Multigene Phylogeny Reveals Haploanthostomella elaeidis gen. et sp. nov. and Familial Replacement of Endocalyx (Xylariales, Sordariomycetes, Ascomycota)
Abstract
1. Introduction
2. Materials and Methods
2.1. Collection, Isolation, and Identification
2.2. DNA Extraction and Amplification (PCR)
2.3. Phylogenetic Analyses
3. Results
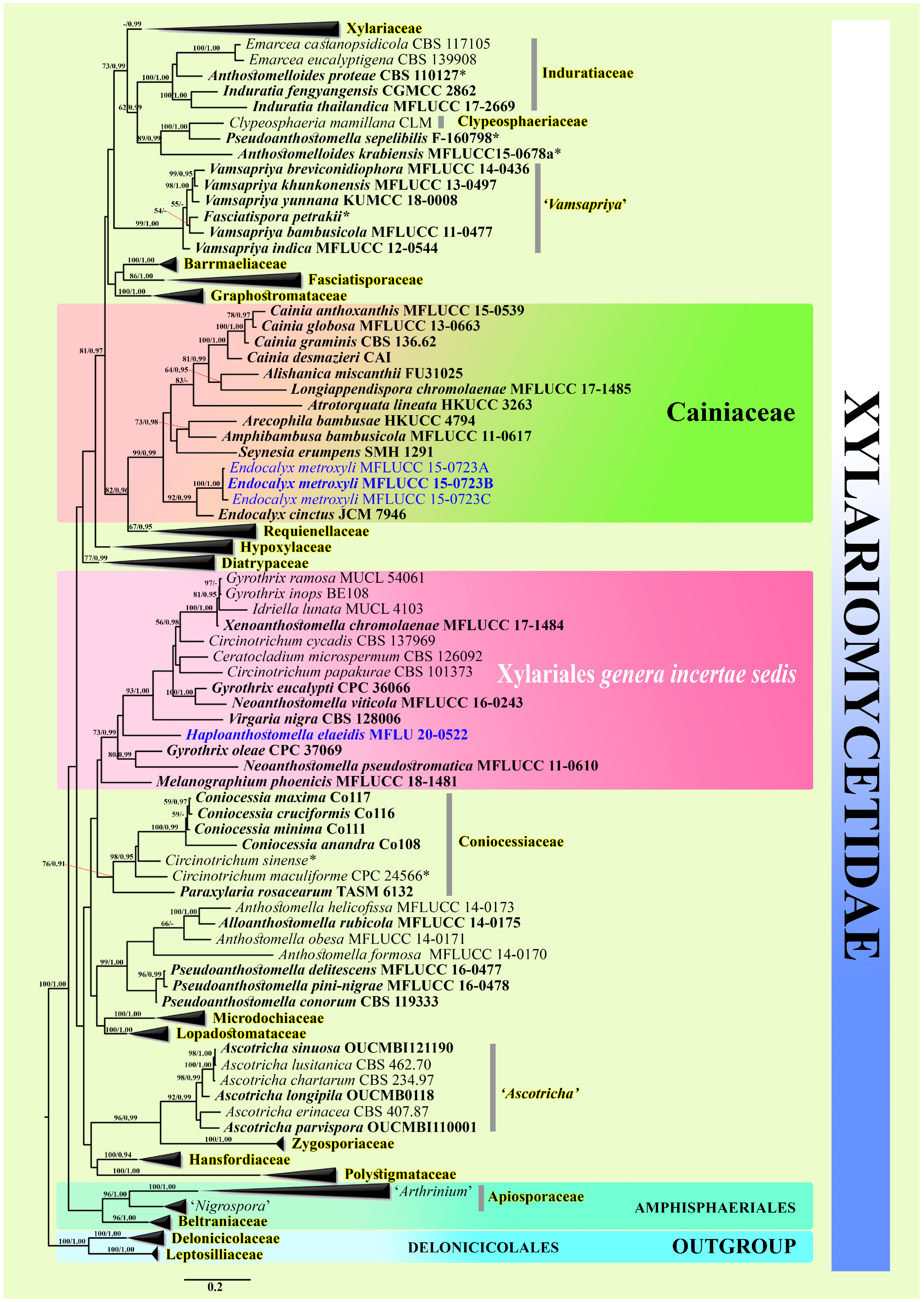
3.1. Morphology and Phylogeny
3.1.1. Haploanthostomella Konta & K.D. Hyde. gen. nov.
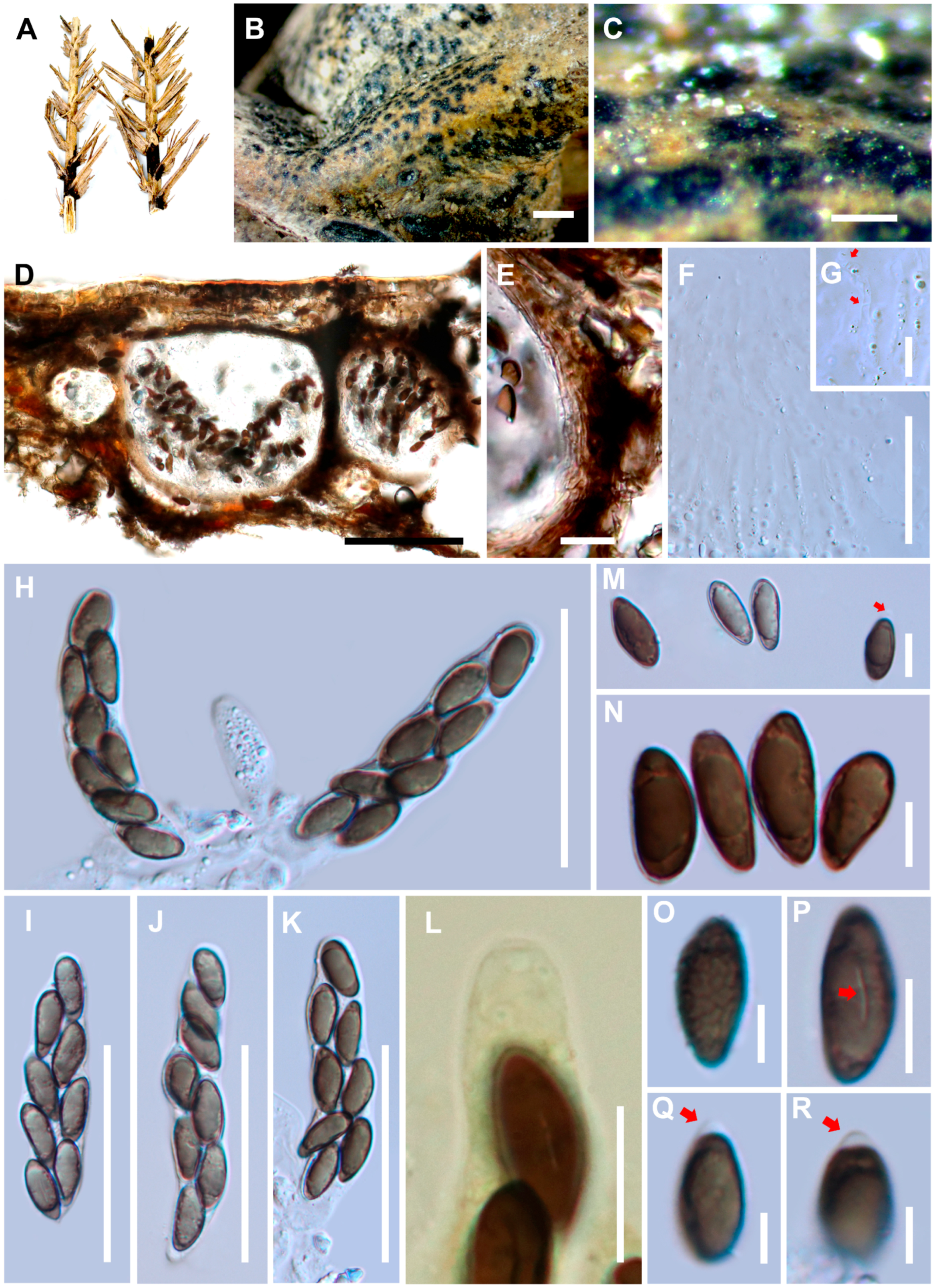
3.1.2. Haploanthostomella elaeidis Konta & K.D. Hyde., sp. nov.
| Key to genera related to Anthostomella-like genera | |
| 1. Hyaline ascospores | Alloanthostomella |
| 1. Brown ascospores | 2 |
| 2. Asci with a J- apical ring | 3 |
| 2. Asci with or without J+ apical ring | 5 |
| 3. Ascospores with or without germ slit | 4 |
| 3. Ascospores with germ slit | Xenoanthostomella |
| 4. Ascospores with a germ slit and the length less than spore length with a mucilaginous cap at the apex | Haploanthostomella |
| 4. Ascospores with or without germ slit, with mucilaginous sheath | Neoanthostomella |
| 5. Asci with a J+ apical ring, ascospores with germ slit, with or without mucilaginous sheath | 6 |
| 5. Asci with J+ or J- apical ring, ascospores with or without germ slit (straight or spiral), and also with or without appendages or mucilaginous sheath | Anthostomella |
| 6. Ascospores with germ slit less than spore length, with or without mucilaginous sheath | 7 |
| 6. Ascospores with germ slit extending over full length with mucilaginous sheath | Pseudoanthostomella |
| 7. Ellipsoid ascospores without mucilaginous sheath | Virgaria |
| 7. Inequilaterally oblong-ellipsoidal ascospores with mucilaginous sheath | Anthostomelloides |
