Population Genomics Insights into the First Wave of COVID-19
Abstract
1. Introduction
2. Materials and Methods
2.1. Mutation Rate Analysis and Estimation of the Time of the Most Recent Common Ancestor between Bat CoV and SARS-CoV-2
2.2. Recombination Analysis of Nonhuman Betacoronaviruses That Have Contributed to the SARS-CoV-2 Evolution and between the Human SARS-CoV-2 Genomes
2.3. Linkage Disequilibrium (LD) in Human SARS-CoV-2 Genomes
2.4. Selective Sweeps and Common Outliers
2.5. Estimation of the Time of the Most Recent Common Ancestor
2.6. Demographic Inference
3. Results
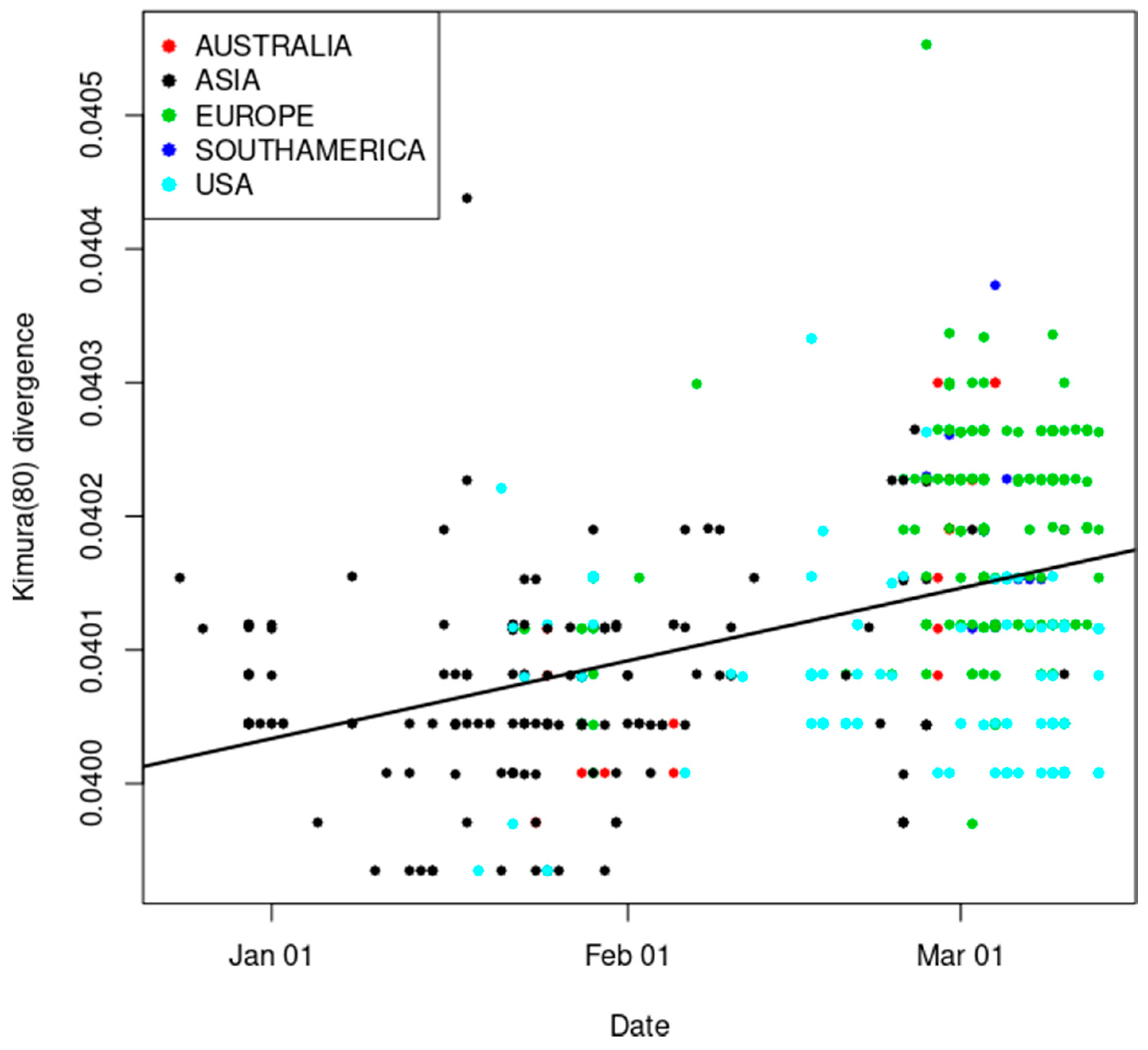
3.1. Estimation of Mutation Rate and Divergence from Bat CoV
3.2. Recombination Events
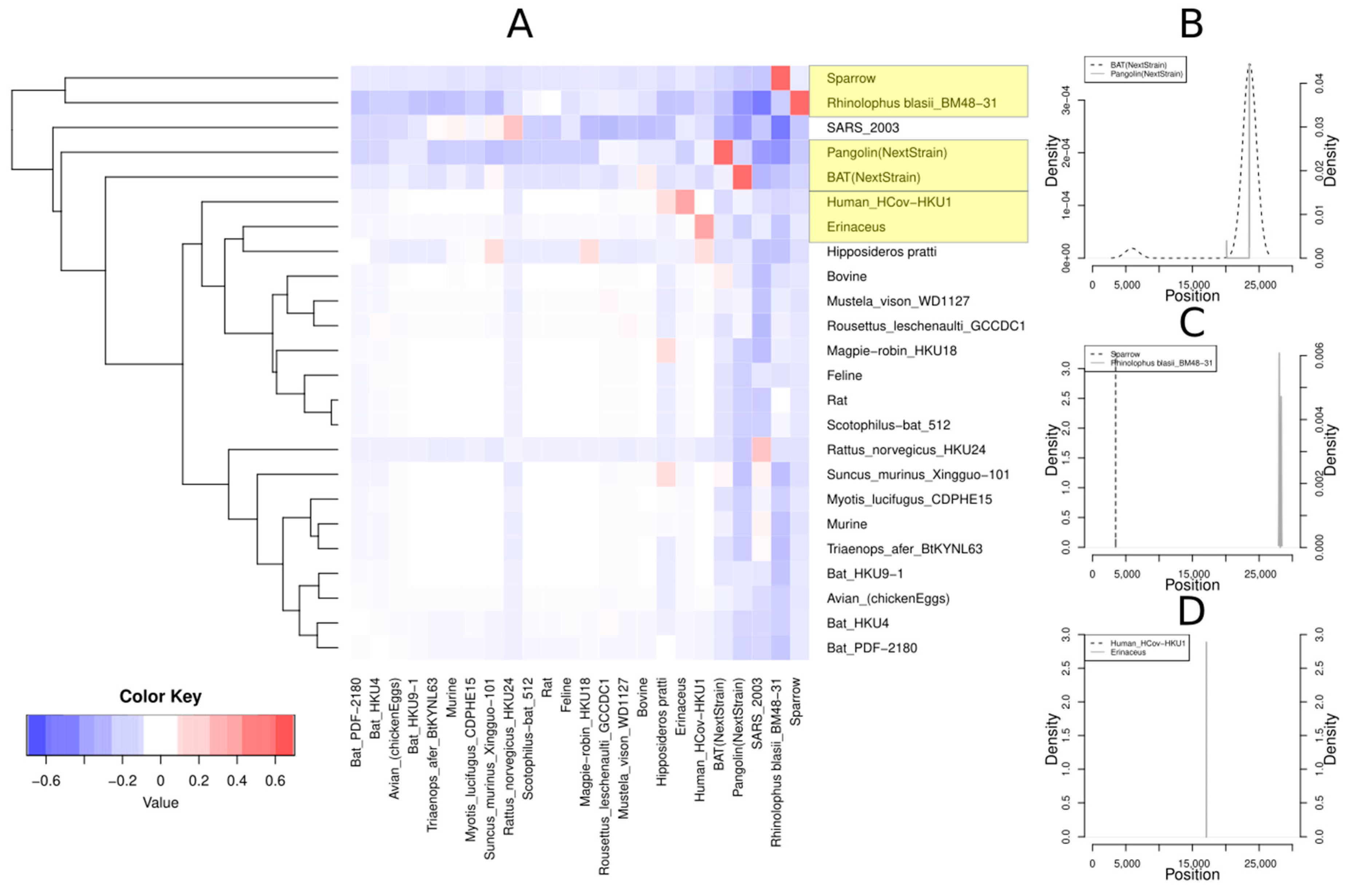
3.2.1. Host Analysis
3.2.2. Localization of Recombination Events
3.3. Detection of Recombination Events amongst Human SARS-CoV-2 Genomes
3.4. Linkage Disequilibrium (LD) Analysis
3.5. Selective Sweeps
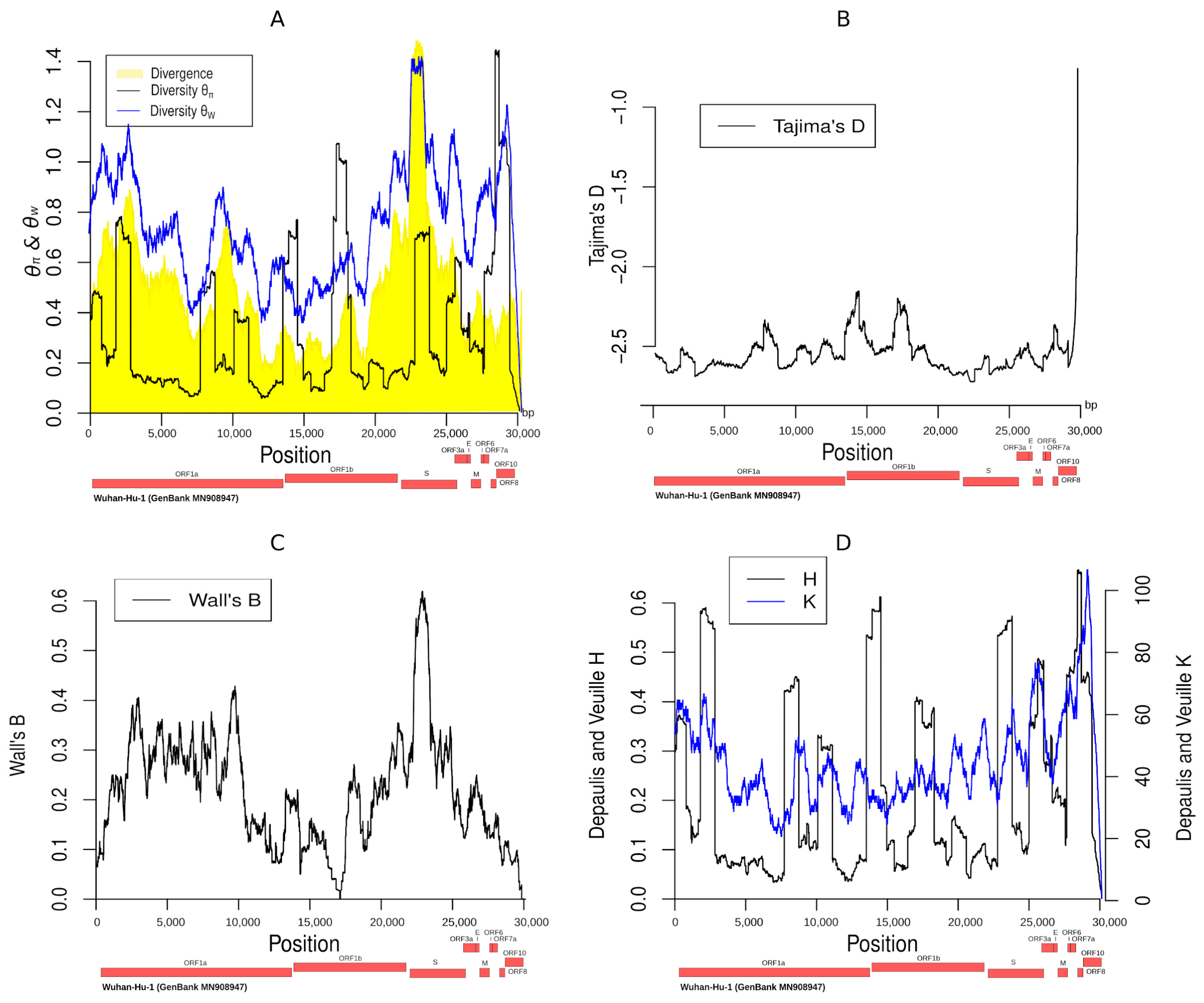
3.6. Summary Statistics along the SARS-CoV-2 Genome
3.7. Estimation of the Time of the Most Recent Common Ancestor
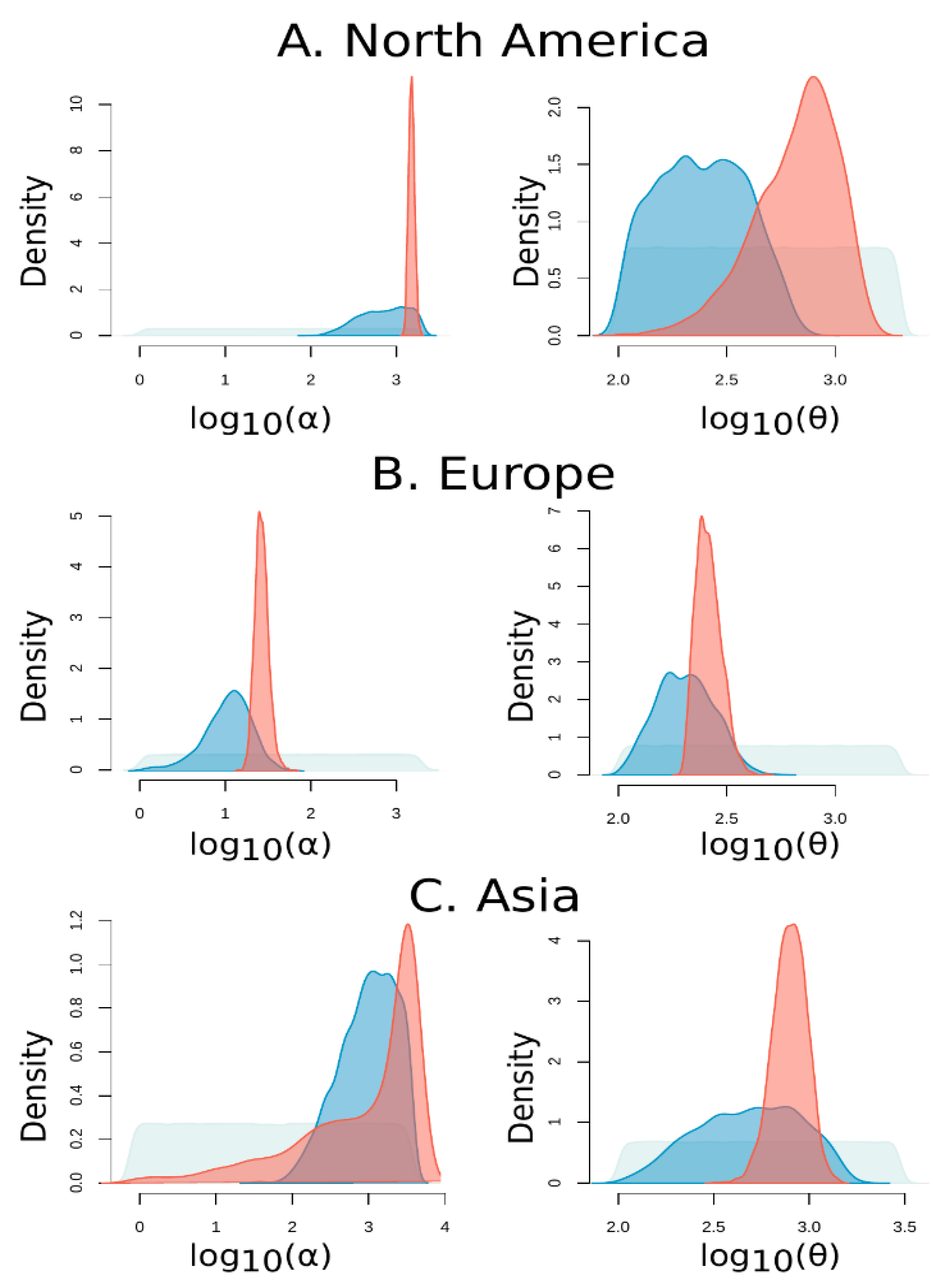
3.8. Demographic Inference
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species Severe acute respiratory syndrome-related coronavirus: Classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 2020, 5, 536–544. [Google Scholar] [CrossRef]
