Bioprocess Control: Current Progress and Future Perspectives
Abstract
1. Introduction
2. Sustainability in Biologic Manufacturing
3. Strategies for Bioprocess Control
3.1. Open Loop Control
3.2. Proportional Integral/Proportional Integral Derivative (PI/PID) Control
3.3. Cascade Control
3.4. Model Predictive Control
3.5. Neural Network-Based Control
3.6. Fuzzy Logic-Based Control
3.7. Model-Based Control
4. Future Scope
4.1. Soft Sensor-Based Control
4.2. PAT Based Control Strategies
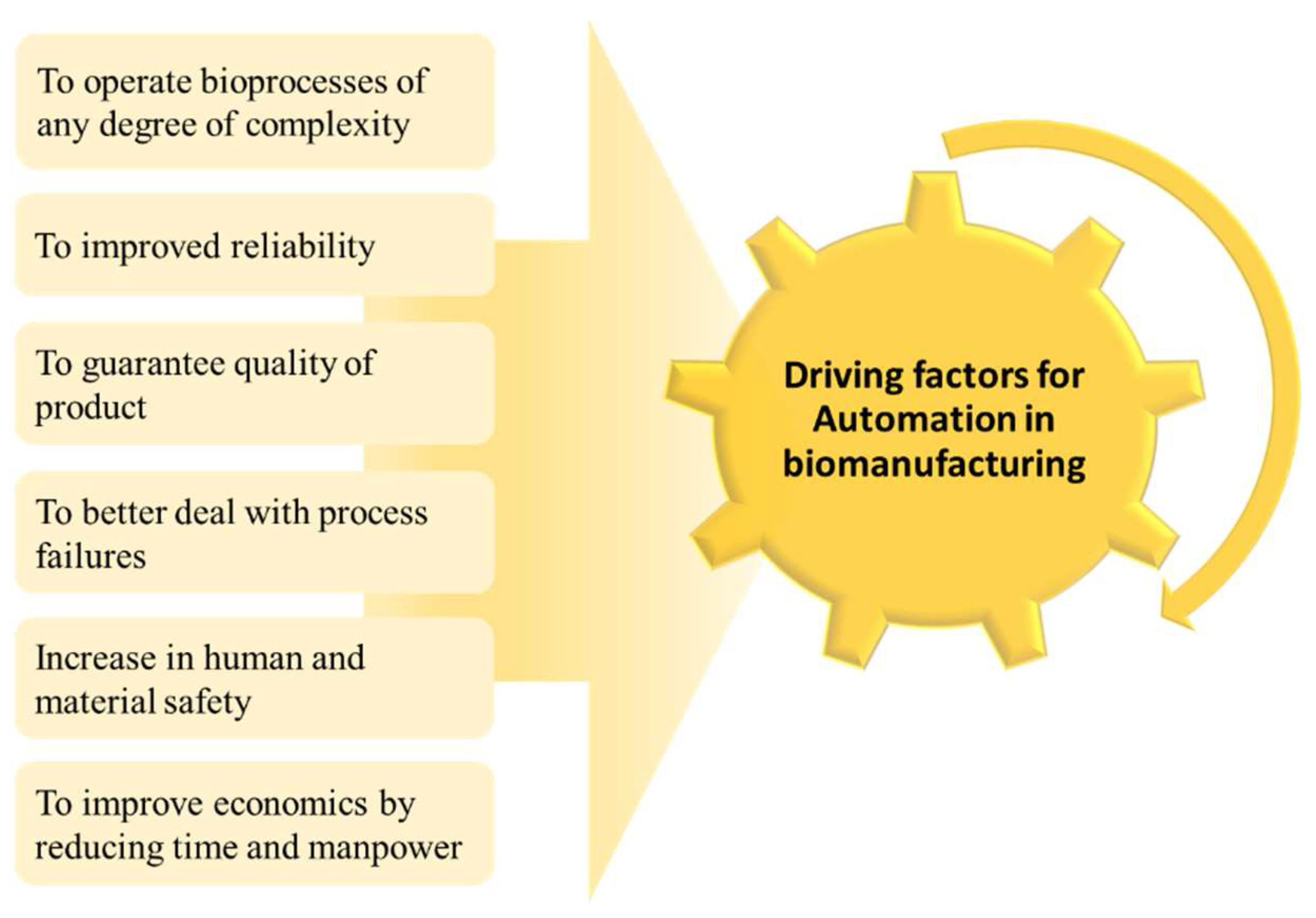
4.3. Automation in Biomanufacturing
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Alford, J.S. Bioprocess control: Advances and challenges. Comput. Chem. Eng. 2006, 30, 1464–1475. [Google Scholar] [CrossRef]
- Available online: https://www.nature.com/articles/d43747-020-00765-2#:~:text=From%20the%20publishers%20of%20Nature,attract%20partners%20and%20dealmaking%20opportunities (accessed on 11 February 2021).
- Available online: https://www.alliedmarketresearch.com/biopharmaceutical-market (accessed on 11 February 2021).