3.1.3. Endocalyx Berk. & Broome, J. Linn. Soc., Bot. 15(1): 84 (1876) [1877]
3.1.4. Endocalyx metroxyli Konta & K.D. Hyde. sp. nov.
| Keys to genera of Cainiaceae | |
| 1. Asexual morph | |
| 1.1 Coelomycetous; 1–3 phialides conidiogenous cells, and elongate fusiform conidia with unicellular or septate, with pointed ends | Cainia |
| 1.1 Coelomycetous; conidiomata with ostiolar opening surrounded by yellow, with unicellular conidia, ellipsoid-globose, pale to dark brown to black, with a straight germ slit extending the full-length | Endocalyx |
| 2. Sexual morph | |
| 2.1 Cylindrical-clavate asci, ascospores with 1-septate | (2.2) |
| 2.1 Cylindrical, or cylindrical to elongate cylindrical asci, ascospores with 1-septate | (2.3) |
| 2.2 Ellipsoidal ascospores, with brown, and sheath | Cainia |
| 2.2 Ellipsoidal to fusiform ascospores, with brown, and sheath | Atrotorquata |
| 2.3 Ellipsoid to broadly fusiform ascospores, longitudinal striations, bristle-like polar appendages from both ends, without a gelatinous sheath | Longiappendispora |
| 2.3 Fusiform to broad-fusiform ascospores with pointed at both ends, striation wall, and sheath | Amphibambusa |
| 2.3 Ellipsoidal or oblong ascospores | (2.4) |
| 2.4 Oblong ascospores with cap-like appendage, germ slits | Seynesia |
| 2.4 Ellipsoidal ascospores | (2.5) |
| 2.5 Ascospores with striation wall, brown, and sheath | Alishanica |
| 2.5 Ascospores with striate or verrucose wall, and subhyaline to brown | Arecophila |
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lewis, C.E.; Baker, W.J.; Asmussen, C.B. DNA and palm evolution. Palms 2000, 44, 19–24. [Google Scholar]
- Hawksworth, D.L.; Lücking, R. Fungal diversity revisited: 2.2 to 3.8 million species. Fungal Kingd. 2017, 4, 79–95. [Google Scholar] [CrossRef]
- Hyde, K.D.; Norphanphoun, C.; Maharachchikumbura, S.S.N.; Bhat, D.J.; Jones, E.B.G.; Bundhun, D.; Chen, Y.J.; Bao, D.F.; Boonmee, S.; Calabon, M.S.; et al. Refined families of Sordariomycetes. Mycosphere 2020, 11, 305–1059. [Google Scholar] [CrossRef]
- Samarakoon, M.C.; Thongbai, B.; Hyde, K.D.; Brönstrup, M.; Beutling, U.; Lambert, C.; Miller, A.N.; Liu, J.K.J.; Promputtha, I.; Stadler, M. Elucidation of the life cycle of the endophytic genus Muscodor and its transfer into the genus Induratia in Induratiaceae fam. nov., based on a polyphasic taxonomic approach. Fungal Divers. 2020, 101, 177–210. [Google Scholar] [CrossRef]
- Senanayake, I.C.; Maharachchikumbura, S.S.N.; Hyde, K.D.; Bhat, J.D.; Jones, E.B.G.; McKenzie, E.H.C.; Dai, D.Q.; Daranagama, D.A.; Dayarathne, M.C.; Goonasekara, I.D.; et al. Towards unraveling relationships in Xylariomycetidae (Sordariomycetes). Fungal Divers. 2015, 73, 73–144. [Google Scholar] [CrossRef]
- Maharachchikumbura, S.S.N.; Hyde, K.D.; Jones, E.G.; McKenzie, E.H.; Huang, S.K.; Abdel-Wahab, M.A.; Daranagama, D.A.; Dayarathne, M.; D’souza, M.J.; Goonasekara, I.D.; et al. Towards a natural classification and backbone tree for Sordariomycetes. Fungal Divers. 2015, 72, 199–301. [Google Scholar] [CrossRef]
- Hongsanan, S.; Maharachchikumbura, S.S.N.; Hyde, K.D.; Samarakoon, M.C.; Jeewon, R.; Zhao, Q.; Al-Sadi, A.M.; Bahkali, A.H. An updated phylogeny of Sordariomycetes based on phylogenetic and molecular clock evidence. Fungal Divers. 2017, 84, 25–41. [Google Scholar] [CrossRef]
- Wijayawardene, N.N.; Hyde, K.D.; Al-Ani, L.K.T.; Tedersoo, L.; Haelewaters, D.; Rajeshkumar, K.C.; Zhao, R.L.; Aptroot, A.; Leontyev, D.V.; Saxena, R.K.; et al. Outline of Fungi and fungus-like taxa. Mycosphere 2020, 11, 1060–1456. [Google Scholar] [CrossRef]
- Krug, J.C. The genus Cainia and a new family, Cainiaceae. Sydowia 1978, 30, 122–133. [Google Scholar]
- Maharachchikumbura, S.S.N.; Hyde, K.D.; Jones, E.G.; McKenzie, E.H.C.; Bhat, J.D.; Dayarathne, M.C.; Huang, S.K.; Norphanphoun, C.; Senanayake, I.C.; Perera, R.H.; et al. Families of Sordariomycetes. Fungal Divers. 2016, 79, 1–317. [Google Scholar] [CrossRef]
- Mapook, A.; Hyde, K.D.; McKenzie, E.H.; Jones, E.G.; Bhat, D.J.; Jeewon, R.; Stadler, M.; Samarakoon, M.C.; Malaithong, M.; Tanunchai, B.; et al. Taxonomic and phylogenetic contributions to fungi associated with the invasive weed Chromolaena odorata (Siam weed). Fungal Divers. 2020, 101, 1–175. [Google Scholar] [CrossRef]
- Hyde, K.D.; Norphanphoun, C.; Chen, J.; Dissanayake, A.J.; Doilom, M.; Hongsanan, S.; Jayawardena, R.S.; Jeewon, R.; Perera, R.H.; Thongbai, B.; et al. Thailand’s amazing diversity—Up to 96% of fungi in northern Thailand are novel. Fungal Divers. 2018, 93, 215–239. [Google Scholar] [CrossRef]
- Hyde, K.D.; Dong, Y.; Phookamsak, R.; Jeewon, R.; Bhat, D.J.; Jones, E.B.; Liu, N.G.; Abeywickrama, P.D.; Mapook, A.; Wei, D.P.; et al. Fungal diversity notes 1151–1276: Taxonomic and phylogenetic contributions on genera and species of fungal taxa. Fungal Divers. 2020, 100, 5–277. [Google Scholar] [CrossRef]
- Hyde, K.D.; Jeewon, R.; Chen, Y.J.; Bhunjun, C.S.; Calabon, M.S.; Jiang, H.B.; Lin, C.G.; Norphanphoun, C.; Sysouphanthong, P.; Pem, D.; et al. The numbers of fungi: Is the descriptive curve flattening? Fungal Divers. 2020, 103, 219–271. [Google Scholar] [CrossRef]
- Konta, S.; Hyde, K.D.; Eungwanichayapant, P.D.; Doilom, M.; Tennakoon, D.S.; Senwanna, C.; Boonmee, S. Fissuroma (Aigialaceae: Pleosporales) appears to be hyperdiverse on Arecaceae: Evidence from two new species from southern Thailand. Acta Bot. Bras. 2020, 34, 384–393. [Google Scholar] [CrossRef]
- Senanayake, I.C.; Rathnayaka, A.R.; Marasinghe, D.S.; Calabon, M.S.; Gentekaki, E.; Lee, H.B.; Hurdeal, V.G.; Pem, D.; Dissanayake, L.S.; Wijesinghe, S.N.; et al. Morphological approaches in studying fungi: Collection, examination, isolation, sporulation and preservation. MYCOSP 2020, 11, 2678–2754. [Google Scholar] [CrossRef]
- Jayasiri, S.C.; Hyde, K.D.; Ariyawansa, H.A.; Bhat, J.; Buyck, B.; Cai, L.; Dai, Y.C.; Abd-Elsalam, K.A.; Ertz, D.; Hidayat, I.; et al. The Faces of Fungi database: Fungal names linked with morphology, phylogeny and human impacts. Fungal Divers. 2015, 74, 3–18. [Google Scholar] [CrossRef]
- Index Fungorum. 2021. Available online: http://www.indexfungorum.org/names/Names.asp (accessed on 30 April 2020).
- Dissanayake, A.J.; Bhunjun, C.S.; Maharachchikumbura, S.S.N.; Liu, J.K. Applied aspects of methods to infer phylogenetic relationships amongst fungi. Mycosphere 2020, 11, 2652–2676. [Google Scholar] [CrossRef]
- O’Donnell, K.; Cigelnik, E. Two divergent intragenomic rDNA ITS2 types within amonophyletic lineage of the fungus Fusarium are nonorthologous. Mol. Phylogenet. Evol. 1997, 7, 103–116. [Google Scholar] [CrossRef] [PubMed]
- Pintos, Á.; Alvarado, P.; Planas, J.; Jarling, R. Six new species of Arthrinium from Europe and notes about A. caricicola and other species found in Carex spp. hosts. MycoKeys 2019, 49, 15–48. [Google Scholar] [CrossRef]
- Crous, P.W.; Groenewald, J.Z. A phylogenetic re-evaluation of Arthrinium. IMA Fungus 2013, 4, 133–154. [Google Scholar] [CrossRef]
- Wang, M.; Liu, F.; Crous, P.W.; Cai, L. Phylogenetic reassessment of Nigrospora: Ubiquitous endophytes, plant and human pathogens. Pers. Mol. Phylogeny Evol. Fungi 2017, 39, 118–142. [Google Scholar] [CrossRef] [PubMed]
- Crous, P.W.; Schumacher, R.K.; Akulov, A.; Thangavel, R.; Hernández-Restrepo, M.; Carnegie, A.J.; Cheewangkoon, R.; Wingfield, M.J.; Summerell, B.A.; Quaedvlieg, W.; et al. New and interesting fungi. 2. Fungal Syst. Evol. 2019, 3, 57–134. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Bonthond, G.; Groenewald, J.Z.; Cai, L.; Crous, P.W. Sporocadaceae, a family of coelomycetous fungi with appendage-bearing conidia. Stud. Mycol. 2019, 92, 287–415. [Google Scholar] [CrossRef] [PubMed]
- Crous, P.W.; Shivas, R.G.; Quaedvlieg, W.; van der Bank, M.; Zhang, Y.; Summerell, B.A.; Guarro, J.; Wingfield, M.J.; Wood, A.R.; Alfenas, A.C.; et al. Fungal Planet description sheets: 214–280. Pers. Mol. Phylogeny Evol. Fungi 2014, 32, 184–306. [Google Scholar] [CrossRef] [PubMed]
- Jeewon, R.; Liew, E.C.; Hyde, K.D. Molecular systematics of the Amphisphaeriaceae based on cladistic analyses of partial LSU rDNA gene sequences. Mycol. Res. 2003, 107, 1392–1402. [Google Scholar] [CrossRef]
- Liu, J.K.; Hyde, K.D.; Jones, E.G.; Ariyawansa, H.A.; Bhat, D.J.; Boonmee, S.; Maharachchikumbura, S.S.N.; McKenzie, E.H.C.; Phookamsak, R.; Phukhamsakda, C.; et al. Fungal diversity notes 1–110: Taxonomic and phylogenetic contributions to fungal species. Fungal Divers. 2015, 72, 1–197. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Gardiennet, A.; Voglmayr, H. Resolution of morphology-based taxonomic delusions: Acrocordiella, Basiseptospora, Blogiascospora, Clypeosphaeria, Hymenopleella, Lepteutypa, Pseudapiospora, Requienella, Seiridium and Strickeria. Pers. Mol. Phylogeny Evol. Fungi 2016, 37, 82–105. [Google Scholar] [CrossRef]
- Tibpromma, S.; Hyde, K.D.; Jeewon, R.; Maharachchikumbura, S.S.; Liu, J.K.; Bhat, D.J.; Jones, E.G.; McKenzie, E.H.; Camporesi, E.; Bulgakov, T.S.; et al. Fungal diversity notes 491–602: Taxonomic and phylogenetic contributions to fungal taxa. Fungal Divers. 2017, 83, 1–261. [Google Scholar] [CrossRef]
- Lumbsch, H.T.; Schmitt, I.; Lindemuth, R.; Miller, A.; Mangold, A.; Fernandez, F.; Huhndorf, S. Performance of four ribosomal DNA regions to infer higher-level phylogenetic relationships of inoperculate euascomycetes (Leotiomyceta). Mol. Phylogenet. Evol. 2005, 34, 512–524. [Google Scholar] [CrossRef]
- Okada, G.; Iida, T.; Ohkuma, M. The DNA Bank, RIKEN Bio Resource Research Center, Japan, 2017. Available online: https://www.jcm.riken.jp/cgi-bin/jcm/jcm_number?JCM=7946 (accessed on 30 April 2020).