- Rota, P.A.; Oberste, M.S.; Monroe, S.S.; Nix, W.A.; Campagnoli, R.; Icenogle, J.P.; Peñaranda, S.; Bankamp, B.; Maher, K.; Chen, M.H.; et al. Characterization of a novel coronavirus associated with severe acute respiratory syndrome. Science 2003, 300, 1394–1399. [Google Scholar] [CrossRef]
- Zaki, A.M.; Van Boheemen, S.; Bestebroer, T.M.; Osterhaus, A.D.M.E.; Fouchier, R.A.M. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 2012, 367, 1814–1820. [Google Scholar] [CrossRef]
- Holshue, M.L.; DeBolt, C.; Lindquist, S.; Lofy, K.H.; Wiesman, J.; Bruce, H.; Spitters, C.; Ericson, K.; Wilkerson, S.; Tural, A.; et al. First Case of 2019 Novel Coronavirus in the United States. N. Engl. J. Med. 2020, 382, 929–936. [Google Scholar] [CrossRef]
- Andersen, K.G.; Rambaut, A.; Lipkin, W.I.; Holmes, E.C.; Garry, R.F. The proximal origin of SARS-CoV-2. Nat. Med. 2020, 26, 450–452. [Google Scholar] [CrossRef]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 2020, 382, 727–733. [Google Scholar] [CrossRef]
- Gwinn, M.; MacCannell, D.; Armstrong, G.L. Next-Generation Sequencing of Infectious Pathogens. JAMA 2019, 321, 893. [Google Scholar] [CrossRef] [PubMed]
- Moya, A.; Holmes, E.C.; González-Candelas, F. The population genetics and evolutionary epidemiology of RNA viruses. Nat. Rev. Microbiol. 2004, 2, 279–288. [Google Scholar] [CrossRef]
