F1099L-CFTR (c.3297C>G) has Impaired Channel Function and Associates with Mild Disease Phenotypes in Two Pediatric Patients
Abstract
1. Introduction
2. Materials and Methods
2.1. Clinical Features
2.2. Antibodies and Reagents
2.3. Generation of F1099L-CFTR Mutation (cDNA Name: c.3297C>G)
2.4. Cell Culture and Plasmids Transfection
2.5. Western Blotting
2.6. Real-Time PCR to Measure CFTR mRNA Levels
2.7. Iodide (I−) Efflux Assay
2.8. Immunofluorescence Labeling and Imaging
2.9. Statistical Analysis
3. Results
3.1. Clinical Features
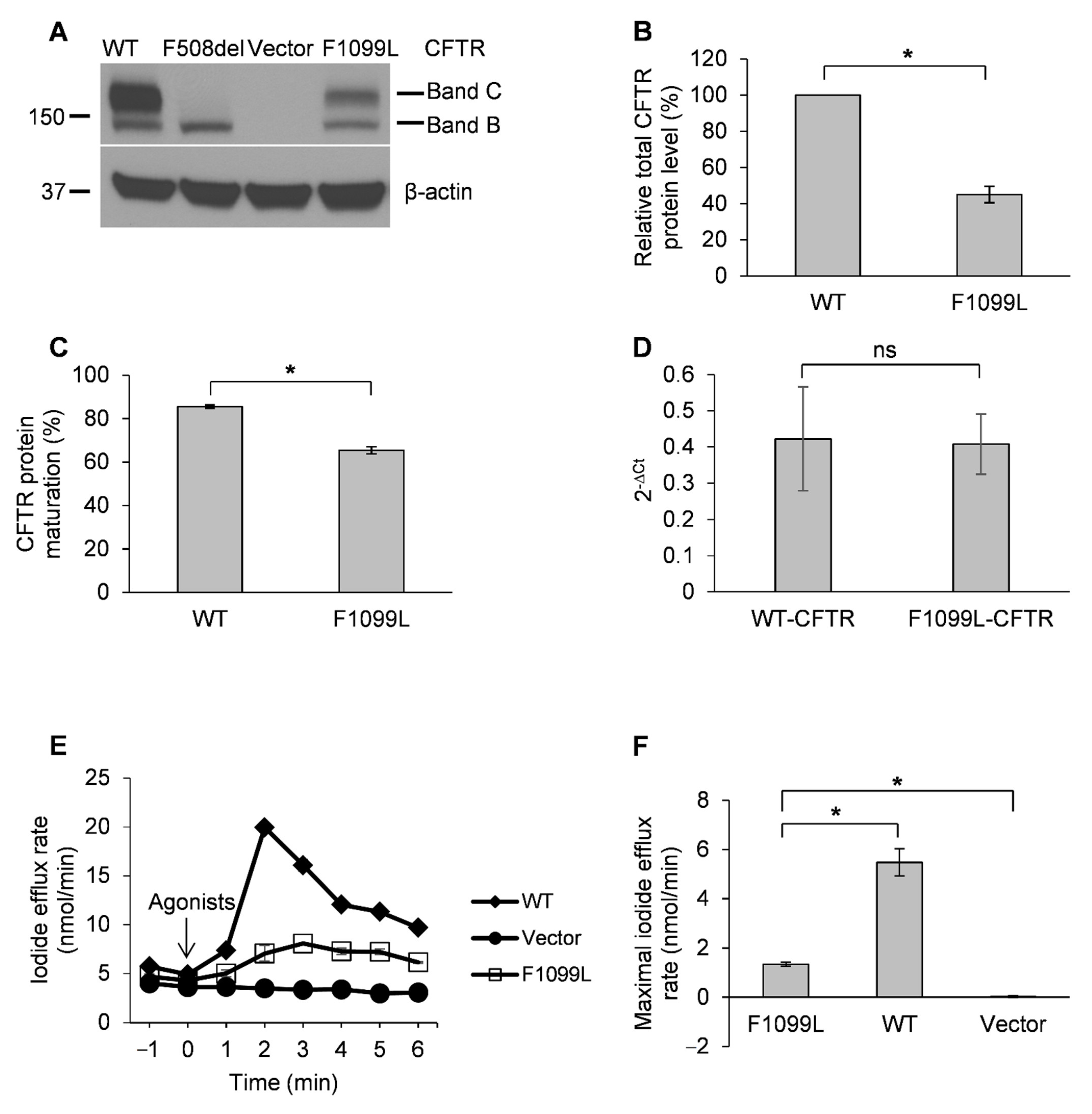
3.2. F1099L-CFTR Has a Defect in Protein Maturation and Exhibits Impaired Channel Function Compared to WT-CFRT
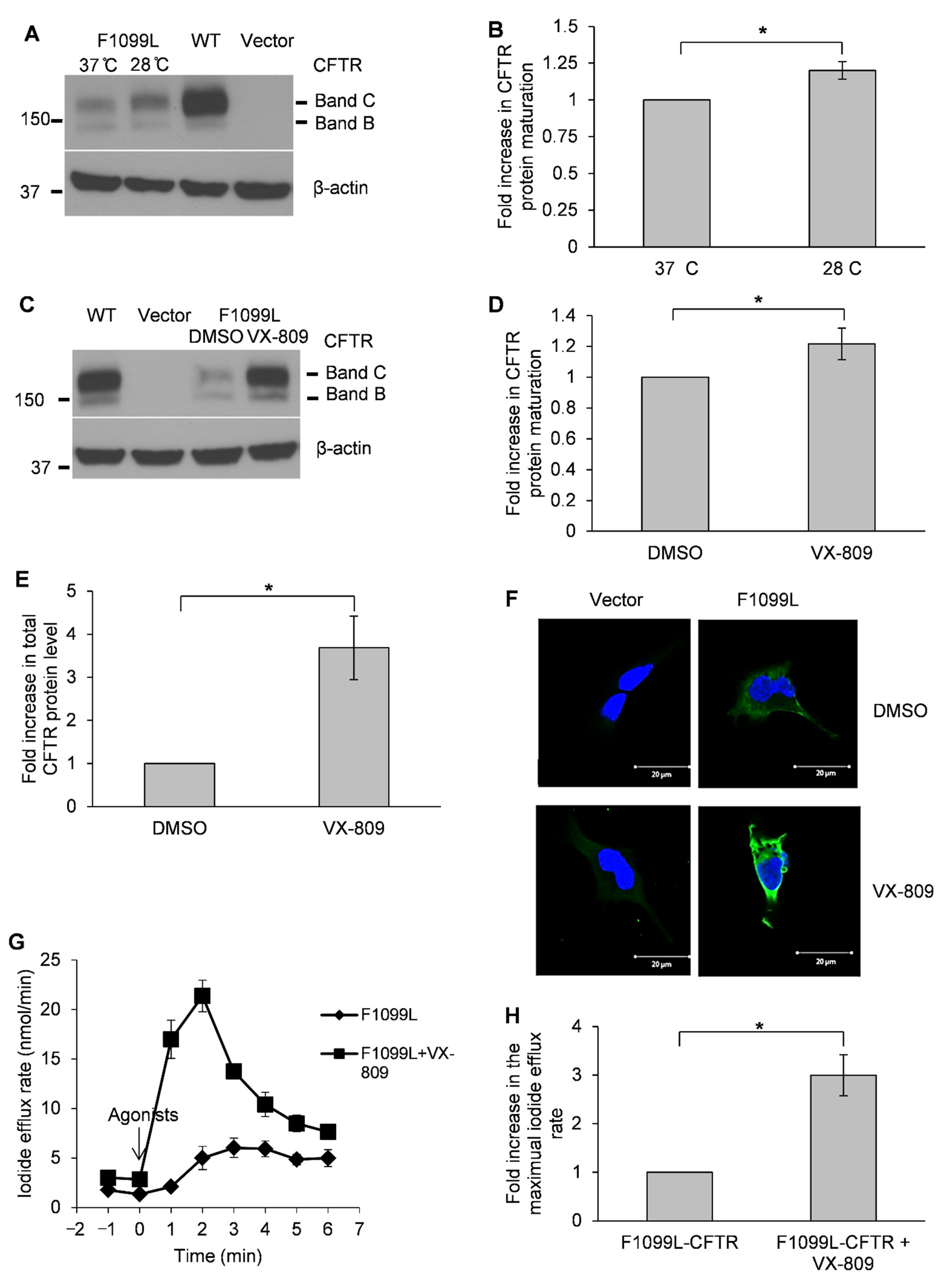
3.3. F1099L-CFTR Is a Temperature-Sensitive Mutation and Can Be Effectively Rescued by Using VX-809
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cheng, S.H.; Gregory, R.J.; Marshall, J.; Paul, S.; Souza, D.W.; White, G.A.; O’Riordan, C.R.; Smith, A.E. Defective intracellular transport and processing of CFTR is the molecular basis of most cystic fibrosis. Cell 1990, 63, 827–834. [Google Scholar] [CrossRef]
