Novel Methylation Patterns Predict Outcome in Uveal Melanoma
Abstract
1. Introduction
2. Methods
2.1. Dataset
2.2. Removal of Poor-Quality Probes
2.3. Normalization of Samples
2.4. Hierarchical Clustering of the Top Differentially Methylated Probes (DMPs)
2.5. Analysis of DMPs between the Two Groups
3. Results
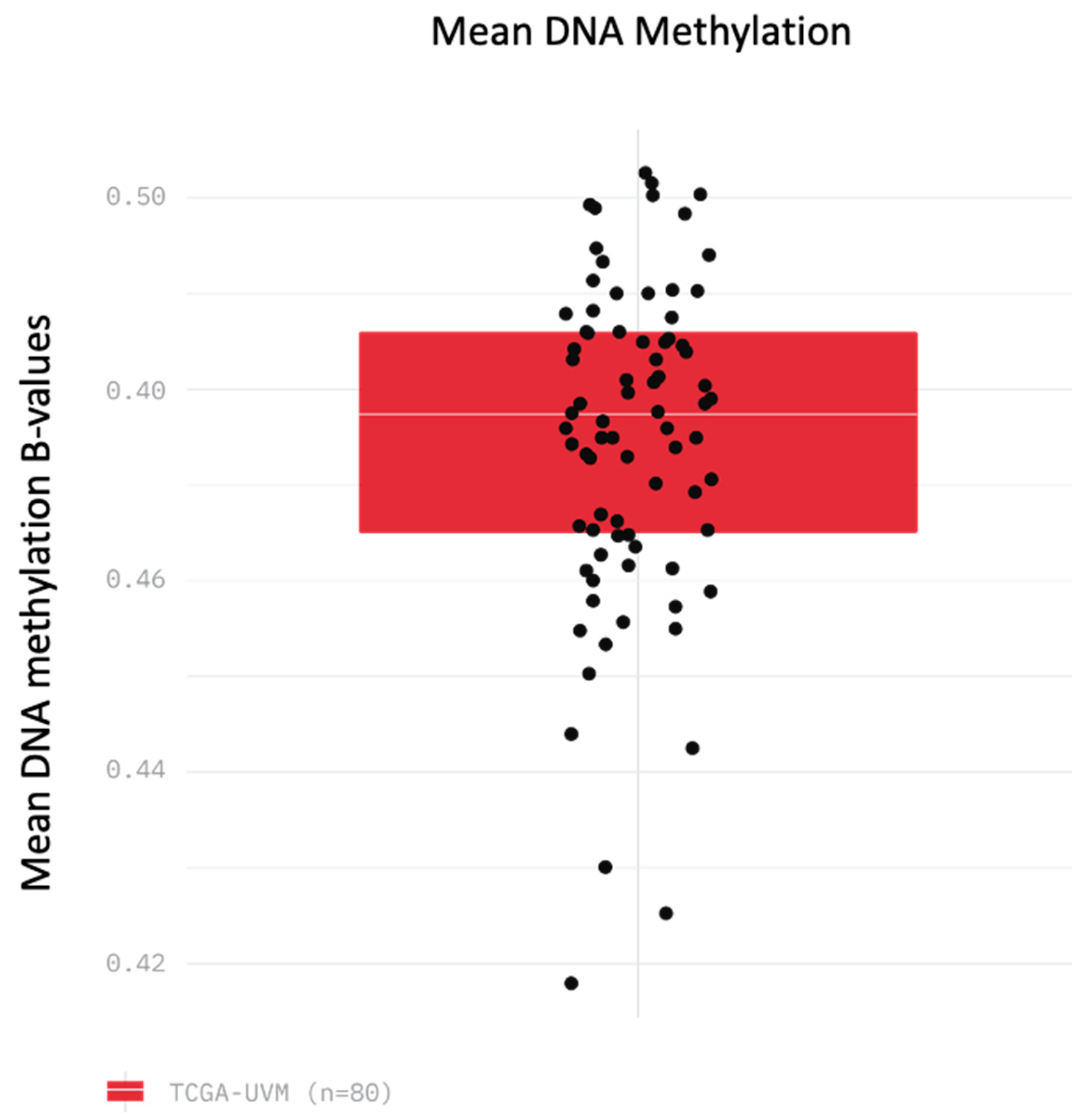
