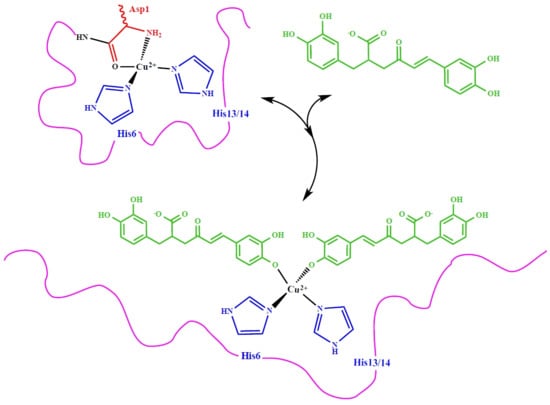
Novel Perspective on Alzheimer’s Disease Treatment: Rosmarinic Acid Molecular Interplay with Copper(II) and Amyloid β
Abstract
1. Introduction
2. Materials and Methods
3. Results
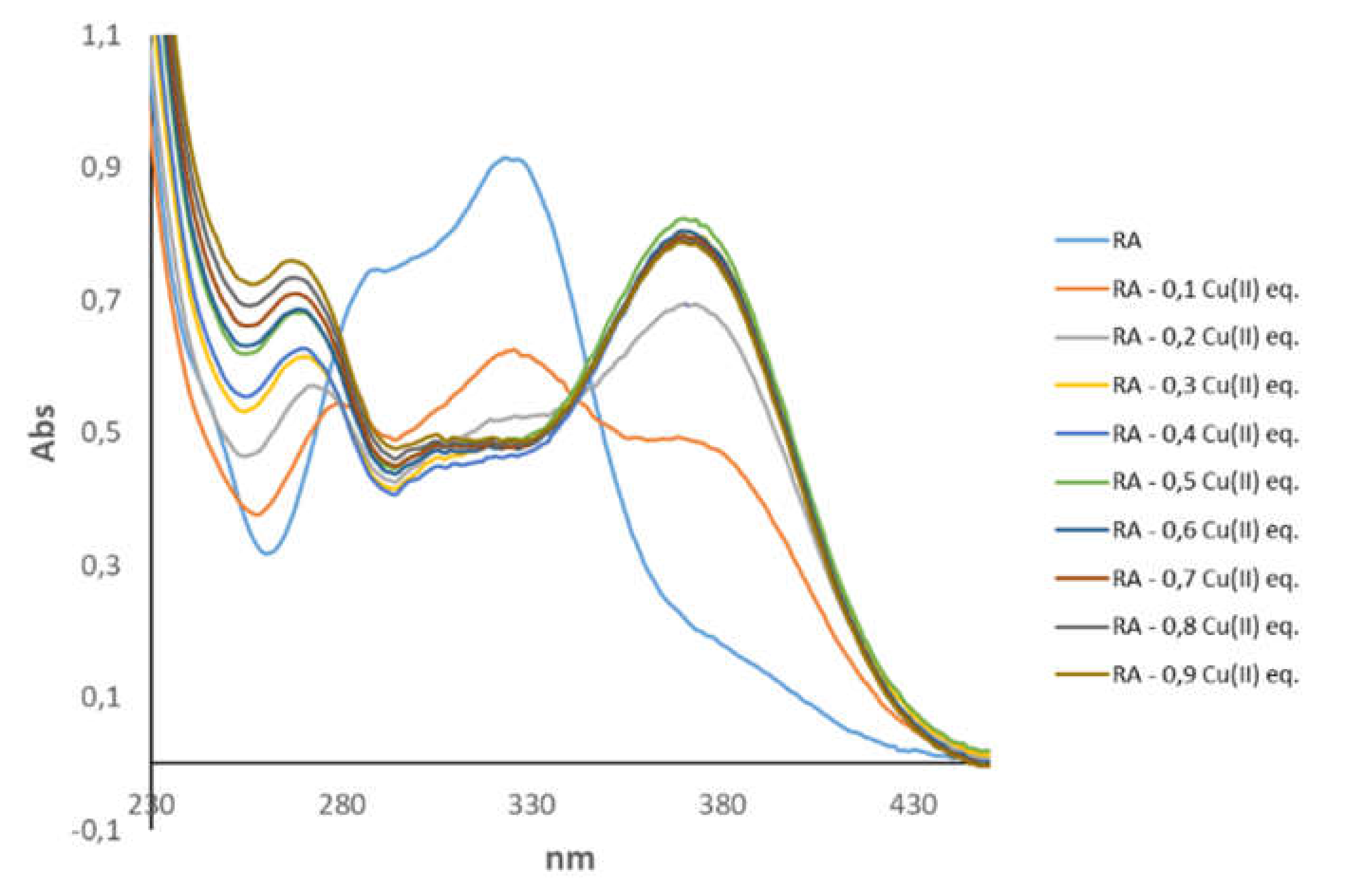
3.1. Spectroscopic Characterization of RA–Cu(II) Complex
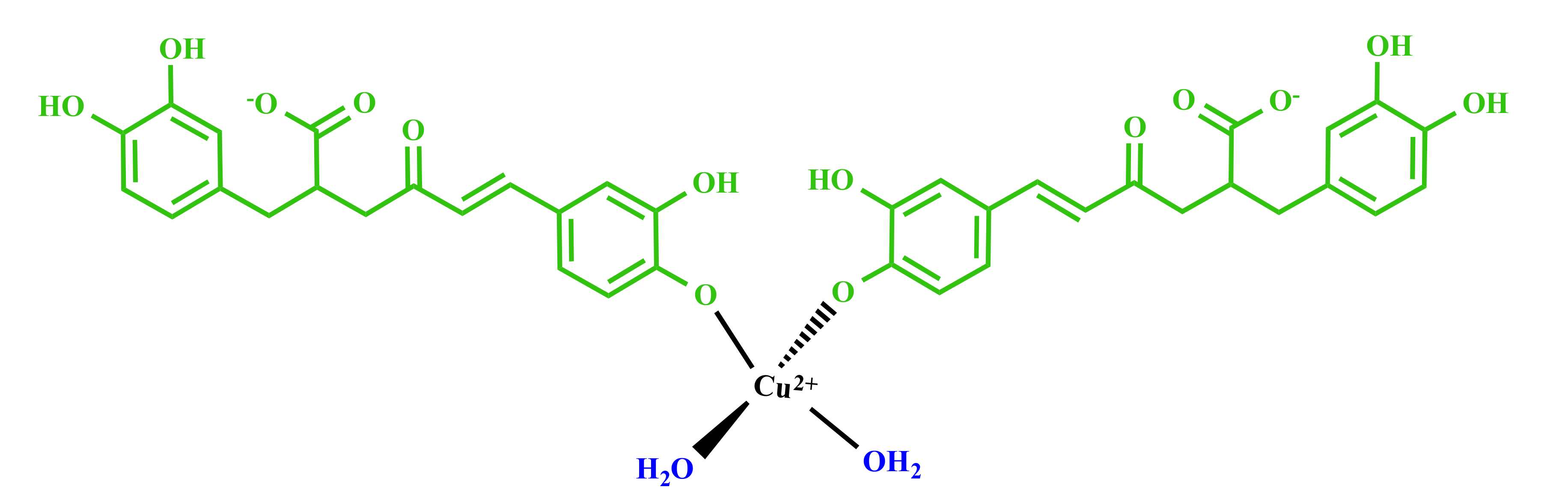
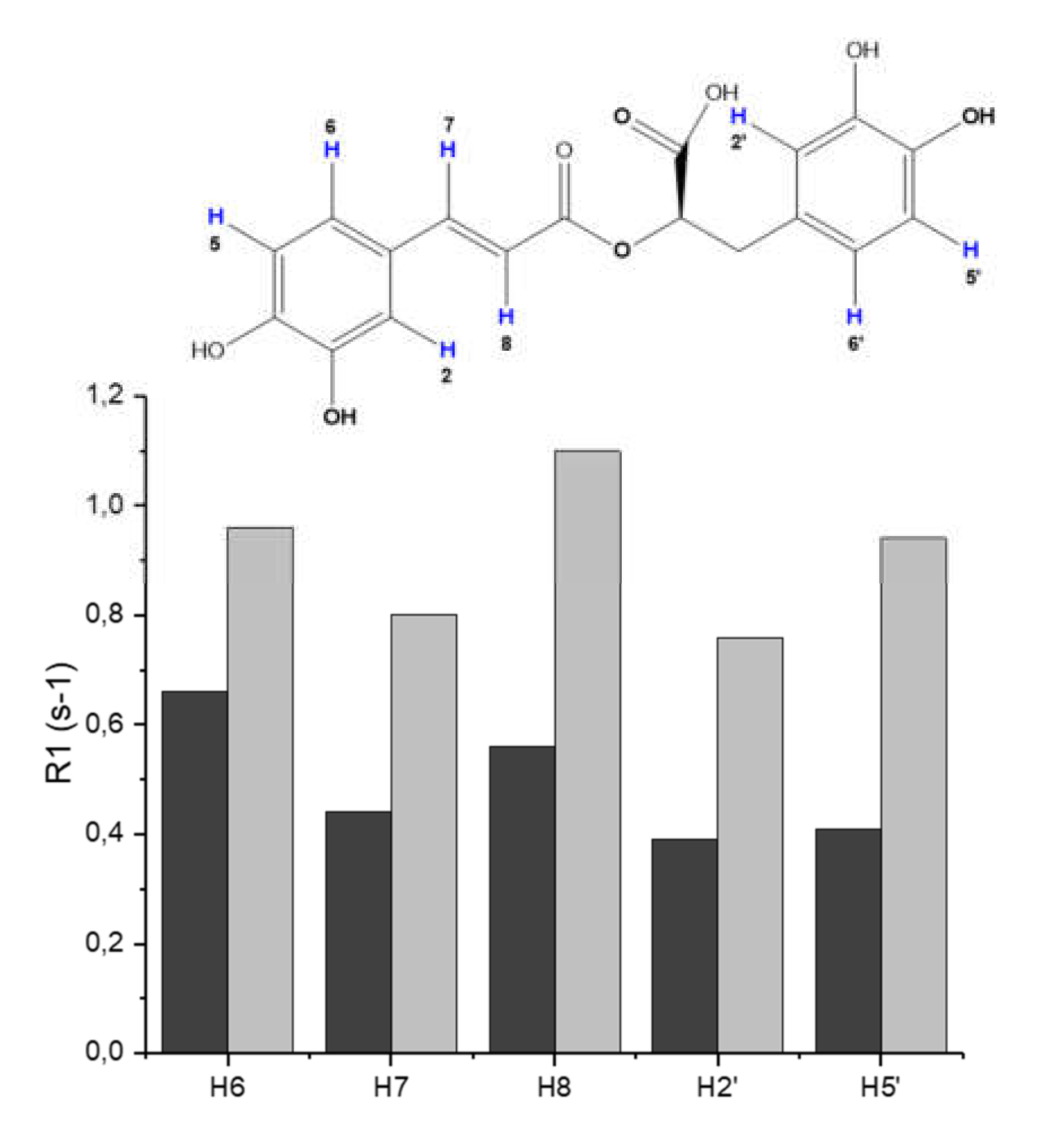
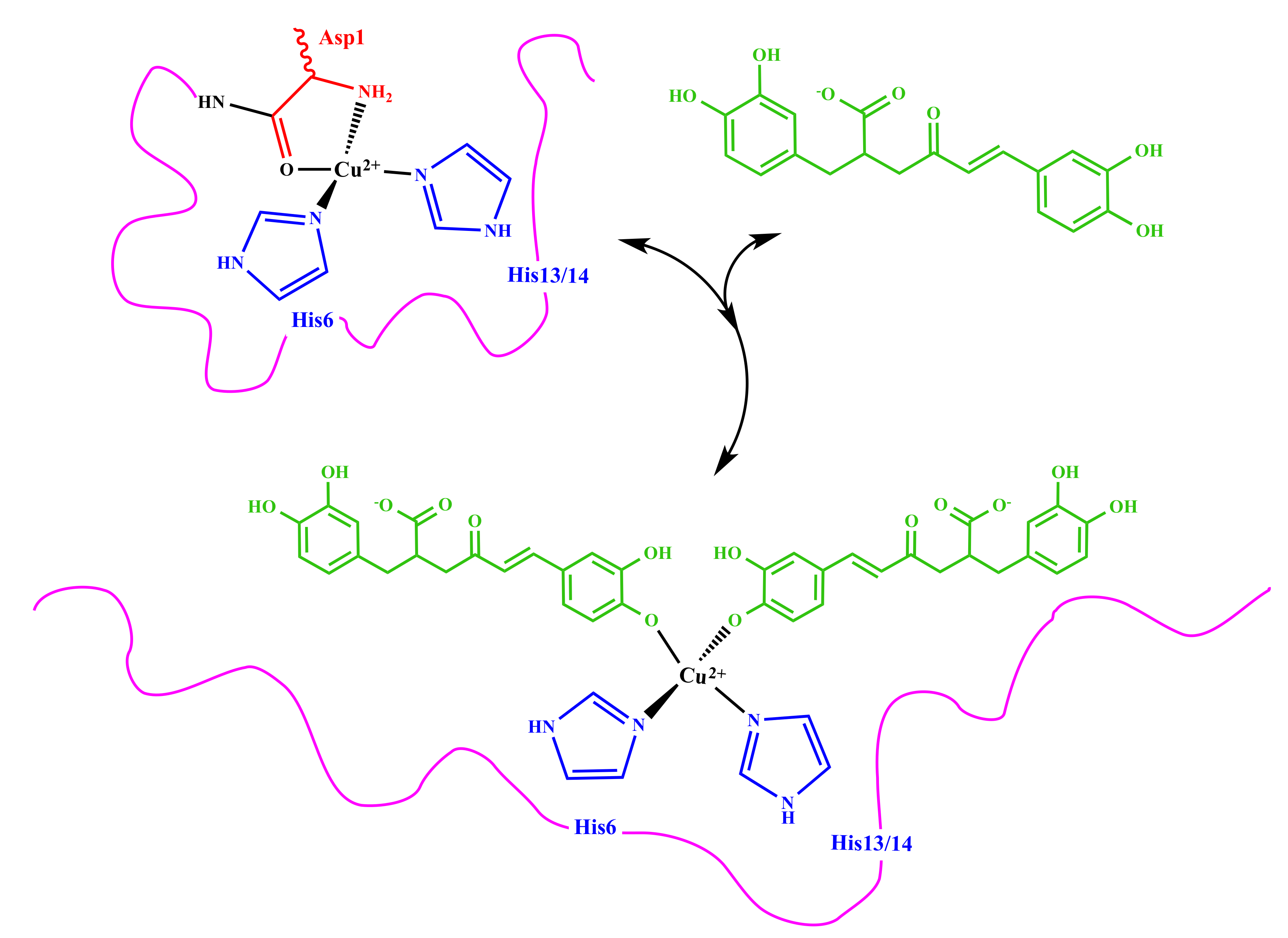
3.2. Molecular Structure of RA–Cu(II) Complex
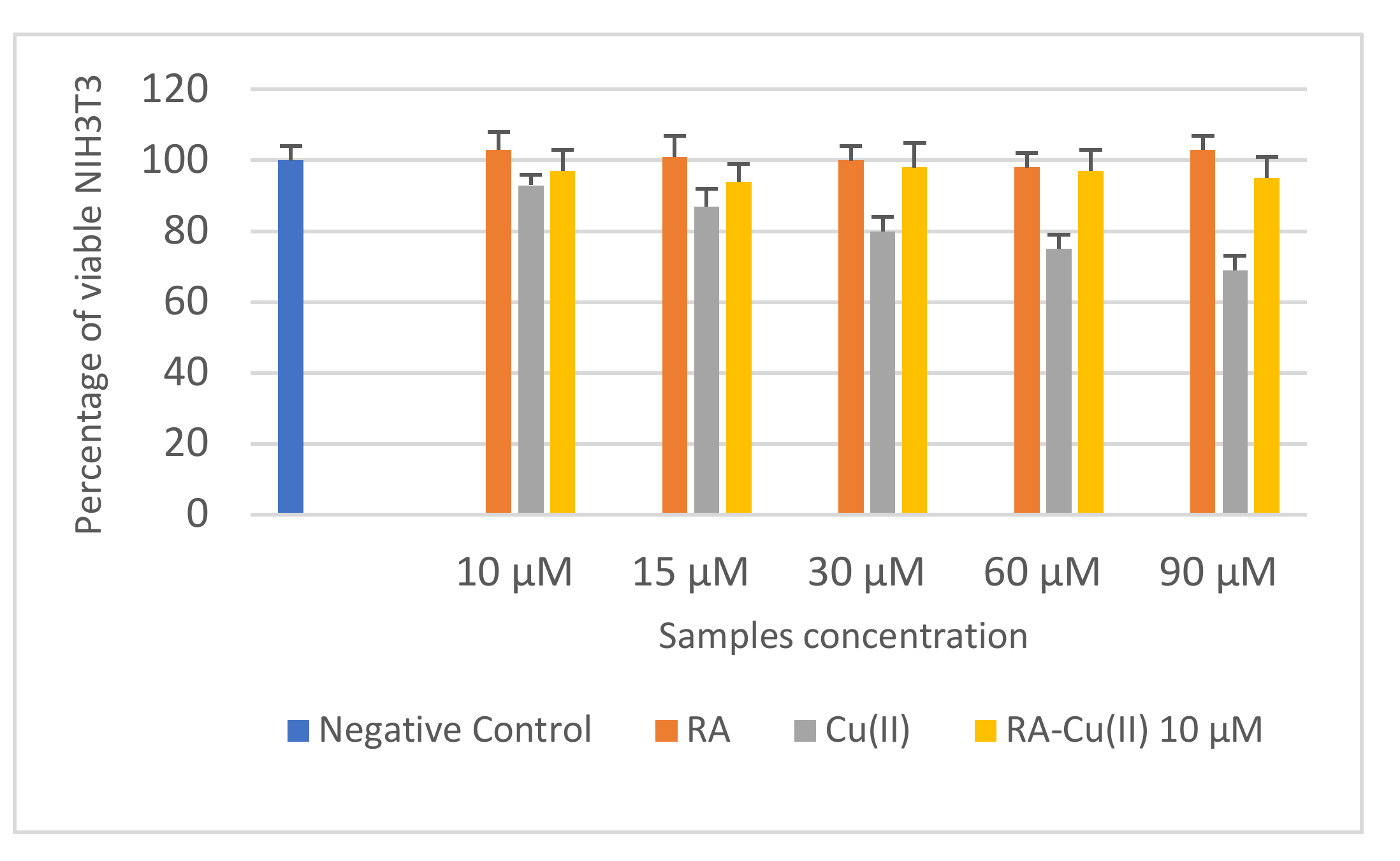
3.3. In Vitro Cytotoxicity and Mutagenicity of RA and RA–Cu(II) Complex
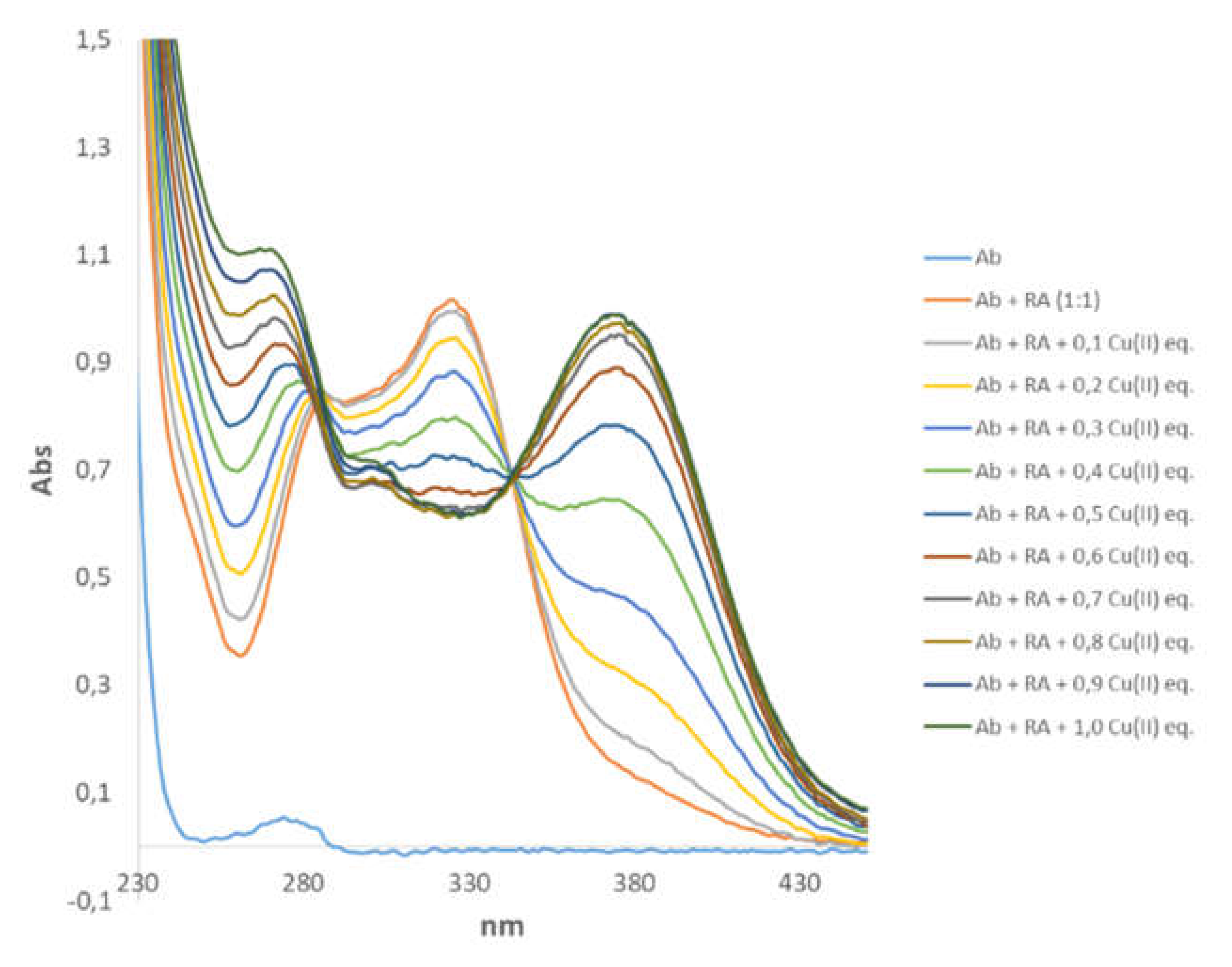
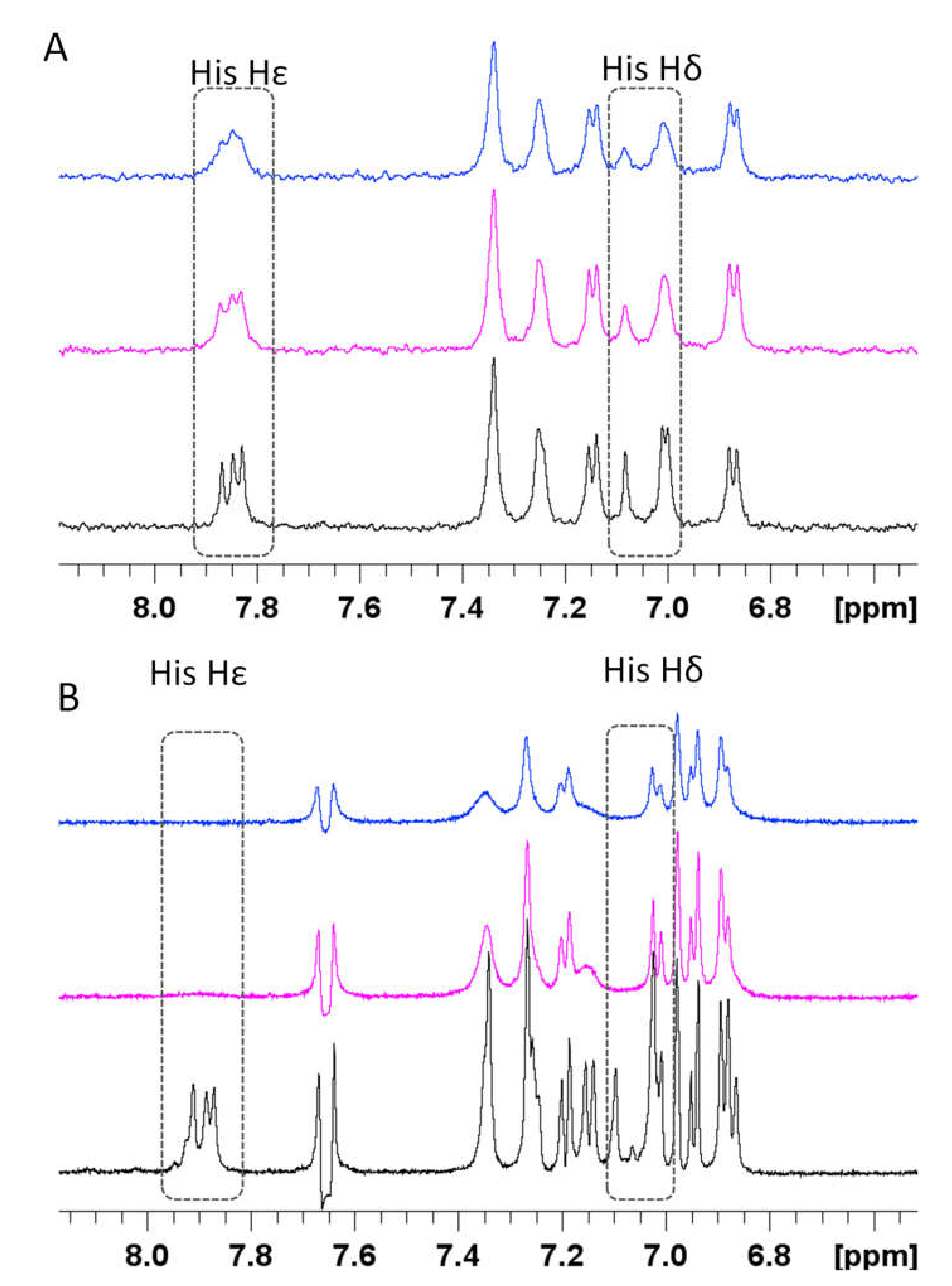
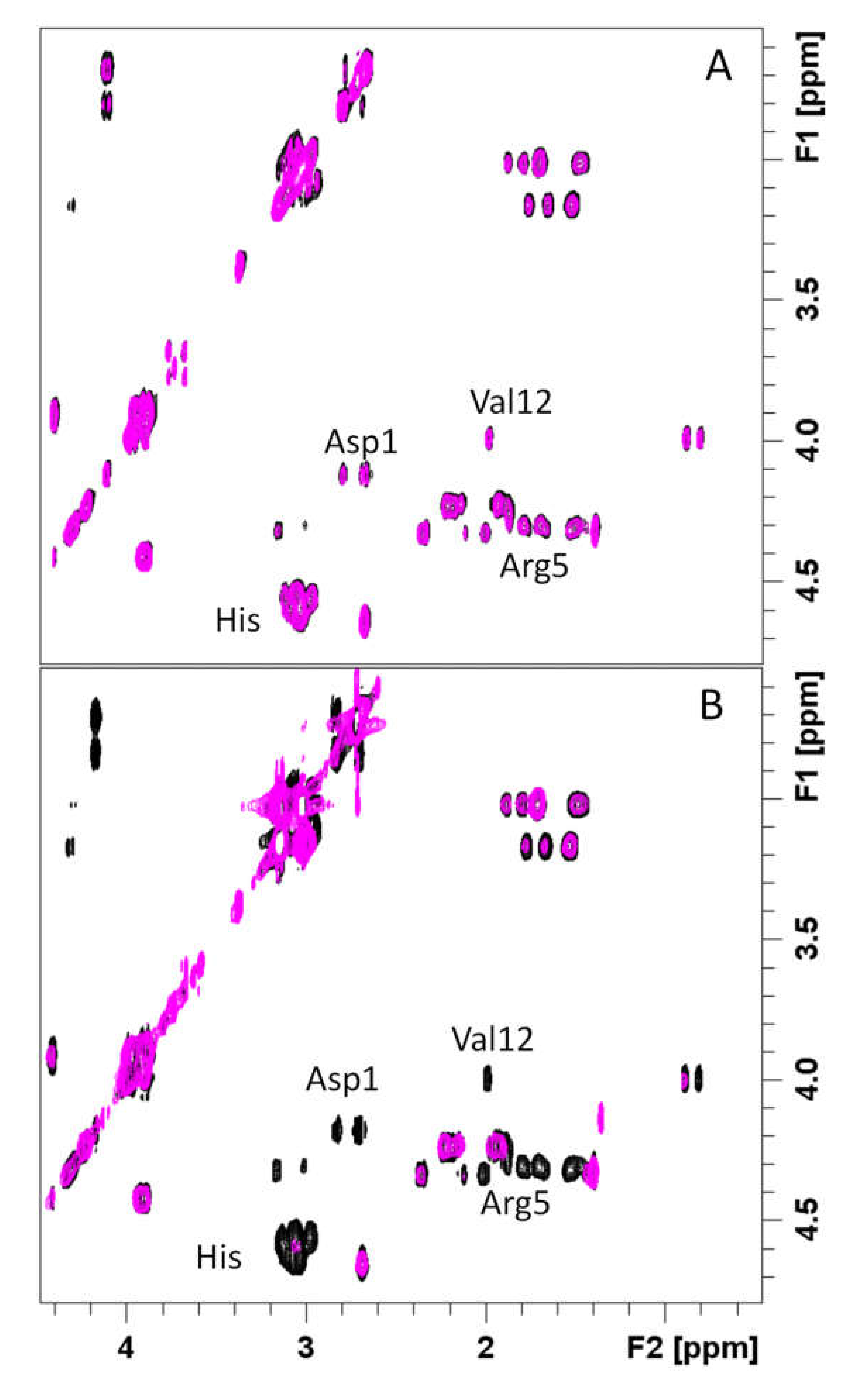
3.4. Spectroscopic Characterization of RA–Cu(II)–Aβ System
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Palamidessi, T. Guarire con le Piante Medicinali; Copyright Alessandro Benassai 2013; Archeosofica Editore: Roma, Italy, 1966; Volume 1, pp. 15–99. [Google Scholar]
- Wang, Y.S.; Shen, C.Y.; Jiang, J.G. Antidepressant Active Ingredients from Herbs and Nutraceuticals Used in TCM: Pharmacological Mechanisms and Prospects for Drug Discovery. Pharmacol. Res. 2019, 150, 104520. [Google Scholar] [CrossRef]
- Anwar, F.; Abbas, A.; Mehmood, T.; Gilani, A.H.; Rehman, U.N. Mentha: A genus rich in vital nutra-pharmaceuticals—A review. Phytother. Res. 2019, 33, 2548–2570. [Google Scholar] [CrossRef] [PubMed]
- Elufioye, T.O.; Habtemariam, S. Hepatoprotective effects of rosmarinic acid: Insight into its mechanisms of action. Biomed. Pharmacother. 2019, 112, 108600. [Google Scholar] [CrossRef] [PubMed]
- Fachel, F.N.S.; Schuh, R.S.; Veras, K.S.; Bassani, V.L.; Koester, L.S.; Henriques, A.T.; Braganhol, E.; Teixeira, H.F. An Overview of the Neuroprotective Potential of Rosmarinic Acid and Its Association with Nanotechnology-Based Delivery Systems: A Novel Approach to Treating Neurodegenerative Disorders. Neurochem. Int. 2019, 122, 47–58. [Google Scholar] [CrossRef] [PubMed]
- Swamy, M.K.; Sinniah, U.R. A Ghasemzadeh Anticancer Potential of Rosmarinic Acid and Its Improved Production Through Biotechnological Interventions and Functional Genomics. Appl. Microbiol. Biotechnol. 2018, 102, 7775–7793. [Google Scholar] [CrossRef] [PubMed]
- Andrade, J.M.; Faustino, C.; Garcia, C.; Ladeiras, D.; Reis, C.P.; Rijo, P. Rosmarinus officinalis L.: An Update Review of Its Phytochemistry and Biological Activity. Future Sci. OA 2018, 4, FSO283. [Google Scholar] [CrossRef] [PubMed]