- Zulkeflee, S.A.; Aziz, N. Control Implementation in Bioprocess System: A Review. In Proceedings of the International Conference on Control, Instrumentation and Mechatronics Engineering (CIM’07), Johor Bahru, Malaysia, 28–29 May 2007; pp. 798–804. [Google Scholar]
- Hong, M.S.; Severson, K.A.; Jiang, M.; Lu, A.E.; Love, J.C.; Braatz, R.D. Challenges and opportunities in biopharmaceutical manufacturing control. Comput. Chem. Eng. 2018, 110, 106–114. [Google Scholar] [CrossRef]
- Baishan, F.; Hongwen, C.; Xiaolan, X.; Ning, W.; Zongding, H. Using genetic algorithms coupling neural networks in a study of xylitol production: Medium optimisation. Process. Biochem. 2003, 38, 979–985. [Google Scholar] [CrossRef]
- Franco-Lara, E.; Link, H.; Weuster-Botz, D. Evaluation of artificial neural networks for modelling and optimization of medium composition with a genetic algorithm. Process. Biochem. 2006, 41, 2200–2206. [Google Scholar] [CrossRef]
- Jenzsch, M.; Gnoth, S.; Beck, M.; Kleinschmidt, M.; Simutis, R.; Lübbert, A. Open-loop control of the biomass concentration within the growth phase of recombinant protein production processes. J. Biotechnol. 2006, 127, 84–94. [Google Scholar] [CrossRef]
- Jenzsch, M.; Gnoth, S.; Kleinschmidt, M.; Simutis, R.; Lübbert, A. Improving the batch-to-batch reproducibility in microbial cultures during recombinant protein production by guiding the process along a predefined total biomass profile. Bioprocess. Biosyst. Eng. 2006, 29, 315–321. [Google Scholar] [CrossRef]
- Gronemeyer, P.; Ditz, R.; Strube, J. Trends in upstream and downstream process development for antibody manufacturing. Bioengineering 2014, 1, 188–212. [Google Scholar] [CrossRef] [PubMed]
- Rathore, A.S.; Sofer, G. Process Validation in Manufacturing of Biopharmaceuticals. In Biotechnology and Bioprocessing; CRC Press: Boca Raton, FL, USA, 2012. [Google Scholar]
- Parenteral drug Association. PDA technical report no. 42: Process validation of protein manufacturing. PDA J. Pharm. Sci. Technol. 2005, 59 (Suppl. S4), 1–28. [Google Scholar]
- Conor, M.; Qi, Z.; Prodromos, D.; Wei-Shou, H. A hybrid mechanistic-empirical model for in silico mammalian cell bioprocess simulation. Metab. Eng. 2021, 66, 31–40. [Google Scholar]
- Oliver, J.F.; Nicholas, J.W.; Laura, P.; Darren, B.; Martin, R.; Rachel, L.G. Multiple target data-driven models to enable sustainable process manufacturing: An industrial bioprocess case study. J. Clean. Prod. 2021, 296, 126242. [Google Scholar]
- Hong, M.S.; Braatz, R.D. Mechanistic modeling and parameter-adaptive nonlinear model predictive control of a microbioreactor. Comput. Chem. Eng. 2021, 147, 107255. [Google Scholar] [CrossRef]
- Frank, N.; Alexandre, T.; Klaus-Robert, M.; Cecilia, C. Machine Learning for Molecular Simulation. Annu. Rev. Phys. Chem. 2020, 71, 361–390. [Google Scholar]
- Sebastian, R.; Benjamin, K. Machine learning and AI-based approaches for bioactive ligand discovery and GPCR-ligand recognition. Methods 2020, 180, 89–110. [Google Scholar]
- Gupta, R.; Srivastava, D.; Sahu, M.; Tiwari, S.; Ambasta, R.K.; Kumar, P. Artificial intelligence to deep learning: Machine intelligence approach for drug discovery. Mol. Divers. 2021, 1–46. [Google Scholar] [CrossRef]
- Rathore, A.S.; Winkle, H. Quality by design for biopharmaceuticals: Regulatory perspective and approach. Nat. Biotechnol. 2009, 27, 26–34. [Google Scholar] [CrossRef]
- Rathore, A.S.; Bhambure, R.; Ghare, V. Process analytical technology (PAT) for biopharmaceutical products. Anal. Bioanal. Chem. 2010, 398, 137–154. [Google Scholar] [CrossRef]
- Rathore, A.S. Roadmap for implementation of quality by design (QbD) for biotechnology products. Trends Biotechnol. 2009, 27, 546–553. [Google Scholar] [CrossRef] [PubMed]
- U.S. Department of Health and Human Services, Food and Drug Administration. Guidance for Industry: PAT—A Framework for Innovative Pharmaceutical Development, Manufacturing Andquality Assurance. 2004. Available online: http://www.fda.gov/downloads/Drugs/GuidanceComplianceRegulatoryInformation/Guidances/ucm070305.pdf (accessed on 11 February 2021).
- Rathore, A.S. QbD/PAT for bioprocessing: Moving from theory to implementation. Curr. Opin. Chem. Eng. 2014, 6, 1–8. [Google Scholar] [CrossRef]
- Ögmundarson, Ó.; Sukumara, S.; Herrgård, M.J.; Fantke, P. Combining environmental and economic performance for bioprocess optimization. Trends Biotechnol. 2020, 38, 1203–1214. [Google Scholar] [CrossRef]
- Narayanan, H.; Luna, M.F.; Von Stosch, M.; Bournazou, M.N.C.; Polotti, G.; Morbidelli, M.; Butté, A.; Sokolov, M. Bioprocessing in the digital age: The role of process models. Biotechnol. J. 2020, 15, e1900172. [Google Scholar] [CrossRef] [PubMed]
- Bunnak, P.; Allmendinger, R.; Ramasamy, S.V.; Lettieri, P.; Titchener-Hooker, N.J. Life-cycle and cost of goods assessment of fed-batch and perfusion-based manufacturing processes for mAbs. Biotechnol. Prog. 2016, 32, 1324–1335. [Google Scholar] [CrossRef]
- Farid, S.S.; Baron, M.; Stamatis, C.; Nie, W.; Coffman, J. Benchmarking biopharmaceutical process development and manufacturing cost contributions to R&D. mAbs 2020, 12, 1754999. [Google Scholar] [CrossRef] [PubMed]
