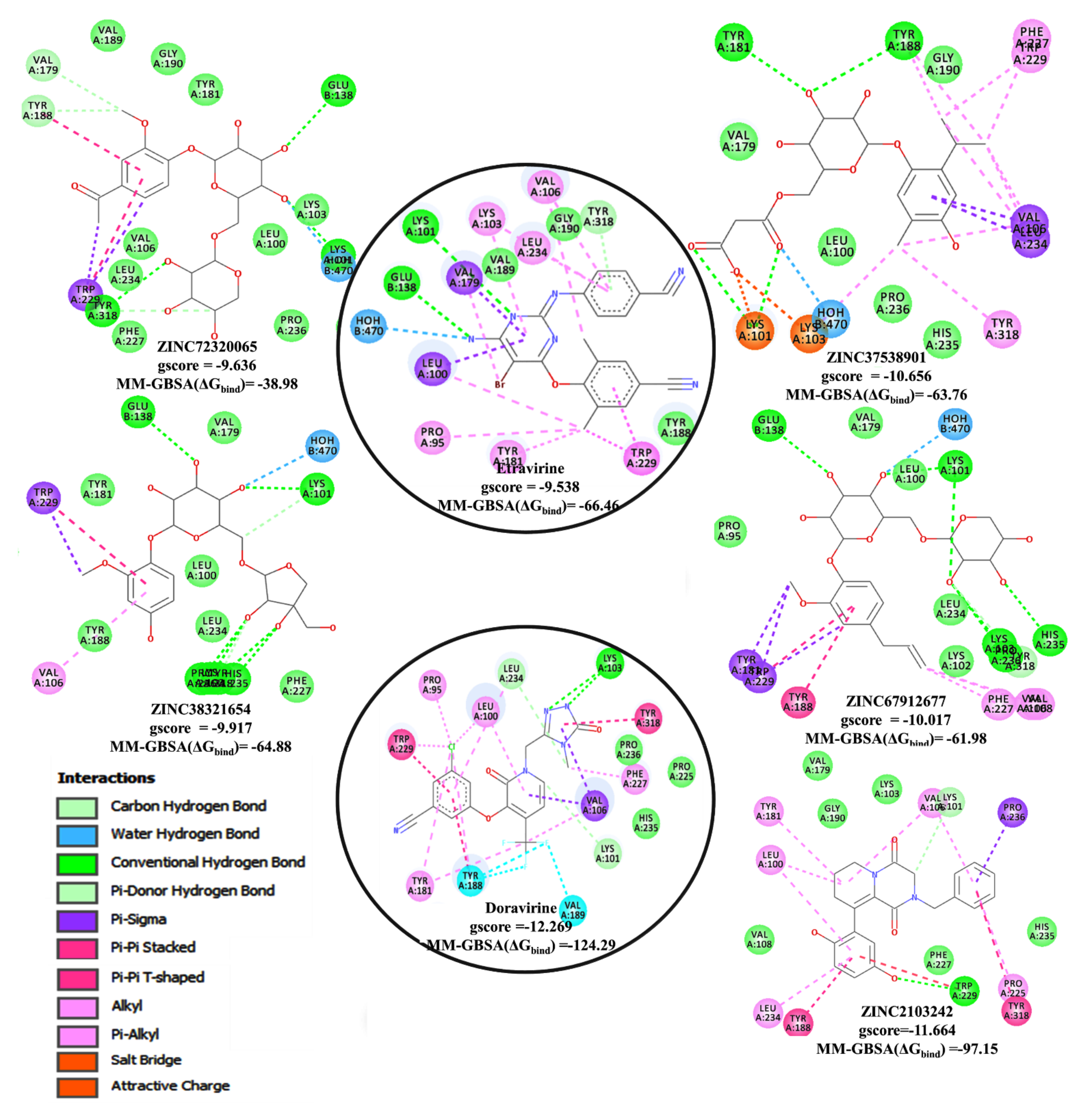
3.6. Docking Analysis
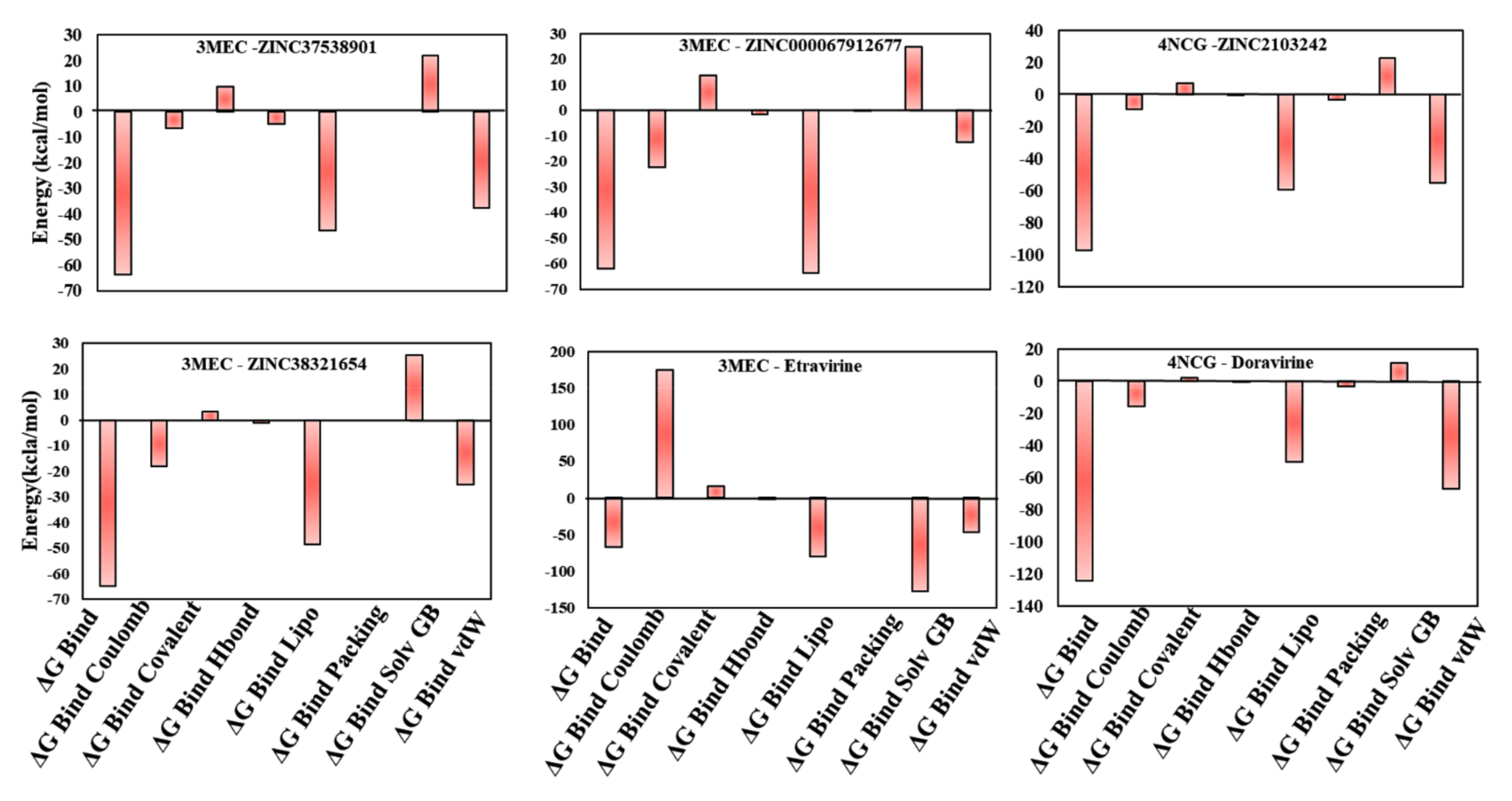
3.7. MM-GBSA Binding Free Energy Analysis
3.8. Known Therapeutic Benefits of the Proposed Natural Products
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Word Health Organization. Available online: https://www.who.int/health-topics/hiv-aids/ (accessed on 19 January 2021).