- Bhattacharya, D.; Lutzoni, F.; Reeb, V.; Simon, D.; Nason, J.; Fernandez, F. Widespread occurrence of spliceosomal introns in the rDNA genes of ascomycetes. Mol. Biol. Evol. 2000, 17, 1971–1984. [Google Scholar] [CrossRef]
- Voglmayr, H.; Friebes, G.; Gardiennet, A.; Jaklitsch, W.M. Barrmaelia and Entosordaria in Barrmaeliaceae (fam. nov., Xylariales) and critical notes on Anthostomella-like genera based on multigene phylogenies. Mycol. Prog. 2018, 17, 155–177. [Google Scholar] [CrossRef] [PubMed]
- Asgari, B.; Zare, R. A contribution to the taxonomy of the genus Coniocessia (Xylariales). Mycol. Prog. 2011, 10, 189–206. [Google Scholar] [CrossRef]
- García, D.; Stchigel, A.M.; Cano, J.; Calduch, M.; Hawksworth, D.L.; Guarro, J. Molecular phylogeny of Coniochaetales. Mycol. Res. 2006, 110, 1271–1289. [Google Scholar] [CrossRef] [PubMed]
- Wanasinghe, D.N.; Phukhamsakda, C.; Hyde, K.D.; Jeewon, R.; Lee, H.B.; Jones, E.G.; Tibpromma, S.; Tennakoon, D.S.; Dissanayake, A.J.; Jayasiri, S.C.; et al. Fungal diversity notes 709–839: Taxonomic and phylogenetic contributions to fungal taxa with an emphasis on fungi on Rosaceae. Fungal Divers. 2018, 89, 1–236. [Google Scholar] [CrossRef]
- Senwanna, C.; Phookamsak, R.; Doilom, M.; Hyde, K.D.; Cheewangkoon, R. Novel taxa of Diatrypaceae from Para rubber (Hevea brasiliensis) in northern Thailand; introducing a novel genus Allocryptovalsa. Mycosphere 2017, 8, 1835–1855. [Google Scholar] [CrossRef]
- Konta, S.; Maharachchikumbura, S.S.N.; Senanayake, I.C.; McKenzie, E.H.C.; Stadler, M.; Boonmee, S.; Phookamsak, R.; Jayawardena, R.S.; Senwanna, C.; Hyde, K.D.; et al. A new genus Allodiatrype, five new species and a new host record of diatrypaceous fungi from palms (Arecaceae). Mycosphere 2020, 11, 239–268. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Fournier, J.; Rogers, J.D.; Voglmayr, H. Phylogenetic and taxonomic revision of Lopadostoma. Pers. Mol. Phylogeny Evol. Fungi 2014, 32, 52–82. [Google Scholar] [CrossRef]
- Spatafora, J.W.; Sung, G.H.; Johnson, D.; Hesse, C.; O’Rourke, B.; Serdani, M.; Spotts, R.; Lutzoni, F.; Hofstetter, V.; Miadlikowska, J.; et al. A five-gene phylogeny of Pezizomycotina. Mycologia 2006, 98, 1018–1028. [Google Scholar] [CrossRef] [PubMed]
- Lynch, S.C.; Eskalen, A.; Zambino, P.J.; Mayorquin, J.S.; Wang, D.H. Identification and pathogenicity of Botryosphaeriaceae species associated with coast live oak (Quercus agrifolia) decline in southern California. Mycologia 2013, 105, 125–140. [Google Scholar] [CrossRef]
- Rolshausen, P.E.; Mahoney, N.E.; Molyneux, R.J.; Gubler, W.D. A reassessment of the species concept in Eutypa lata, the causal agent of Eutypa dieback of grapevine. Phytopathology 2006, 96, 369–377. [Google Scholar] [CrossRef] [PubMed]
- Vu, D.; Groenewald, M.; De Vries, M.; Gehrmann, T.; Stielow, B.; Eberhardt, U.; Al-Hatmi, A.; Groenewald, J.Z.; Cardinali, G.; Houbraken, J.; et al. Large-scale generation and analysis of filamentous fungal DNA barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation. Stud. Mycol. 2019, 92, 135–154. [Google Scholar] [CrossRef] [PubMed]
- Acero, F.J.; González, V.; Sánchez-Ballesteros, J.; Rubio, V.; Checa, J.; Bills, G.F.; Salazar, O.; Platas, G.; Peláez, F. Molecular phylogenetic studies on the Diatrypaceae based on rDNA-ITS sequences. Mycologia 2004, 96, 249–259. [Google Scholar] [CrossRef] [PubMed]
- Dayarathne, M.C.; Phookamsak, R.; Hyde, K.D.; Manawasinghe, I.S.; To-Anun, C.; Jones, G.E. Halodiatrype, a novel diatrypaceous genus from mangroves with H. salinicola and H. avicenniae spp. nov. Mycosphere 2016, 7, 612–627. [Google Scholar] [CrossRef]
- Klaysuban, A.; Sakayaroj, J.; Jones, E.G. An additional marine fungal lineage in the Diatrypaceae, Xylariales: Pedumispora rhizophorae. Bot. Mar. 2014, 57, 413–420. [Google Scholar] [CrossRef]
- Hyde, K.D.; Tennakoon, D.S.; Jeewon, R.; Bhat, D.J.; Maharachchikumbura, S.S.N.; Rossi, W.; Leonardi, M.; Lee, H.B.; Mun, H.Y.; Houbraken, J.; et al. Fungal diversity notes 1036–1150: Taxonomic and phylogenetic contributions on genera and species of fungal taxa. Fungal Divers. 2019, 96, 1–242. [Google Scholar] [CrossRef]
- Hyde, K.D.; Norphanphoun, C.; Abreu, V.P.; Bazzicalupo, A.; Chethana, K.T.; Clericuzio, M.; Dayarathne, M.C.; Dissanayake, A.J.; Ekanayaka, A.H.; He, M.Q.; et al. Fungal diversity notes 603–708: Taxonomic and phylogenetic notes on genera and species. Fungal Divers. 2017, 87, 1–235. [Google Scholar] [CrossRef]
- Wendt, L.; Sir, E.B.; Kuhnert, E.; Heitkämper, S.; Lambert, C.; Hladki, A.I.; Romero, A.I.; Luangsa-Ard, J.J.; Srikitikulchai, P.; Peršoh, D.; et al. Resurrection and emendation of the Hypoxylaceae, recognised from a multigene phylogeny of the Xylariales. Mycol. Prog. 2018, 17, 115–154. [Google Scholar] [CrossRef]
- Zhang, N.; Castlebury, L.A.; Miller, A.N.; Huhndorf, S.M.; Schoch, C.L.; Seifert, K.A.; Rossman, A.Y.; Rogers, J.D.; Kohlmeyer, J.; Volkmann-Kohlmeyer, B.; et al. An overview of the systematics of the Sordariomycetes based on a four-gene phylogeny. Mycologia 2006, 98, 1076–1087. [Google Scholar] [CrossRef]
- Stadler, M.; Læssøe, T.; Fournier, J.; Decock, C.; Schmieschek, B.; Tichy, H.V.; Peršoh, D. A polyphasic taxonomy of Daldinia (Xylariaceae). Stud. Mycol. 2014, 77, 1–143. [Google Scholar] [CrossRef]
- Koukol, O.; Kelnarová, I.; Černý, K. Recent observations of sooty bark disease of sycamore maple in Prague (Czech Republic) and the phylogenetic placement of Cryptostroma corticale. For. Pathol. 2015, 45, 21–27. [Google Scholar] [CrossRef]
- Pažoutová, S.; Šrůtka, P.; Holuša, J.; Chudíčková, M.; Kolařík, M. The phylogenetic position of Obolarina dryophila (Xylariales). Mycol. Prog. 2010, 9, 501–507. [Google Scholar] [CrossRef]
- Kuhnert, E.; Sir, E.B.; Lambert, C.; Hyde, K.D.; Hladki, A.I.; Romero, A.I.; Rohde, M.; Stadler, M. Phylogenetic and chemotaxonomic resolution of the genus Annulohypoxylon (Xylariaceae) including four new species. Fungal Divers. 2017, 85, 1–43. [Google Scholar] [CrossRef]
- Triebel, D.; Peršoh, D.; Wollweber, H.; Stadler, M. Phylogenetic relationships among Daldinia, Entonaema, and Hypoxylon as inferred from ITS nrDNA analyses of Xylariales. Nova Hedwig 2005, 80, 25–43. [Google Scholar] [CrossRef]
- Kuhnert, E.; Fournier, J.; Peršoh, D.; Luangsa-Ard, J.J.D.; Stadler, M. New Hypoxylon species from Martinique and new evidence on the molecular phylogeny of Hypoxylon based on ITS rDNA and β-tubulin data. Fungal Divers. 2014, 64, 181–203. [Google Scholar] [CrossRef]
- Læssøe, T.; Srikitikulchai, P.; Jennifer, J.; Luangsa-Ard, D.; Stadler, M. Theissenia reconsidered, including molecular phylogeny of the type species T. pyrenocrata and a new genus Durotheca (Xylariaceae, Ascomycota). IMA Fungus 2013, 4, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Daranagama, D.A.; Liu, X.; Chamyuang, S.; Stadler, M.; Bahkali, A.; Hyde, K.D. Rhopalostroma brevistipitatum sp. nov. from Thailand with an extended generic description for Rhopalostroma. Phytotaxa 2015, 227, 229–242. [Google Scholar] [CrossRef][Green Version]
- Stadler, M.; Kuhnert, E.; Peršoh, D.; Fournier, J. The Xylariaceae as model example for a unified nomenclature following the “One Fungus-One Name” (1F1N) concept. Mycology 2013, 4, 5–21. [Google Scholar] [CrossRef]
- Bitzer, J.; Læssøe, T.; Fournier, J.; Kummer, V.; Decock, C.; Tichy, H.V.; Piepenbring, M.; Peršoh, D.; Stadler, M. Affinities of Phylacia and the daldinoid Xylariaceae, inferred from chemotypes of cultures and ribosomal DNA sequences. Mycol. Res. 2008, 112, 251–270. [Google Scholar] [CrossRef]
- Miller, A.N.; Vasilyeva, L.N.; Rogers, J.D. Chlorostroma subcubisporum gen. et sp. nov. and notes on the systematic position of Thuemenella cubispora. Sydowia 2007, 59, 138–147. [Google Scholar]
- Stadler, M.; Flessa, F.; Rambold, G.; Peršoh, D.; Fournier, J.; Læssøe, T.; Chlebicki, A.; Lechat, C. Chemotaxonomic and phylogenetic studies of Thamnomyces (Xylariaceae). Mycoscience 2010, 51, 189–207. [Google Scholar] [CrossRef]
