Comparative Analysis of the Intermolt and Postmolt Hepatopancreas Transcriptomes Provides Insight into the Mechanisms of Procambarus clarkii Molting Process
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Animals
2.2. Sample Collection and Preparation
2.3. RNA Isolation and RNA-Seq Library Preparation
2.4. Transcriptome Assembly and Annotation
2.5. Analysis of Simple-Sequence Repeats (SSR), SNP, InDells, and Genebody Coverage
2.6. Analysis of Differentially Expressed Genes (DEGs)
2.7. Molting Pathway Analysis
2.8. qRT–PCR
3. Results
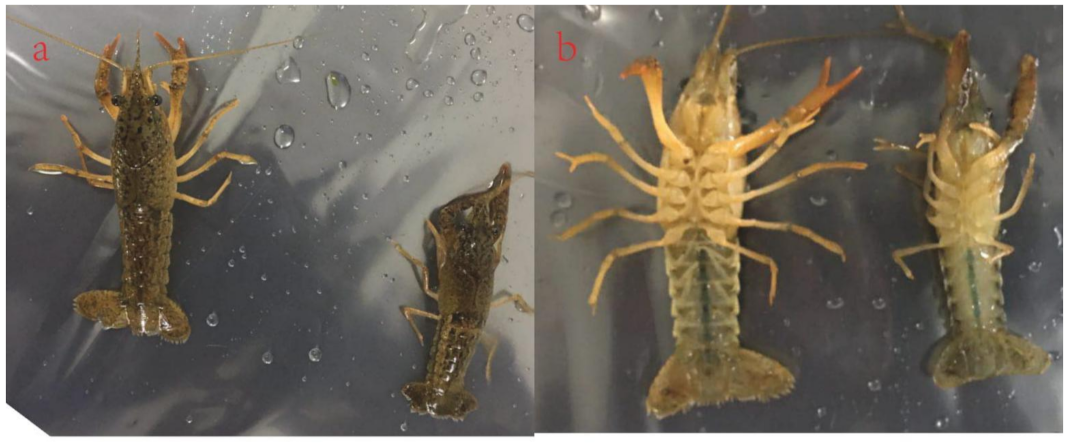
3.1. Ecdysteroid Changes in Crayfish Morphology
3.2. Read Sequencing, Assembly, and Mapping
3.3. Functional Annotation
3.4. Characterization of SSR, SNPs, and InDels
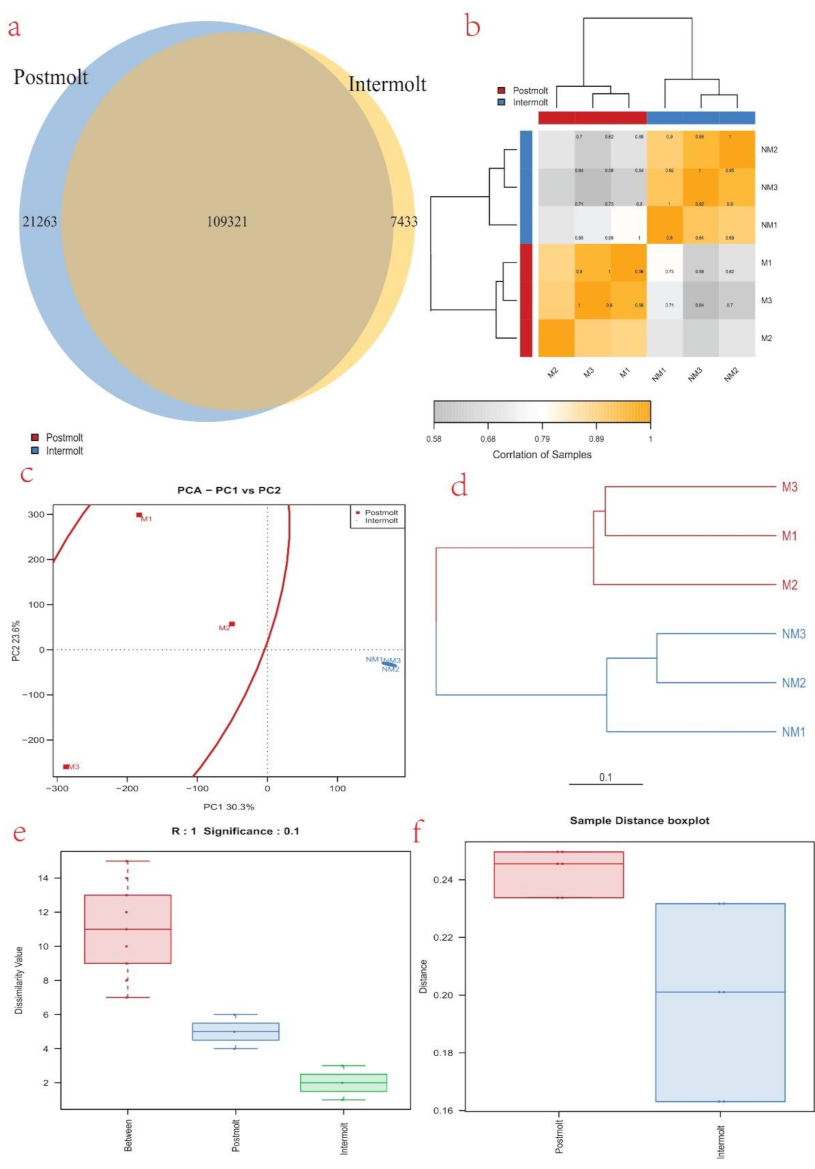
3.5. Differentially Expressed Genes (DEGs) Analysis
3.6. GO and KEGG Analysis of Differentially Expressed Genes
3.7. Analysis of Transcriptome Data by qRT–PCR
4. Discussion
4.1. Energy Demand during the Molting Process
4.2. Immunoregulation Process during Molting
4.3. Exoskeleton Calcification and Sclerotization-Related Activities
4.4. Hormone Regulation Process during Molting
4.5. DEGs and SNPs Probe for Molting-Related Factors
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bondad-Reantaso, M.G.; Subasinghe, R.P.; Josupeit, H.; Cai, J.; Zhou, X. The Role of Crustacean Fisheries and Aquaculture in Global Food Security: Past, Present and Future. J. Invertebr. Pathol. 2012, 110, 158–165. [Google Scholar] [CrossRef] [PubMed]
- Hosamani, N.; Reddy, B.S.; Reddy, P.R. Crustacean Molting: Regulation and Effects of Environmental Toxicants. J. Mar. Sci. Res. Dev. 2017, 7, 1–8. [Google Scholar] [CrossRef]
- Venugopal, V.; Gopakumar, K. Shellfish: Nutritive Value, Health Benefits, and Consumer Safety. Compr. Rev. Food Sci. Food Saf. 2017, 16, 1219–1242. [Google Scholar] [CrossRef] [PubMed]
- Stentiford, G.D. Diseases in Aquatic Crustaceans: Problems and Solutions for Global Food Security. J. Invertebr. Pathol. 2012, 110, 139. [Google Scholar] [CrossRef] [PubMed]
- Skinner, D.M. Molting and Regeneration; Bliss, D., Crustacea, T.B., Eds.; Academic Press: New York, NY, USA, 1985. [Google Scholar] [CrossRef]
- Smith, G.; Ritar, A. Reproduction and Growth of Decapod Crustaceans in Relation to Aquaculture. Reprod. Biol. Crustac. 2008, 456–490. [Google Scholar] [CrossRef]
- Wenne, R.; Boudry, P.; Hemmer-Hansen, J.; Lubieniecki, K.P.; Was, A.; Kause, A. What Role for Genomics in Fisheries Management and Aquaculture? Aquat. Living Resour. 2007, 20, 241–255. [Google Scholar] [CrossRef]
- McAndrew, B.; Napier, J. Application of Genetics and Genomics to Aquaculture Development: Current and Future Directions. J. Agric. Sci. 2011, 149, 143–151. [Google Scholar] [CrossRef]
- Aiken, D.E.; Waddy, S.L. Molting and Growth in Crayfish, a Review. Can. Tech. Rep. Fish. Aquat. Sci. 1987, 587, 1–34. [Google Scholar]
- Chang, E.S. Physiological and Biochemical Changes during the Molt Cycle in Decapod Crustaceans: An Overview. J. Exp. Mar. Bio. Ecol. 1995, 193, 1–14. [Google Scholar] [CrossRef]
- Naya, Y.; Ohnishi, M.; Ikeda, M.; Miki, W.; Nakanishi, K. What Is Molt-Inhibiting Hormone? The Role of an Ecdysteroidogenesis Inhibitor in the Crustacean Molting Cycle. Proc. Natl. Acad. Sci. USA 1989, 86, 6826–6829. [Google Scholar] [CrossRef]
- Lachaise, F.; Le Roux, A.; Hubert, M.; Lafont, R. The Molting Gland of Crustaceans: Localization, Activity, And Endocrine Control (A Review). J. Crustac. Biol. 1993, 13, 198–234. [Google Scholar] [CrossRef]