- Habtemariam, S. Molecular Pharmacology of Rosmarinic and Salvianolic Acids: Potential Seeds for Alzheimer’s and Vascular Dementia Drugs. Int. J. Mol. Sci. 2018, 19, 458. [Google Scholar] [CrossRef]
- Alagawany, M.; El-Hack, M.E.A.; Farag, M.R.; Gopi, M.; Karthik, K.; Malik, Y.S.; Dhama, K. Rosmarinic Acid: Modes of Action, Medicinal Values and Health Benefits. Anim. Health Res. Rev. 2017, 18, 167–176. [Google Scholar] [CrossRef] [PubMed]
- Bulgakov, V.P.; Vereshchagina, Y.V.; Veremeichik, G.N. Anticancer Polyphenols From Cultured Plant Cells: Production and New Bioengineering Strategies. Curr. Med. Chem. 2018, 25, 4671–4692. [Google Scholar] [CrossRef]
- Moore, J.; Yousef, M.; Tsiani, E. Anticancer Effects of Rosemary (Rosmarinus officinalis L.) Extract and Rosemary Extract Polyphenols. Nutrients 2016, 8, 731. [Google Scholar] [CrossRef]
- Shen, C.Y.; Jiang, J.G.; Yang, L.; Wang, D.W.; Zhu, W. Anti-ageing Active Ingredients from Herbs and Nutraceuticals Used in Traditional Chinese Medicine: Pharmacological Mechanisms and Implications for Drug Discovery. Br. J. Pharmacol. 2017, 174, 1395–1425. [Google Scholar] [CrossRef] [PubMed]
- Lamaison, J.L.; Petitjean-Freytet, C.; Carnat, A. Rosmarinic Acid, Total Hydroxycinnamic Derivatives and Antioxidant Activity of Apiaceae, Borraginaceae and Lamiceae Medicinals. Ann. Pharm. Fr. 1990, 48, 103–108. [Google Scholar] [PubMed]
- Lamaison, J.L.; Petitjean-Freytet, C.; Carnat, A. Medicinal Lamiaceae with Antioxidant Properties, a Potential Source of Rosmarinic Acid. Pharm. Acta Helv. 1991, 66, 185. [Google Scholar]
- Nabavi, S.F.; Tenore, G.C.; Daglia, M.; Tundis, R.; Loizzo, M.R.; Mohammad, N.S. The Cellular Protective Effects of Rosmarinic Acid: From Bench to Bedside. Curr. Neurovascular Res. 2015, 12, 98–105. [Google Scholar] [CrossRef] [PubMed]
- Kelsey, N.A.; Wilkins, H.M.; Linseman, D.A. Nutraceutical Antioxidants as Novel Neuroprotective Agents. Molecules 2010, 15, 7792–7814. [Google Scholar] [CrossRef] [PubMed]
- Orhan, I.E.; Senol, F.S.; Ozturk, N.; Akaydin, G.; Sener, B. Profiling of in Vitro Neurobiological Effects and Phenolic Acids of Selected Endemic Salvia Species. Food Chem. 2012, 132, 1360–1367. [Google Scholar] [CrossRef] [PubMed]
- Bigford, G.E.; del Rossi, G. Supplemental Substances Derived from Foods as Adjunctive Therapeutic Agents for Treatment of Neurodegenerative Diseases and Disorders. Adv. Nutr. 2014, 14, 394–403. [Google Scholar] [CrossRef]
- James, B.D.; Leurgans, S.E.; Hebert, L.E.; Scherr, P.A.; Yaffe, K.; Bennett, D.A. Contribution of Alzheimer Disease to Mortality in the United States. Neurology 2014, 82, 1045–1050. [Google Scholar] [CrossRef]