- Combelas, N.; Holmblat, B.; Joffret, M.-L.; Colbère-Garapin, F.; Delpeyroux, F. Recombination between Poliovirus and Coxsackie A Viruses of Species C: A Model of Viral Genetic Plasticity and Emergence. Viruses 2011, 3, 1460–1484. [Google Scholar] [CrossRef]
- Pérez-Losada, M.; Arenas, M.; Galán, J.C.; Palero, F.; González-Candelas, F. Recombination in viruses: Mechanisms, methods of study, and evolutionary consequences. Infect. Genet. Evol. 2015, 30, 296–307. [Google Scholar] [CrossRef]
- Makarenkov, V.; Mazoure, B.; Rabusseau, G.; Legendre, P. Horizontal gene transfer and recombination analysis of SARS-CoV-2 genes helps discover its close relatives and shed light on its origin. BMC Ecol. Evol. 2021, 21, 5. [Google Scholar]
- Boni, M.F.; Lemey, P.; Jiang, X.; Lam, T.T.-Y.; Perry, B.; Castoe, T.; Rambaut, A.; Robertson, D.L. Evolutionary origins of the SARS-CoV-2 sarbecovirus lineage responsible for the COVID-19 pandemic. bioRxiv 2020. [Google Scholar] [CrossRef]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef]
- Liu, P.; Chen, W.; Chen, J.P. Viral metagenomics revealed sendai virus and coronavirus infection of malayan pangolins (manis javanica). Viruses 2019, 11, 979. [Google Scholar] [CrossRef]
- Lam, T.T.-Y.; Shum, M.H.-H.; Zhu, H.-C.; Tong, Y.-G.; Ni, X.-B.; Liao, Y.-S.; Wei, W.; Cheung, W.Y.-M.; Li, W.-J.; Li, L.-F.; et al. Identification of 2019-nCoV related coronaviruses in Malayan pangolins in southern China. bioRxiv 2020. [Google Scholar] [CrossRef]