- Welsh, M.J.; Smith, A.E. Molecular mechanisms of CFTR chloride channel dysfunction in cystic fibrosis. Cell 1993, 73, 1251–1254. [Google Scholar] [CrossRef]
- Davies, J.C.; Alton, E.W.; Bush, A. Cystic fibrosis. BMJ 2007, 335, 1255–1259. [Google Scholar] [CrossRef]
- O’Sullivan, B.P.; Freedman, S.D. Cystic fibrosis. Lancet 2009, 373, 1891–1904. [Google Scholar] [CrossRef]
- Rohlfs, E.M.; Zhou, Z.; Heim, R.A.; Nagan, N.; Rosenblum, L.S.; Flynn, K.; Scholl, T.; Akmaev, V.R.; Sirko-Osadsa, D.A.; Allitto, B.A.; et al. Cystic fibrosis carrier testing in an ethnically diverse US population. Clin. Chem. 2011, 57, 841–848. [Google Scholar] [CrossRef] [PubMed]
- Sugarman, E.A.; Rohlfs, E.M.; Silverman, L.M.; Allitto, B.A. CFTR mutation distribution among Hispanic US and African American individuals: Evaluation in cystic fibrosis patient and carrier screening populations. Genet. Med. 2004, 6, 392–399. [Google Scholar] [CrossRef]