- Zürcher, P.; Shirahata, H.; Badr, S.; Sugiyama, H. Multi-stage and multi-objective decision-support tool for biopharmaceutical drug product manufacturing: Equipment technology evaluation. Chem. Eng. Res. Des. 2020, 161, 240–252. [Google Scholar] [CrossRef]
- Ramasamy, S.V.; Titchener-Hooker, N.J.; Lettieri, P. Life cycle assessment as a tool to support decision making in the biopharmaceutical industry: Considerations and challenges. Food Bioprod. Process. 2015, 94, 297–305. [Google Scholar] [CrossRef]
- Mears, L.; Stocks, S.M.; Albaek, M.O.; Sin, G.; Gernaey, K.V. Mechanistic fermentation models for process design, monitoring, and control. Trends Biotechnol. 2017, 35, 914–924. [Google Scholar] [CrossRef]
- Yee, J.C.; Rehmann, M.S.; Yao, G.; Sowa, S.W.; Aron, K.L.; Tian, J.; Borys, M.C.; Li, Z.J. Advances in process control strategies for mammalian fed-batch cultures. Curr. Opin. Chem. Eng. 2018, 22, 34–41. [Google Scholar] [CrossRef]
- Mears, L.; Stocks, S.M.; Sin, G.; Gernaey, K.V. A review of control strategies for manipulating the feed rate in fed-batch fermentation processes. J. Biotechnol. 2017, 245, 34–46. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.-C.; Inwood, S.; Gong, T.; Sharma, A.; Yu, L.-Y.; Zhu, P. Fed-batch high-cell-density fermentation strategies for Pichia pastoris growth and production. Crit. Rev. Biotechnol. 2019, 39, 258–271. [Google Scholar] [CrossRef]
- Julia, R.; Xavier, G.; José, L.M.; Denise, F.; Francisco, V. Continuous operation, a realistic alternative to fed-batch fermentation for the production of recombinant lipase B from Candida antarctica under the constitutive promoter PGK in Pichia pastoris. Biochem. Eng. J. 2019, 147, 39–47. [Google Scholar]
- Khan, O.; Madhuranthakam, C.M.R.; Douglas, P.; Lau, H.; Sun, J.; Farrell, P. Optimized PID controller for an industrial biological fermentation process. J. Process. Control. 2018, 71, 75–89. [Google Scholar] [CrossRef]
- De Freitas, H.F.S.; Olivo, J.E.; Andrade, C.M.G. Optimization of bioethanol in silico production process in a fed-batch bioreactor using non-linear model predictive control and evolutionary computation techniques. Energies 2017, 10, 1763. [Google Scholar] [CrossRef]
- García, C.; Alcaraz, W.; Acosta-Cárdenas, A.; Ochoa, S. Application of process system engineering tools to the fed-batch production of poly(3-hydroxybutyrate-co-3-hydroxyvalerate) from a vinasses–molasses Mixture. Bioprocess. Biosyst. Eng. 2019, 42, 1023–1037. [Google Scholar] [CrossRef]
- Scomparin, A.; Bureik, M. A convenient new method for reproducible fed-batch fermentation of fission yeast Schizosaccharomyces pombe. Biotechnol. Lett. 2020, 42, 937–943. [Google Scholar] [CrossRef]
- Pantano, M.N.; Serrano, M.E.; Fernández, M.C.; Rossomando, F.G.; Ortiz, O.A.; Scaglia, G.J.E. Multivariable control for tracking optimal profiles in a nonlinear fed-batch bioprocess integrated with state estimation. Ind. Eng. Chem. Res. 2017, 56, 6043–6056. [Google Scholar] [CrossRef]
- Joanofarc, X.; Nivedhika, D.; Patnaik, S.; Panda, R. Closed-loop Performance and Analysis of a Real Time Non-linear Bioreactor Process. In Proceedings of the 2019 2nd International Conference on Power and Embedded Drive Control (ICPEDC), Chennai, India, 21–23 August 2019; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2019; pp. 406–412. [Google Scholar]
- Kumar, M.; Prasad, D.; Giri, B.S.; Singh, R.S. Temperature control of fermentation bioreactor for ethanol production using IMC-PID controller. Biotechnol. Rep. 2019, 22, e00319. [Google Scholar] [CrossRef]
- Abadli, M.; Dewasme, L.; Tebbani, S.; Dumur, D.; Wouwer, A.V. Generic Model Control Applied to E. coli BL21(DE3) Fed-Batch Cultures. Process 2020, 8, 772. [Google Scholar] [CrossRef]
- Carredano, E.N.; Nordberg, R.; Westin, S.; Busson, K.; Karlsson, T.M.; Blank, T.S.; Sandegren, H.; Jagschies, G. Simplification of Buffer formulation and improvement of buffer control with in-line conditioning (IC). Biopharm. Process. 2018, 513–525. [Google Scholar] [CrossRef]
- Fabbrini, D.; Simonini, C.; Lundkvist, J.; Carredano, E.; Otero, D. Addressing the challenge of complex buffer management an in-line conditioning collaboration. BioProcess Int. 2017, 15, 43–46. [Google Scholar]
- Sendrescu, D.; Petre, E.; Selisteanu, D. Nonlinear PID controller for a Bacterial Growth Bioprocess. In Proceedings of the 2017 18th International Carpathian Control Conference (ICCC), Sinaia, Romania, 28–31 May 2017; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2017; pp. 151–155. [Google Scholar]
- Simutis, R.; Lübbert, A. Bioreactor control improves bioprocess performance. Biotechnol. J. 2015, 10, 1115–1130. [Google Scholar] [CrossRef] [PubMed]
- Schlembach, I.; Grünberger, A.; Rosenbaum, M.A.; Regestein, L. Measurement techniques to resolve and control population dynamics of mixed-culture processes. Trends Biotechnol. 2021. [Google Scholar] [CrossRef] [PubMed]
- Kager, J.; Tuveri, A.; Ulonska, S.; Kroll, P.; Herwig, C. Experimental verification and comparison of model predictive, PID and model inversion control in a Penicillium chrysogenum fed-batch process. Process. Biochem. 2020, 90, 1–11. [Google Scholar] [CrossRef]