- Okoy, A.A.; Picker, L.J. CD4(+) T-cell depletion in HIV infection: Mechanisms of immunological failure. Immunol. Rev. 2013, 254, 54–64. [Google Scholar] [CrossRef]
- Panel on Antiretroviral Guidelines for Adults and Adolescents. In Guidelines for the Use of Antiretroviral Agents in Adults and Adolescents with HIV; Department of Health and Human Services: Washington, DC, USA, 2019. Available online: http://www.aidsinfo.nih.gov/ContentFiles/AdultandAdolescentGL.pdf (accessed on 19 January 2021).
- FDA Approved HIV Medicines. Available online: https://hivinfo.nih.gov/understanding-hiv/fact-sheets/fda-approved-hiv-medicines (accessed on 19 January 2021).
- De Clercq, E. The role of non-nucleoside reverse transcriptase inhibitors (NNRTIs) in the therapy of HIV-1 infection. Antivir. Res. 1998, 38, 153–179. [Google Scholar] [CrossRef]
- De Clercq, E. New developments in anti-HIV chemotherapy. Curr. Med. Chem. 2001, 8, 1543–1572. [Google Scholar] [CrossRef] [PubMed]
- Sluis-Cremer, N.; Temiz, N.A.; Bahar, I. Conformational changes in HIV-1 reverse transcriptase induced by nonnucleoside reverse transcriptase inhibitor binding. Curr. HIV Res. 2004, 2, 323–332. [Google Scholar] [CrossRef] [PubMed]
- Platten, M.; Fätkenheuer, G. Lersivirine-a new drug for HIV infection therapy. Expert Opin. Investig. Drugs 2013, 22, 1687–1694. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.Q.; Flavin, M.T.; Jenta, T.R. Calanolides, the naturally occurring anti-HIV agents. Curr. Opin. Drug Discov. Dev. 2000, 3, 155–166. [Google Scholar]
- Wensing, A.M.; Calvez, V.; Ceccherini-Silberstein, F.; Charpentier, C.; Günthard, H.F.; Paredes, R.; Shafer, R.W.; Richman, D. 2019 update of the drug resistance mutations in HIV-1. Top. Antivir. Med. 2019, 27, 111–121. [Google Scholar]
- Talwani, R.; Temesgen, Z. Doravirine: A new non-nucleoside reverse transcriptase inhibitor for the treatment of HIV infection. Drugs Today 2020, 56, 113–124. [Google Scholar] [CrossRef]
- Ivan, D.; Crisan, L.; Funar-Timofei, S.; Mracec, M. A quantitative structure–activity relationships study for the anti-HIV-1 activities of 1-[(2-hydroxyethoxy)methyl]-6-(phenylthio)thymine derivatives using the multiple linear regression and partial least squares methodologies. J. Serb. Chem. Soc. 2013, 78, 495–506. [Google Scholar] [CrossRef]
- Petric, M.; Crisan, L.; Crisan, M.; Micle, A.; Maranescu, B.; Ilia, G. Synthesis and QSRR study for a series of phosphoramidic acid derivatives. Heteroat. Chem. 2013, 24, 138–145. [Google Scholar] [CrossRef]
- Hussain, W.; Majeed, A.; Akhtar, A.; Rasool, N. Computational Studies of 3D-QSAR on a Highly Active Series of Naturally Occurring Nonnucleoside Inhibitors of HIV-1 RT (NNRTI). J. Comput. Biophys. Chem. 2021, 20, 3–11. [Google Scholar] [CrossRef]
