A Single Faecal Microbiota Transplantation Altered the Microbiota of Weaned Pigs
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Design and Sample Collection
- FMT: Faecal transplant administered two days post weaning (n = 21; FMT)
- Control: Saline administered two days post weaning (n = 21; CON)
2.2. DNA Extraction and 16S rRNA Amplicon Sequencing
2.3. Statistical Methods
3. Results
3.1. Production Characteristics
3.2. Age Related Effects
3.3. Donor Sample Composition
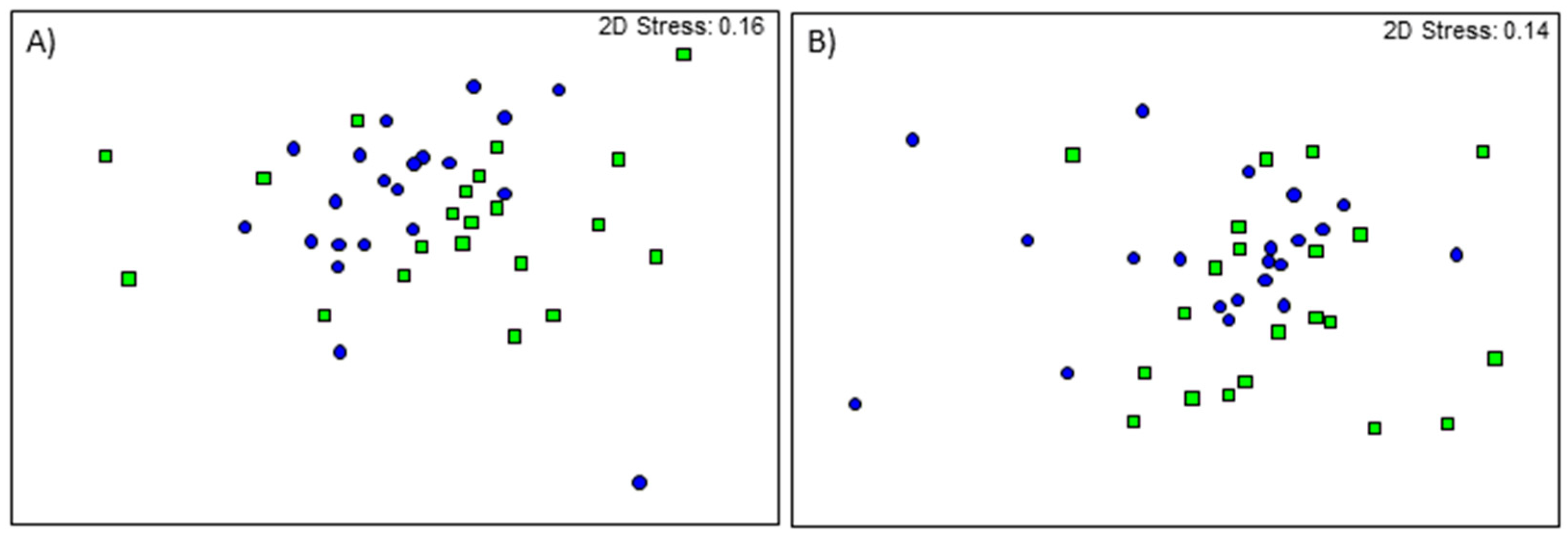
3.4. Treatment by Age Effects
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Gresse, R.; Chaucheyras-Durand, F.; Fleury, M.A.; Van de Wiele, T.; Forano, E.; Blanquet-Diot, S. Gut microbiota dysbiosis in postweaning piglets: Understanding the keys to health. Trends Microbiol. 2017, 25, 851–873. [Google Scholar] [CrossRef]
- Boudry, G.; Péron, V.; Huërou-Luron, I.L.; Lallès, J.P.; Sève, B. Weaning induces both transient and long-lasting modifcations of absorptive, secretory, and barrier properties of piglet intestine. Nutr. Metab. 2004, 134, 2256–2262. [Google Scholar]
- Pluske, J.R.; Kim, J.C.; Hansen, C.F.; Mullan, B.P.; Payne, H.G.; Hampson, D.J.; Callesen, J.; Wilson, R.H. Piglet growth before and after weaning in relation to a qualitative estimate of solid (creep) feed intake during lactation: A pilot study. Arch. Anim. Nutr. 2007, 61, 469–480. [Google Scholar] [CrossRef]
- Dong, X.; Xu, Q.; Wang, C.; Zou, X.; Lu, J. Supplemental-coated zinc oxide relieves diarrhoea by decreasing intestinal permeability in weanling pigs. J. Appl. Anim. Res. 2019, 47, 362–368. [Google Scholar] [CrossRef]
- Reid, G.; Friendship, R. Alternatives to antibiotic use: Probiotics for the gut. Anim. Biotechnol. 2002, 13, 97–112. [Google Scholar] [CrossRef]
- Jensen, J.; Larsen, M.M.; Bak, J. National monitoring study in denmark finds increased and critical levels of copper and zinc in arable soils fertilized with pig slurry. Environ. Pollut. 2016, 214, 334–340. [Google Scholar] [CrossRef] [PubMed]
- Slifierz, M.J.; Friendship, R.; Weese, J.S. Zinc oxide therapy increases prevalence and persistence of methicillin-resistant staphylococcus aureus in pigs: A randomized controlled trial. Zoonoses Public Health 2015, 62, 301–308. [Google Scholar] [CrossRef]
- Abeles, S.R.; Pride, D.T. Molecular bases and role of viruses in the human microbiome. J. Mol. Biol. 2014, 426, 3892–3906. [Google Scholar] [CrossRef]