- Voglmayr, H.; Aguirre-Hudson, M.B.; Wagner, H.G.; Tello, S.; Jaklitsch, W.M. Lichens or endophytes? The enigmatic genus Leptosillia in the Leptosilliaceae fam. nov. (Xylariales), and Furfurella gen. nov. (Delonicicolaceae). Pers. Mol. Phylogeny Evol. Fungi 2019, 42, 228–260. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.L.; Wang, G.P.; Mao, L.J.; Komon-Zelazowska, M.; Yuan, Z.L.; Lin, F.C.; Druzhinina, I.S.; Kubicek, C.P. Muscodor fengyangensis sp. nov. from southeast China: Morphology, physiology and production of volatile compounds. Fungal Biol. Rev. 2010, 114, 797–808. [Google Scholar] [CrossRef]
- Becerra-Hernández, C.I.; González, D.; De Luna, E.; Mena-Portales, J. First report of pleoanamorphy in Gyrothrix verticiclada with an Idriella-like synanamorph. Cryptogam. Mycol. 2016, 37, 241–252. [Google Scholar] [CrossRef]
- Hernández-Restrepo, M.; Groenewald, J.Z.; Crous, P.W. Taxonomic and phylogenetic re-evaluation of Microdochium, Monographella and Idriella. Pers. Mol. Phylogeny Evol. Fungi 2016, 36, 57–82. [Google Scholar] [CrossRef]
- Bundhun, D.; Jeewon, R.; Dayarathne, M.C.; Bulgakov, T.S.; Khramtsov, A.K.; Aluthmuhandiram, J.V.; Pem, D.; To-Anun, C.; Hyde, K.D. A morpho-molecular re-appraisal of Polystigma fulvum and P. rubrum (Polystigma, Polystigmataceae). Phytotaxa 2019, 422, 209–224. [Google Scholar] [CrossRef]
- Maharachchikumbura, S.S.N.; Hyde, K.D.; Perera, R.H.; Al-Sadi, A.M. Acrocordiella omanensis sp. nov. (Requienellaceae, Xylariales) from the Sultanate of Oman. Phytotaxa 2018, 338, 294–300. [Google Scholar] [CrossRef]
- Park, J.H.; Choi, G.J.; Lee, H.B.; Kim, K.M.; Jung, H.S.; Lee, S.W.; Jang, K.S.; Cho, K.Y.; Kim, J.C. Griseofulvin from Xylaria sp. strain F0010, an endophytic fungus of Abies holophylla and its antifungal activity against plant pathogenic fungi. J. Microbiol. Biotechnol. 2005, 15, 112–117. [Google Scholar]
- Hsieh, H.M.; Lin, C.R.; Fang, M.J.; Rogers, J.D.; Fournier, J.; Lechat, C.; Ju, Y.M. Phylogenetic status of Xylaria subgenus Pseudoxylaria among taxa of the subfamily Xylarioideae (Xylariaceae) and phylogeny of the taxa involved in the subfamily. Mol. Phylogenet. Evol. 2010, 54, 957–969. [Google Scholar] [CrossRef]
- Dai, D.Q.; Phookamsak, R.; Wijayawardene, N.N.; Li, W.J.; Bhat, D.J.; Xu, J.C.; Taylor, J.E.; Hyde, K.D.; Chukeatirote, E. Bambusicolous fungi. Fungal Divers. 2017, 82, 1–105. [Google Scholar] [CrossRef]
- Marincowitz, S.; Crous, P.W.; Groenewald, J.Z.; Wingfield, M.J. Microfungi Occurring on Proteaceae in the Fynbos; CBS Biodiversity Series; CBS-KNAW Fungal Biodiversity Centre: Utrecht, The Netherlands, 2008. [Google Scholar]
- Jaklitsch, W.M.; Voglmayr, H. Phylogenetic relationships of five genera of Xylariales and Rosasphaeria gen. nov. (Hypocreales). Fungal Divers. 2012, 52, 75–98. [Google Scholar] [CrossRef]
- Johnston, P.R.; Rogers, J.D.; Park, D.; Martin, N.A. Entalbostroma erumpens gen. et sp. nov. (Xylariaceae) from Phormium in New Zealand. Mycotaxon 2016, 131, 765–771. [Google Scholar] [CrossRef]
- Soca-Chafre, G.; Rivera-Orduña, F.N.; Hidalgo-Lara, M.E.; Hernandez-Rodriguez, C.; Marsch, R.; Flores-Cotera, L.B. Molecular phylogeny and paclitaxel screening of fungal endophytes from Taxus globosa. Fungal Biol. Rev. 2011, 115, 143–156. [Google Scholar] [CrossRef]
- Jayanetti, D.R.; Yue, Q.; Bills, G.F.; Gloer, J.B. Hypocoprins A–C: New sesquiterpenoids from the coprophilous fungus Hypocopra rostrata. J. Nat. Prod. 2015, 78, 396–401. [Google Scholar] [CrossRef]
- Ju, Y.M.; Hsieh, H.M.; Ho, M.C.; Szu, D.H.; Fang, M.J. Theissenia rogersii sp. nov. and phylogenetic position of Theissenia. Mycologia 2007, 99, 612–621. [Google Scholar] [CrossRef] [PubMed]
- Voglmayr, H.; Beenken, L. Linosporopsis, a new leaf-inhabiting scolecosporous genus in Xylariaceae. Mycol. Prog. 2020, 19, 205–222. [Google Scholar] [CrossRef]
- U’Ren, J.M.; Miadlikowska, J.; Zimmerman, N.B.; Lutzoni, F.; Stajich, J.E.; Arnold, A.E. Contributions of North American endophytes to the phylogeny, ecology, and taxonomy of Xylariaceae (Sordariomycetes, Ascomycota). Mol. Phylogenet. Evol. 2016, 98, 210–232. [Google Scholar] [CrossRef]
- Konta, S.; Hyde, K.D.; Phookamsak, R.; Xu, J.C.; Maharachchikumbura, S.S.N.; Daranagama, D.A.; McKenzie, E.H.C.; Boonmee, S.; Tibpromma, S.; Eungwanichayapant, P.D.; et al. Polyphyletic genera in Xylariaceae (Xylariales): Neoxylaria gen. nov. and Stilbohypoxylon. Mycosphere 2020, 11, 2629–2651. [Google Scholar] [CrossRef]
- Tang, A.M.C.; Jeewon, R.; Hyde, K.D. A re-evaluation of the evolutionary relationships within the Xylariaceae based on ribosomal and protein-coding gene sequences. Fungal Divers. 2009, 34, 127–155. [Google Scholar]
- Fournier, J.; Stadler, M.; Hyde, K.D.; Duong, M.L. The new genus Rostrohypoxylon and two new Annulohypoxylon species from Northern Thailand. Fungal Divers. 2010, 40, 23–36. [Google Scholar] [CrossRef]
- Dai, D.Q.; Bahkali, A.H.; Li, Q.R.; Bhat, D.J.; Wijayawardene, N.N.; Li, W.J.; Chukeatirote, E.; Zhao, R.L.; Xu, J.C.; Hyde, K.D. Vamsapriya (Xylariaceae) re-described, with two new species and molecular sequence data. Cryptogam. Mycol. 2014, 35, 339–357. [Google Scholar] [CrossRef]
- Jiang, H.B.; Phookamsak, R.; Bhat, D.J.; Khan, S.; Bahkali, A.; Elgorban, A.; Hyde, K.D. Vamsapriya yunnana, a new species of Vamsapriya (Xylariaceae, Xylariales) associated with bamboo from Yunnan, China. Phytotaxa 2018, 356, 61–70. [Google Scholar] [CrossRef]
- Nonaka, K.; Ishii, T.; Shiomi, K.; Ōmura, S.; Masuma, R. Virgaria boninensis, a new hyphomycete (Xylariaceae) from soils in the Bonin Islands, Japan. Mycoscience 2013, 54, 394–399. [Google Scholar] [CrossRef]
- Sir, E.B.; Kuhnert, E.; Surup, F.; Hyde, K.D.; Stadler, M. Discovery of new mitorubrin derivatives from Hypoxylon fulvo-sulphureum sp. nov. (Ascomycota, Xylariales). Mycol. Prog. 2015, 14, 28. [Google Scholar] [CrossRef]
- Cheng, X.; Li, W.; Cai, L. Molecular phylogeny of Ascotricha, including two new marine algae-associated species. Mycologia 2015, 107, 490–504. [Google Scholar] [CrossRef]
- Daranagama, D.A.; Hyde, K.D.; Sir, E.B.; Thambugala, K.M.; Tian, Q.; Samarakoon, M.C.; McKenzie, E.H.; Jayasiri, S.C.; Tibpromma, S.; Bhat, J.D. Towards a natural classification and backbone tree for Graphostromataceae, Hypoxylaceae, Lopadostomataceae and Xylariaceae. Fungal Divers. 2018, 88, 1–165. [Google Scholar] [CrossRef]
- Crous, P.W.; Schumacher, R.K.; Wingfield, M.J.; Lombard, L.; Giraldo, A.; Christensen, M.; Gardiennet, A.; Nakashima, C.; Pereira, O.L.; Smith, A.J. Fungal systematics and evolution: FUSE 1. Sydowia 2015, 67, 81–118. [Google Scholar] [CrossRef]
- Li, D.W.; Schultes, N.P.; Chen, J.Y.; Wang, Y.X.; Castañeda-Ruiz, R.F. Circinotrichum sinense, a new asexual fungus from Hubei, China. Botany 2017, 95, 1099–1108. [Google Scholar] [CrossRef]
- Crous, P.W.; Wingfield, M.J.; Lombard, L.; Roets, F.; Swart, W.J.; Alvarado, P.; Carnegie, A.J.; Moreno, G.; Luangsaard, J.; Thangavel, R.; et al. Fungal Planet description sheets: 951–1041. Pers. Mol. Phylogeny Evol. Fungi 2019, 43, 223–425. [Google Scholar] [CrossRef]
- Li, J.F.; Phookamsak, R.; Jeewon, R.; Tibpromma, S.; Maharachchikumbura, S.S.N.; Bhat, D.J.; Chukeatirote, E.; Lumyong, S.; Hyde, K.D.; McKenzie, E.H.C. Establishment of Zygosporiaceae fam. nov. (Xylariales, Sordariomycetes) based on rDNA sequence data to accommodate Zygosporium. MYCOAP 2017, 8, 1855–1868. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, K. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Glez-Peña, D.; Gómez-Blanco, D.; Reboiro-Jato, M.; Fdez-Riverola, F.; Posada, D. ALTER: Program-oriented conversion of DNA and protein alignments. Nucleic Acid. Res. 2010, 38, 14–18. [Google Scholar] [CrossRef] [PubMed]
- Nylander, J.A.A. MrModeltest, Version 2; Program Distributed by the Author; Evolutionary Biology Centre, Uppsala University: Uppsala, Sweden, 2004; Available online: https://github.com/Nylander (accessed on 30 April 2020).