- Touati, R.; Haddad-Boubaker, S.; Ferchichi, I.; Messaoudi, I.; Ouesleti, A.E.; Triki, H.; Lachiri, Z.; Kharrat, M. Comparative genomic signature representations of the emerging COVID-19 coronavirus and other coronaviruses: High identity and possible recombination between Bat and Pangolin coronaviruses. Genomics 2020. [Google Scholar] [CrossRef]
- Flores-Alanis, A.; Sandner-Miranda, L.; Delgado, G.; Cravioto, A.; Morales-Espinosa, R. The receptor binding domain of SARS-CoV-2 spike protein is the result of an ancestral recombination between the bat-CoV RaTG13 and the pangolin-CoV MP789. BMC Res. Notes 2020, 13, 398. [Google Scholar] [CrossRef]
- Karamitros, T.; Papadopoulou, G.; Bousali, M.; Mexias, A.; Tsiodras, S.; Mentis, A. SARS-CoV-2 exhibits intra-host genomic plasticity and low-frequency polymorphic quasispecies. J. Clin. Virol. 2020, 131, 104585. [Google Scholar] [CrossRef] [PubMed]
- Paraskevis, D.; Kostaki, E.G.; Magiorkinis, G.; Panayiotakopoulos, G.; Sourvinos, G.; Tsiodras, S. Full-genome evolutionary analysis of the novel corona virus (2019-nCoV) rejects the hypothesis of emergence as a result of a recent recombination event. Infect. Genet. Evol. 2020, 79, 104212. [Google Scholar] [CrossRef]
- Gallaher, W.R. A palindromic RNA sequence as a common breakpoint contributor to copy-choice recombination in SARS-COV-2. Arch. Virol. 2020, 165, 2341–2348. [Google Scholar] [CrossRef]