- Riordan, J.R.; Rommens, J.M.; Kerem, B.; Alon, N.; Rozmahel, R.; Grzelczak, Z.; Zielenski, J.; Lok, S.; Plavsic, N.; Chou, J.L.; et al. Identification of the cystic fibrosis gene: Cloning and characterization of complementary DNA. Science 1989, 245, 1066–1073. [Google Scholar] [CrossRef] [PubMed]
- Anderson, M.P.; Gregory, R.J.; Thompson, S.; Souza, D.W.; Paul, S.; Mulligan, R.C.; Smith, A.E.; Welsh, M.J. Demonstration that CFTR is a chloride channel by alteration of its anion selectivity. Science 1991, 253, 202–205. [Google Scholar] [CrossRef] [PubMed]
- Bear, C.E.; Li, C.H.; Kartner, N.; Bridges, R.J.; Jensen, T.J.; Ramjeesingh, M.; Riordan, J.R. Purification and functional reconstitution of the cystic fibrosis transmembrane conductance regulator (CFTR). Cell 1992, 68, 809–818. [Google Scholar] [CrossRef]
- Hadida, S.; Van Goor, F.; Grootenhuis, P.D.J. CFTR modulators for the treatment of cystic fibrosis. In Annual Reports in Medicinal Chemistry; Macor, J.E., Ed.; Elsevier Inc.: Amsterdam, The Netherlands, 2010; Volume 45, pp. 157–173. [Google Scholar]
- Cystic Fibrosis Mutation Database. Available online: http://www.genet.sickkids.on.ca/StatisticsPage.html (accessed on 14 December 2020).