- Pamuru, R.R.; Rosen, O.; Manor, R.; Chung, J.S.; Zmora, N.; Glazer, L.; Aflalo, E.D.; Weil, S.; Tamone, S.L.; Sagi, A. Stimulation of Molt by RNA Interference of the Molt-Inhibiting Hormone in the Crayfish Cherax Quadricarinatus. Gen. Comp. Endocrinol. 2012, 178, 227–236. [Google Scholar] [CrossRef] [PubMed]
- Fingerman, M. The Endocrine Mechanisms of Crustaceans. J. Crustac. Biol. 1987, 7, 1. [Google Scholar] [CrossRef]
- Taketomi, Y.; Miyawaki, M.; Katakura, Y. Endocrine System of Crustaceans. Atlas Endocr. Organs 1992, 207–226. [Google Scholar] [CrossRef]
- Shyamal, S.; Sudha, K.; Gayathri, N.; Anilkumar, G. The Y-Organ Secretory Activity Fluctuates in Relation to Seasons of Molt and Reproduction in the Brachyuran Crab, Metopograpsus Messor (Grapsidae): Ultrastructural and Immunohistochemical Study. Gen. Comp. Endocrinol. 2014, 196, 81–90. [Google Scholar] [CrossRef]
- Keller, R. Crustacean Neuropeptides: Structures, Functions and Comparative Aspects. Experientia 1992, 48, 439–448. [Google Scholar] [CrossRef]
- Jiang, H.; Yin, Y.; Zhang, X.; Hu, S.; Wang, Q. Chasing Relationships between Nutrition and Reproduction: A Comparative Transcriptome Analysis of Hepatopancreas and Testis from Eriocheir Sinensis. Comp. Biochem. Physiol. Part D Genom. Proteom. 2009, 4, 227–234. [Google Scholar] [CrossRef]
- Wang, W.; Wu, X.; Liu, Z.; Zheng, H.; Cheng, Y. Insights into Hepatopancreatic Functions for Nutrition Metabolism and Ovarian Development in the Crab Portunus Trituberculatus: Gene Discovery in the Comparative Transcriptome of Different Hepatopancreas Stages. PLoS ONE 2014, 9, e84921. [Google Scholar] [CrossRef] [PubMed]
- Kleinholz, L.H. Hormones in Crustacea. Biol. Rev. 1942, 17, 91–119. [Google Scholar] [CrossRef]
- Lee, J.H.; Suryaningtyas, I.T.; Yoon, T.H.; Shim, J.M.; Park, H.; Kim, H.W. Transcriptomic Analysis of the Hepatopancreas Induced by Eyestalk Ablation in Shrimp, Litopenaeus Vannamei. Comp. Biochem. Physiol. Part D Genom. Proteom. 2017, 24, 99–110. [Google Scholar] [CrossRef]
- Venkitaraman, P.R.; Jayalakshmy, K.V.; Balasubramanian, T. Effect of Eyestalk Ablation on Moulting and Growth in the Penaeid Shrimp Metapenaeus Monoceros (Fabricius, 1798). Indian J. Fish. 2010, 57, 25–32. [Google Scholar]
- Rana, S. Eyestalk Ablation of Freshwater Crab, Barytelphusa Lugubris: An Alternative Approach of Hormonal Induced Breeding. Int. J. Pure Appl. Zool. 2018, 6, 30–34. [Google Scholar]
- Diwan, A. Neuroendocrine Control of Moulting and Growth. Biotechnol. Penaeid Shrimps 2020, 383–410. [Google Scholar] [CrossRef]
- Tom, M.; Manfrin, C.; Chung, S.J.; Sagi, A.; Gerdol, M.; De Moro, G.; Pallavicini, A.; Giulianini, P.G. Expression of Cytoskeletal and Molt-Related Genes Is Temporally Scheduled in the Hypodermis of the Crayfish Procambarus Clarkii during Premolt. J. Exp. Biol. 2014, 217, 4193–4202. [Google Scholar] [CrossRef] [PubMed]
- Kuballa, A.V.; Holton, T.A.; Paterson, B.; Elizur, A. Moult Cycle Specific Differential Gene Expression Profiling of the Crab Portunus Pelagicus. BMC Genom. 2011, 12, 147. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Zhang, X.; Wei, J.; Sun, X.; Yuan, J.; Li, F.; Xiang, J. Whole Transcriptome Analysis Provides Insights into Molecular Mechanisms for Molting in Litopenaeus Vannamei. PLoS ONE 2015, 10, e0144350. [Google Scholar] [CrossRef]
- Huang, S.; Wang, J.; Yue, W.; Chen, J.; Gaughan, S.; Lu, W.; Lu, G.; Wang, C. Transcriptomic Variation of Hepatopancreas Reveals the Energy Metabolism and Biological Processes Associated with Molting in Chinese Mitten Crab, Eriocheir Sinensis. Sci. Rep. 2015, 5, 1–11. [Google Scholar] [CrossRef]