- WHO. Global Action Plan on the Public Health Response to Dementia 2017–2025; WHO Document Production Services: Geneva, Switzerland, 2017; ISBN 978-92-4-151348-7. [Google Scholar]
- Takeda, S. Progression of Alzheimer’s Disease, Tau Propagation, and Its Modifiable Risk Factors. Neurosci. Res. 2019, 141, 36–42. [Google Scholar] [CrossRef]
- Dorszewska, J.; Prendecki, M.; Oczkowska, A.; Dezor, M.; Kozubski, W. Molecular Basis of Familial and Sporadic Alzheimer’s Disease. Curr. Alzheimer Res. 2016, 13, 952–962. [Google Scholar] [CrossRef]
- Rollo, J.L.; Banihashemi, N.; Vafaee, F.; Crawford, J.W.; Kuncic, Z.; Holsinger, R.M. Unraveling the Mechanistic Complexity of Alzheimer’s Disease Through Systems Biology. Alzheimers Dement. 2016, 12, 708–718. [Google Scholar] [CrossRef]
- Penke, B.; Bogár, F.; Paragi, G.; Gera, J.; Fülöp, L. Key Peptides and Proteins in Alzheimer’s Disease. Curr. Protein Pept. Sci. 2019, 20, 577–599. [Google Scholar] [CrossRef] [PubMed]
- Guix, F.X. He Interplay Between Aging-Associated Loss of Protein Homeostasis and Extracellular Vesicles in Neurodegeneration. J. Neurosci. Res. 2020, 98, 262–283. [Google Scholar] [CrossRef] [PubMed]
- Taylor, J.P.; Hardy, J.; Fischbeck, K.H. Toxic Proteins in Neurodegenerative Disease. Science 2002, 296, 1991–1995. [Google Scholar] [CrossRef] [PubMed]
- Hardy, J.; Selkoe, D.J. The Amyloid Hypothesis of Alzheimer’s Disease: Progress and Problems on the Road to Therapeutics. Science 2002, 297, 353–356. [Google Scholar] [CrossRef] [PubMed]
- Karran, E.; de Strooper, B. The Amyloid Cascade Hypothesis: Are We Poised for Success or Failure? J. Neurochem. 2016, 139, 237–252. [Google Scholar] [CrossRef]
- Chun, W.; Johnson, G.V. The Role of Tau Phosphorylation and Cleavage in Neuronal Cell Death. Front. Biosci. 2007, 12, 733–756. [Google Scholar] [CrossRef] [PubMed]
- Mehta, D.; Jackson, R.; Paul, G.; Shi, J.; Sabbagh, M. Why do trials for Alzheimer’s disease drugs keep failing? A discontinued drug perspective for 2010–2015. Expert Opin. Investig. Drugs 2017, 26, 735–739. [Google Scholar] [CrossRef]
- Morris, G.P.; Clark, I.A.; Vissel, B. Questions concerning the role of amyloid-beta in the definition, aetiology and diagnosis of Alzheimer’s disease. Acta Neuropathol. 2018, 136, 663–689. [Google Scholar] [CrossRef]
- Sensi, S.L.; Granzotto, A.; Siotto, M.; Squitti, R. Copper and Zinc Dysregulation in Alzheimer’s Disease. Trends Pharmacol. Sci. 2018, 39, 1049–1063. [Google Scholar] [CrossRef]