- Bolton, W. Controllers. In Instrumentation and Control Systems, 3rd ed.; Elsevier Science: Amsterdam, The Netherlands, 2021; pp. 297–327, Chapter 13. [Google Scholar]
- Raja, G.L.; Ali, A. Modified parallel cascade control structure for integrating processes. In Proceedings of the 2015 International Conference on Recent Developments in Control, Automation and Power Engineering (RDCAPE), Noida, India, 12–13 March 2015; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2015; pp. 90–95. [Google Scholar]
- Campos-Rodríguez, A.; García-Sandoval, J.; González-Álvarez, V. Hybrid cascade control for a class of nonlinear dynamical systems. J. Process. Control. 2019, 76, 141–154. [Google Scholar] [CrossRef]
- Raja, G.L.; Ali, A. Series cascade control: An outline survey. In Proceedings of the 2017 Indian Control Conference (ICC), Guwahati, India, 4–6 January 2017; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2017; pp. 409–414. [Google Scholar]
- Velez-Suberbie, M.L.; Betts, J.P.J.; Walker, K.L.; Robinson, C.; Zoro, B.; Keshavarz-Moore, E. High throughput automated microbial bioreactor system used for clone selection and rapid scale-down process optimization. Biotechnol. Prog. 2018, 34, 58–68. [Google Scholar] [CrossRef]
- Hoshan, L.; Jiang, R.; Moroney, J.; Bui, A.; Zhang, X.; Hang, T.-C.; Xu, S. Effective bioreactor pH control using only sparging gases. Biotechnol. Prog. 2019, 35, e2743. [Google Scholar] [CrossRef] [PubMed]
- Gehan, O.; Pigeon, E.; Menard, T.; Mosrati, R.; Pouliquen, M.; Fall, L.; Reuter, J. Dissolved oxygen level output feedback control based on discrete-time measurements during a Pseudomonas putida mt-2 fermentation. J. Process. Control. 2019, 79, 29–40. [Google Scholar] [CrossRef]
- Kuprijanov, A.; Schaepe, S.; Aehle, M.; Simutis, R.; Lübbert, A. Improving cultivation processes for recombinant protein production. Bioprocess. Biosyst. Eng. 2011, 35, 333–340. [Google Scholar] [CrossRef]
- Sommeregger, W.; Sissolak, B.; Kandra, K.; Von Stosch, M.; Mayer, M.; Striedner, G. Quality by control: Towards model predictive control of mammalian cell culture bioprocesses. Biotechnol. J. 2017, 12. [Google Scholar] [CrossRef]
- Sansana, J.; Joswiak, M.N.; Castillo, I.; Wang, Z.; Rendall, R.; Chiang, L.H.; Reis, M.S. Recent trends on hybrid modeling for Industry 4.0. Comput. Chem. Eng. 2021, 2021, 107365. [Google Scholar] [CrossRef]
- Hille, R.; Brandt, H.; Colditz, V.; Classen, J.; Hebing, L.; Langer, M.; Kreye, S.; Neymann, T.; Krämer, S.; Tränkle, J.; et al. Application of model-based online monitoring and robust optimizing control to fed-batch bioprocesses. IFAC-PapersOnLine 2020, 53, 16846–16851. [Google Scholar] [CrossRef]
- Sha, S.; Huang, Z.; Wang, Z.; Yoon, S. Mechanistic modeling and applications for CHO cell culture development and production. Curr. Opin. Chem. Eng. 2018, 22, 54–61. [Google Scholar] [CrossRef]
- Jabarivelisdeh, B.; Carius, L.; Findeisen, R.; Waldherr, S. Adaptive predictive control of bioprocesses with constraint-based modeling and estimation. Comput. Chem. Eng. 2020, 135, 106744. [Google Scholar] [CrossRef]
- Sridhar, L.N. Multiobjective optimization and nonlinear model predictive control of the continuous fermentation process involving Saccharomyces Cerevisiae. Biofuels 2019, 1, 1–16. [Google Scholar] [CrossRef]
- Chang, L.; Liu, X.; Henson, M.A. Nonlinear model predictive control of fed-batch fermentations using dynamic flux balance models. J. Process. Control. 2016, 42, 137–149. [Google Scholar] [CrossRef]
- Santos, L.; Dewasme, L.; Coutinho, D.; Wouwer, A.V. Nonlinear model predictive control of fed-batch cultures of micro-organisms exhibiting overflow metabolism: Assessment and robustness. Comput. Chem. Eng. 2012, 39, 143–151. [Google Scholar] [CrossRef]
- Gorrini, F.; Biagiola, S.; Figueroa, J.L.; Wouwer, A.V. Reaction rate estimation and model predictive control of hybridoma cell cultures. IFAC-PapersOnLine 2019, 52, 715–720. [Google Scholar] [CrossRef]
- Dewasme, L.; Fernandes, S.; Amribt, Z.; Santos, L.; Bogaerts, P.; Wouwer, A.V. State estimation and predictive control of fed-batch cultures of hybridoma cells. J. Process. Control. 2015, 30, 50–57. [Google Scholar] [CrossRef]
- Lyon, D. Achieving active control of cell culture performance with the aid of machine learning techniques, Abstracts. In Proceedings of the 74th Northwest Regional Meeting of the American Chemical Society, NORM-298. Portland, OR, USA, 16–19 June 2019. [Google Scholar]
- Papathanasiou, M.M.; Burnak, B.; Katz, J.; Shah, N.; Pistikopoulos, E.N. Assisting continuous biomanufacturing through advanced control in downstream purification. Comput. Chem. Eng. 2019, 125, 232–248. [Google Scholar] [CrossRef]
- Noll, P.; Henkel, M. History and evolution of modeling in biotechnology: Modeling & simulation, application and hardware performance. Comput. Struct. Biotechnol. J. 2020, 18, 3309–3323. [Google Scholar] [CrossRef]
- Lauri, D.; Lennox, B.; Camacho, J. Model predictive control for batch processes: Ensuring validity of predictions. J. Process. Control. 2014, 24, 239–249. [Google Scholar] [CrossRef]
- Kotidis, P.; Demis, P.; Goey, C.H.; Correa, E.; McIntosh, C.; Trepekli, S.; Shah, N.; Klymenko, O.V.; Kontoravdi, C. Constrained global sensitivity analysis for bioprocess design space identification. Comput. Chem. Eng. 2019, 125, 558–568. [Google Scholar] [CrossRef]