- Lau, S.K.P.; Luk, H.K.H.; Wong, A.C.P.; Li, K.S.M.; Zhu, L.; He, Z.; Fung, J.; Chan, T.T.Y.; Fung, K.S.C.; Woo, P.C.Y. Possible Bat Origin of Severe Acute Respiratory Syndrome Coronavirus 2. Emerg. Infect. Dis. 2020, 26, 1542–1547. [Google Scholar] [CrossRef]
- Li, X.; Giorgi, E.E.; Marichannegowda, M.H.; Foley, B.; Xiao, C.; Kong, X.-P.; Chen, Y.; Gnanakaran, S.; Korber, B.; Gao, F. Emergence of SARS-CoV-2 through recombination and strong purifying selection. Sci. Adv. 2020, 6. [Google Scholar] [CrossRef]
- Beaumont, M.A.; Zhang, W.; Balding, D.J. Approximate Bayesian computation in population genetics. Genetics 2002, 162, 2025–2035. [Google Scholar]
- Vijaykrishna, D.; Smith, G.J.D.; Zhang, J.X.; Peiris, J.S.M.; Chen, H.; Guan, Y. Evolutionary insights into the ecology of coronaviruses. J. Virol. 2007, 81, 4012–4020. [Google Scholar] [CrossRef]
- Maynard Smith, J.; Haigh, J. The hitch-hiking effect of a favourable gene. Genet. Res. 1974, 89, 391–403. [Google Scholar] [CrossRef]
- Stephan, W.; Wiehe, T.H.E.; Lenz, M.W. The effect of strongly selected substitutions on neutral polymorphism: Analytical results based on diffusion theory. Theor. Popul. Biol. 1992, 41, 237–254. [Google Scholar] [CrossRef]
- Kim, Y.; Stephan, W. Detecting a local signature of genetic hitchhiking along a recombining chromosome. Genetics 2002, 160, 765–777. [Google Scholar]
- GISAID—Initiative. Available online: https://www.gisaid.org/ (accessed on 4 February 2020).
- Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Kimura, M. The number of heterozygous nucleotide sites maintained in a finite population due to steady flux of mutations. Genetics 1969, 61, 893–903. [Google Scholar] [CrossRef] [PubMed]
- Hill, W.G.; Robertson, A. Linkage disequilibrium in finite populations. Theor. Appl. Genet. 1968, 38, 226–231. [Google Scholar] [CrossRef]
- Pavlidis, P.; Živković, D.; Stamatakis, A.; Alachiotis, N. SweeD: Likelihood-based detection of selective sweeps in thousands of genomes. Mol. Biol. Evol. 2013, 30, 2224–2234. [Google Scholar] [CrossRef]
- Alachiotis, N.; Pavlidis, P. RAiSD detects positive selection based on multiple signatures of a selective sweep and SNP vectors. Commun. Biol. 2018, 1, 79. [Google Scholar] [CrossRef]
- Pavlidis, P.; Hutter, S.; Stephan, W. A population genomic approach to map recent positive selection in model species. Mol. Ecol. 2008, 17, 3585–3598. [Google Scholar] [CrossRef] [PubMed]
- Csilléry, K.; Blum, M.G.B.; Gaggiotti, O.E.; François, O. Approximate Bayesian Computation (ABC) in practice. Trends Ecol. Evol. 2010, 25, 410–418. [Google Scholar] [CrossRef]
- Pavlidis, P.; Laurent, S.; Stephan, W. msABC: A modification of Hudson’s ms to facilitate multi-locus ABC analysis. Mol. Ecol. Resour. 2010, 10, 723–727. [Google Scholar] [CrossRef]
- Watterson, G.A. On the number of segregating sites in genetical models without recombination. Theor. Popul. Biol. 1975, 7, 256–276. [Google Scholar] [CrossRef]
- Tajima, F. Evolutionary relationship of DNA sequences in finite populations. Genetics 1983, 105, 437–460. [Google Scholar] [CrossRef]
- Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 1989, 123, 585–595. [Google Scholar] [CrossRef]
- Kelly, J.K. A test of neutrality based on interlocus associations. Genetics 1997, 146, 1197–1206. [Google Scholar] [CrossRef]
- Depaulis, F.; Veuille, M. Neutrality tests based on the distribution of haplotypes under an infinite-site model. Mol. Biol. Evol. 1998, 15, 1788–1790. [Google Scholar] [CrossRef][Green Version]
- Thomson, R.; Pritchard, J.K.; Shen, P.; Oefner, P.J.; Feldman, M.W. Recent common ancestry of human Y chromosomes: Evidence from DNA sequence data. Proc. Natl. Acad. Sci. USA 2000, 97, 7360–7365. [Google Scholar] [CrossRef]
- Hudson, R.R. The variance of coalescent time estimates from DNA sequences. J. Mol. Evol. 2007, 64, 702–705. [Google Scholar] [CrossRef]
- Hein, J.; Schierup, M.; Wiuf, C. Gene Genalogies Variation and Evolution; Oxford University Press: New York, NY, USA, 2005; ISBN 9780198529958. [Google Scholar]
- Csilléry, K.; François, O.; Blum, M.G.B. abc: An R package for approximate Bayesian computation (ABC). Methods Ecol. Evol. 2012, 3, 475–479. [Google Scholar] [CrossRef]
- Blum, M.G.B.; François, O. Non-linear regression models for Approximate Bayesian Computation. Stat. Comput. 2010, 20, 63–73. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Thornton, K. libsequence: A C++ class library for evolutionary genetic analysis. Bioinformatics 2003, 19, 2325–2327. [Google Scholar] [CrossRef] [PubMed]
- NCBI. Available online: https://www.ncbi.nlm.nih.gov/genomes/GenomesGroup.cgi?taxid=11118 (accessed on 4 February 2020).