- Rambaut, A. FigTree Version 1.4.0. 2012. Available online: http://tree.bio.ed.ac.uk/software/figtree/ (accessed on 30 April 2020).
- Daranagama, D.A.; Camporesi, E.; Tian, Q.; Liu, X.; Chamyuang, S.; Stadler, M.; Hyde, K.D. Anthostomella is polyphyletic comprising several genera in Xylariaceae. Fungal Divers. 2015, 73, 203–238. [Google Scholar] [CrossRef]
- Petch, T. The genus Endocalyx, Berkeley and Broome. Ann. Bot. 1908, 22, 389–400. [Google Scholar] [CrossRef]
- Okada, G.; Tubaki, K. A new species and a new variety of Endocalyx (Deuteromycotina) from Japan. Mycologia 1984, 76, 300–313. [Google Scholar] [CrossRef]
- Vitoria, N.S.; Cavalcanti, M.A.Q.; Luz, E.D.M.N.; Bezerra, J.L. Endocalyx melanoxanthus var. melanoxanthus (Ascomycota): New to Brazil and three new hosts. Mycotaxon 2011, 117, 109–113. [Google Scholar] [CrossRef]
- Patel, U.S.; Pandey, A.K.; Rajak, R.C. Two new hyphomycetes. J. Mycol. Plant Pathol. 2002, 32, 70–71. [Google Scholar]
- Kobayashi, T. Index of Fungi Inhabiting Woody Plants in Japan: Host, Distribution and Literature; Zenkoku-Noson-Kyoiku Kyokai Publishing Co., Ltd.: Tokyo, Japan, 2007; pp. 1–1227. [Google Scholar]
- Lu, B.; Hyde, K.D.; Ho, W.H.; Tsui, K.M.; Taylor, J.E.; Wong, K.M.; Yanna; Zhou, D. Checklist of Hong Kong Fungi; Fungal Diversity Press: Hong Kong, China, 2000; pp. 1–207. [Google Scholar]
- Portales, J.M.; Sierra, A.M. Nuevas especies de Endocalyx y Stachylidium (Hyphomycetes, Deuteromycotina) de Cuba. Rev. Jard. Bot. Nac. 1984, 5, 53–60. [Google Scholar]
- Kapoor, J.N.; Munjal, R.L. Indian species of Stilbaceae. Indian Phytopathol. 1966, 19, 348–350. [Google Scholar]
- Heredia, G.; Arias, R.M.; Reyes, M. Contribucion al conocimiento de los hongos Hyphomycetes de Mexico. Acta Bot. Mex. 2000, 51, 39–51. [Google Scholar] [CrossRef]
- Taylor, J.E.; Hyde, K.D. Microfungi of Tropical and Temperate Palms; Fungal Diversity Press: Hong Kong, China, 2003; pp. 1–459. [Google Scholar]
- Hughes, S.J. Fungi from the gold coast. I. Mycol. Pap. 1952, 48, 1–91. [Google Scholar]
- Anonymous. Index of Plant Diseases in the United States. USDA Agric. Handb. 1960, 165, 1–531. [Google Scholar]
- Raabe, R.D.; Conners, I.L.; Martinez, A.P. Checklist of Plant Diseases in Hawaii; Information Text Series No. 22; College of Tropical Agriculture and Human Resources, University of Hawaii: Honolulu, HI, USA, 1981; pp. 1–313. [Google Scholar]
- Turner, P.D. Microorganisms associated with oil palm (Elaeis guineensis Jacq.). Phytopathol. Pap. 1971, 14, 1–58. [Google Scholar]
- Shaw, D.E. Microorganisms in Papua New Guinea. Dept. Prim. Ind. Res. Bull. 1984, 33, 1–344. [Google Scholar]
- Urtiaga, R. [Host index of plant diseases and disorders from Venezuela—Addendum]. 2004; pp. 1–268. Available online: https://nt.ars-grin.gov/fungaldatabases/fungushost/new_rptOneLit.cfm?fungRec=39196&thisError= (accessed on 30 April 2020).
- Urtiaga, R. Indice de enfermedades en plantas de Venezuela y Cuba, Second Edition. 2004; pp. 1–301. Available online: https://nt.ars-grin.gov/fungaldatabases/fungushost/new_rptOneLit.cfm?fungRec=39195&thisError= (accessed on 30 April 2020).
- Thaung, M.M. A list of hypomycetes (and agonomycetes) in Burma. Australas. Mycol. 2008, 27, 149–172. [Google Scholar]
- Pinruan, U.; Hyde, K.D.; Lumyong, S.; McKenzie, E.H.C.; Jones, E.B.G. Occurrence of fungi on tissues of the peat swamp palm Licuala longicalycata. Fungal Divers. 2007, 25, 157–173. [Google Scholar]
- Matsushima, T. Matsushima Mycological Memoirs No. 1. Saprophytic Microfungi from Taiwán, Part1; Matsushima Fungus collection: Kobe, Japan, 1980; pp. 1–82. [Google Scholar]
- Ahmad, S. Fungi of West Pakistan. Biol. Soc. Pak. Monogr. 1969, 5, 1–110. [Google Scholar]
- Zhuang, W.Y. Higher Fungi of Tropical China; Mycotaxon Limited: Ithaca, NY, USA, 2001; pp. 1–485. [Google Scholar]
- Hughes, S.J. New Zealand Fungi. 25. Miscellaneous species. N. Z. J. Bot. 1978, 16, 311–370. [Google Scholar] [CrossRef]
- Delgado, G. South Florida microfungi: New records of saprophytic hyphomycetes on plant debris. FLA Sci. 2008, 71, 76–89. [Google Scholar]
- Arnold, G.R.W. Lista de Hongos Fitopatogenos de Cuba; Ministerio de Cultura Editorial Cientifico-Tecnica: Havana, Cuba, 1986; pp. 1–207. [Google Scholar]
- Sierra, A.M. Hifomicetes Demaciaceos de Sierra del Rosario, Cuba; Editorial Academica: Havana, Cuba, 1984; pp. 1–81. [Google Scholar]
- Urtiaga, R. Indice de Enfermedades en Plantas de Venezuela y Cuba; Impresos en Impresos Nuevo Siglo S.R.L.: Barquisimeto, Venezuela, 1986; pp. 1–202. [Google Scholar]
- Delgado-Rodriguez, G.; Mena-Portales, J.; Calduch, M.; Decock, C. Hyphomycetes (hongos mitosporicos) del area protegida mil cumbres, Cuba Occidental. Cryptog. Mycol. 2002, 23, 277–293. [Google Scholar]
- Petrak, F. Ein Beitrag zur Pilzflora Floridas. Sydowia 1953, 7, 103–116. [Google Scholar]
- Sutton, B.C. New and interesting Hyphomycetes from Tampa, Florida. Mycologia 1978, 70, 784–801. [Google Scholar] [CrossRef]
- Capdeet, M.; Romero, A.I. Fungi from palms in Argentina. 1. Mycotaxon 2010, 112, 339–355. [Google Scholar] [CrossRef]
- Vitoria, N.S.; Fortes, N.G.S.; dos Santos, M.A.L.; Barbosa, R.L. Mycota (Ascomycota) of Syagrus coronata (Mart.) Becc., Raso da Catarina Ecological Station, Brazil: New records. Acta Bras. 2020, 4, 110–120. [Google Scholar] [CrossRef]
- Hughes, S.J. Fungi from the gold coast. II. Mycol. Pap. 1953, 50, 1–104. [Google Scholar]
- Ellis, M.B. Dematiaceous Hyphomycetes; Commonwealth Mycological Institute: London, UK, 1971; pp. 1–608. [Google Scholar]
- Species Fungorum. 2021. Available online: http://www.speciesfungorum.org/Names/Names.asp (accessed on 30 April 2020).