- Bharadwaj, S.; Dubey, A.; Kamboj, N.K.; Amaresh Kumar Sahoo, A.K.; Kang, S.G.; Umesh Yadava, U. Drug repurposing for ligand-induced rearrangement of Sirt2 active site-based inhibitors via molecular modelling and quantum mechanics calculations. Sci. Rep. 2021, 11, 10169. [Google Scholar] [CrossRef]
- Wang, W.; Tian, Y.; Wan, Y.; Gu, S.; Ju, X.; Luo, X.; Liu, G. Insights into the key structural features of N1-ary-benzimidazols as HIV-1 NNRTIs using molecular docking, molecular dynamics, 3D-QSAR, and pharmacophore modelling. Struct. Chem. 2018, 30, 385–397. [Google Scholar] [CrossRef]
- Wang, W.; Tian, Y.; Wan, Y.; Gu, S.; Ju, X.; Luo, X.; Liu, G. In silico studies of diarylpyridine derivatives as novel HIV-1 NNRTIs using docking-based 3D-QSAR, molecular dynamics, and pharmacophore modeling approaches. RSC Adv. 2018, 8, 40529–40543. [Google Scholar] [CrossRef]
- Chen, Y.; Tian, Y.; Gao, Y.; Wu, F.; Luo, X.; Ju, X.; Liu, G. In silico design of novel HIV-1 NNRTIs based on combined modeling studies of dihydrofuro[3,4-d]pyrimidines. Front. Chem. 2020, 8, 164. [Google Scholar] [CrossRef] [PubMed]
- Hdoufane, I.; Stoycheva, J.; Tadjer, A.; Villemin, D.; Najdoska-Bogdanov, M.; Bogdanov, J.; Cherqaoui, D. QSAR and molecular docking studies of indole-based analogs as HIV-1 attachment inhibitors. J. Mol. Struct. 2019, 1193, 429–443. [Google Scholar] [CrossRef]
- Singh, V.K.; Srivastava, R.; Gupta, P.S.S.; Naaz, F.; Chaurasia, H.; Mishra, R.; Rana, M.K.; Singh, R.K. Anti-HIV potential of diarylpyrimidine derivatives as non-nucleoside reverse transcriptase inhibitors: Design, synthesis, docking, TOPKAT analysis and molecular dynamics simulations. J. Biomol. Struct. Dyn. 2021, 39, 2430–2446. [Google Scholar] [CrossRef]
- Chu, H.; He, Q.; Wang, J.; Deng, Y.; Wang, J.; Hu, Y.; Wang, Y.; Lin, Z. 3D-QSAR, molecular docking, and molecular dynamics simulation of a novel thieno[3,4-d]pyrimidine inhibitor targeting human immunodeficiency virus type 1 reverse transcriptase. J. Biomol. Struct. Dyn. 2020, 38, 4567–4578. [Google Scholar] [CrossRef] [PubMed]
- Santos, L.H.; Ferreira, R.S.; Caffarena, E.R. Computational drug design strategies applied to the modelling of human immunodeficiency virus-1 reverse transcriptase inhibitors. Mem. Inst. Oswaldo Cruz 2015, 110, 847–864. [Google Scholar] [CrossRef]
- Kirchmair, J.; Distinto, S.; Liedl, K.R.; Markt, P.; Rollinger, J.M.; Schuster, D.; Spitzer, G.M.; Wolber, G. Development of anti-viral agents using molecular modeling and virtual screening techniques. Infect. Disord. Drug Targets 2011, 11, 64–93. [Google Scholar] [CrossRef] [PubMed]
- Frey, K.M. Structure-enhanced methods in the development of non-nucleoside inhibitors targeting HIV reverse transcriptase variants. Future Microbiol. 2015, 10, 1767–1772. [Google Scholar] [CrossRef] [PubMed]
- Fischl, M.A.; Richman, D.D.; Grieco, M.H.; Gottlieb, M.S.; Volberding, P.A.; Laskin, O.L.; Leedom, J.M.; Groopman, J.E.; Mildvan, D.; Schooley, R.T.; et al. The efficacy of azidothymidine (AZT) in the treatment of patients with AIDS and AIDS-related complex. N. Engl. J. Med. 1987, 317, 185–191. [Google Scholar] [CrossRef] [PubMed]