- Adlard, P.A.; Bush, A.I. Metals and Alzheimer’s Disease: How Far Have We Come in the Clinic? J. Alzheimers Dis. 2018, 62, 1369–1379. [Google Scholar] [CrossRef] [PubMed]
- Mathys, Z.K.; White, A.R. Copper and Alzheimer’s Disease. Adv. Neurobiol. 2017, 18, 199–216. [Google Scholar] [CrossRef] [PubMed]
- Avan, A.; Hoogenraad, T.U. Zinc and Copper in Alzheimer’s Disease. J. Alzheimers Dis. 2015, 46, 89–92. [Google Scholar] [CrossRef]
- Bulcke, F.; Dringen, R.; Scheiber, I.F. Neurotoxicity of Copper. Adv. Neurobiol. 2017, 18, 313–343. [Google Scholar] [CrossRef] [PubMed]
- Mezzaroba, L.; Alfieri, D.F.; Simão, A.N.C.; Reiche, E.M.V. The Role of Zinc, Copper, Manganese and Iron in Neurodegenerative Diseases. Neurotoxicology 2019, 74, 230–241. [Google Scholar] [CrossRef] [PubMed]
- Gaggelli, E.; Kozlowski, H.; Valensin, D.; Valensin, G. Copper Homeostasis and Neurodegenerative Disorders (Alzheimer’s, Prion, and Parkinson’s Diseases and Amyotrophic Lateral Sclerosis). Chem. Rev. 2006, 106, 1995–2044. [Google Scholar] [CrossRef]
- Kozlowski, H.; Luczkowski, M.; Remelli, M.; Valensin, D. Copper, zinc and iron in neurodegenerative diseases (Alzheimer’s, Parkinso’s and prion diseases). Coord. Chem. Rev. 2012, 256, 2129–2141. [Google Scholar] [CrossRef]
- Hecel, A.; de Ricco, R.; Valensin, D. Influence of membrane environments and copper ions on the structural features of amyloidogenic proteins correlated to neurodegeneration. Coord. Chem. Rev. 2016, 327, 8–19. [Google Scholar] [CrossRef]
- Atrian-Blasco, E.; Gonzalez, P.; Santoro, A.; Alies, B.; Faller, P.; Hureau, C. Cu and Zn coordination to amyloid peptides: From fascinating chemistry to debated pathological relevance. Coord. Chem. Rev. 2018, 371, 38–55. [Google Scholar] [CrossRef]
- Hureau, C. Coordination of redox active metal ions to the amyloid precursor protein and to amyloid-β peptides involved in Alzheimer disease. Part 1: An overview. Coord. Chem. Rev. 2012, 256, 2164–2174. [Google Scholar] [CrossRef]
- Drew, S.C.; Barnham, K. The Heterogeneous Nature of Cu2+ Interactions with Alzheimer’s Amyloid-β Peptide. Acc. Chem. Res. 2011, 44, 1146–1155. [Google Scholar] [CrossRef] [PubMed]
- Migliorini, C.; Porciatti, E.; Luczkowski, M.; Valensin, D. Structural characterization of Cu2+, Ni2+ and Zn2+ binding sites of model peptides associated with neurodegenerative diseases. Coord. Chem. Rev. 2012, 256, 352–368. [Google Scholar] [CrossRef]
- Rana, M.; Sharma, A.K. Cu and Zn Interactions with Aβ Peptides: Consequence of Coordination on Aggregation and Formation of Neurotoxic Soluble Aβ Oligomers. Metallomics 2019, 11, 64–84. [Google Scholar] [CrossRef] [PubMed]