- Hallen-Adams, H.E.; Suhr, M.J. Fungi in the healthy human gastrointestinal tract. Virulence 2017, 8, 352–358. [Google Scholar] [CrossRef]
- Gensollen, T.; Iyer, S.S.; Kasper, D.L.; Blumberg, R.S. How colonization by microbiota in early life shapes the immune system. Science 2016, 352, 539–544. [Google Scholar] [CrossRef]
- Ruczizka, U.; Metzler-Zebeli, B.; Unterweger, C.; Mann, E.; Schwarz, L.; Knecht, C.; Hennig-Pauka, I. Early parenteral administration of ceftiofur has gender-specific short and long-term effects on the fecal microbiota and growth in pigs from the suckling to growing phase. Animals 2019, 10, 17. [Google Scholar] [CrossRef]
- Niederwerder, M.C.; Constance, L.A.; Rowland, R.R.R.; Abbas, W.; Fernando, S.C.; Potter, M.L.; Sheahan, M.A.; Burkey, T.E.; Hesse, R.A.; Cino-Ozuna, A.G. Fecal microbiota transplantation is associated with reduced morbidity and mortality in porcine circovirus associated disease. Front. Microbiol. 2018, 9, 1631. [Google Scholar] [CrossRef] [PubMed]
- Sangild, P.T.; Siggers, R.H.; Schmidt, M.; Elnif, J.; Bjornvad, C.R.; Thymann, T.; Grondahl, M.L.; Hansen, A.K.; Jensen, S.K.; Boye, M.; et al. Diet- and colonization-dependent intestinal dysfunction predisposes to necrotizing enterocolitis in preterm pigs. Gastroenterology 2006, 130, 1776–1792. [Google Scholar] [CrossRef] [PubMed]
- Cahenzli, J.; Köller, Y.; Wyss, M.; Geuking, M.B.; McCoy, K.D. Intestinal microbial diversity during early-life colonization shapes long-term ige levels. Cell Host Microbe 2013, 14, 559–570. [Google Scholar] [CrossRef]
- Bakken, J.S.; Borody, T.; Brandt, L.J.; Brill, J.V.; Demarco, D.C.; Franzos, M.A.; Kelly, C.; Khoruts, A.; Louie, T.; Martinelli, L.P.; et al. Treating clostridium difficile infection with fecal microbiota transplantation. Clin. Gastroenterol. Hepatol. 2011, 9, 1044–1049. [Google Scholar] [CrossRef] [PubMed]
- Brandt, L.J.; Aroniadis, O.C. An overview of fecal microbiota transplantation: Techniques, indications, and outcomes. Gastrointest. Endosc. 2013, 78, 240–249. [Google Scholar] [CrossRef]
- Hu, L.S.; Geng, S.J.; Li, Y.; Cheng, S.S.; Fu, X.F.; Yue, X.J.; Han, X.Y. Exogenous fecal microbiota transplantation from local adult pigs to crossbred newborn piglets. Front. Microbiol. 2018, 8, 2663. [Google Scholar] [CrossRef]
- Seyfang, J.; Plush, K.J.; Kirkwood, R.N.; Tilbrook, A.J.; Ralph, C.R. The sex ratio of a litter affects the behaviour of its female pigs until at least 16 weeks of age. Appl. Anim. Behav. Sci. 2018, 200, 45–50. [Google Scholar] [CrossRef]