3.1. Similar Levels of Overall Methylation Are Seen in 80 Cases of UM through Analysis of TCGA Data
3.2. Unsupervised Clustering Analysis Reveals Two Main Methylation Patterns in This Cohort
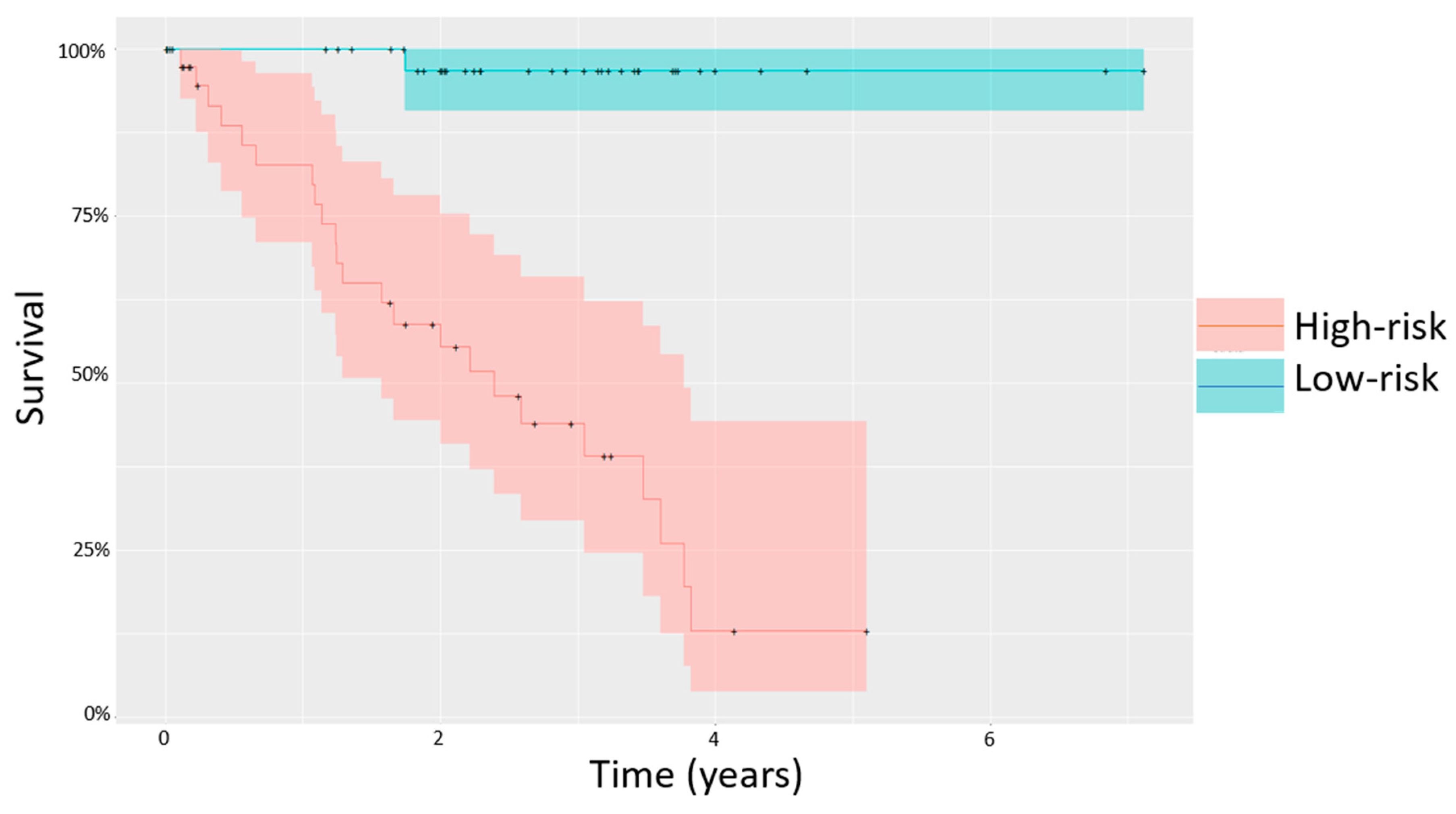
3.3. Methylation Patterns Are Significantly Associated with Outcome and Effectively Stratify Patients into High and Low Risk Groups