- Zumla, A.; Chan, J.F.; Azhar, E.I.; Hui, D.S.; Yuen, K.Y. Coronaviruses-drug discovery and therapeutic options. Nat. Rev. Drug Discov. 2016, 15, 327–347. [Google Scholar] [CrossRef] [PubMed]
- Cao, B.; Wang, Y.; Wen, D.; Liu, W.; Wang, J.; Fan, G.; Ruan, L.; Song, B.; Cai, Y.; Wei, M.; et al. A Trial of Lopinavir–Ritonavir in Adults Hospitalized with Severe Covid-19. N. Engl. J. Med. 2020, 382, 1787–1799. [Google Scholar] [CrossRef] [PubMed]
- Sterling, T.; Irwin, J.J. ZINC 15–Ligand discovery for everyone. J. Chem. Inf. Model. 2015, 55, 2324–2337. [Google Scholar] [CrossRef]
- Hawkins, P.C.D.; Skillman, A.G.; Nicholls, A. Comparison of shape-matching and docking as virtual screening tools. J. Med. Chem. 2007, 50, 74–82. [Google Scholar] [CrossRef]
- ROCS Version 3.2.1.4; OpenEye Scientific Software: Santa Fe, NM, USA, 2013; Available online: http://www.eyesopen.com (accessed on 8 June 2021).
- Crisan, L.; Istrate, D.; Bora, A.; Pacureanu, L. Virtual screening and drug repurposing experiments to identify potential novel selective MAO-B inhibitors for Parkinson’s disease treatment. Mol. Divers. 2020. [Google Scholar] [CrossRef] [PubMed]
- LigPrep; Schrödinger Release 2018-4; Schrödinger, LLC: New York, NY, USA, 2018.
- Hawkins, P.C.D.; Skillman, A.G.; Warren, G.L.; Ellingson, B.A.; Stahl, M.T. Conformer generation with OMEGA: Algorithm and validation using high quality structures from the Protein Databank and Cambridge structural database. J. Chem. Inf. Model. 2010, 50, 572–584. [Google Scholar] [CrossRef]
- OMEGA version 2.5.1.4; OpenEye Scientific Software: Santa Fe, NM, USA, 2013; Available online: http://www.eyesopen.com (accessed on 8 June 2021).
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef]
- Pires, D.E.V.; Blundell, T.L.; Ascher, D.B. pkCSM: Predicting small-molecule pharmacokinetic properties using graph-based signatures. J. Med. Chem. 2015, 58, 4066–4072. [Google Scholar] [CrossRef]
- Qureshi, A.; Rajput, A.; Kaur, G.; Kumar, M. HIVprotI: An integrated web based platform for prediction and design of HIV proteins inhibitors. J. Cheminform. 2018, 10, 12. [Google Scholar] [CrossRef] [PubMed]
- Halgren, T.A.; Murphy, R.B.; Friesner, R.A.; Beard, H.S.; Frye, L.L.; Pollard, W.T.; Banks, J.L. Glide: A new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J. Med. Chem. 2004, 47, 1750–1759. [Google Scholar] [CrossRef] [PubMed]
- Friesner, R.A.; Banks, J.L.; Murphy, R.B.; Halgren, T.A.; Klicic, J.J.; Mainz, D.T.; Repasky, M.P.; Knoll, E.H.; Shelley, M.; Perry, J.K.; et al. Glide: A new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 2004, 47, 1739–1749. [Google Scholar] [CrossRef]
- Glide; Schrödinger Release 2018-4; Schrödinger, LLC: New York, NY, USA, 2018.