- Samarakoon, B.C.; Wanasinghe, D.N.; Samarakoon, M.C.; Phookamsak, R.; McKenzie, E.H.; Chomnunti, P.; Hyde, K.D.; Lumyong, S.; Karunarathna, S.C. Multi-gene phylogenetic evidence suggests Dictyoarthrinium belongs in Didymosphaeriaceae (Pleosporales, Dothideomycetes) and Dictyoarthrinium musae sp. nov. on Musa from Thailand. MycoKeys 2020, 71, 101–118. [Google Scholar] [CrossRef]

| Order | Family | Species | Strain No. | GenBank Accession No. | References | |||
|---|---|---|---|---|---|---|---|---|
| ITS | LSU | rpb2 | tub2 | |||||
| Amphisphaeriales | Apiosporaceae | Arthrinium balearicum | AP24118 | MK014869 | MK014836 | - | MK017946 | [21] |
| Amphisphaeriales | Apiosporaceae | Arthrinium caricicola | CBS 145127 | MK014871 | MK014838 | - | MK017948 | [21] |
| Amphisphaeriales | Apiosporaceae | Arthrinium hydei | CBS 114990 | KF144890 | KF144936 | - | KF144982 | [22] |
| Amphisphaeriales | Apiosporaceae | Arthrinium phragmitis | CBS 135458 | KF144909 | KF144956 | - | KF145001 | [22] |
| Amphisphaeriales | Apiosporaceae | Arthrinium pseudospegazzinii | CBS 102052 | KF144911 | KF144958 | - | KF145002 | [22] |
| Amphisphaeriales | Apiosporaceae | Nigrospora aurantiaca | CGMCC 3.18130 | NR_153477 | NG_069394 | - | KY019465 | [23] |
| Amphisphaeriales | Apiosporaceae | Nigrospora brasiliensis | CMM 1214 | KY569629 | - | - | MK720816 | [24] |
| Amphisphaeriales | Apiosporaceae | Nigrospora zimmermanii | CBS 290.62 | KY385309 | - | KY806276 | KY385317 | [23] |
| Amphisphaeriales | Beltraniaceae | Beltrania rhombica | CBS 123.58 = IMI 072432 | MH553990 | MH554209 | MH554899 | MH704631 | [25] |
| Amphisphaeriales | Beltraniaceae | Beltraniella endiandrae | CBS 137976 | KJ869128 | KJ869185 | - | - | [26] |
| Amphisphaeriales | Beltraniaceae | Beltraniopsis neolitseae | CBS 137974 | KJ869126 | KJ869183 | - | - | [26] |
| Amphisphaeriales | Beltraniaceae | Arecophila bambusae | HKUCC 4794 | - | AF452038 | - | - | [27] |
| Xylariales | Cainiaceae | Alishanica miscanthii | FU31025 | MK503821 | MK503827 | - | - | [3] |
| Xylariales | Cainiaceae | Amphibambusa bambusicola | MFLUCC 11-0617 | KP744433 | KP744474 | - | - | [28] |
| Xylariales | Cainiaceae | Atrotorquata lineata | HKUCC 3263 | AF009807 | - | - | - | Unpublished |
| Xylariales | Cainiaceae | Cainia anthoxanthis | MFLUCC 15-0539 | KR092787 | KR092777 | - | - | [5] |
| Xylariales | Cainiaceae | Cainia desmazieri | CAI | KT949896 | KT949896 | - | - | [29] |
| Xylariales | Cainiaceae | Cainia globosa | MFLUCC 13-0663 | KX822127 | KX822123 | - | - | [30] |
| Xylariales | Cainiaceae | Cainia graminis | CBS 136.62 | KR092793 | AF431949 | - | - | [5,31] |
| Xylariales | Cainiaceae | Longiappendispora chromolaenae | MFLUCC 17-1485 | MT214370 | MT214464 | - | - | [11] |
| Xylariales | Cainiaceae | Endocalyx cinctus | JCM 7946 | LC228648 | LC228704 | - | - | [32] |
| Xylariales | Cainiaceae | Endocalyx metroxyli | MFLUCC 15-0723A | MT929162 | MT929313 | - | - | This study |
| Xylariales | Cainiaceae | Endocalyx metroxyli | MFLUCC 15-0723B | MT929163 | MT929314 | - | MT928155 | This study |
| Xylariales | Cainiaceae | Endocalyx metroxyli | MFLUCC 15-0723C | - | MT929315 | - | - | This study |
| Xylariales | Cainiaceae | Seynesia erumpens | SMH 1291 | - | AF279410 | - | - | [33] |
| Xylariales | Clypeosphaeriaceae | Clypeosphaeria mamillana | CBS 140735 | KT949897 | KT949897 | MF489001 | MH704637 | [29,34] |
| Xylariales | Coniocessiaceae | Coniocessia anandra | Co108 | GU553338 | GU553349 | - | - | [35] |
| Xylariales | Coniocessiaceae | Coniocessia cruciformis | Co116 | GU553336 | GU553347 | - | - | [35] |
| Xylariales | Coniocessiaceae | Coniocessia maxima | Co117 | GU553332 | GU553344 | - | - | [35] |
| Xylariales | Coniocessiaceae | Coniocessia minima | Co111 | GU553334 | GU553345 | - | - | [35] |
| Xylariales | Coniocessiaceae | Coniocessia nodulisporioides | CBS 281.77T | - | AJ875224 | - | - | [36] |
| Xylariales | Coniocessiaceae | Paraxylaria rosacearum | TASM 6132 | MG828941 | MG829050 | - | - | [37] |
| Xylariales | Diatrypaceae | Allocryptovalsa polyspora | MFLUCC 17-0364 | MF959500 | MF959503 | - | MG334556 | [38] |
| Xylariales | Diatrypaceae | Allodiatrype arengae | MFLUCC 15-0713 | MN308411 | MN308402 | MN542886 | MN340297 | [39] |
| Xylariales | Diatrypaceae | Cryptovalsa rabenhorstii | CreI = CBS 125574 | KC774567 | KC774567 | - | - | [40] |
| Xylariales | Diatrypaceae | Diatrype disciformis | CBS 197.49 | - | DQ470964 | DQ470915 | - | [41] |
| Xylariales | Diatrypaceae | Diatrypella verruciformis | UCROK1467 | JX144793 | - | - | JX174093 | [42] |
| Xylariales | Diatrypaceae | Eutypa lata | CBS 208.87 | DQ006927 | MH873755 | - | DQ006969 | [43,44] |
| Xylariales | Diatrypaceae | Eutypella caricae | EL5C | AJ302460 | - | - | - | [45] |
| Xylariales | Diatrypaceae | Halodiatrype salinicola | MFLUCC 15-1277 | KX573915 | - | - | KX573932 | [46] |
| Xylariales | Diatrypaceae | Monosporascus cannonballus | CMM3646 | JX971617 | - | - | - | Unpublished |
| Xylariales | Diatrypaceae | Neoeutypella baoshanensis | EL51C, CBS 274.87 | AJ302460 | - | - | - | [45] |
| Xylariales | Diatrypaceae | Pedumispora rhizophorae | BCC44877 | KJ888853 | KJ888850 | - | - | [47] |
| Xylariales | Diatrypaceae | Peroneutypa longiasca | MFLUCC 17-0371 | MF959502 | MF959505 | - | MG334558 | [38] |
| Xylariales | Fasciatisporaceae | Fasciatispora arengae | MFLUCC 15-0326a | MK120275 | MK120300 | MK890794 | MK890793 | [48] |
| Xylariales | Fasciatisporaceae | Fasciatispora calami | MFLUCC 15-0294 | - | MF459055 | - | MF459056 | [49] |
| Xylariales | Fasciatisporaceae | Fasciatispora cocoes | MFLUCC 18-1445 | MN482680 | MN482675 | MN481517 | MN505154 | [13] |
| Xylariales | Fasciatisporaceae | Fasciatispora nypae | MFLUCC 11-0382 | - | KP744484 | - | - | [28] |
| Xylariales | Fasciatisporaceae | Fasciatispora petrakii | - | AY083828 | - | - | Unpublished | |
| Xylariales | Graphostromataceae | Biscogniauxia nummularia | MUCL 51395 | KY610382 | KY610427 | KY624236 | KX271241 | [50] |
| Xylariales | Graphostromataceae | Camillea obularia | ATCC 28093 | KY610384 | KY610429 | KY624238 | KX271243 | [50] |
| Xylariales | Graphostromataceae | Graphostroma platystomum | CBS 270.87 | JX658535 | DQ836906 | KY624296 | HG934108 | [50,51,52,53] |
| Xylariales | Graphostromataceae | Obolarina dryophila | MUCL 49882 | GQ428316 | GQ428316 | KY624284 | GQ428322 | [50,54] |
| Xylariales | Hansfordiaceae | Hansfordia pulvinate | CBS 194.56 | MK442585 | MH869122 | KU684307 | - | [24] |
| Xylariales | Hansfordiaceae | Hansfordia pulvinate | CBS 144422 | MK442587 | MK442527 | - | - | [24] |
| Xylariales | Hypoxylaceae | Annulohypoxylon truncatum | CBS 140778 | KY610419 | KY610419 | KY624277 | KX376352 | [50,55] |
| Xylariales | Hypoxylaceae | Anthocanalis sparti | MFLUCC 14-0010 | KP297394 | KP340536 | KP340522 | KP406605 | [54] |
| Xylariales | Hypoxylaceae | Anthostoma decipiens | CD = CBS 133221 | KC774565 | KC774565 | - | - | [40] |
| Xylariales | Hypoxylaceae | Daldinia concentrica | CBS 113277 | AY616683 | KY610434 | KY624243 | KC977274 | [50,56,57] |
| Xylariales | Hypoxylaceae | Durotheca depressa | BCC28073 | - | - | - | GQ160492 | [58] |
| Xylariales | Hypoxylaceae | Entonaema liquescens | ATCC 46302 | KY610389 | KY610443 | KY624253 | KX271248 | [50] |
| Xylariales | Hypoxylaceae | Hypomontagnella monticulosa | MUCL 54604 | KY610404 | KY610487 | KY624305 | KX271273 | [50] |
| Xylariales | Hypoxylaceae | Hypoxylon fragiforme | MUCL 51264 | KC477229 | KM186295 | KM186296 | KX271282 | [50,59,60] |
| Xylariales | Hypoxylaceae | Jackrogersella multiformis | CBS 119016 | KC477234 | KY610473 | KY624290 | KX271262 | [50,55,57] |
| Xylariales | Hypoxylaceae | Pyrenomyxa morganii | CBS 116990T | AM749920 | - | - | - | [61] |
| Xylariales | Hypoxylaceae | Pyrenomyxa picea | ILLS 58257 | - | EF562506 | - | - | [62] |
| Xylariales | Hypoxylaceae | Pyrenopolyporus hunteri | MUCL 52673 | KY610421 | KY610472 | KY624309 | KU159530 | [50,55] |
| Xylariales | Hypoxylaceae | Rhopalostroma indicum | CBS 113035 | MH862909 | MH874483 | - | - | [44] |
| Xylariales | Hypoxylaceae | Thamnomyces dendroidea | CBS 123578 | FN428831 | KY610467 | KY624232 | KY624313 | [50,63] |
| Xylariales | Hypoxylaceae | Thuemenella cubispora | CBS 119807 | JX658531 | EF562508 | - | - | [62] |
| Xylariales | Hypoxylaceae | Phylacia sagrana | CBS 119992 | AM749919 | - | - | - | [61] |
| Xylariales | Hypoxylaceae | Pyrenopolyporus symphyon | TBRC:8873 | MH938529 | MH938538 | MK165428 | MK165419 | [64] |