- Shyamal, S.; Das, S.; Guruacharya, A.; Mykles, D.L.; Durica, D.S. Transcriptomic Analysis of Crustacean Molting Gland (Y-Organ) Regulation via the MTOR Signaling Pathway. Sci. Rep. 2018, 8, 1–17. [Google Scholar] [CrossRef]
- Moss, C.G. Soft-Shell Crab Fishery in Texas. In Proceedings of the National Symposium on the Soft-Shelled Blue Crab Fishery, Ocean Springs, MS, USA, 12–13 February 1985. [Google Scholar]
- Ye, Y.; An, Y.; Li, R.; Mu, C.; Wang, C. Strategy of Metabolic Phenotype Modulation in Portunus Trituberculatus Exposed to Low Salinity. J. Agric. Food Chem. 2014, 62, 3496–3503. [Google Scholar] [CrossRef]
- Bukowski, R.; Sun, Q. De Novo Transcriptome Assembly Using Trinity. Available online: https://biohpc.cornell.edu/lab/doc/trinity_workshop_part1.pdf (accessed on 7 March 2020).
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-Length Transcriptome Assembly from RNA-Seq Data without a Reference Genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and Memory-Efficient Alignment of Short DNA Sequences to the Human Genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Dewey, C.N. RSEM: Accurate Transcript Quantification from RNA-Seq Data with or without a Reference Genome. BMC Bioinf. 2011, 12, 323. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and Accurate Long-Read Alignment with Burrows Wheeler Transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Aligment/Map Format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed]
- BioBam Bioinformatics, S.L. Blast2GO Command Line User Manual. 2014. Available online: https://www.blast2go.com/images/b2g_pdfs/blast2go_cli_manual.pdf (accessed on 7 March 2020).
- Conesa, A.; Gotz, S.; Garcia-Gomez, J.M.; Terol, J.; Talon, M.; Robles, M. Blast2GO: A Universal Tool for Annotation, Visualization and Analysis in Functional Genomics Research. Bioinformatics 2005, 21, 3674–3676. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Huber, W. Differential Expression Analysis for Sequence Count Data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef]
- Yu, G.; Wang, L.G.; Han, Y.; He, Q.Y. ClusterProfiler: An R Package for Comparing Biological Themes among Gene Clusters. Omi. A J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef]
- Young, M.D.; Wakefield, M.J.; Smyth, G.K.; Oshlack, A. Gene Ontology Analysis for RNA-Seq: Accounting for Selection Bias. Genome Biol. 2010, 11, R14. [Google Scholar] [CrossRef]
- Saito, R.; Smoot, M.E.; Ono, K.; Ruscheinski, J.; Wang, P.L.; Lotia, S.; Pico, A.R.; Bader, G.D.; Ideker, T. A Travel Guide to Cytoscape Plugins. Nat. Methods 2012, 1069–1076. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Real-Time Quantitative PCR and the 2-ΔΔCT Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- De Hoon, M.J.L.; Imoto, S.; Nolan, J.; Miyano, S. Open Source Clustering Software. Bioinformatics 2004, 20, 1453–1454. [Google Scholar] [CrossRef] [PubMed]
- Saldanha, A.J. Java Treeview—Extensible Visualization of Microarray Data. Bioinformatics 2004, 20, 3246–3248. [Google Scholar] [CrossRef]
- Cirillo, E.; Parnell, L.D.; Evelo, C.T. A Review of Pathway-Based Analysis Tools That Visualize Genetic Variants. Front. Genet. 2017, 8, 174. [Google Scholar] [CrossRef]
- Liao, F.; Zeng, G.; Yuan, X.; Li, S.; Wang, C.; Liang, Z. Complete Mitochodrial Genome of Elopichthys Bambusa (Cypriniformes, Cyprinidae). Mitochondrial DNA 2016, 27, 1387–1388. [Google Scholar] [CrossRef] [PubMed]
- Raviv, S.; Parnes, S.; Sagi, A. Coordination of Reproduction and Molt in Decapods. Available online: http://lifeserv.bgu.ac.il/wb/sagia/media/Book%20chapters/15%20%20Reprod%20Biol%20Crust_2008_Coordination%20of%20reproduction-2008.pdf (accessed on 15 August 2020).