- Lansdon, E.B.; Brendza, K.M.; Hung, M.; Wang, R.; Mukund, S.; Jin, D.; Birkus, G.; Kutty, N.; Liu, X. Crystal structures of HIV-1 reverse transcriptase with etravirine (TMC125) and Rilpivirine (TMC278): Implications for drug design. J. Med. Chem. 2010, 53, 4295–4299. [Google Scholar] [CrossRef]
- Ren, J.; Milton, J.; Weaver, K.L.; Short, S.A.; Stuart, D.I.; Stammers, D.K. Structural basis for the resilience of efavirenz (DMP-266) to drug resistance mutations in HIV-1 reverse transcriptase. Structure 2000, 8, 1089–1094. [Google Scholar] [CrossRef]
- Cote, B.; Burch, J.D.; Asante-Appiah, E.; Bayly, C.; Bedard, L.; Blouin, M.; Campeau, L.C.; Cauchon, E.; Chan, M.; Chefson, A.; et al. Discovery of MK-1439, an orally bioavailable non-nucleoside reverse transcriptase inhibitor potent against a wide range of resistant mutant HIV viruses. Bioorg. Med. Chem. Lett. 2014, 24, 917–922. [Google Scholar] [CrossRef]
- Sastry, G.M.; Adzhigirey, M.; Day, T.; Annabhimoju, R.; Sherman, W. Protein and ligand preparation: Parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided Mol. Des. 2013, 27, 221–234. [Google Scholar] [CrossRef] [PubMed]
- Maestro; Schrödinger Release 2018-4; Schrödinger, LLC: New York, NY, USA, 2018.
- Jacobson, M.P.; Pincus, D.L.; Rapp, C.S.; Day, T.J.F.; Honig, B.; Shaw, D.E.; Friesner, R.A. A hierarchical approach to all-atom protein loop prediction. Proteins 2004, 55, 351–367. [Google Scholar] [CrossRef]
- Prime; Schrödinger Release: 2018-4; Schrödinger, LLC: New York, NY, USA, 2018.
- Trivedi, J.; Tripathi, A.; Chattopadhyay, D.; Mitra, D. Plant-Derived Molecules in Managing HIV Infection, Chapter 11. In New Look to Phytomedicine; Khan, M.S.A., Ahmad, I., Chattopadhyay, D., Eds.; Academic Press: Cambridge, MA, USA; Elsevier: Amsterdam, The Netherlands, 2019; pp. 273–298. [Google Scholar]
- Kurapati, K.R.; Atluri, V.S.; Samikkannu, T.; Garcia, G.; Nair, M.P. Natural products as anti-HIV agents and role in HIV-associated neurocognitive disorders (HAND): A brief overview. Front. Microbiol. 2016, 6, 1444. [Google Scholar] [CrossRef] [PubMed]
- Singh, I.P.; Bodiwala, H.S. Recent advances in anti-HIV natural products. Nat. Prod. Rep. 2010, 27, 1781–1800. [Google Scholar] [CrossRef]
- FILTER; OpenEye Scientific Software: Santa Fe, NM, USA; Available online: http://www.eyesopen.com (accessed on 8 June 2021).
- Sorokina, M.; Steinbeck, C. Review on natural products databases: Where to find data in 2020. J. Cheminform. 2020, 12, 20. [Google Scholar] [CrossRef]
- Choy, Y.B.; Prausnitz, M.R. The rule of five for non-oral routes of drug delivery: Ophthalmic, inhalation and transdermal. Pharm. Res. 2011, 28, 943–948. [Google Scholar] [CrossRef]
- Crisan, L.; Pacureanu, L.; Avram, S.; Bora, A.; Avram, S.; Kurunczi, L. PLS and shape-based similarity analysis of maleimides - GSK-3 inhibitors. J. Enzyme. Inhib. Med. Chem. 2014, 29, 599–610. [Google Scholar] [CrossRef][Green Version]
- Sheridan, R.P.; McGaughey, G.B.; Cornell, W.D. Multiple protein structures and multiple ligands: Effects on the apparent goodness of virtual screening results. J. Comput. Aided Mol. Des. 2008, 22, 257–265. [Google Scholar] [CrossRef]