- Armstrong, A.; Horry, K.; Cui, T.; Hulley, M.; Turner, R.; Farid, S.S.; Goldrick, S.; Bracewell, D.G. Advanced control strategies for bioprocess chromatography: Challenges and opportunities for intensified processes and next generation products. J. Chromatogr. A 2021, 1639, 461914. [Google Scholar] [CrossRef]
- Abiodun, O.I.; Jantan, A.; Omolara, A.E.; Dada, K.V.; Mohamed, N.A.; Arshad, H. State-of-the-art in artificial neural network applications: A survey. Heliyon 2018, 4, e00938. [Google Scholar] [CrossRef] [PubMed]
- Del Rio-Chanona, E.A.; Manirafasha, E.; Zhang, D.; Yue, Q.; Jing, K. Dynamic modeling and optimization of cyanobacterial C-phycocyanin production process by artificial neural network. Algal Res. 2016, 13, 7–15. [Google Scholar] [CrossRef]
- Natarajan, P.; Moghadam, R.; Jagannathan, S. Online deep neural network-based feedback control of a Lutein bioprocess. J. Process. Control. 2021, 98, 41–51. [Google Scholar] [CrossRef]
- Rómoli, S.; Serrano, M.; Rossomando, F.; Vega, J.; Ortiz, O.; Scaglia, G. Neural network-based state estimation for a closed-loop control strategy applied to a fed-batch bioreactor. Complex 2017, 2017, 9391879. [Google Scholar] [CrossRef]
- Murugan, C.; Natarajan, P. Estimation of fungal biomass using multiphase artificial neural network based dynamic soft sensor. J. Microbiol. Methods 2019, 159, 5–11. [Google Scholar] [CrossRef] [PubMed]
- López, M.E.; Rene, E.R.; Boger, Z.; Veiga, M.C.; Kennes, C. Modelling the removal of volatile pollutants under transient conditions in a two-stage bioreactor using artificial neural networks. J. Hazard. Mater. 2017, 324, 100–109. [Google Scholar] [CrossRef] [PubMed]
- Sena, H.J.; da Silva, F.V.; Fileti, A.M.F. ANN model adaptation algorithm based on extended Kalman filter applied to pH control using MPC. J. Process. Control. 2021, 102, 15–23. [Google Scholar] [CrossRef]
- Tavasoli, T.; Arjmand, S.; Siadat, S.O.R.; Shojaosadati, S.A.; Lotfi, A.S. A robust feeding control strategy adjusted and optimized by a neural network for enhancing of alpha 1-antitrypsin production in Pichia pastoris. Biochem. Eng. J. 2019, 144, 18–27. [Google Scholar] [CrossRef]
- Kahraman, C.; Öztayşi, B.; Onar, S.; Çevik, A. Comprehensive Literature Review of 50 Years of Fuzzy Set Theory. Int. J. Comput. Intell. Syst. 2016, 9, 3–24. [Google Scholar] [CrossRef]
- Abyad, M.; Karama, A.; Khallouq, A. Fuzzy Takagi-Sugeno based modelling and control for an alcoholic fermentation process. In Proceedings of the 2017 International Conference on Electrical and Information Technologies (ICEIT), Rabat, Morocco, 15–18 November 2017; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2017; pp. 1–6. [Google Scholar]
- Abyad, M.; Karama, A.; Khallouq, A. Takagi-Sugeno tracking control of a fermentation process with respect to asymmetric constraints. Int. J. Adapt. Control. Signal. Process. 2019, 34, 266–282. [Google Scholar] [CrossRef]
- Khamparia, A.; Pandey, B.; Pandey, D.; Gupta, D.; Khanna, A.; de Albuquerque, V.H.C. Comparison of RSM, ANN and fuzzy logic for extraction of oleonolic acid from ocimum sanctum. Comput. Ind. 2020, 117, 103200. [Google Scholar] [CrossRef]
- Fonseca, R.R.; Franco, I.C.; Da Silva, F.V. Bioreactor temperature control using a generic fuzzy feedforward control system. In Proceedings of the 15th IASTED International Conference Intelligent Systems and Control (ISC 2016), Campinas, Brazil, 16–18 August 2016. [Google Scholar] [CrossRef]
- Fonseca, R.R.; Sencio, R.R.; Franco, I.C.; Da Silva, F.V. An adaptive fuzzy feedforward-feedback control system applied to a saccharification process. Chem. Prod. Process. Model. 2018, 13, 13. [Google Scholar] [CrossRef]
- Escalante-Sánchez, A.; Barrera-Cortés, J.; Poggi-Varaldo, H.M.; Ponce-Noyola, T.; Baruch, I.S. A soft sensor based on online biomass measurements for the glucose estimation and control of fed-batch cultures of Bacillus thuringiensis. Bioprocess Biosyst. Eng. 2018, 41, 1471–1484. [Google Scholar] [CrossRef]
- Humbird, D.; Fei, Q. Scale-Up Considerations for Biofuels. In Biotechnology for Biofuel Production and Optimization; Eckert, C.A., Trinh, C.T., Eds.; Elsevier: Amsterdam, The Netherlands, 2016; pp. 513–537. [Google Scholar]
- Benattia, S.E.; Tebbani, S.; Dumur, D. Linearized min-max robust model predictive control: Application to the control of a bioprocess. Int. J. Robust Nonlinear Control. 2020, 30, 100–120. [Google Scholar] [CrossRef]
- Pistikopoulos, E.N.; Diangelakis, N.A.; Oberdieck, R.; Papathanasiou, M.M.; Nascu, I.; Sun, M. PAROC—An integrated framework and software platform for the optimisation and advanced model-based control of process systems. Chem. Eng. Sci. 2015, 136, 115–138. [Google Scholar] [CrossRef]
- Papathanasiou, M.M.; Avraamidou, S.; Oberdieck, R.; Mantalaris, A.; Steinebach, F.; Morbidelli, M.; Mueller-Spaeth, T.; Pistikopoulos, S. Advanced control strategies for the multicolumn countercurrent solvent gradient purification process. AIChE J. 2016, 62, 2341–2357. [Google Scholar] [CrossRef]