- Chang, E.S.; Mykles, D.L. Regulation of Crustacean Molting: A Review and Our Perspectives. Gen. Comp. Endocrinol. 2011, 323–330. [Google Scholar] [CrossRef] [PubMed]
- Shen, H.; Zang, Y.; Song, K.; Ma, Y.; Dai, T.; Serwadda, A. A Meta-Transcriptomics Survey Reveals Changes in the Microbiota of the Chinese Mitten Crab Eriocheir Sinensis Infected with Hepatopancreatic Necrosis Disease. Front. Microbiol. 2017, 8, 732. [Google Scholar] [CrossRef]
- Devaraj, H.; Natarajan, A. Molecular Mechanisms Regulating Molting in a Crustacean. FEBS J. 2006, 273, 839–846. [Google Scholar] [CrossRef]
- Kuballa, A.V.; Elizur, A. Differential Expression Profiling of Components Associated with Exoskeletal Hardening in Crustaceans. BMC Genom. 2008, 9, 575. [Google Scholar] [CrossRef]
- Wenner, A.; Gore, R.H. Molting and Growth in Decapod Larvae. In Crustacean Issues 2; Routledge: London, UK, 2018; pp. 1–65. [Google Scholar] [CrossRef]
- Wassenberg, T.J.; Hill, B.J. Moulting Behaviour of the Tiger Prawn Penaeus Esculentus (Haswell). Mar. Freshw. Res. 1984, 35, 561–571. [Google Scholar] [CrossRef]
- Young, J.H. An Enzymological Approach to Mitochondrial Energy Transduction. In Membrane Structure and Mechanisms of Biological Energy Transduction; Springer: New York, NY, USA, 1972; pp. 137–146. [Google Scholar] [CrossRef]
- Vasudevan, D.; Sreekumari, S.; Vaidyanathan, M. Biological Oxidation and Electron Transport Chain. In Textbook of Biochemistry for Medical Students; Jaypee Brothers Medical Publishers (P) Ltd.: New Delhi, India, 2007; p. 225. [Google Scholar] [CrossRef]
- Navarro, A.; Boveris, A. The Mitochondrial Energy Transduction System and the Aging Process. Am. J. Phys. Cell Phys. 2007, 670–686. [Google Scholar] [CrossRef]
- Enríquez-Freire, E.; López, R.; Peña, A. Potassium Ion Efflux Induced by Cationic Compounds in Yeast. Biochim. Biophys. Acta Biomembr. 1999, 1418, 147–157. [Google Scholar] [CrossRef][Green Version]
- Eggleton, M.G. A Physiological Study of Phosphagen in Plain Muscle. J. Physiol. 1934, 82, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, T.; Furukohri, T. Evolution of Phosphagen Kinase. Primary Structure of Glycocyamine Kinase and Arginine Kinase from Invertebrates. J. Mol. Biol. 1994, 237, 353–357. [Google Scholar] [CrossRef] [PubMed]
- Ellington, W.R. Evolution and Physiological Roles of Phosphagen Systems. Annu. Rev. Physiol. 2001, 63, 289–325. [Google Scholar] [CrossRef]
- Kotlyar, S.; Weihrauch, D.; Paulsen, R.; Towle, D. Expression of Arginine Kinase MRNA and Protein in Gills of the Green Shore Crab Carcinus Maenas. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 1999, 124, S80. [Google Scholar] [CrossRef]
- Hoffmann, J.A.; Kafatos, F.C.; Janeway, C.A.; Ezekowitz, R.A.B. Phylogenetic Perspectives in Innate Immunity. Science 1999, 1313–1318. [Google Scholar] [CrossRef]
- Iwanaga, S.; Bok, L.L. Recent Advances in the Innate Immunity of Invertebrate Animals. J. Biochem. Mol. Biol. 2005, 38, 128–150. [Google Scholar] [CrossRef]
- Vazquez, L.; Alpuche, J.; Maldonado, G.; Agundis, C.; Pereyra-Morales, A.; Zenteno, E. Immunity Mechanisms in Crustaceans. Innate Immun. 2009, 179–188. [Google Scholar] [CrossRef]
- Xu, Z.; Liu, A.; Li, S.; Wang, G.; Ye, H. Hepatopancreas Immune Response during Molt Cycle in the Mud Crab, Scylla paramamosain. Sci. Rep. 2020, 10, 1–14. [Google Scholar] [CrossRef]
- Van Herreweghe, J.M.; Michiels, C.W. Invertebrate Lysozymes: Diversity and Distribution, Molecular Mechanism and In Vivo Function. J. Biosci. 2012, 327–348. [Google Scholar] [CrossRef]
- Rowley, A.F. The Immune System of Crustaceans. In Encyclopedia of Immunobiology; Elsevier Inc.: Amsterdam, The Netherlands, 2016; Volume 1, pp. 437–453. [Google Scholar] [CrossRef]
- Chen, T.-T.; Tan, L.-R.; Hu, N.; Dong, Z.-Q.; Hu, Z.; Jiang, Y.-M.; Chen, P.; Pan, M.-H.; Lu, C. C-Lysozyme Contributes to Antiviral Immunity in Bombyx Mori against Nucleopolyhedrovirus Infection. J. Insect Physiol. 2018, 108, 54–60. [Google Scholar] [CrossRef]