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.R.; Bender, D.; Maller, J.; Sklar, P.; De Bakker, P.I.W.; Daly, M.J.; et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef]
- Tai, W.; He, L.; Zhang, X.; Pu, J.; Voronin, D.; Jiang, S.; Zhou, Y.; Du, L. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: Implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell. Mol. Immunol. 2020. [Google Scholar] [CrossRef]
- Wall, J.D. Recombination and the power of statistical tests of neutrality. Genet. Res. 1999, 74, 65–79. [Google Scholar] [CrossRef]
- Li, X.; Zai, J.; Zhao, Q.; Nie, Q.; Li, Y.; Foley, B.T.; Chaillon, A. Evolutionary history, potential intermediate animal host, and cross-species analyses of SARS-CoV-2. J. Med. Virol. 2020, 92, 602–611. [Google Scholar] [CrossRef]
- Liu, Q.; Zhao, S.; Shi, C.-M.; Song, S.; Zhu, S.; Su, Y.; Zhao, W.; Li, M.; Bao, Y.; Xue, Y.; et al. Population Genetics of SARS-CoV-2: Disentangling Effects of Sampling Bias and Infection Clusters. Genom. Proteom. Bioinform. 2020. [Google Scholar] [CrossRef]
- Zhao, Z.; Li, H.; Wu, X.; Zhong, Y.; Zhang, K.; Zhang, Y.P.; Boerwinkle, E.; Fu, Y.X. Moderate mutation rate in the SARS coronavirus genome and its implications. BMC Evol. Biol. 2004, 4, 1–9. [Google Scholar] [CrossRef]
- Bar-On, Y.M.; Flamholz, A.; Phillips, R.; Milo, R. SARS-CoV-2 (COVID-19) by the numbers. Elife 2020, 9. [Google Scholar] [CrossRef]
- Shen, Z.; Xiao, Y.; Kang, L.; Ma, W.; Shi, L.; Zhang, L.; Zhou, Z.; Yang, J.; Zhong, J.; Yang, D.; et al. Genomic diversity of SARS-CoV-2 in Coronavirus Disease 2019 patients. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Benvenuto, D.; Giovanetti, M.; Salemi, M.; Prosperi, M.; De Flora, C.; Junior Alcantara, L.C.; Angeletti, S.; Ciccozzi, M. The global spread of 2019-nCoV: A molecular evolutionary analysis. Pathog. Glob. Health 2020, 114, 64–67. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Li, Z.; Cui, X.; Wu, C. Bayesian phylodynamic inference on the temporal evolution and global transmission of SARS-CoV-2. J. Infect. 2020, 81, 318–356. [Google Scholar] [CrossRef] [PubMed]
- Graham, R.L.; Baric, R.S. Recombination, reservoirs, and the modular spike: Mechanisms of coronavirus cross-species transmission. J. Virol. 2010, 84, 3134–3146. [Google Scholar] [CrossRef]
- Mirza, M.U.; Froeyen, M. Structural elucidation of SARS-CoV-2 vital proteins: Computational methods reveal potential drug candidates against main protease, Nsp12 polymerase and Nsp13 helicase. J. Pharm. Anal. 2020, 10, 320–328. [Google Scholar] [CrossRef]
- Forni, D.; Cagliani, R.; Mozzi, A.; Pozzoli, U.; Al-Daghri, N.; Clerici, M.; Sironi, M. Extensive Positive Selection Drives the Evolution of Nonstructural Proteins in Lineage C Betacoronaviruses. J. Virol. 2016, 90, 3627–3639. [Google Scholar] [CrossRef]
- Cagliani, R.; Forni, D.; Clerici, M.; Sironi, M. Computational Inference of Selection Underlying the Evolution of the Novel Coronavirus, Severe Acute Respiratory Syndrome Coronavirus 2. J. Virol. 2020, 94. [Google Scholar] [CrossRef] [PubMed]
- Neuman, B.W. Bioinformatics and functional analyses of coronavirus nonstructural proteins involved in the formation of replicative organelles. Antiviral Res. 2016, 135, 97–107. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhai, X.; Zhou, J. Snapshot of the evolution and mutation patterns of SARS-CoV-2. bioRxiv 2020. [Google Scholar] [CrossRef]