- Venhorst, J.; Núñez, S.; Terpstra, J.W.; Kruse, C.G. Assessment of scaffold hopping efficiency by use of molecular interaction fingerprints. J. Med. Chem. 2008, 51, 3222–3229. [Google Scholar] [CrossRef] [PubMed]
- Muchmore, S.W.; Debe, D.A.; Metz, J.T.; Brown, S.P.; Martin, Y.C.; Hajduk, P.J. Application of belief theory to similarity data fusion for use in analog searching and lead hopping. J. Chem. Inf. Model. 2008, 48, 941–948. [Google Scholar] [CrossRef]
- Bortolato, A.; Perruccio, F.; Moro, S. Successful Applications of in Silico Approaches for Lead/Drug Discovery; Miteva, M.A., Ed.; Bentham Science Publishers: Emirate of Sharjah, UAE, 2011. [Google Scholar]
- Daly, A.K.; Rettie, A.E.; Fowler, D.M.; Miners, J.O. Pharmacogenomics of CYP2C9: Functional and clinical considerations. J. Pers. Med. 2017, 8, 1. [Google Scholar] [CrossRef] [PubMed]
- Crisan, L.; Pacureanu, L.; Bora, A.; Avram, S.; Kurunczi, L.; Simon, Z. QSAR study and molecular docking on indirubin inhibitors of Glycogen Synthase Kinase-3. Cent. Eur. J. Chem. 2013, 11, 63–77. [Google Scholar] [CrossRef]
- Tarasova, O.; Poroikov, V.; Veselovsky, A. Molecular docking studies of HIV-1 resistance to reverse transcriptase inhibitors: Mini-rReview. Molecules 2018, 23, 1233. [Google Scholar] [CrossRef] [PubMed]
- Kroeger Smith, M.B.; Rouzer, C.A.; Taneyhill, L.A.; Smith, N.A.; Hughes, S.H.; Boyer, P.L.; Janssen, P.A.; Moereels, H.; Koymans, L.; Arnold, E.; et al. Molecular modeling studies of HIV-1 reverse transcriptase nonnucleoside inhibitors: Total energy of complexation as a predictor of drug placement and activity. Protein Sci. 1995, 4, 2203–2222. [Google Scholar] [CrossRef]
- Hwang, S.J.; Lee, H.J. Phenyl-β-D-Glucopyranoside exhibits anti-inflammatory activity in lipopolysaccharide-activated RAW 264.7 cells. Inflammation 2015, 38, 1071–1079. [Google Scholar] [CrossRef] [PubMed]
- Ghorab, H.; Khettaf, A.; Lehbili, M.; Kabouche, A.; Magid, A.A.; Harakat, D.; Voutquenne-Nazabadioko, L.; Kabouche, Z. A new cardenolide and other compounds from Salsola tetragona. Nat. Prod. Comm. 2017, 12, 3–5. [Google Scholar] [CrossRef]
- Gonzalez, C.; Tapia, M.; Perez, E.; Pallet, D.; Dornier, M. Main properties of steviol glycosides and their potential in the food industry: A review. Fruits 2014, 69, 127–141. [Google Scholar] [CrossRef]
- Lunt, E.; Newton, C.G. Pyridodiazines and Their Benzo Derivatives in Comprehensive Heterocyclic Chemistry; Katritzky, A.R., Rees, C.W., Eds.; Oxford [Oxfordshire]; Pergamon Press: Elmsford, NY, USA, 1984; pp. 199–262. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Crisan, L.; Bora, A. Small Molecules of Natural Origin as Potential Anti-HIV Agents: A Computational Approach. Life 2021, 11, 722. https://doi.org/10.3390/life11070722
Crisan L, Bora A. Small Molecules of Natural Origin as Potential Anti-HIV Agents: A Computational Approach. Life. 2021; 11(7):722. https://doi.org/10.3390/life11070722
Chicago/Turabian StyleCrisan, Luminita, and Alina Bora. 2021. "Small Molecules of Natural Origin as Potential Anti-HIV Agents: A Computational Approach" Life 11, no. 7: 722. https://doi.org/10.3390/life11070722
APA StyleCrisan, L., & Bora, A. (2021). Small Molecules of Natural Origin as Potential Anti-HIV Agents: A Computational Approach. Life, 11(7), 722. https://doi.org/10.3390/life11070722