- Zhou, J.; Zhao, S.; Fang, W.H.; Zhou, J.F.; Zhang, J.X.; Ma, H.; Lan, J.F.; Li, X.C. Newly Identified Invertebrate-Type Lysozyme (Splys-i) in Mud Crab (Scylla paramamosain) Exhibiting Muramidase-Deficient Antimicrobial Activity. Dev. Comp. Immunol. 2017, 74, 154–166. [Google Scholar] [CrossRef] [PubMed]
- Kawai, T.; Akira, S. Pathogen Recognition with Toll-like Receptors. Curr. Opin. Immunol. 2005, 338–344. [Google Scholar] [CrossRef] [PubMed]
- Marion, J. Toll-Like Receptors: Pathogen Recognition and Signaling. In Molecular Life Sciences; Springer: New York, NY, USA, 2018; pp. 1198–1204. [Google Scholar] [CrossRef]
- Sánchez-Paz, A.; Muhlia-Almazán, A. Uncovering and Defragmenting the Role of the Toll Pathway in the Innate Immune Responses of Cultured Crustaceans against Viral Pathogens. Rev. Aquac. 2020, 12, 12411. [Google Scholar] [CrossRef]
- Nakajima, H.; Iwamoto, I. Interleukin 15, Interleukin 17, and Interleukin 25. In Inflammation and Allergy Drug Design; Wiley-Blackwell: Oxford, UK, 2011; pp. 197–204. [Google Scholar] [CrossRef]
- Gu, W.-B.; Zhou, Y.L.; Tu, D.D.; Zhou, Z.K.; Zhu, Q.H.; Chen, Y.Y.; Shu, M.A. Identification and Characterization of Pro-Interleukin-16 from Mud Crab Scylla paramamosain: The First Evidence of Proinflammatory Cytokine in Crab Species. Fish Shellfish Immunol. 2017, 70, 701–709. [Google Scholar] [CrossRef]
- Franchi, L.; Warner, N.; Viani, K.; Nuñez, G. Function of Nod-like Receptors in Microbial Recognition and Host Defense. Immunol. Rev. 2009, 106–128. [Google Scholar] [CrossRef] [PubMed]
- Shi, X.Z.; Zhao, X.F.; Wang, J.X. Molecular Cloning and Expression Analysis of Chymotrypsin-like Serine Protease from the Chinese Shrimp, Fenneropenaeus Chinensis. Fish Shellfish Immunol. 2008, 25, 589–597. [Google Scholar] [CrossRef] [PubMed]
- Fang, D.A.; Huang, X.M.; Zhang, Z.Q.; Xu, D.P.; Zhou, Y.F.; Zhang, M.Y.; Liu, K.; Duan, J.R.; Shi, W.G. Molecular Cloning and Expression Analysis of Chymotrypsin-like Serine Protease from the Redclaw Crayfish (Cherax quadricarinatus): A Possible Role in the Junior and Adult Innate Immune Systems. Fish Shellfish Immunol. 2013, 34, 1546–1552. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.-X.; Liu, H.-P.; Chen, R.-Y.; Shen, K.-L.; Wang, K.-J. Identification of a Serine Proteinase Homolog (Sp-SPH) Involved in Immune Defense in the Mud Crab Scylla paramamosain. PLoS ONE 2013, 8, e63787. [Google Scholar] [CrossRef]
- Alcaraz, M.; Sardá, F. Oxygen Consumption by Nephrops norvegicus (L.), (Crustacea: Decapoda) in Relationship with Its Moulting Stage. J. Exp. Mar. Biol. Ecol. 1981, 54, 113–118. [Google Scholar] [CrossRef]
- Penkoff, S.J.; Thurberg, F.P. Changes in Oxygen Consumption of the American Lobster, Homarus American Us, during the Molt Cycle. Comp. Biochem. Physiol. Part A Physiol. 1982, 72, 621–622. [Google Scholar] [CrossRef]
- Cockcroft, A.C.; Wooldridge, T. The Effects of Mass, Temperature and Molting on the Respiration of Macropetasma Africanus Balss (Decapoda: Penaeoidea). Comp. Biochem. Physiol. Part A Physiol. 1985, 81, 143–148. [Google Scholar] [CrossRef]
- Wilhelm Filho, D.; Torres, M.A.; Tribess, T.B.; Pedrosa, R.C.; Soares, C.H.L. Influence of Season and Pollution on the Antioxidant Defenses of the Cichlid Fish Acará (Geophagus Brasiliensis). Braz. J. Med. Biol. Res. 2001, 34, 719–726. [Google Scholar] [CrossRef]
- Liu, H.P.; Chen, F.Y.; Gopalakrishnan, S.; Qiao, K.; Bo, J.; Wang, K.J. Antioxidant Enzymes from the Crab Scylla paramamosain: Gene Cloning and Gene/Protein Expression Profiles against LPS Challenge. Fish Shellfish Immunol. 2010, 28, 862–871. [Google Scholar] [CrossRef] [PubMed]
- Koháryová, M.; Kollárová, M. Thioredoxin System—A Novel Therapeutic Target. Gen. Physiol. Biophys. 2015, 34, 221–233. [Google Scholar] [CrossRef] [PubMed]
- Cunningham, G.M.; Roman, M.G.; Flores, L.C.; Hubbard, G.B.; Salmon, A.B.; Zhang, Y.; Gelfond, J.; Ikeno, Y. The Paradoxical Role of Thioredoxin on Oxidative Stress and Aging. Arch. Biochem. Biophys. 2015, 32–38. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Zhang, F.; Jiang, K.; Fang, Y.; Wang, J.; Zhao, M.; Qiao, Z.; Ma, L. Molecular Characterization of Thioredoxin-1 and Thioredoxin Reductase Activity in Mud Crab Scylla paramamosain. Genet. Mol. Res. 2014, 13, 10241–10255. [Google Scholar] [CrossRef] [PubMed]