- Stirling, G.R.; Griffin, D.; Ophel-Keller, K.; McKay, A.; Hartley, A.; Curran, J.; Stirling, A.M.; Monsour, C.; Winch, J.; Hardie, B. Combining an initial risk assessment process with DNA assays to improve prediction of soilborne diseases caused by root-knot nematode (meloidogyne spp.) and fusarium oxysporum f. Sp. Lycopersici in the queensland tomato industry. Australas. Plant Path. 2004, 33, 285–293. [Google Scholar] [CrossRef]
- Torok, V.A.; Ophel-Keller, K.; Loo, M.; Hughes, R.J. Application of methods for identifying broiler chicken gut bacterial species linked with increased energy metabolism. Appl. Environ. Microbiol. 2008, 74, 783–791. [Google Scholar] [CrossRef]
- Torok, V.A.; Percy, N.J.; Moate, P.J.; Ophel-Keller, K. Influence of dietary docosahexaenoic acid supplementation on the overall rumen microbiota of dairy cows and linkages with production parameters. Can. J. Microbiol. 2014, 60, 267–275. [Google Scholar] [CrossRef]
- Zhang, J.; Kobert, K.; Flouri, T.; Stamatakis, A. Pear: A fast and accurate illumina paired-end read merger. Bioinformatics 2014, 30, 614–620. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kucztnski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. Qiime allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Search and clustering orders of magnitude faster than blast. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. Uchime improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. Uparse: Highly accurate otu sequences from microbial amplicon reads. Nat. Methods 2013, 10, 996–998. [Google Scholar] [CrossRef]
- DeSantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a chimera-checked 16s rrna gene database and workbench compatible with arb. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef]
- Bray, J.R.; Curtis, J.T. An ordination of the upland forest communities of southern wisconsin. Ecol. Monogr. 1957, 27, 325–349. [Google Scholar] [CrossRef]
- Clarke, K.R. Non-parametric multivariate analyses of changes in community structure. J. Ecol. 1993, 18, 117–143. [Google Scholar] [CrossRef]
- Kruskal, J.B. Multidimensional scaling by optimizing a goodness of fit to a nonmetric hypothesis. Phsychometrics 1964, 29, 1–28. [Google Scholar] [CrossRef]
- Shepard, R.N. The analysis of proximities: Multidimensional scaling with an unknown distance function. I. Psychometrika 1962, 27, 125–140. [Google Scholar] [CrossRef]