| Xylariales | Induratiaceae | Emarcea castanopsidicola | CBS 117105 | MK762710 | MK762717 | MK791285 | MK776962 | [64] |
| Xylariales | Induratiaceae | Emarcea eucalyptigena | CBS 139908 | MK762711 | MK762718 | MK791286 | MK776963 | [64] |
| Xylariales | Induratiaceae | Induratia fengyangensis | CGMCC 2862 | HM034856 | HM034859 | HM034849 | HM034843 | [65] |
| Xylariales | Induratiaceae | Induratia thailandica | MFLUCC 17-2669 | MK762707 | MK762714 | MK791283 | MK776960 | [64] |
| Xylariales | Lopadostomataceae | Creosphaeria sassafras | STMA 14087 | KY610411 | KY610468 | KY624265 | KX271258 | [50] |
| Xylariales | Lopadostomataceae | Lopadostoma turgidum | CBS 133207 | KC774618 | KC774618 | KC774563 | MF489024 | [29,40] |
| Xylariales | Microdochiaceae | Idriella lunata | MUCL 4103 | KC775734 | KC775709 | - | - | [66] |
| Xylariales | Microdochiaceae | Idriella lunata | CBS 204.56 | KP859044 | KP858981 | - | - | [67] |
| Xylariales | Microdochiaceae | Microdochium phragmitis | CBS 423.78 | KP859012 | KP858948 | KP859121 | KP859076 | [67] |
| Xylariales | Polystigmataceae | Polystigma fulvum | MFLU 18-0261 | MK429738 | MK429727 | - | - | [68] |
| Xylariales | Polystigmataceae | Polystigma rubrum | MFLU 15-3091 | KY594023 | MF981079 | - | - | [68] |
| Xylariales | Requienellaceae | Acrocordiella occulta | RS9 | KT949893 | KT949893 | - | - | [29] |
| Xylariales | Requienellaceae | Acrocordiella omanensis | SQUCC 15091 | MG584568 | MG584570 | - | - | [69] |
| Xylariales | Requienellaceae | Requienella fraxini | RS2 | KT949909 | KT949909 | - | - | [29] |
| Xylariales | Requienellaceae | Requienella seminuda | RS12 = CBS 140502 | KT949912 | KT949912 | MK523300 | - | [29,64] |
| Xylariales | Xylariaceae | Abieticola koreana | EML-F0010-1 | JN977612 | JQ014618 | KP792128 | KP792126 | [70] |
| Xylariales | Xylariaceae | Amphirosellinia nigrospora | HAST 91092308 | GU322457 | - | GQ848340 | GQ495951 | [71] |
| Xylariales | Xylariaceae | Anthostomella formosa | MFLUCC 14-0170 | KP297403 | KP340544 | KP340531 | KP406614 | [59] |
| Xylariales | Xylariaceae | Anthostomella helicofissa | MFLUCC 14-0173 | KP297406 | KP340547 | KP340534 | KP406617 | [59] |
| Xylariales | Xylariaceae | Anthostomella obesa | MFLUCC 14-0171 | KP297405 | KP340546 | KP340533 | KP406616 | [59] |
| Xylariales | Xylariaceae | Anthostomella pseudobambusicola | MFLUCC 15-0192 | KU940153 | KU863141 | - | - | [72] |
| Xylariales | Xylariaceae | Anthostomelloides brabeji | CBS 110128 | EU552098 | EU552098 | - | - | [73] |
| Xylariales | Xylariaceae | Anthostomelloides forlicesenica | MFLUCC 14-0558 | KP297397 | KP340539 | - | KP406608 | [66] |
| Xylariales | Xylariaceae | Anthostomelloides krabiensis | MFLUCC 15-0678 | KX305927 | KX305928 | KX305929 | - | [30] |
| Xylariales | Xylariaceae | Anthostomelloides leucospermi | CBS:110126 | EU552100 | - | - | - | [73] |
| Xylariales | Xylariaceae | Anthostomelloides proteae | CBS 110127 | EU552101 | - | - | - | [73] |
| Xylariales | Xylariaceae | Astrocystis mirabilis | 94070803 HAST | GU322448 | - | GQ844835 | GQ495941 | [71] |
| Xylariales | Xylariaceae | Brunneiperidium gracilentum | MFLUCC 14-0011 Ex-type | KP297400 | KP340542 | KP340528 | KP406611 | [66] |
| Xylariales | Xylariaceae | Collodiscula japonica | CBS 124266 | JF440974 | JF440974 | KY624273 | KY624316 | [50,74] |
| Xylariales | Xylariaceae | Coniolariella gamsii | Co27IRAN 842C, CBS114379 (T) | GU553325 | GU553329 | - | - | [35] |
| Xylariales | Xylariaceae | Entalbostroma erumpens | ICMP 21152 | KX258206 | - | KX258204 | KX258205 | [75] |
| Xylariales | Xylariaceae | Entoleuca mammata | J.D.R. 100 | GU300072 | - | GQ844782 | GQ470230 | [71] |
| Xylariales | Xylariaceae | Euepixylon sphaeriostomum | J.D.R. 261 | GU292821 | - | GQ844774 | GQ470224 | [71] |
| Xylariales | Xylariaceae | Halorosellinia oceanica | SGLAf82 | EU715635 | - | - | - | [76] |
| Xylariales | Xylariaceae | Hypocopra rostrata | NRRL 66178 | KM067909 | - | - | - | [77] |
| Xylariales | Xylariaceae | Hypocreodendron sanguineum | J.D.R. 169 | GU322433 | - | GQ844819 | GQ487710 | [71] |
| Xylariales | Xylariaceae | Kretzschmaria clavus | YMJ 114 | EF026126 | - | GQ844789 | EF025611 | [71,78] |
| Xylariales | Xylariaceae | Linosporopsis ischnotheca | LIF1 = CBS 145761 | MN818952 | MN818952 | MN820708 | MN820715 | [79] |
| Xylariales | Xylariaceae | Lunatiannulus irregularis | MFLUCC 14-0014 | KP297398 | KP340540 | KP340526 | KP406609 | [57] |
| Xylariales | Xylariaceae | Nemania serpens | CBS 679.86 | KU683765 | - | KU684284 | KU684188 | [80] |
| Xylariales | Xylariaceae | Neoxylaria arengae | MFLUCC 15-0292 | MT496747 | - | MT502418 | - | [81] |
| Xylariales | Xylariaceae | Podosordaria mexicana | WSP 176 | GU324762 | - | GQ853039 | GQ844840 | [71] |
| Xylariales | Xylariaceae | Poronia punctata | CBS 656.78 | KT281904 | KY610496 | KY624278 | KX271281 | [5,50] |
| Xylariales | Xylariaceae | Rosellinia aquila | MUCL 51703 | KY610392 | KY610460 | KY624285 | KX271253 | [50] |
| Xylariales | Xylariaceae | Rostrohypoxylon terebratum | CBS 119137 | DQ631943 | DQ840069 | DQ631954 | DQ840097 | [82,83] |
| Xylariales | Xylariaceae | Ruwenzoria pseudoannulata | MUCL 51394 | KY610406 | KY610494 | KY624286 | KX271278 | [50] |
| Xylariales | Xylariaceae | Sarcoxylon compunctum | CBS 359.61 | KT281903 | KY610462 | KY624230 | KX271255 | [5,50] |
| Xylariales | Xylariaceae | Stilbohypoxylon elaeicola | Y.M.J. 173 | EF026148 | - | GQ844826 | EF025616 | [71] |
| Xylariales | Xylariaceae | Stilbohypoxylon elaeidis | MFLUCC 15-0295a | MT496745 | MT496755 | MT502416 | MT502420 | [81] |
| Xylariales | Xylariaceae | Stilbohypoxylon quisquiliarum | Y.M.J. 172 | EF026119 | - | GQ853020 | EF025605 | [71] |
| Xylariales | Xylariaceae | Vamsapriya bambusicola | MFLUCC 11-0477 | KM462835 | KM462836 | KM462834 | KM462833 | [84] |
| Xylariales | Xylariaceae | Vamsapriya breviconidiophora | MFLUCC 14-0436 | MF621584 | MF621588 | - | - | [39] |
| Xylariales | Xylariaceae | Vamsapriya indica | MFLUCC 12-0544 | KM462839 | KM462840 | KM462841 | KM462838 | [84] |
| Xylariales | Xylariaceae | Vamsapriya khunkonensis | MFLUCC 11-0475 | KM462830 | KM462831 | KM462829 | KM462828 | [84] |
| Xylariales | Xylariaceae | Vamsapriya yunnana | KUMCC 18-0008 | MG833874 | MG833873 | MG833875 | - | [85] |
| Xylariales | Xylariaceae | Virgaria boninensis | JCM 18624 | AB740956 | AB740960 | - | - | [86] |
| Xylariales | Xylariaceae | Virgaria nigra | CBS 128006 | MH864744 | MH876180 | - | - | [44] |
| Xylariales | Xylariaceae | Xylaria hypoxylon | CBS 122620 | KY610407 | KY610495 | KY624231 | KX271279 | [50,87] |
| Sordariomycetes genera incertae sedis | Xylariales genera incertae sedis | Melanographium phoenicis | MFLUCC 18-1481 | MN482677 | MN482678 | - | - | [13] |
| Sordariomycetes genera incertae sedis | Xylariales genera incertae sedis | Ceratocladium microspermum | CBS126092 | MH864077 | MH875534 | - | - | [44] |
| Xylariales | Xylariales genera incertae sedis | Ascotricha chartarum | CBS 234.97 | KF893284 | - | - | KF893271 | [88] |
| Xylariales | Xylariales genera incertae sedis | Ascotricha longipila | OUCMBI110118 (T) | KC503896 | - | - | KF893265 | [88] |
| Xylariales | Xylariales genera incertae sedis | Ascotricha lusitanica | CBS 462.70 (IT) | KF893289 | - | - | KF893275 | [88] |
| Xylariales | Xylariales genera incertae sedis | Ascotricha parvispora | OUCMBI110001 (T) | JX014298 | - | - | KF893267 | [88] |
| Xylariales | Xylariales genera incertae sedis | Ascotricha sinuosa | OUCMBI101190 (T) | JX014299 | - | - | KF893266 | [88] |
| Xylariales | Xylariales genera incertae sedis | Alloanthostomella rubicola | MFLUCC 14-0175 | KP297407 | KP340548 | KP340535 | KP406618 | [89] |
| Xylariales | Xylariales genera incertae sedis | Circinotrichum cycadis | CPC 17285 | KJ869121 | KJ869178 | - | - | [26] |
| Xylariales | Xylariales genera incertae sedis | Circinotrichum maculiforme | CPC 24566 | KR611874 | KR611895 | - | - | [90] |
| Xylariales | Xylariales genera incertae sedis | Circinotrichum papakurae | CBS 101373 | KR611876 | KR611897 | - | - | [90] |
| Xylariales | Xylariales genera incertae sedis | Circinotrichum sinense | KY994106 | KY994107 | - | - | [91] | |
| Xylariales | Xylariales genera incertae sedis | Gyrothrix eucalypti | CPC 36066 | MN562109 | MN567617 | - | - | [92] |
| Xylariales | Xylariales genera incertae sedis | Gyrothrix inops | BE108 | KC775746 | KC775721 | - | - | [66] |
| Xylariales | Xylariales genera incertae sedis | Gyrothrix oleae | CPC 37069 | MN562136 | MN567643 | - | - | [92] |