- Clancy, J.P.; Jain, M. Personalized medicine in cystic fibrosis: Dawning of a new era. Am. J. Respir. Crit. Care Med. 2012, 186, 593–597. [Google Scholar] [CrossRef]
- Bell, S.C.; De Boeck, K.; Amaral, M.D. New pharmacological approaches for cystic fibrosis: Promises, progress, pitfalls. Pharm. Ther. 2015, 145, 19–34. [Google Scholar] [CrossRef]
- Veit, G.; Avramescu, R.G.; Chiang, A.N.; Houck, S.A.; Cai, Z.; Peters, K.W.; Hong, J.S.; Pollard, H.B.; Guggino, W.B.; Balch, W.E.; et al. From CFTR biology toward combinatorial pharmacotherapy: Expanded classification of cystic fibrosis mutations. Mol. Biol. Cell 2016, 27, 424–433. [Google Scholar] [CrossRef]
- Lopes-Pacheco, M. CFTR Modulators: The Changing Face of Cystic Fibrosis in the Era of Precision Medicine. Front. Pharmacol. 2020, 10, 1662. [Google Scholar] [CrossRef]
- Cutting, G.R. Cystic fibrosis genetics: From molecular understanding to clinical application. Nat. Rev. Genet. 2015, 16, 45–56. [Google Scholar] [CrossRef]
- U.S. Cystic Fibrosis Foundation. Drug Development Pipeline. Available online: https://www.cff.org/trials/pipeline (accessed on 14 December 2020).
- Ikpa, P.T.; Bijvelds, M.J.; de Jonge, H.R. Cystic fibrosis: Toward personalized therapies. Int. J. Biochem. Cell Biol. 2014, 52, 192–200. [Google Scholar] [CrossRef] [PubMed]
- Clinical and Functional Translation of CFTR (CFTR2). Available online: https://cftr2.org/mutation/general/F1099L/ (accessed on 20 December 2020).
- McGinniss, M.J.; Chen, C.; Redman, J.B.; Buller, A.; Quan, F.; Peng, M.; Giusti, R.; Hantash, F.M.; Huang, D.; Sun, W.; et al. Extensive sequencing of the CFTR gene: Lessons learned from the first 157 patient samples. Hum. Genet. 2005, 118, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Degrugillier, F.; Simon, S.; Aissat, A.; Remus, N.; Mekki, C.; Decrouy, X.; Hatton, A.; Hinzpeter, A.; Hoffmann, B.; Sermet-Gaudelus, I.; et al. Unsolved severe chronic rhinosinusitis elucidated by extensive CFTR genotyping. Clin. Case Rep. 2019, 7, 2128–2134. [Google Scholar] [CrossRef] [PubMed]
- Raraigh, K.S.; Han, S.T.; Davis, E.; Evans, T.A.; Pellicore, M.J.; McCague, A.F.; Joynt, A.T.; Lu, Z.; Atalar, M.; Sharma, N.; et al. Functional Assays Are Essential for Interpretation of Missense Variants Associated with Variable Expressivity. Am. J. Hum. Genet. 2018, 102, 1062–1077. [Google Scholar] [CrossRef]
- Ren, H.Y.; Grove, D.E.; De La Rosa, O.; Houck, S.A.; Sopha, P.; Van Goor, F.; Hoffman, B.J.; Cyr, D.M. VX-809 corrects folding defects in cystic fibrosis transmembrane conductance regulator protein through action on membrane-spanning domain 1. Mol. Biol. Cell 2013, 24, 3016–3024. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Hothi, J.S.; Zhang, Y.H.; Srinivasan, S.; Stokes, D.C.; Zhang, W. c.3623G>A mutation encodes a CFTR protein with impaired channel function. Respir. Res. 2016, 17, 8. [Google Scholar] [CrossRef]
- Cystic Fibrosis Mutation Database. Available online: http://www.genet.sickkids.on.ca/CftrDomainPage.html (accessed on 26 January 2021).