- Faller, P.; Hureau, C. Bioinorganic Chemistry of Copper and Zinc Ions Coordinated to Amyloid-Beta Peptide. Dalton Trans. 2009, 1080–1094. [Google Scholar] [CrossRef] [PubMed]
- Esmieu, C.; Guettas, D.; Conte-Daban, A.; Sabater, L.; Faller, P.; Hureau, C. Copper-Targeting Approaches in Alzheimer’s Disease: How to Improve the Fallouts Obtained From in Vitro Studies. Inorg. Chem. 2019, 58, 13509–13527. [Google Scholar] [CrossRef]
- Stefaniak, E.; Bal, W. CuII Binding Properties of N-Truncated Aβ Peptides: In Search of Biological Function. Inorg. Chem. 2019, 58, 13561–13577. [Google Scholar] [CrossRef]
- Bacchella, C.; Gentili, S.; Bellotti, D.; Quartieri, E.; Draghi, S.; Baratto, M.C.; Remelli, M.; Valensin, D.; Monzani, E.; Nicolis, S.; et al. Binding and Reactivity of Copper to R 1 and R 3 Fragments of Tau Protein. Inorg. Chem. 2020, 59, 274–286. [Google Scholar] [CrossRef]
- de Ricco, R.; Potocki, S.; Kozlowski, H.; Valensin, D. NMR investigations of metal interactions with unstructured soluble protein domains. Coord. Chem. Rev. 2014, 269, 1–12. [Google Scholar] [CrossRef]
- de Gregorio, G.; Biasotto, F.; Hecel, A.; Luczkowski, M.; Kozlowski, H.; Valensin, D. Structural Analysis of copper(I) Interaction with Amyloid β Peptide. J. Inorg. Biochem. 2019, 195, 31–38. [Google Scholar] [CrossRef]
- Giampietro, R.; Spinelli, F.; Contino, M.; Colabufo, N.A. The Pivotal Role of Copper in Neurodegeneration: A New Strategy for the Therapy of Neurodegenerative Disorders. Mol. Pharm. 2018, 15, 808–820. [Google Scholar] [CrossRef]
- Savelieff, M.G.; Nam, G.; Kang, J.; Lee, H.J.; Lee, M.; Lim, M. Development of Multifunctional Molecules as Potential Therapeutic Candidates for Alzheimer’s Disease, Parkinson’s Disease, and Amyotrophic Lateral Sclerosis in the Last Decade. Chem. Rev. 2019, 119, 1221–1322. [Google Scholar] [CrossRef] [PubMed]
- Nesi, G.; Sestito, S.; Digiacomo, M.; Rapposelli, S. Oxidative Stress, Mitochondrial Abnormalities and Proteins Deposition: Multitarget Approaches in Alzheimer’s Disease. Curr. Top. Med. Chem. 2017, 17, 3062–3079. [Google Scholar] [CrossRef] [PubMed]
- Tegoni, M.; Valensin, D.; Loso, L.; Remelli, M. Copper Chelators: Chemical Properties and Bio-Medical Applications. Curr. Med. Chem. 2014, 21, 3785–3818. [Google Scholar] [CrossRef] [PubMed]
- Valensin, D.; Gabbiani, C.; Messori, L. Metal compounds as inhibitors of β-amyloid aggregation. Perspectives for an innovative metallotherapeutics on Alzheimer’s disease. Coord. Chem. Rev. 2012, 256, 2357–2366. [Google Scholar] [CrossRef]