- Wood, Z.A.; Poole, L.B.; Karplus, P.A. Peroxiredoxin Evolution and the Regulation of Hydrogen Peroxide Signaling. Science 2003, 300, 650–653. [Google Scholar] [CrossRef] [PubMed]
- Sue, G.R.; Ho, Z.C.; Kim, K. Peroxiredoxins: A Historical Overview and Speculative Preview of Novel Mechanisms and Emerging Concepts in Cell Signaling. Free Radic. Biol. Med. 2005, 1543–1552. [Google Scholar] [CrossRef]
- Tu, D.D.; Zhou, Y.L.; Gu, W.B.; Zhu, Q.H.; Xu, B.P.; Zhou, Z.K.; Liu, Z.P.; Wang, C.; Chen, Y.Y.; Shu, M.A. Identification and Characterization of Six Peroxiredoxin Transcripts from Mud Crab Scylla paramamosain: The First Evidence of Peroxiredoxin Gene Family in Crustacean and Their Expression Profiles under Biotic and Abiotic Stresses. Mol. Immunol. 2018, 93, 223–235. [Google Scholar] [CrossRef]
- Wu, L.; Zhou, Y.; Abbas, M.N.; Kausar, S.; Chen, Q.; Jiang, C.X.; Dai, L.S. Molecular Structure and Functional Characterization of the Peroxiredoxin 5 in Procambarus Clarkii Following LPS and Poly I:C Challenge. Fish Shellfish Immunol. 2017, 71, 28–34. [Google Scholar] [CrossRef]
- Decker, H.; Hellmann, N.; Jaenicke, E.; Lieb, B.; Meissner, U.; Markl, J. Minireview: Recent Progress in Hemocyanin Research. Integr. Comp. Biol. 2007, 47, 631–644. [Google Scholar] [CrossRef] [PubMed]
- Richards, A.G. The Integument of Arthropods Minneapolis; University of Minnesota Press: Minneapolis, MN, USA, 1951. [Google Scholar]
- Neville, A.C. Biology of the Arthropod Cuticle Berlin and New York; Springer: Berlin, Germany, 1975. [Google Scholar]
- Matsushita, M.; Fujita, T. The Lectin Pathway. Res. Immunol. 1996, 147, 115–118. [Google Scholar] [CrossRef]
- Shafer, T.H.; Roer, R.D.; Midgette-Luther, C.; Brookins, T.A. Postecdysial Cuticle Alteration in the Blue Crab, Callinectes Sapidus: Synchronous Changes in Glycoproteins and Mineral Nucleation. J. Exp. Zool. 1995, 271, 171–182. [Google Scholar] [CrossRef]
- Tweedie, E.P.; Coblentz, F.E.; Shafer, T.H. Purification of a Soluble Glycoprotein from the Uncalcified Ecdysial Cuticle of the Blue Crab Callinectes Sapidus and Its Possible Role in Initial Mineralization. J. Exp. Biol. 2004, 207, 2589–2598. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Akita, H.; Fukae, M.; Shimoda, S.; Aoba, T. Localization of Glycosylated Matrix Proteins in Secretory Porcine Enamel and Their Possible Functional Roles in Enamel Mineralization. Arch. Oral Biol. 1992, 37, 953–962. [Google Scholar] [CrossRef]
- Merzendorfer, H.; Zimoch, L. Chitin Metabolism in Insects: Structure, Function and Regulation of Chitin Synthases and Chitinases. J. Exp. Biol. 2003, 4393–4412. [Google Scholar] [CrossRef]
- Arakane, Y.; Hogenkamp, D.G.; Zhu, Y.C.; Kramer, K.J.; Specht, C.A.; Beeman, R.W.; Kanost, M.R.; Muthukrishnan, S. Characterization of Two Chitin Synthase Genes of the Red Flour Beetle, Tribolium Castaneum, and Alternate Exon Usage in One of the Genes during Development. Insect Biochem. Mol. Biol. 2004, 34, 291–304. [Google Scholar] [CrossRef]
- Hogenkamp, D.G.; Arakane, Y.; Zimoch, L.; Merzendorfer, H.; Kramer, K.J.; Beeman, R.W.; Kanost, M.R.; Specht, C.A.; Muthukrishnan, S. Chitin Synthase Genes in Manduca Sexta: Characterization of a Gut-Specific Transcript and Differential Tissue Expression of Alternately Spliced MRNAs during Development. Insect Biochem. Mol. Biol. 2005, 35, 529–540. [Google Scholar] [CrossRef]
- Terwilliger, N.B.; Ryan, M.C.; Towle, D. Evolution of Novel Functions: Cryptocyanin Helps Build New Exoskeleton in Cancer Magister. J. Exp. Biol. 2005, 208, 2467–2474. [Google Scholar] [CrossRef]
- Terra, W.R. The Origin and Functions of the Insect Peritrophic Membrane and Peritrophic Gel. Arch. Insect Biochem. Physiol. 2001, 47, 47–61. [Google Scholar] [CrossRef] [PubMed]
- Chung, J.S.; Webster, S.G. Moult Cycle-Related Changes in Biological Activity of Moult-Inhibiting Hormone (MIH) and Crustacean Hyperglycaemic Hormone (CHH) in the Crab, Carcinus Maenas. From Target to Transcript. Eur. J. Biochem. 2003, 270, 3280–3288. [Google Scholar] [CrossRef] [PubMed]