- Pluske, J.R.; Turpin, D.L.; Kim, J.-C. Gastrointestinal tract (gut) health in the young pig. Anim. Nutr. 2018, 4, 187–196. [Google Scholar] [CrossRef] [PubMed]
- Nowland, T.L.; Plush, K.J.; Barton, M.; Kirkwood, R.N. Development and function of the intestinal microbiome and potential implications for pig production. Animals 2019, 9, 76. [Google Scholar] [CrossRef] [PubMed]
- Brunse, A.; Martin, L.; Rasmussen, T.S.; Christensen, L.; Skovsted Cilieborg, M.; Wiese, M.; Khakimov, B.; Pieper, R.; Nielsen, D.S.; Sangild, P.T.; et al. Effect of fecal microbiota transplantation route of administration on gut colonization and host response in preterm pigs. ISME J. 2019, 13, 720–733. [Google Scholar] [CrossRef] [PubMed]
- Geng, S.; Cheng, S.; Li, Y.; Wen, Z.; Ma, X.; Jiang, X.; Wang, Y.; Han, X. Faecal microbiota transplantation reduces susceptibility to epithelial injury and modulates tryptophan metabolism of the microbial community in a piglet model. J. Crohns. Colitis 2018, 12, 1359–1374. [Google Scholar] [CrossRef]
- Sylvia, K.E.; Demas, G.E. A gut feeling: Microbiome-brain-immune interactions modulate social and affective behaviors. Horm. Behav. 2018, 99, 41–49. [Google Scholar] [CrossRef]
- Cheng, C.S.; Wei, H.K.; Wang, P.; Yu, H.C.; Zhang, X.M.; Jiang, S.W.; Peng, J. Early intervention with faecal microbiota transplantation: An effective means to improve growth performance and the intestinal development of suckling piglets. Animal 2018, 13, 533–541. [Google Scholar] [CrossRef]
- Grabig, A.; Paclik, D.; Guzy, C.; Dankof, A.; Baumgart, D.C.; Erckenbrecht, J.; Raupach, B.; Sonnenborn, U.; Eckert, J.; Schumann, R.R.; et al. Escherichia coli strain nissle 1917 ameliorates experimental colitis via toll-like receptor 2- and toll-like receptor 4-dependent pathways. Infect. Immunol. 2006, 74, 4075–4082. [Google Scholar] [CrossRef] [PubMed]
- Grozdanov, L.; Raasch, C.; Schulze, J.; Sonnenborn, U.; Gottschalk, G.; Hacker, J.; Dobrindt, U. Analysis of the genome structure of the nonpathogenic probiotic escherichia coli strain nissle 1917. J. Bacteriol. 2004, 186, 5432–5441. [Google Scholar] [CrossRef]
- Lee, J.H.; Valeriano, V.D.; Shin, Y.R.; Chae, J.P.; Kim, G.B.; Ham, J.S.; Chun, J.; Kang, D.K. Genome sequence of lactobacillus mucosae lm1, isolated from piglet feces. J. Bacteriol. 2012, 194, 4766. [Google Scholar] [CrossRef]