- Piemontese, L.; Vitucci, G.; Catto, M.; Laghezza, A.; Rullo, F.M.P.M.; Loiodice, F.; Capriati, V.; Solfrizzo, M. Natural Scaffolds with Multi-Target Activity for the Potential Treatment of Alzheimer’s Disease. Molecules 2018, 29, 2182. [Google Scholar] [CrossRef] [PubMed]
- Alkam, T.; Nitta, A.; Mizoguchi, H.; Itoh, A.; Nabeshima, T. A Natural Scavenger of Peroxynitrites, Rosmarinic Acid, Protects Against Impairment of Memory Induced by Abeta(25–35). Behav. Brain Res. 2007, 180, 139–145. [Google Scholar] [CrossRef]
- Taguchi, R.; Hatayama, K.; Takahashi, T.; Hayashi, T.; Sato, Y.; Sato, D.; Ohta, K.; Nakano, H.; Seki, C.; Endo, Y.; et al. Structure-activity Relations of Rosmarinic Acid Derivatives for the Amyloid β Aggregation Inhibition and Antioxidant Properties. Eur. J. Med. Chem. 2017, 138, 1066–1075. [Google Scholar] [CrossRef]
- Hase, T.; Shishido, S.; Yamamoto, S.; Yamashita, R.; Nukima, H.; Taira, S.; Toyoda, T.; Abe, K.; Hamaguchi, T.; Ono, K.; et al. Rosmarinic Acid Suppresses Alzheimer’s Disease Development by Reducing Amyloid β Aggregation by Increasing Monoamine Secretion. Sci. Rep. 2019, 9, 1–13. [Google Scholar] [CrossRef]
- Ono, K.; Hasegawa, K.; Naiki, H.; Yamada, M. Curcumin Has Potent Anti-Amyloidogenic Effects for Alzheimer’s Beta-Amyloid Fibrils in Vitro. J. Neurosci. Res. 2004, 75, 742–750. [Google Scholar] [CrossRef]
- Hamaguchi, T.; Ono, K.; Murase, A.; Yamada, M. Phenolic Compounds Prevent Alzheimer’s Pathology Through Different Effects on the Amyloid-Beta Aggregation Pathway. Am. J. Pathol. 2009, 175, 2557–2565. [Google Scholar] [CrossRef]
- Ono, K.; Li, L.; Takamura, Y.; Yoshiike, Y.; Zhu, L.; Han, F.; Mao, X.; Ikeda, T.; Takasaki, J.; Nishijo, H.; et al. Phenolic Compounds Prevent Amyloid β-Protein Oligomerization and Synaptic Dysfunction by Site-Specific Binding. J. Biol. Chem. 2012, 287, 14631–14643. [Google Scholar] [CrossRef] [PubMed]
- Iuvone, T.; de Filippis, D.; Esposito, G.; D′Amico, A.; Izzo, A.A. The Spice Sage and Its Active Ingredient Rosmarinic Acid Protect PC12 Cells from Amyloid-Beta Peptide-Induced Neurotoxicity. J. Pharmacol. Exp. Ther. 2006, 317, 1143–1149. [Google Scholar] [CrossRef] [PubMed]
- Rong, H.; Liang, Y.; Niu, Y. Rosmarinic Acid Attenuates β-Amyloid-Induced Oxidative Stress via Akt/GSK-3β/Fyn-mediated Nrf2 Activation in PC12 Cells. Free Radic. Biol. Med. 2018, 120, 114–123. [Google Scholar] [CrossRef] [PubMed]
- Hwang, T.L.; Shaka, A.J.J. Water suppression that works. Excitation sculpting using arbitrary waveforms and pulsed field gradients. J. Magn. Reson. Ser. A 1995, 112, 275–279. [Google Scholar] [CrossRef]