| Xylariales | Xylariales genera incertae sedis | Gyrothrix ramosa | MUCL54061 | KC775747 | KC775722 | - | - | [66] |
| Xylariales | Xylariales genera incertae sedis | Haploanthostomella elaeidis | MFLU 20-0522 | MT929161 | MT929312 | MT928154 | - | This study |
| Xylariales | Xylariales genera incertae sedis | Neoanthostomella pseudostromatica | MFLUCC 11-0610 | KU940158 | KU863146 | - | - | [72] |
| Xylariales | Xylariales genera incertae sedis | Neoanthostomella viticola | MFLUCC 16-0243 | KX505957 | KX505958 | KX789496 | KX789495 | [89] |
| Xylariales | Xylariales genera incertae sedis | Pseudoanthostomella conorum | CBS 119333 | EU552099 | - | - | - | [73] |
| Xylariales | Xylariales genera incertae sedis | Pseudoanthostomella delitescens | MFLUCC 16-0477 | KX533451 | KX533452 | KX789491 | KX789490 | [89] |
| Xylariales | Xylariales genera incertae sedis | Pseudoanthostomella pini-nigrae | MFLUCC 16-0478 | KX533453 | KX533454 | KX789492 | - | [89] |
| Xylariales | Xylariales genera incertae sedis | Pseudoanthostomella sepelibilis | AY908989 | AY875645 | - | - | Unpublished | |
| Xylariales | Xylariales genera incertae sedis | Xenoanthostomella chromolaenae | MFLUCC 17-1484 | MN638863 | MN638848 | - | - | [3] |
| Xylariales | Zygosporiaceae | Zygosporium oscheoides | MFLUCC 14-0402 | MF621585 | MF621589 | - | - | [93] |
| Xylariales | Zygosporiaceae | Zygosporium minus | HKAS99625 | MF621586 | MF621590 | - | - | [93] |
| No. | Species | Host | Country | Reference | |
|---|---|---|---|---|---|
| Eudicots | Monocots | ||||
| 1 | Endocalyx amarkantakensis | Shorea robusta (Dipterocarpaceae) | India (Holotype) | [103] | |
| 2 | E. cinctus * | Livistona chinensis var. boninensis (Arecaceae; solitary palm) | Japan | [104] | |
| Oncosperma fasciculatum (Arecaceae; clustering, rarely solitary palm) | Japan | [101] | |||
| Oncosperma sp. (Arecaceae; clustering, rarely solitary palm) | Sri Lanka (Holotype) | [100] | |||
| Phoenix canariensis (Arecaceae; solitary palm) | Japan | [101] | |||
| Phoenix hanceana (Arecaceae; solitary palm) | Hong Kong | [105] | |||
| Trachycarpus fortunei (Arecaceae; solitary palm) | Japan | [101] | |||
| 3 | E. collantesis | Smilax sp. (Smilacaceae) | Cuba (Holotype) | [106] | |
| 4 | E. indicus | twigs of woody | India (Holotype) | [107] | |
| 5 | E. indumentum | Livistona chinensis var. boninensis (Arecaceae; solitary palm) | Japan (Holotype) | [101,104] | |
| Phoenix canariensis (Arecaceae; solitary palm) | Japan | [104] | |||
| 6 | E. melanoxanthus | Acrocomia mexicana (Arecaceae) | Mexico | [108] | |
| Archontophoenix alexandrae (Arecaceae; solitary palm) | Australia | [109] | |||
| Hong Kong | [105,109] | ||||
| Malaysia | [109] | ||||
| Singapore | [109] | ||||
| Arecaceae | Mexico | [108] | |||
| Arenga engleri (Arecaceae; clustering palm) | Hong Kong | [105] | |||
| Japan | [104] | ||||
| Dypsis lutescens (=Chrysalidocarpus lutescens) (Arecaceae; clustering palm) | Japan | [104] | |||
| Caryota urens (Arecaceae; solitary palm) | Sri Lanka (Holotype) | [100] | |||
| Cocos nucifera (Arecaceae; solitary palm) | Australia | [109] | |||
| Ghana | [110] | ||||
| Hawaii | [111,112] | ||||
| Japan | [104] | ||||
| Malaysia | [109,113] | ||||
| Papua New Guinea | [114] | ||||
| Seychelles | [109] | ||||
| Singapore | [109] | ||||
| Coffea arabica (Rubiaceae) | Venezuela | [115] | |||
| Dracaena fragrans (Asparagaceae) | Cuba | [116] | |||
| Venezuela | [115] | ||||
| Elaeis guineensis (Arecaceae; solitary palm) | Ghana | [110] | |||
| Myanmar | [117] | ||||
| Sierra Leone | [113] | ||||
| Elaeis sp. (Arecaceae; solitary palm) | Japan | [104] | |||
| Licuala longicalycata (Arecaceae; solitary palm) | Thailand | [118] | |||
| Livistona chinensis (Arecaceae; solitary palm) | Hong Kong | [105] | |||
| Livistona chinensis var. boninensis (Arecaceae; solitary palm) | Japan | [104] | |||
| Livistona rotundifolia (Arecaceae; solitary palm) | Taiwan | [119] | |||
| Livistona speciosa (Arecaceae; solitary palm) | Myanmar | [117] | |||
| Nannorrhops ritchieana (Arecaceae; clustering palm) | Pakistan | [120] | |||
| Phoenix canariensis (Arecaceae; solitary palm) | Japan | [104] | |||
| Phoenix hanceana (Arecaceae; solitary palm) | Hong Kong | [105,121] | |||
| Phoenix reclinata (Arecaceae; solitary palm) | Ghana | [110] | |||
| Phoenix roebelenii (Arecaceae; solitary palm) | Japan | [104] | |||
| Phoenix roebelenii-senegalensis (Arecaceae; solitary palm) | Japan | [104] | |||
| Ravenala madagascariensis (Strelitziaceae) | Japan | [104] | |||
| Taiwan | [119] | ||||
| Ripogonum scandens (Ripogonaceae) | New Zealand | [122] | |||
| Roystonea borinquena (Arecaceae; solitary palm) | USA (Florida) | [123] | |||
| Roystonea regia (Arecaceae; solitary palm) | Cuba | [124,125,126,127] | |||
| Sabal palmetto (Arecaceae; solitary palm) | USA (Florida) | [128] | |||
| Serenoa serrulata (Arecaceae; clustering and solitary palm) | USA (Florida) | [129] | |||
| Smilax sp. (Smilacaceae) | USA (Florida) | [128] | |||
| Trachycarpus fortunei (Arecaceae; solitary palm) | China | [109] | |||
| unknown, palm | Australia | [109] | |||
| China | [109] | ||||
| Hong Kong | [109] | ||||
| Malaysia | [109] | ||||
| Seychelles | [109] | ||||
| Singapore | [109] | ||||
| Wodyetia bifurcata (Arecaceae; solitary palm) | Florida | [123] | |||
| E. melanoxanthus (=E. melanoxanthus var. grossus) | Trachycarpus fortunei (Arecaceae; solitary palm) | Japan | [101] | ||
| E. melanoxanthus (=E. melanoxanthus var. melanoxanthus) | Acrocomia intumescens (Arecaceae; solitary palm) | Brazil | [102] | ||
| Butia yatay (Arecaceae; solitary palm) | Argentina | [130] | |||
| Cocos nucifera (Arecaceae; solitary palm) | Ghana | [101] | |||
| Euterpe edulis (Arecaceae; solitary, or rarely clustering palm (growing in dense tufts or clumps) and then with few stems) | Argentina | [130] | |||
| Brazil | [102] | ||||
| Euterpe oleracea (Arecaceae; clustering palm) | Brazil | [102] | |||
| Livistona chinensis var. boninensis (Arecaceae; solitary palm) | Japan | [101] | |||
| Livistona chinensis var. subglobosa (Arecaceae; solitary palm) | Japan | [101] | |||
| Phoenix canariensis (Arecaceae; solitary palm) | Japan | [101] | |||
| Phoenix roebelenii (Arecaceae; solitary palm) | Japan | [101] | |||
| Satakentia liukiuensis (Arecaceae; solitary palm) | Japan | [101] | |||
| Syagrus coronata (Arecaceae; solitary palm) | Brazil | [131] | |||
| Syagrus romanzoffiana (Arecaceae; solitary palm) | Argentina | [130] | |||
| Trachycarpus fortunei (Arecaceae; solitary palm) | Japan | [101] | |||
| Washingtonia robusta (Arecaceae; solitary palm) | Japan | [101] | |||
| 7 | E. thwaitesii (Type species) | Cissus oreophila (Vitaceae) | Ghana | [132] | |
| Cissus sp. (Vitaceae) | Ghana | [133] | |||
| Sri Lanka | [133] | ||||
| Oncosperma sp. (Arecaceae; clustering, rarely solitary palm) | Ghana | [133] | |||
| Sri Lanka (Holotype) | [133] | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Konta, S.; Hyde, K.D.; Eungwanichayapant, P.D.; Karunarathna, S.C.; Samarakoon, M.C.; Xu, J.; Dauner, L.A.P.; Aluthwattha, S.T.; Lumyong, S.; Tibpromma, S. Multigene Phylogeny Reveals Haploanthostomella elaeidis gen. et sp. nov. and Familial Replacement of Endocalyx (Xylariales, Sordariomycetes, Ascomycota). Life 2021, 11, 486. https://doi.org/10.3390/life11060486
Konta S, Hyde KD, Eungwanichayapant PD, Karunarathna SC, Samarakoon MC, Xu J, Dauner LAP, Aluthwattha ST, Lumyong S, Tibpromma S. Multigene Phylogeny Reveals Haploanthostomella elaeidis gen. et sp. nov. and Familial Replacement of Endocalyx (Xylariales, Sordariomycetes, Ascomycota). Life. 2021; 11(6):486. https://doi.org/10.3390/life11060486
Chicago/Turabian StyleKonta, Sirinapa, Kevin D. Hyde, Prapassorn D. Eungwanichayapant, Samantha C. Karunarathna, Milan C. Samarakoon, Jianchu Xu, Lucas A. P. Dauner, Sasith Tharanga Aluthwattha, Saisamorn Lumyong, and Saowaluck Tibpromma. 2021. "Multigene Phylogeny Reveals Haploanthostomella elaeidis gen. et sp. nov. and Familial Replacement of Endocalyx (Xylariales, Sordariomycetes, Ascomycota)" Life 11, no. 6: 486. https://doi.org/10.3390/life11060486
APA StyleKonta, S., Hyde, K. D., Eungwanichayapant, P. D., Karunarathna, S. C., Samarakoon, M. C., Xu, J., Dauner, L. A. P., Aluthwattha, S. T., Lumyong, S., & Tibpromma, S. (2021). Multigene Phylogeny Reveals Haploanthostomella elaeidis gen. et sp. nov. and Familial Replacement of Endocalyx (Xylariales, Sordariomycetes, Ascomycota). Life, 11(6), 486. https://doi.org/10.3390/life11060486