- McCormack, U.M.; Curiao, T.; Metzler-Zebeli, B.U.; Wilkinson, T.; Reyer, H.; Crispie, F.; Cotter, P.D.; Creevey, C.J.; Gardiner, G.E.; Lawlor, P.G. Improvement of feed efficiency in pigs through microbial modulation via fecal microbiota transplantation in sows and dietary supplementation of inulin in offspring. Appl. Environ. Microbiol. 2019, 85, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Dou, S.; Gadonna-Widehem, P.; Rome, V.; Hamoudi, D.; Rhazi, L.; Lakhal, L.; Larcher, T.; Bahi-Jaber, N.; Pinon-Quintana, A.; Guyonvarch, A.; et al. Characterisation of early-life fecal microbiota in susceptible and healthy pigs to post-weaning diarrhoea. PLoS ONE 2017, 12, e0169851. [Google Scholar] [CrossRef] [PubMed]
- McCormack, U.M.; Curiao, T.; Buzoianu, S.G.; Prieto, M.L.; Ryan, T.; Varley, P.; Crispie, F.; Magowan, E.; Metzler-Zebeli, B.U.; Berry, D.; et al. Exploring a possible link between the intestinal microbiota and feed efficiency in pigs. Appl. Environ. Microbiol. 2017, 83, 1–16. [Google Scholar] [CrossRef]
- Rees, A.S.; Lysons, R.J.; Stokes, C.R.; Bourne, F.J. Antibody production by the pig colon during infection with treponema hyodyseneriae. Res. Vet. Sci. 1989, 47, 263–269. [Google Scholar] [CrossRef]
- McCormack, U.M.; Curiao, T.; Wilkinson, T.; Metzler-Zebeli, B.U.; Reyer, H.; Ryan, T.; Calderon-Diaz, J.A.; Crispie, F.; Cotter, P.D.; Creevey, C.J.; et al. Fecal microbiota transplantation in gestating sows and neonatal offspring alters lifetime intestinal microbiota and growth in offspring. Am. Soc. Microbiol. 2018, 3. [Google Scholar] [CrossRef]
- Nowland, T.L.; Torok, V.A.; Low, W.Y.; Barton, M.D.; Plush, K.J.; Kirkwood, R.N. Faecal microbiota analysis of piglets during lactation. Animals 2020, 10, 762. [Google Scholar] [CrossRef]
- Lallès, J.-P.; Bosi, P.; Smidt, H.; Stokes, C.R. Weaning—Achallenge to gut physiologists. Livest. Sci. 2007, 108, 82–93. [Google Scholar] [CrossRef]



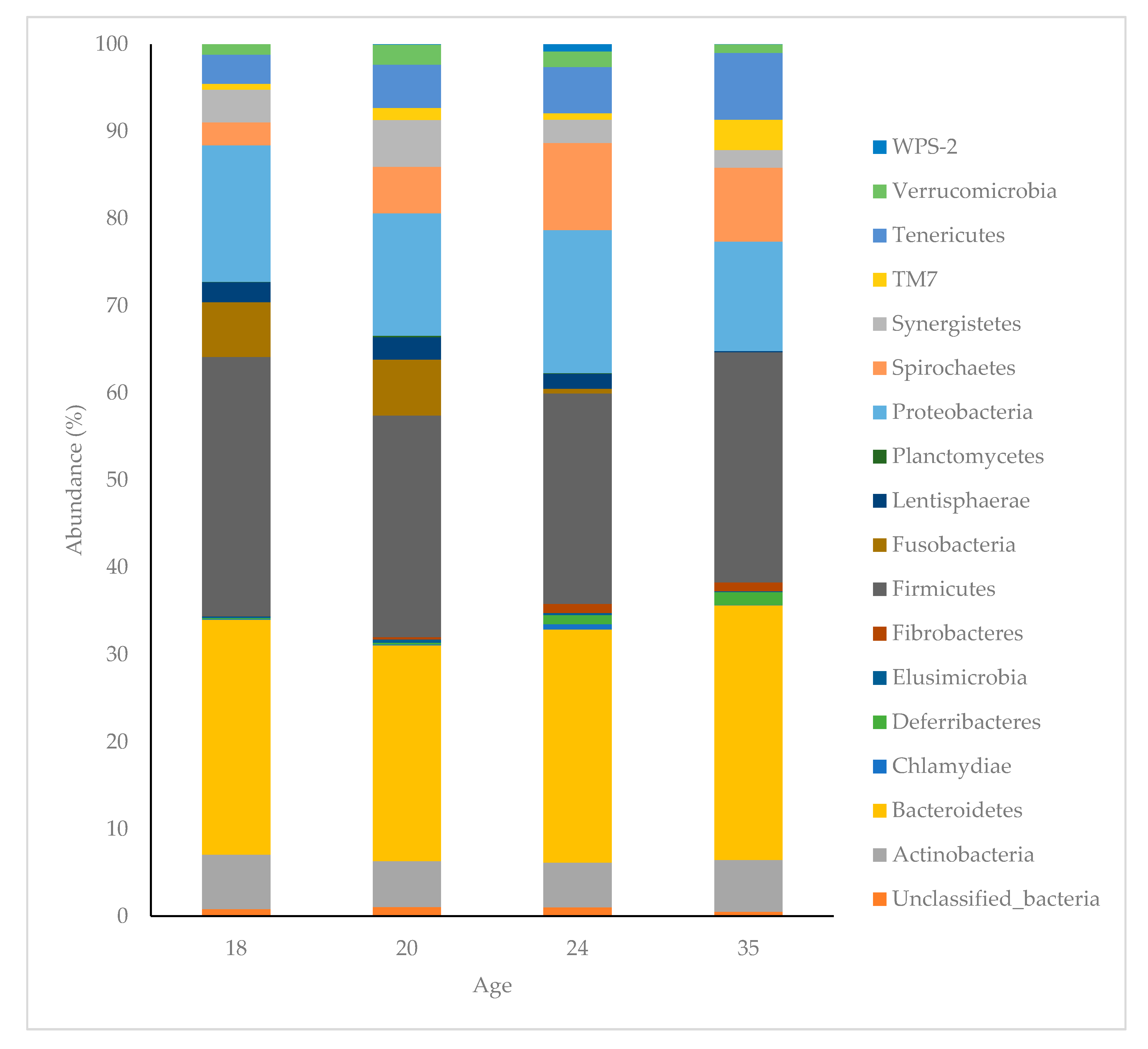

| Group FMT | Group CON | ||||||
|---|---|---|---|---|---|---|---|
| Phyla | Class | Order | Family | Genus | Average Abundance | Average Abundance | % |
| Actinobacteria | Coriobacteriia | Coriobacteriales | Coriobacteriaceae | Collinsella | 0.26 | 0.27 | 0.69 |
| Bacteroidetes | Bacteroidia | Bacteroidales | Bacteroidaceae | Bacteroides | 0.82 | 0.68 | 1.24 |
| Odoribacteraceae | Butyricimonas | 0.27 | 0.25 | 0.75 | |||
| p-2534-18B5 | 0.46 | 0.29 | 1.09 | ||||
| Paraprevotellaceae | YRC22 | 0.61 | 0.38 | 0.94 | |||
| CF231 | 0.9 | 0.9 | 0.81 | ||||
| Porphyromonadaceae | Parabacteroides | 1.02 | 0.79 | 1.25 | |||
| Paludibacter | 0.37 | 0.24 | 1.18 | ||||