- Akoury, E. Isolation and Structural Elucidation of Rosmarinic Acid by Nuclear Magnetic Resonance Spectroscopy. Am. Res. J. Chem. 2017, 2, 17–23. [Google Scholar]
- Bertini, I.; Luchinat, C. NMR of Paramagnetic Substances. Coord. Chem. Rev. 1996, 150, 1–296. [Google Scholar]
- Solomon, I. Relaxation Processes in a System of Two Spins. Phys. Rev. 1955, 99, 559–565. [Google Scholar] [CrossRef]
- ISO. ISO 10995-5:2009, Biological evaluation of medical devices—Part 5: Tests for cytotoxicity: In vitro methods; ISO: Geneva, Switzerland, 2009. [Google Scholar]
- Lamponi, S.; Leone, G.; Consumi, M.; Nelli, N.; Magnani, A. Porous multi-layered composite hydrogel as cell substrate for in vitro culture of chondrocytes. Int. J. Polym. Mater. Polym. Biomater. 2020. [Google Scholar] [CrossRef]
- Peisach, J.; Blumberg, W.E. Structural Implications Derived From the Analysis of Electron Paramagnetic Resonance Spectra of Natural and Artificial Copper Proteins. Arch. Biochem. Biophys. 1974, 165, 691–708. [Google Scholar] [CrossRef]
- Bennett, B.; Kowalski, J.M. EPR Methods for Biological Cu(II): L-Band CW and NARS. Methods Enzymol. 2015, 563, 341–361. [Google Scholar] [CrossRef]
- Danafetal, N.A. Photophysical properties of neutral and dissociated forms of rosmarinic acid. J. Lumin. 2016, 175, 50–56. [Google Scholar] [CrossRef]
- Valensin, D.; Gajda, K.; Gralka, E.; Valensin, G.; Kamysz, W.H. Kozlowski Copper Binding to Chicken and Human Prion Protein Amylodogenic Regions: Differences and Similarities Revealed by Ni2+ as a Diamagnetic Probe. J. Inorg. Biochem. 2010, 104, 71–78. [Google Scholar] [CrossRef] [PubMed]
- Valensin, D.; Camponeschi, F.; Luczkowski, M.; Baratto, M.C.; Remelli, M.; Valensin, G.; Kozlowski, H. The Role of His-50 of α-Synuclein in Binding Cu(II): pH Dependence, Speciation, Thermodynamics and Structure. Metallomics 2011, 3, 292–302. [Google Scholar] [CrossRef]
- Remelli, M.; Ceciliato, C.; Guerrini, R.; Kolkowska, P.; Krzywoszynska, K.; Salvadori, S.; Valensin, D.; Watly, J.; Kozlowski, H. DOES Hemopressin Bind Metal Ions in Vivo? Dalton Trans. 2016, 45, 18267–18280. [Google Scholar] [CrossRef] [PubMed]
- Remelli, M.; Valensin, D.; Toso, L.; Gralka, E.; Guerrini, R.; Marzola, E.; Kozłowski, H. Thermodynamic and Spectroscopic Investigation on the Role of Met Residues in Cu(II) Binding to the Non-Octarepeat Site of the Human Prion Protein. Metallomics 2012, 4, 794–806. [Google Scholar] [CrossRef] [PubMed]