- Papathanasiou, M.M.; Burnak, B.; Katz, J.; Shah, N.; Pistikopoulos, S. Control of a dual mode separation process via multi-parametric Model Predictive Control. IFAC-PapersOnLine 2019, 52, 988–993. [Google Scholar] [CrossRef]
- Papathanasiou, M.M.; Burnak, B.; Katz, J.; Müller-Späth, T.; Morbidelli, M.; Shah, N.; Pistikopoulos, E.N. Control of small-scale chromatographic systems under disturbances. In Computer Aided Chemical Engineering; Salvador Garcia Muñoz, S.G., Laird, C.D., Realff, M.J., Eds.; Elsevier: Amsterdam, The Netherlands, 2019; Volume 47, pp. 269–274. [Google Scholar]
- Papathanasiou, M.M.; Steinebach, F.; Morbidelli, M.; Mantalaris, A.; Pistikopoulos, E.N. Intelligent, model-based control towards the intensification of downstream processes. Comput. Chem. Eng. 2017, 105, 173–184. [Google Scholar] [CrossRef]
- Steinebach, F.; Angarita, M.; Karst, D.J.; Müller-Späth, T.; Morbidelli, M. Model based adaptive control of a continuous capture process for monoclonal antibodies production. J. Chromatogr. A 2016, 1444, 50–56. [Google Scholar] [CrossRef]
- Lim, H.-K.; Choi, S.-J.; Kim, K.-Y.; Jung, K.-H. Dissolved-oxygen-stat controlling two variables for methanol induction of rGuamerin in Pichia pastoris and its application to repeated fed-batch. Appl. Microbiol. Biotechnol. 2003, 62, 342–348. [Google Scholar] [CrossRef]
- Potvin, G.; Ahmad, A.; Zhang, Z. Bioprocess engineering aspects of heterologous protein production in Pichia pastoris: A review. Biochem. Eng. J. 2012, 64, 91–105. [Google Scholar] [CrossRef]
- Zheng, R.; Pan, F. On-Line Tendency Control of Dissolved Oxygen Concentration during Aerobic Fed-Batch Fermentations. Appl. Sci. 2019, 9, 5232. [Google Scholar] [CrossRef]
- Ding, J.; Gao, M.J.; Hou, G.L.; Liang, K.X.; Yu, R.S.; Li, Z.; Shi, Z.P. Stabilizing porcine interferon-α production by Pichia pastoris with an ethanol on-line measurement based DO-Stat glycerol feeding strategy. J. Chem. Technol. Biotechnol. 2015, 89, 1948–1953. [Google Scholar] [CrossRef]
- García-Arrazola, R.; Siu, S.C.; Chan, G.; Buchanan, I.; Doyle, B.; Titchener-Hooker, N.; Baganz, F. Evaluation of a pH-stat feeding strategy on the production and recovery of Fab’ fragments from E. coli. Biochem. Eng. J. 2005, 23, 221–230. [Google Scholar] [CrossRef]
- Li, K.-T.; Liu, D.-H.; Chu, J.; Wang, Y.-H.; Zhuang, Y.-P.; Zhang, S.-L. An effective and simplified pH-stat control strategy for the industrial fermentation of vitamin B12 by Pseudomonas denitrificans. Bioprocess. Biosyst. Eng. 2008, 31, 605–610. [Google Scholar] [CrossRef]
- Bastin, G.; Nešić, D.; Tan, Y.; Mareels, I. On extremum seeking in bioprocesses with multivalued cost functions. Biotechnol. Prog. 2009, 25, 683–689. [Google Scholar] [CrossRef] [PubMed]
- Daaou, B.; Dochain, D.; Bachir, D. High order sliding mode observer based extremum seeking controller for a continuous stirred tank bioreactor. In Proceedings of the 2015 3rd International Conference on Control, Engineering & Information Technology (CEIT), Tlemcen, Algeria, 25–27 May 2015; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2015; pp. 1–6. [Google Scholar]
- Johnsson, O.; Sahlin, D.; Linde, J.; Lidén, G.; Hägglund, T. A mid-ranging control strategy for non-stationary processes and its application to dissolved oxygen control in a bioprocess. Control. Eng. Pr. 2015, 42, 89–94. [Google Scholar] [CrossRef]
- Gomis-Fons, J.; Schwarz, H.; Zhang, L.; Andersson, N.; Nilsson, B.; Castan, A.; Solbrand, A.; Stevenson, J.; Chotteau, V. Model-based design and control of a small-scale integrated continuous end-to-end mAb platform. Biotechnol. Prog. 2020, 36, e2995. [Google Scholar] [CrossRef]
- Jiang, M.; Braatz, R.D. Integrated control of continuous (bio)pharmaceutical manufacturing. Am. Pharm Rev. 2016, 19, 110–115. [Google Scholar]
- Birle, S.; Hussein, M.A.; Becker, T. Management of Uncertainty by Statistical Process Control and a Genetic Tuned Fuzzy System. Discret. Dyn. Nat. Soc. 2016, 2016, 1–11. [Google Scholar] [CrossRef]
- Bose, B.K. Artificial intelligence applications in renewable energy systems and smart grid—Some novel applications. In Power Electronics in Renewable Energy Systems and Smart Grid; Wiley: Hoboken, NJ, USA, 2019; pp. 625–675. [Google Scholar]
- Precup, R.-E.; David, R.-C. Nature-inspired algorithms for the optimal tuning of fuzzy controllers. In Nature-inspired Optimization Algorithms for Fuzzy Controlled Servo Systems; Elsevier: Amsterdam, The Netherlands, 2019; pp. 55–80. [Google Scholar]
- Rios, J.D.; Alanis, A.Y.; Arana-Daniel, N.; Lopez-Franco, C. Neural Networks Modeling and Control. In Neural Networks Modeling and Control; Elsevier BV: Amsterdam, The Netherlands, 2020. [Google Scholar]
- Ahmad, S.K.; Guenard, R.; Romero-Torres, S.; Antoniou, C. Hybrid model identification for monoclonal antibody production bioreactor—A digital twin. Am. Pharm Rev. 2019, 22, 1–12. [Google Scholar]
- Randek, J.; Mandenius, C.-F. On-line soft sensing in upstream bioprocessing. Crit. Rev. Biotechnol. 2017, 38, 106–121. [Google Scholar] [CrossRef]
- Guerra, A.; Von Stosch, M.; Glassey, J. Toward biotherapeutic product real-time quality monitoring. Crit. Rev. Biotechnol. 2019, 39, 289–305. [Google Scholar] [CrossRef] [PubMed]