- Chaw, S.-M.; Tai, J.-H.; Chen, S.-L.; Hsieh, C.-H.; Chang, S.-Y.; Yeh, S.-H.; Yang, W.-S.; Chen, P.-J.; Wang, H.-Y. The origin and underlying driving forces of the SARS-CoV-2 outbreak. J. Biomed. Sci. 2020, 27, 73. [Google Scholar] [CrossRef] [PubMed]
- Dearlove, B.; Lewitus, E.; Bai, H.; Li, Y.; Reeves, D.B.; Joyce, M.G.; Scott, P.T.; Amare, M.F.; Vasan, S.; Michael, N.L.; et al. A SARS-CoV-2 vaccine candidate would likely match all currently circulating variants. Proc. Natl. Acad. Sci. USA 2020, 117, 23652–23662. [Google Scholar] [CrossRef] [PubMed]
- Yu, W.-B.; Tang, G.-D.; Zhang, L.; Corlett, R.T. Decoding the evolution and transmissions of the novel pneumonia coronavirus (SARS-CoV-2 / HCoV-19) using whole genomic data. Zool. Res. 2020, 41, 247–257. [Google Scholar] [CrossRef]
| Parameter | Prior Distribution | ||
|---|---|---|---|
| Minimum | Maximum | Distribution | |
| Northern American and European population | |||
| θ | 100 | 2000 | Log Uniform |
| α | 1 | 2000 | Log Uniform |
| Asian population | |||
| θ | 100 | 3000 | Log Uniform |
| α | 1 | 5000 | Log Uniform |
| Population (No. of Sequences) | SweeD Outlier Region | RAiSD Outlier Region | Description |
|---|---|---|---|
| All (1601) | 5276–5463 | 5048–5679 | Non-structural protein 3 (nsp3) |
| 7102–7470 | 7380–7842 | Non-structural protein 3 (nsp3) | |
| 23,043–23,073 | 23,440–23,469 | Spike protein S1/RBD | |
| 24,749–24,773 | 24,762–24,814 | Spike protein S2 | |
| Europe (811) | 5094–5352 | 4695–5089 | Non-structural protein 3 (nsp3) |
| 21,520–21,550 | 21,157–21,339 | Non-structural protein 16 (nsp16) | |
| 22,463–22,967 | 22,687–22,852 | Spike protein S1 | |
| Asia (385) | 7356–7815 | 7578–8182 | Non-structural protein 3 (nsp3) |
| 16,057–17,022 | 15,925–16,714 | nsp12/nsp13 | |
| North America (600) | 8187–8337 | 7969–8220 | nsp3 |
| 12,523 | 12,450–12,923 | nsp8/nsp9 | |
| 13,566–13,691 | 13,192–13,297 | nsp12/nsp10 | |
| 15,147–15,201 | 15,248–15,383 | nsp12 | |
| 23,166 | 23,398–23,427 | Spike protein S1 |
| North America | Asia | Europe | ||||
|---|---|---|---|---|---|---|
| α | θ | α | θ | α | θ | |
| Weighted Mean 2.5% Percentile | 1299.12 | 235.37 | 12.91 | 523.95 | 19.26 | 211.95 |
| Weighted Median | 1500.96 | 709.24 | 2310.46 | 794.61 | 26.21 | 255.9 |
| Weighted Mean | 1509.05 | 717.89 | 2508.8 | 807.84 | 26.85 | 260.3 |
| Weighted Mode | 1507.6 | 733.87 | 4869.54 | 738.66 | 24.81 | 241.78 |
| Weighted Mean 97.5% Percentile | 1766.13 | 1300.2 | 4999.94 | 1147.67 | 38.63 | 333.24 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vasilarou, M.; Alachiotis, N.; Garefalaki, J.; Beloukas, A.; Pavlidis, P. Population Genomics Insights into the First Wave of COVID-19. Life 2021, 11, 129. https://doi.org/10.3390/life11020129
Vasilarou M, Alachiotis N, Garefalaki J, Beloukas A, Pavlidis P. Population Genomics Insights into the First Wave of COVID-19. Life. 2021; 11(2):129. https://doi.org/10.3390/life11020129
Chicago/Turabian StyleVasilarou, Maria, Nikolaos Alachiotis, Joanna Garefalaki, Apostolos Beloukas, and Pavlos Pavlidis. 2021. "Population Genomics Insights into the First Wave of COVID-19" Life 11, no. 2: 129. https://doi.org/10.3390/life11020129
APA StyleVasilarou, M., Alachiotis, N., Garefalaki, J., Beloukas, A., & Pavlidis, P. (2021). Population Genomics Insights into the First Wave of COVID-19. Life, 11(2), 129. https://doi.org/10.3390/life11020129