| Prevotellaceae | Prevotella | 2.12 | 2.31 | 1.13 | |||
| Prevotella | 1.19 | 1.03 | 1.09 | ||||
| Prevotellaceae | 0.69 | 0.58 | 0.82 | ||||
| RF16 | 0.47 | 0.36 | 0.81 | ||||
| Rikenellaceae | 0.25 | 0.2 | 0.73 | ||||
| S24-7 | 1.77 | 1.64 | 0.99 | ||||
| Deferribacteres | Deferribacteres | Deferribacterales | Deferribacteraceae | Mucispirillum | 0.4 | 0.21 | 0.8 |
| Elusimicrobia | Elusimicrobia | Elusimicrobiales | Elusimicrobiaceae | 0.26 | 0.12 | 0.77 | |
| Fibrobacteres | Fibrobacteria | Fibrobacterales | Fibrobacteraceae | Fibrobacter | 0.36 | 0.22 | 0.7 |
| Firmicutes | Bacilli | Lactobacillales | Lactobacillaceae | Lactobacillus | 0.46 | 0.5 | 0.98 |
| Streptococcaceae | Streptococcus | 0.41 | 0.32 | 0.78 | |||
| Clostridia | Clostridiales | Christensenellaceae | 1.1 | 0.9 | 1.27 | ||
| Clostridiaceae | Sarcina | 0.82 | 0.67 | 1.02 | |||
| Clostridiaceae | 0.49 | 0.41 | 0.81 | ||||
| SMB53 | 0.48 | 0.37 | 0.72 | ||||
| Lachnospiraceae | 1.43 | 1.2 | 1.16 | ||||
| Shuttleworthia | 0.37 | 0.13 | 1.02 | ||||
| Dorea | 0.75 | 0.63 | 0.96 | ||||
| Butyrivibrio | 0.54 | 0.4 | 0.94 | ||||
| Lachnobacterium | 0.38 | 0.25 | 0.73 | ||||
| Roseburia | 0.84 | 0.93 | 0.69 | ||||
| Lachnospira | 0.71 | 0.65 | 0.69 | ||||
| Mogibacteriaceae | 0.82 | 0.72 | 0.8 | ||||
| Peptococcaceae | rc4-4 | 0.33 | 0.26 | 0.66 | |||
| Ruminococcaceae | Faecalibacterium | 1.01 | 1.14 | 0.91 | |||
| Oscillospira | 1.52 | 1.45 | 0.73 | ||||
| Ruminococcaceae | 1.69 | 1.56 | 0.66 | ||||
| Veillonellaceae | Mitsuokella | 0.43 | 0.53 | 1.07 | |||
| Dialister | 0.54 | 0.2 | 1.06 | ||||
| Veillonellaceae | 0.63 | 0.61 | 1 | ||||
| Acidaminococcus | 0.57 | 0.43 | 0.98 | ||||
| Megasphaera | 0.63 | 0.64 | 0.9 | ||||
| Anaerovibrio | 0.95 | 0.91 | 0.75 | ||||
| Clostridiales | 1.07 | 0.91 | 1.01 | ||||
| Clostridiales | 1.17 | 1.02 | 0.66 | ||||
| Firmicutes | Erysipelotrichi | Erysipelotrichales | Erysipelotrichaceae | p-75-a5 | 0.77 | 0.67 | 0.82 |
| Catenibacterium | 0.65 | 0.71 | 0.8 | ||||
| Proteobacteria | Alphaproteobacteria | RF32 | 0.33 | 0.21 | 0.82 | ||
| Alphaproteobacteria | 0.3 | 0.16 | 0.76 | ||||
| Deltaproteobacteria | GMD14H09 | 0.72 | 0.68 | 1.1 | |||
| Epsilonproteobacteria | Campylobacterales | Campylobacteraceae | Campylobacter | 0.91 | 0.75 | 1.08 | |
| Gammaproteobacteria | Aeromonadales | Succinivibrionaceae | Succinivibrio | 1.18 | 1.44 | 1.85 | |
| Succinivibrionaceae | Ruminobacter | 0.63 | 0.39 | 1.51 | |||