- Airoldi, C.; Sironi, E.; Dias, C.; Marcelo, F.; Martins, A.; Rauter, A.P.; Nicotra, F.; Jimenez-Barbero, J. Natural Compounds Against Alzheimer’s Disease: Molecular Recognition of Aβ1-42 Peptide by Salvia Sclareoides Extract and Its Major Component, Rosmarinic Acid, as Investigated by NMR. Chem. Asian J. 2013, 8, 596–602. [Google Scholar] [CrossRef]
- Shenberger, Y.; Marciano, O.; Gottlieb, H.E.; Ruthsteinet, S. Insights into the N-terminal Cu(II) and Cu(I) binding sites of the human copper transporter CTR1. J. Coord. Chem. 2018, 71, 1985–2002. [Google Scholar] [CrossRef]
- Rasia, R.; Bertoncini, C.W.; Marsh, D.; Hoyer, W.; Cherny, D.; Zweckstetter, M.; Griesinger, C.; Jovin, T.M.; Fernández, C.O. Structural Characterization of copper(II) Binding to Alpha-Synuclein: Insights Into the Bioinorganic Chemistry of Parkinson’s Disease. Proc. Natl. Acad. Sci. USA 2005, 102, 4294–4299. [Google Scholar] [CrossRef]
- Kowalik-Jankowska, T.; Ruta, M.; Wiśniewska, K.; Lankiewicz, L. Coordination Abilities of the 1–16 and 1–28 Fragments of Beta-Amyloid Peptide Towards copper(II) Ions: A Combined Potentiometric and Spectroscopic Study. J. Inorg. Biochem. 2003, 95, 270–282. [Google Scholar] [CrossRef]
- Karr, J.W.; Kaupp, L.J.; Szalai, V.A. Amyloid-beta Binds Cu2+ in a Mononuclear Metal Ion Binding Site. J. Am. Chem. Soc. 2004, 126, 13534–13538. [Google Scholar] [CrossRef]
- Guilloreau, L.; Damian, L.; Coppel, Y.; Mazarguil, H.; Winterhalter, M.; Faller, P. Structural and Thermodynamical Properties of CuII amyloid-beta16/28 Complexes Associated with Alzheimer’s Disease. J. Biol. Inorg. Chem. 2006, 11, 1024–1038. [Google Scholar] [CrossRef] [PubMed]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kola, A.; Hecel, A.; Lamponi, S.; Valensin, D. Novel Perspective on Alzheimer’s Disease Treatment: Rosmarinic Acid Molecular Interplay with Copper(II) and Amyloid β. Life 2020, 10, 118. https://doi.org/10.3390/life10070118
Kola A, Hecel A, Lamponi S, Valensin D. Novel Perspective on Alzheimer’s Disease Treatment: Rosmarinic Acid Molecular Interplay with Copper(II) and Amyloid β. Life. 2020; 10(7):118. https://doi.org/10.3390/life10070118
Chicago/Turabian StyleKola, Arian, Aleksandra Hecel, Stefania Lamponi, and Daniela Valensin. 2020. "Novel Perspective on Alzheimer’s Disease Treatment: Rosmarinic Acid Molecular Interplay with Copper(II) and Amyloid β" Life 10, no. 7: 118. https://doi.org/10.3390/life10070118
APA StyleKola, A., Hecel, A., Lamponi, S., & Valensin, D. (2020). Novel Perspective on Alzheimer’s Disease Treatment: Rosmarinic Acid Molecular Interplay with Copper(II) and Amyloid β. Life, 10(7), 118. https://doi.org/10.3390/life10070118