- Cotton, J.F.; Ostedgaard, L.S.; Carson, M.R.; Welsh, M.J. Effect of cystic fibrosis-associated mutations in the fourth intracellular loop of cystic fibrosis transmembrane conductance regulator. J. Biol. Chem. 1996, 271, 21279–21284. [Google Scholar] [CrossRef]
- Van Goor, F.; Yu, H.; Burton, B.; Hoffman, B.J. Effect of ivacaftor on CFTR forms with missense mutations associated with defects in protein processing or function. J. Cyst. Fibros. 2014, 13, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Hwang, T.C.; Yeh, J.T.; Zhang, J.; Yu, Y.C.; Yeh, H.I.; Destefano, S. Structural mechanisms of CFTR function and dysfunction. J. Gen. Physiol. 2018, 150, 539–570. [Google Scholar] [CrossRef]
- Ramsey, B.W.; Banks-Schlegel, S.; Accurso, F.J.; Boucher, R.C.; Cutting, G.R.; Engelhardt, J.F.; Guggino, W.B.; Karp, C.L.; Knowles, M.R.; Kolls, J.K.; et al. Future directions in early cystic fibrosis lung disease research: An NHLBI workshop report. Am. J. Respir. Crit. Care Med. 2012, 185, 887–892. [Google Scholar] [CrossRef] [PubMed]
- Corvol, H.; Blackman, S.M.; Boëlle, P.Y.; Gallins, P.J.; Pace, R.G.; Stonebraker, J.R.; Accurso, F.J.; Clement, A.; Collaco, J.M.; Dang, H.; et al. Genome-wide association meta-analysis identifies five modifier loci of lung disease severity in cystic fibrosis. Nat. Commun. 2015, 6, 8382. [Google Scholar] [CrossRef] [PubMed]
- Schechter, M.S.; Shelton, B.J.; Margolis, P.A.; Fitzsimmons, S.C. The association of socioeconomic status with outcomes in cystic fibrosis patients in the United States. Am. J. Respir. Crit. Care Med. 2001, 163, 1331–1337. [Google Scholar] [CrossRef]
- Kopp, B.T.; Sarzynski, L.; Khalfoun, S.; Hayes, D., Jr.; Thompson, R.; Nicholson, L.; Long, F.; Castile, R.; Groner, J. Detrimental effects of secondhand smoke exposure on infants with cystic fibrosis. Pediatr. Pulmonol. 2015, 50, 25–34. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, X.; Hothi, J.S.; Zhang, Y.H.; Ren, A.; Rock, M.J.; Srinivasan, S.; Stokes, D.C.; Naren, A.P.; Zhang, W. F1099L-CFTR (c.3297C>G) has Impaired Channel Function and Associates with Mild Disease Phenotypes in Two Pediatric Patients. Life 2021, 11, 131. https://doi.org/10.3390/life11020131
Zhang X, Hothi JS, Zhang YH, Ren A, Rock MJ, Srinivasan S, Stokes DC, Naren AP, Zhang W. F1099L-CFTR (c.3297C>G) has Impaired Channel Function and Associates with Mild Disease Phenotypes in Two Pediatric Patients. Life. 2021; 11(2):131. https://doi.org/10.3390/life11020131
Chicago/Turabian StyleZhang, Xiaoying, Jaspal S. Hothi, Yanhui H. Zhang, Aixia Ren, Michael J. Rock, Saumini Srinivasan, Dennis C. Stokes, Anjaparavanda P. Naren, and Weiqiang Zhang. 2021. "F1099L-CFTR (c.3297C>G) has Impaired Channel Function and Associates with Mild Disease Phenotypes in Two Pediatric Patients" Life 11, no. 2: 131. https://doi.org/10.3390/life11020131
APA StyleZhang, X., Hothi, J. S., Zhang, Y. H., Ren, A., Rock, M. J., Srinivasan, S., Stokes, D. C., Naren, A. P., & Zhang, W. (2021). F1099L-CFTR (c.3297C>G) has Impaired Channel Function and Associates with Mild Disease Phenotypes in Two Pediatric Patients. Life, 11(2), 131. https://doi.org/10.3390/life11020131