| Succinivibrionaceae | 0.52 | 0.15 | 1.5 | ||||
| Enterobacteriales | Enterobacteriaceae | Escherichia | 0.76 | 0.54 | 1.33 | ||
| Pasteurellales | Pasteurellaceae | Actinobacillus | 0.37 | 0.3 | 0.76 | ||
| Spirochaetes | Spirochaetes | Spirochaetales | Spirochaetaceae | Treponema | 1.31 | 1.16 | 1.14 |
| Synergistetes | Synergistia | Synergistales | Dethiosulfovibrionaceae | Pyramidobacter | 0.36 | 0.26 | 0.87 |
| Dethiosulfovibrionaceae | 0.33 | 0.27 | 0.66 | ||||
| Tenericutes | RF3 | ML615J-28 | 0.38 | 0.27 | 0.75 | ||
| TM7 | TM7-3 | CW040 | F16 | 0.33 | 0.17 | 0.8 | |
| Verrucomicrobia | Verruco-5 | WCHB1-41 | RFP12 | 0.33 | 0.18 | 0.66 | |
| WPS-2 | 0.31 | 0.21 | 0.74 |
| Group FMT | Group CON | ||||||
|---|---|---|---|---|---|---|---|
| Phyla | Class | Order | Family | Genus | Average Abundance | Average Abundance | % |
| Actinobacteria | Coriobacteriia | Coriobacteriales | Coriobacteriaceae | Collinsella | 0.37 | 0.28 | 0.77 |
| Bacteroidetes | Bacteroidia | Bacteroidales | Paraprevotellaceae | YRC22 | 0.21 | 0.28 | 0.9 |
| CF231 | 0.75 | 0.88 | 0.74 | ||||
| 0.7 | 0.55 | 0.69 | |||||
| Porphyromonadaceae | Parabacteroides | 0.52 | 0.59 | 0.94 | |||
| Prevotellaceae | Prevotella | 1.19 | 1.3 | 1.12 | |||
| S24-7 | 1.32 | 1.3 | 0.82 | ||||
| Bacteroidales | 0.43 | 0.41 | 0.9 | ||||
| Deferribacteres | Deferribacteres | Deferribacterales | Deferribacteraceae | Mucispirillum | 0.35 | 0.26 | 0.99 |
| Elusimicrobia | Elusimicrobia | Elusimicrobiales | Elusimicrobiaceae | 0.21 | 0.07 | 0.79 | |
| Fibrobacteres | Fibrobacteria | Fibrobacterales | Fibrobacteraceae | Fibrobacter | 0.31 | 0.2 | 0.78 |
| Streptococcaceae | Streptococcus | 0.45 | 0.32 | 1.01 | |||
| Firmicutes | Clostridia | Clostridiales | Clostridiaceae | Sarcina | 0.53 | 0.73 | 1.31 |
| SMB53 | 0.86 | 0.88 | 0.84 | ||||
| Clostridiaceae | 0.94 | 0.95 | 0.82 | ||||
| Lachnospiraceae | Shuttleworthia | 0.53 | 0.41 | 1.64 | |||
| Lachnobacterium | 0.56 | 0.79 | 1.63 | ||||
| Lachnospiraceae | 1.32 | 1.4 | 1.39 | ||||
| Roseburia | 1.47 | 1.55 | 1.1 | ||||
| Anaerostipes | 0.4 | 0.32 | 0.87 | ||||
| Lachnospira | 0.95 | 1.08 | 0.81 | ||||
| Ruminococcus | 1.06 | 1.03 | 0.79 | ||||
| Dorea | 0.68 | 0.71 | 0.78 | ||||
| Coprococcus | 0.94 | 0.96 | 0.72 | ||||
| Peptococcaceae | rc4-4 | 0.11 | 0.28 | 0.87 | |||
| Ruminococcaceae | Faecalibacterium | 1.18 | 1.08 | 0.82 | |||
| Oscillospira | 1.28 | 1.23 | 0.77 | ||||
| Veillonellaceae | Dialister | 0.91 | 0.34 | 2.29 | |||