- Qiao, H.; Jiang, F.; Xiong, Y.; Jiang, S.; Fu, H.; Li, F.; Zhang, W.; Sun, S.; Jin, S.; Gong, Y.; et al. Characterization, Expression Patterns of Molt-Inhibiting Hormone Gene of Macrobrachium Nipponense and Its Roles in Molting and Growth. PLoS ONE 2018, 13, e0198861. [Google Scholar] [CrossRef]
- Huang, H.; Fu, C.; Chen, X.; Gong, J.; Huang, X.; Ye, H. Molt-Inhibiting Hormone (MIH) Gene from the Green Mud Crab Scylla Paramamosain and Its Expression during the Molting and Ovarian Cycle. Aquac. Res. 2015, 46, 2665–2675. [Google Scholar] [CrossRef]
- Nakagawa, Y.; Henrich, V.C. Arthropod Nuclear Receptors and Their Role in Molting. FEBS J. 2009, 6128–6157. [Google Scholar] [CrossRef] [PubMed]
- Souty-Grosset, C. Vitellogenin Synthesis in Marine Invertebrates Catherine Souty-Grosset. Endocrinol. Reprod. 1997, 1, 215–260. [Google Scholar]
- Girardie, J.; Boureme, D.; Couillaud, F.; Tamarelle, M.; Girardie, A. Anti-Juvenile Effect of Neuroparsin A, a Neuroprotein Isolated from Locust Corpora Cardiaca. Insect Biochem. 1987, 17, 977–983. [Google Scholar] [CrossRef]
- Girardie, J.; Girardie, A.; Huet, J.-C.; Pernollet, J.-C. Amino Acid Sequence of Locust Neuroparsins. FEBS Lett. 1989, 245, 4–8. [Google Scholar] [CrossRef]
- Badisco, L.; Marchal, E.; Van Wielendaele, P.; Verlinden, H.; Vleugels, R.; Vanden Broeck, J. RNA Interference of Insulin-Related Peptide and Neuroparsins Affects Vitellogenesis in the Desert Locust Schistocerca Gregaria. Peptides 2011, 32, 573–580. [Google Scholar] [CrossRef]
- Jiang, T.; Liu, H.; Shen, X.; Shimasaki, Y.; Oshima, Y.; Yang, J. Life History Variations among Different Populations of Coilia Nasus along the Chinese Coast Inferred from Otolith Microchemistry. J. Fac. Agric. Kyushu Univ. 2014, 59, 383–389. [Google Scholar] [CrossRef]
- Zou, X.T.; Qiao, X.J.; Xu, Z.R. Effect of β-Mannanase (Hemicell) on Growth Performance and Immunity of Broilers. Poult. Sci. 2006, 85, 2176–2179. [Google Scholar] [CrossRef] [PubMed]
- Clemmons, D.R. Insulin-like Growth Factor Binding Proteins and Their Role in Controlling IGF Actions. Cytokine Growth Factor Rev. 1997, 8, 45–62. [Google Scholar] [CrossRef]
- Zhou, Y.; Luo, W.; Yu, X.; Liu, Q.; Tong, J. Brain and Intestine Transcriptome Analyses and Identification of Genes Involved in Feed Conversion Efficiency of Yellow River Carp (Cyprinus Carpio Haematopterus). Comp. Biochem. Physiol. Part D Genom. Proteom. 2019, 29, 221–227. [Google Scholar] [CrossRef]
- Terwilliger, N.B.; Dangott, L.; Ryan, M. Cryptocyanin, a Crustacean Molting Protein: Evolutionary Link with Arthropod Hemocyanins and Insect Hexamerins. Proc. Natl. Acad. Sci. USA 1999, 96, 2013–2018. [Google Scholar] [CrossRef] [PubMed]
- Grieb, G.; Merk, M.; Bernhagen, J.; Bucala, R. Macrophage Migration Inhibitory Factor (MIF): A Promising Biomarker. Drug News Perspect. 2010, 257–264. [Google Scholar] [CrossRef] [PubMed]
- Iwama, G.K.; Thomas, P.T.; Forsyth, R.B.; Vijayan, M.M. Heat Shock Protein Expression in Fish. Rev. Fish. Biol. Fish. 1998, 8, 35–56. [Google Scholar] [CrossRef]
- Wang, G.Q.; Bonkovsky, H.L.; De Lemos, A.; Burczynski, F.J. Recent Insights into the Biological Functions of Liver Fatty Acid Binding Protein 1. J. Lipid Res. 2015, 2238–2247. [Google Scholar] [CrossRef]
- Cao, Y.; Traer, E.; Zimmerman, G.A.; McIntyre, T.M.; Prescott, S.M. Cloning, Expression, and Chromosomal Localization of Human Long-Chain Fatty Acid-CoA Ligase 4 (FACL4). Genomics 1998, 49, 327–330. [Google Scholar] [CrossRef]
- Smith, S.; Witkowski, A.; Joshi, A.K. Structural and Functional Organization of the Animal Fatty Acid Synthase. Prog. Lipid Res. 2003, 289–317. [Google Scholar] [CrossRef]
- Inpanathan, S.; Botelho, R.J. The Lysosome Signaling Platform: Adapting with the Times. Front. Cell Dev. Biol. 2019, 113. [Google Scholar] [CrossRef] [PubMed]
- Blomhoff, R.; Blomhoff, H.K. Overview of Retinoid Metabolism and Function. J. Neurobiol. 2006, 606–630. [Google Scholar] [CrossRef] [PubMed]