- Biechele, P.; Busse, C.; Solle, D.; Scheper, T.; Reardon, K. Sensor systems for bioprocess monitoring. Eng. Life Sci. 2015, 15, 469–488. [Google Scholar] [CrossRef]
- Steinwandter, V.; Borchert, D.; Herwig, C. Data science tools and applications on the way to Pharma 4.0. Drug Discov. Today 2019, 24, 1795–1805. [Google Scholar] [CrossRef]
- Zhang, A.; Zhu, K.; Zhuang, X.; Liao, L.; Huang, S.; Yao, C.; Fang, B. A robust soft sensor to monitor 1,3-propanediol fermentation process by Clostridium butyricum based on artificial neural network. Biotechnol. Bioeng. 2020, 117, 3345–3355. [Google Scholar] [CrossRef]
- Brunner, V.; Klöckner, L.; Kerpes, R.; Geier, D.U.; Becker, T. Online sensor validation in sensor networks for bioprocess monitoring using swarm intelligence. Anal. Bioanal. Chem. 2019, 412, 2165–2175. [Google Scholar] [CrossRef] [PubMed]
- Grigs, O.; Bolmanis, E.; Galvanauskas, V. Application of in-situ and soft-sensors for estimation of recombinant P. pastoris GS115 biomass concentration: A case analysis of HBcAg (Mut+) and HBsAg (MutS) production processes under varying conditions. Sensors 2021, 21, 1268. [Google Scholar] [CrossRef] [PubMed]
- Pappenreiter, M.; Sissolak, B.; Sommeregger, W.; Striedner, G. Oxygen uptake rate soft-sensing via dynamic kLa computation: Cell volume and metabolic transition prediction in mammalian bioprocesses. Front. Bioeng. Biotechnol. 2019, 7, 195. [Google Scholar] [CrossRef]
- Bayer, B.; Von Stosch, M.; Melcher, M.; Duerkop, M.; Striedner, G. Soft sensor based on 2D-fluorescence and process data enabling real-time estimation of biomass in Escherichia coli cultivations. Eng. Life Sci. 2020, 20, 26–35. [Google Scholar] [CrossRef] [PubMed]
- Gopakumar, V.; Tiwari, S.; Rahman, I. A deep learning based data driven soft sensor for bioprocesses. Biochem. Eng. J. 2018, 136, 28–39. [Google Scholar] [CrossRef]
- Kornecki, M.; Strube, J. Process analytical technology for advanced process control in biologics manufacturing with the aid of macroscopic kinetic modeling. Bioengineering 2018, 5, 25. [Google Scholar] [CrossRef] [PubMed]
- Steinwandter, V.; Zahel, T.; Sagmeister, P.; Herwig, C. Propagation of measurement accuracy to biomass soft-sensor estimation and control quality. Anal. Bioanal. Chem. 2016, 409, 693–706. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zabadaj, M.; Chreptowicz, K.; Mierzejewska, J.; Ciosek, P. Two-dimensional fluorescence as soft sensor in the monitoring of biotransformation performed by yeast. Biotechnol. Prog. 2016, 33, 299–307. [Google Scholar] [CrossRef] [PubMed]
- Baradez, M.-O.; Biziato, D.; Hassan, E.; Marshall, D. Application of Raman Spectroscopy and Univariate Modelling as a Process Analytical Technology for Cell Therapy Bioprocessing. Front. Med. 2018, 5, 47. [Google Scholar] [CrossRef]
- Bhatia, H.; Mehdizadeh, H.; Drapeau, D.; Yoon, S. In-line monitoring of amino acids in mammalian cell cultures using raman spectroscopy and multivariate chemometrics models. Eng. Life Sci. 2018, 18, 55–61. [Google Scholar] [CrossRef]
- Brestrich, N.; Briskot, T.; Osberghaus, A.; Hubbuch, J. A tool for selective inline quantification of co-eluting proteins in chromatography using spectral analysis and partial least squares regression. Biotechnol. Bioeng. 2014, 111, 1365–1373. [Google Scholar] [CrossRef] [PubMed]
- Capito, F.; Skudas, R.; Kolmar, H.; Hunzinger, C. At-line mid infrared spectroscopy for monitoring downstream processing unit operations. Process. Biochem. 2015, 50, 997–1005. [Google Scholar] [CrossRef]
- Krishna, V.V.S.V.; Pappa, N.; Rani, S.P.J.V. Deep Learning based Soft Sensor for Bioprocess Application. In Proceedings of the 2021 IEEE Second International Conference on Control, Measurement and Instrumentation (CMI), Kolkata, India, 8–10 January 2021; Institute of Electrical and Electronics Engineers (IEEE): Piscataway, NJ, USA, 2021; pp. 155–159. [Google Scholar]
- Hrnčiřík, P.; Moucha, T.; Mareš, J.; Náhlík, J.; Janáčová, D. Software sensors for biomass concentration estimation in filamentous microorganism cultivation process. Chem. Biochem. Eng. Q. 2019, 33, 141–151. [Google Scholar] [CrossRef]
- Hausmann, R.; Henkel, M.; Hecker, F.; Hitzmann, B. Present Status of automation for industrial bioprocesses. Curr. Dev. Biotechnol. Bioeng. 2017, 2017, 725–757. [Google Scholar] [CrossRef]
- Alarcon, C.; Shene, C. Fermentation 4.0, a case study on computer vision, soft sensor, connectivity, and control applied to the fermentation of a thraustochytrid. Comput. Ind. 2021, 128, 103431. [Google Scholar] [CrossRef]
- Das, P.; Mishra, S.; Nayak, B. The colossal role of QbD and PAT tools in the pharmaceutical process automation. Int. J. Pharma Res. Health Sci. 2017, 5, 1909–1923. [Google Scholar]
- Jiang, M.; Severson, K.A.; Love, J.; Madden, H.; Swann, P.; Zang, L.; Braatz, R.D. Opportunities and challenges of real-time release testing in biopharmaceutical manufacturing. Biotechnol. Bioeng. 2017, 114, 2445–2456. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Fu, H.-Y.; Zhou, W.; Hu, W.-S. Advances in process monitoring tools for cell culture bioprocesses. Eng. Life Sci. 2015, 15, 459–468. [Google Scholar] [CrossRef]