| Veillonellaceae | 0.71 | 0.66 | 1.43 | ||||
| Acidaminococcus | 0.53 | 0.34 | 1.4 | ||||
| Mitsuokella | 0.65 | 0.53 | 1.35 | ||||
| Megasphaera | 0.7 | 0.53 | 1.28 | ||||
| Phascolarctobacterium | 0.72 | 0.96 | 0.99 | ||||
| Clostridia | 0.24 | 0.04 | 0.86 | ||||
| Erysipelotrichi | Erysipelotrichales | Erysipelotrichaceae | Catenibacterium | 0.75 | 0.58 | 1.13 | |
| Bulleidia | 0.98 | 0.91 | 1.09 | ||||
| p-75-a5 | 0.64 | 0.68 | 0.77 | ||||
| L7A_E11 | 0.33 | 0.25 | 0.74 | ||||
| Proteobacteria | Alphaproteobacteria | RF32 | 0.36 | 0.45 | 1.09 | ||
| Betaproteobacteria | Burkholderiales | Oxalobacteraceae | Oxalobacter | 0.14 | 0.15 | 0.73 | |
| Tremblayales | 0.32 | 0.26 | 1.44 | ||||
| Deltaproteobacteria | GMD14H09 | 0.52 | 0.45 | 1.36 | |||
| Epsilonproteobacteria | Campylobacterales | Campylobacteraceae | Campylobacter | 0.77 | 0.67 | 1.34 | |
| Gammaproteobacteria | Aeromonadales | Succinivibrionaceae | Succinivibrio | 0.93 | 0.98 | 1.84 | |
| Succinivibrionaceae | 0.47 | 0.2 | 1.7 | ||||
| Enterobacteriales | Enterobacteriaceae | Escherichia | 0.34 | 0.27 | 0.81 | ||
| Pasteurellales | Pasteurellaceae | Actinobacillus | 0.32 | 0.15 | 0.93 | ||
| Spirochaetes | Spirochaetes | Spirochaetales | Spirochaetaceae | Treponema | 0.91 | 1.01 | 1.65 |
| Synergistetes | Synergistia | Synergistales | Dethiosulfovibrionaceae | 0.22 | 0.27 | 0.7 | |
| TM7 | TM7-3 | CW040 | F16 | 0.38 | 0.41 | 0.93 | |
| Unclassified Bacteria | 0.26 | 0.14 | 0.81 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nowland, T.L.; Torok, V.A.; Low, W.Y.; Plush, K.J.; Barton, M.D.; Kirkwood, R.N. A Single Faecal Microbiota Transplantation Altered the Microbiota of Weaned Pigs. Life 2020, 10, 203. https://doi.org/10.3390/life10090203
Nowland TL, Torok VA, Low WY, Plush KJ, Barton MD, Kirkwood RN. A Single Faecal Microbiota Transplantation Altered the Microbiota of Weaned Pigs. Life. 2020; 10(9):203. https://doi.org/10.3390/life10090203
Chicago/Turabian StyleNowland, Tanya L., Valeria A. Torok, Wai Y. Low, Kate J. Plush, Mary D. Barton, and Roy N. Kirkwood. 2020. "A Single Faecal Microbiota Transplantation Altered the Microbiota of Weaned Pigs" Life 10, no. 9: 203. https://doi.org/10.3390/life10090203
APA StyleNowland, T. L., Torok, V. A., Low, W. Y., Plush, K. J., Barton, M. D., & Kirkwood, R. N. (2020). A Single Faecal Microbiota Transplantation Altered the Microbiota of Weaned Pigs. Life, 10(9), 203. https://doi.org/10.3390/life10090203