- Younis, S.; Schönke, M.; Massart, J.; Hjortebjerg, R.; Sundström, E.; Gustafson, U.; Björnholm, M.; Krook, A.; Frystyk, J.; Zierath, J.R.; et al. The ZBED6-IGF2 Axis Has a Major Effect on Growth of Skeletal Muscle and Internal Organs in Placental Mammals. Proc. Natl. Acad. Sci. USA 2018, 115, E2048–E2057. [Google Scholar] [CrossRef] [PubMed]




| NM2 | NM3 | NM1 | M1 | M3 | M2 | |
|---|---|---|---|---|---|---|
| Total Reads Count (#) | 52,358,598 | 50,324,500 | 43,018,686 | 46,972,334 | 46,798,082 | 68,136,198 |
| Total Bases Count (bp) | 7.61 × 109 | 7.27 × 109 | 6.23 × 109 | 6.79 × 109 | 6.81 × 109 | 9.9 × 109 |
| Average Read Length (bp) | 145.3 | 144.43 | 144.87 | 144.58 | 145.59 | 145.35 |
| Q10 Bases Count (bp) | 7.61 × 109 | 7.27 × 109 | 6.23 × 109 | 6.79 × 109 | 6.81 × 109 | 9.9 × 109 |
| Q10 Bases Ratio (%) | 100.00% | 100.00% | 100.00% | 100.00% | 100.00% | 100.00% |
| Q20 Bases Count (bp) | 7.49 × 109 | 7.16 × 109 | 6.14 × 109 | 6.69 × 109 | 6.75 × 109 | 9.77 × 109 |
| Q20 Bases Ratio (%) | 98.48% | 98.48% | 98.45% | 98.48% | 99.07% | 98.64% |
| Q30 Bases Count (bp) | 7.2 × 109 | 6.88 × 109 | 5.89 × 109 | 6.43 × 109 | 6.6 × 109 | 9.41 × 109 |
| Q30 Bases Ratio (%) | 94.63% | 94.64% | 94.54% | 94.61% | 96.88% | 95.03% |
| N Bases Count (bp) | 2331 | 2248 | 1972 | 2022 | 52894 | 5878 |
| N Bases Ratio (%) | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% | 0.00% |
| GC Bases Count (bp) | 3.72 × 109 | 3.59 × 109 | 2.97 × 109 | 3.17 × 109 | 3.19 × 109 | 4.65 × 109 |
| GC Bases Ratio (%) | 48.86% | 49.35% | 47.62% | 46.67% | 46.83% | 46.91% |
| No. | ≥500 bp | ≥1000 bp | N50 | N90 | Max Len | Min Len | Total Len | Average Len | |
|---|---|---|---|---|---|---|---|---|---|
| Transcript | 272,346 | 112,989 | 66,293 | 1806 | 307 | 20,532 | 201 | 2.41 × 108 | 884.82 |
| Unigene | 139.100 | 46,048 | 21,701 | 1111 | 263 | 20,532 | 201 | 93,974,306 | 675.59 |
| Factor Name | Number of Genes |
|---|---|
| NADH dehydrogenase | 3 |
| Cytochrome P450 | 8 |
| Thyroid hormone signaling pathway | 16 |
| Creatine kinase | 1 |
| Arginine kinase | 1 |
| C-type lectin-like domain | 1 |
| Factor Name | Number of Genes |
|---|---|
| Innate immune response | 3 |
| Lysosome | 1 |
| Toll-like receptors | 4 |
| Caspace | 3 |
| Trypsin-like serine protease | 3 |
| Peroxiredoxin | 1 |
| Thioredoxin | 3 |
| Glutaredoxin | 1 |
| C-type lectin-like | 6 |
| Factor Name | Number of Genes |
|---|---|
| MIH | 2 |
| Crustacean hyperglycemic hormone (CHH) | 2 |
| Hyperglycemic hormone-like peptide 2 precursor | 2 |
| Neuroparsin | 2 |
| Ecdysteroid regulated-like protein | 1 |
| Ecdysteroid kinase | 5 |
| Vitellogenin | 1 |
| 17-Beta-dehydrogenase 8-like isoform X1 | 3 |
| E75 Nuclear receptor | 1 |
| Factor Name | Number of Genes |
|---|---|
| C-type lectin-like | 6 |
| Manose-Binding Protein | 1 |
| Eriocheir sinensis chitin synthase gene | 1 |
| CDA like 2 | 1 |
| CBM 14 | 3 |
| Cryptocyanin 1 | 5 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Su, S.; Munganga, B.P.; Tian, C.; Li, J.; Yu, F.; Li, H.; Wang, M.; He, X.; Tang, Y. Comparative Analysis of the Intermolt and Postmolt Hepatopancreas Transcriptomes Provides Insight into the Mechanisms of Procambarus clarkii Molting Process. Life 2021, 11, 480. https://doi.org/10.3390/life11060480
Su S, Munganga BP, Tian C, Li J, Yu F, Li H, Wang M, He X, Tang Y. Comparative Analysis of the Intermolt and Postmolt Hepatopancreas Transcriptomes Provides Insight into the Mechanisms of Procambarus clarkii Molting Process. Life. 2021; 11(6):480. https://doi.org/10.3390/life11060480
Chicago/Turabian StyleSu, Shengyan, Brian Pelekelo Munganga, Can Tian, Jianlin Li, Fan Yu, Hongxia Li, Meiyao Wang, Xinjin He, and Yongkai Tang. 2021. "Comparative Analysis of the Intermolt and Postmolt Hepatopancreas Transcriptomes Provides Insight into the Mechanisms of Procambarus clarkii Molting Process" Life 11, no. 6: 480. https://doi.org/10.3390/life11060480
APA StyleSu, S., Munganga, B. P., Tian, C., Li, J., Yu, F., Li, H., Wang, M., He, X., & Tang, Y. (2021). Comparative Analysis of the Intermolt and Postmolt Hepatopancreas Transcriptomes Provides Insight into the Mechanisms of Procambarus clarkii Molting Process. Life, 11(6), 480. https://doi.org/10.3390/life11060480