- Chopda, V.R.; Gomes, J.; Rathore, A.S. Bridging the gap between PAT concepts and implementation: An integrated software platform for fermentation. Biotechnol. J. 2016, 11, 164–171. [Google Scholar] [CrossRef]
- Priyanka; Kumar, J.; Gomes, J.; Rathore, A.S.; Dalal, P. Implementing process analytical technology for the production of recombinant proteins in Escherichia coli using an advanced controller scheme. Biotechnol. J. 2019, 14, e1800556. [Google Scholar] [CrossRef]
- Jenzsch, M.; Bell, C.; Buziol, S.; Kepert, F.; Wegele, H.; Hakemeyer, C. Trends in process analytical technology: Present state in bioprocessing. Blue Biotechnol. 2017, 165, 211–252. [Google Scholar] [CrossRef]
- Ryder, A.G. Cell culture media analysis using rapid spectroscopic methods. Curr. Opin. Chem. Eng. 2018, 22, 11–17. [Google Scholar] [CrossRef]
- Schmid, I.; Aschoff, J. A scalable software framework for data integration in bioprocess development. Eng. Life Sci. 2016, 17, 1159–1165. [Google Scholar] [CrossRef]
- Whitford. The era of digital biomanufacturing. BioProcess Int. 2017, 15(3), 12–18. [Google Scholar]
- Tung, G.; Morris, K.; Perrone, P.; Reinbigler, R.; Miller, S.; Lai, C. The value of plug-and-play automation in single-use technology. BioProcess Int. 2019, 17, 12–19. [Google Scholar]
- Feidl, F.; Vogg, S.; Wolf, M.; Podobnik, M.; Ruggeri, C.; Ulmer, N.; Wälchli, R.; Souquet, J.; Broly, H.; Butté, A.; et al. Process-wide control and automation of an integrated continuous manufacturing platform for antibodies. Biotechnol. Bioeng. 2020, 117, 1367–1380. [Google Scholar] [CrossRef]
- CFR 21 Part 11—Food and Drug Administration (FDA) Guidelines on Electronic Records and Electronic Signatures; US FDA: Montgomery, MD, USA, 2003.
- Charan, H.Y.; Gupta, N.V. Gamp 5 a Quality Risk Management Approach to Computer System Validation. Int. J. Pharm. Sci. Rev. Res. 2016, 36, 195–198. [Google Scholar]

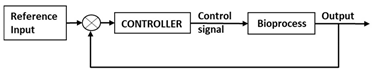
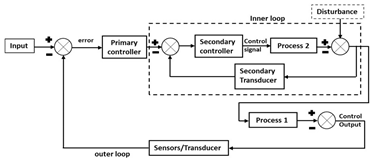
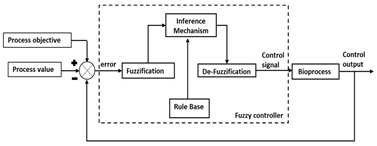
| Control Strategy | Description | Control Structure |
|---|---|---|
| Open loop control |
|  |
| Closed loop control/ feedback control with PI/PID controller |
|  |
| Cascade control |
|  |
| Fuzzy logic-based control |
|  |
| Artificial neural network-based control |
| 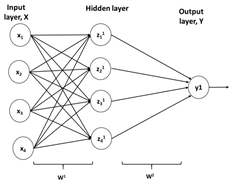 |
| Model predictive control |
| 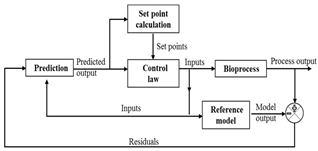 |
| Model based control |
| 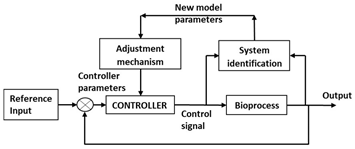 |
| Technique | Application | Reference |
|---|---|---|
| Artificial neural network | Monitor fermentation process—measurement of glycerol, 1,2 propanediol and biomass | [116] |
| Multiple linear regression in combination with mechanistic model | Prediction of biomass concentration | [117] |
| Empirical models | Biomass concentration | [118] |
| Empirical modelling of oxygen uptake rate | Predicted viable biomass | [119] |
| Multivariate adaptive regression spline algorithm in combination with 2D fluorescence spectra and process data | Biomass concentration | [120] |
| Artificial neural network | Glucose estimation in fed batch culture | [78] |
| Deep neural network | Parameter estimation in penicillin and streptokinase fermentation process | [121] |
| Partial least square regression with turbidity and Raman measurements in combination with Monod kinetics model | Predict substrate concentration | [122] |
| Error propagation method | Estimate error in predicted biomass fermentation rate and substrate consumption rate | [123] |
| Partial least square regression in combination with fluorescence fingerprinting | Monitor biotransformation production of 2 phenyl ethanol by yeast | [124] |
| Partial least squares regression model in combination with Raman spectroscopy | In line monitoring of the nutrient consumption and production of markers associated with cell metabolism | [125] |
| Partial least squares regression model in combination with Raman spectroscopy | In line monitoring on amino acid | [126] |
| Partial least squares regression with the help of mid-UV absorption spectra | Monitor chromatography steps | [127] |
| Partial least squares regression with Fourier transform mid infrared spectroscopy | Monitor HCP and aggregates at line | [128] |
| Deep neural network | Lactose and ethanol concentration measurement | [129] |
| Linear regression on data obtained from biocalorimetry in combination with bioreactor off gas analysis | Biomass concentration measurement | [130] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rathore, A.S.; Mishra, S.; Nikita, S.; Priyanka, P. Bioprocess Control: Current Progress and Future Perspectives. Life 2021, 11, 557. https://doi.org/10.3390/life11060557
Rathore AS, Mishra S, Nikita S, Priyanka P. Bioprocess Control: Current Progress and Future Perspectives. Life. 2021; 11(6):557. https://doi.org/10.3390/life11060557
Chicago/Turabian StyleRathore, Anurag S., Somesh Mishra, Saxena Nikita, and Priyanka Priyanka. 2021. "Bioprocess Control: Current Progress and Future Perspectives" Life 11, no. 6: 557. https://doi.org/10.3390/life11060557
APA StyleRathore, A. S., Mishra, S., Nikita, S., & Priyanka, P. (2021). Bioprocess Control: Current Progress and Future Perspectives. Life, 11(6), 557. https://doi.org/10.3390/life11060557