3.4. The High and Low-Risk Groups Differed in Clinical and Histopathological Features of the Ocular Tumors
3.5. Methylation Stratification Highly Correlated with Genomic Factors Associated with Metastasis
3.6. Gene Ontology (GO) Analysis Reveals Enrichment for Genes Involved in Signal Transduction Pathways in the High-Risk Group
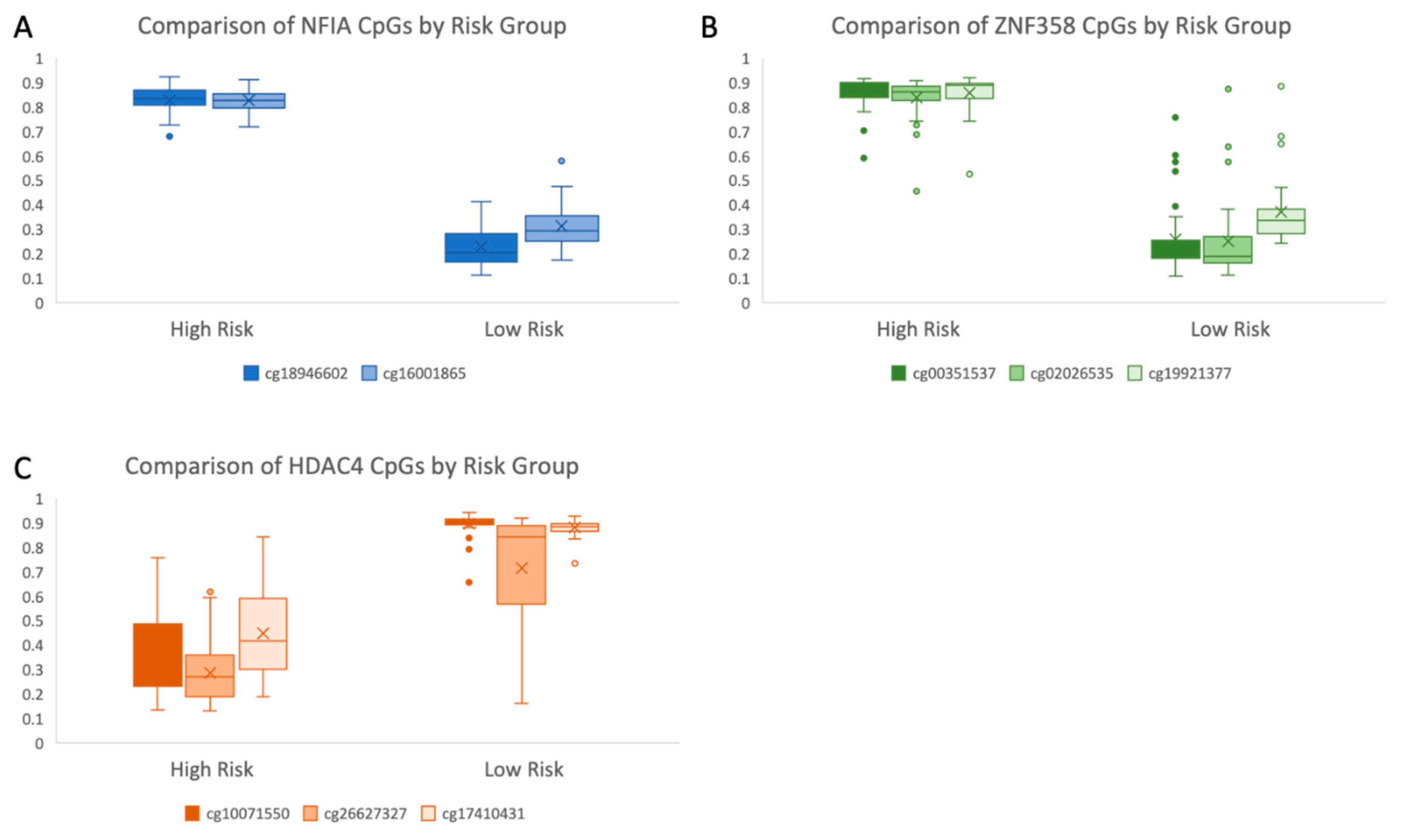
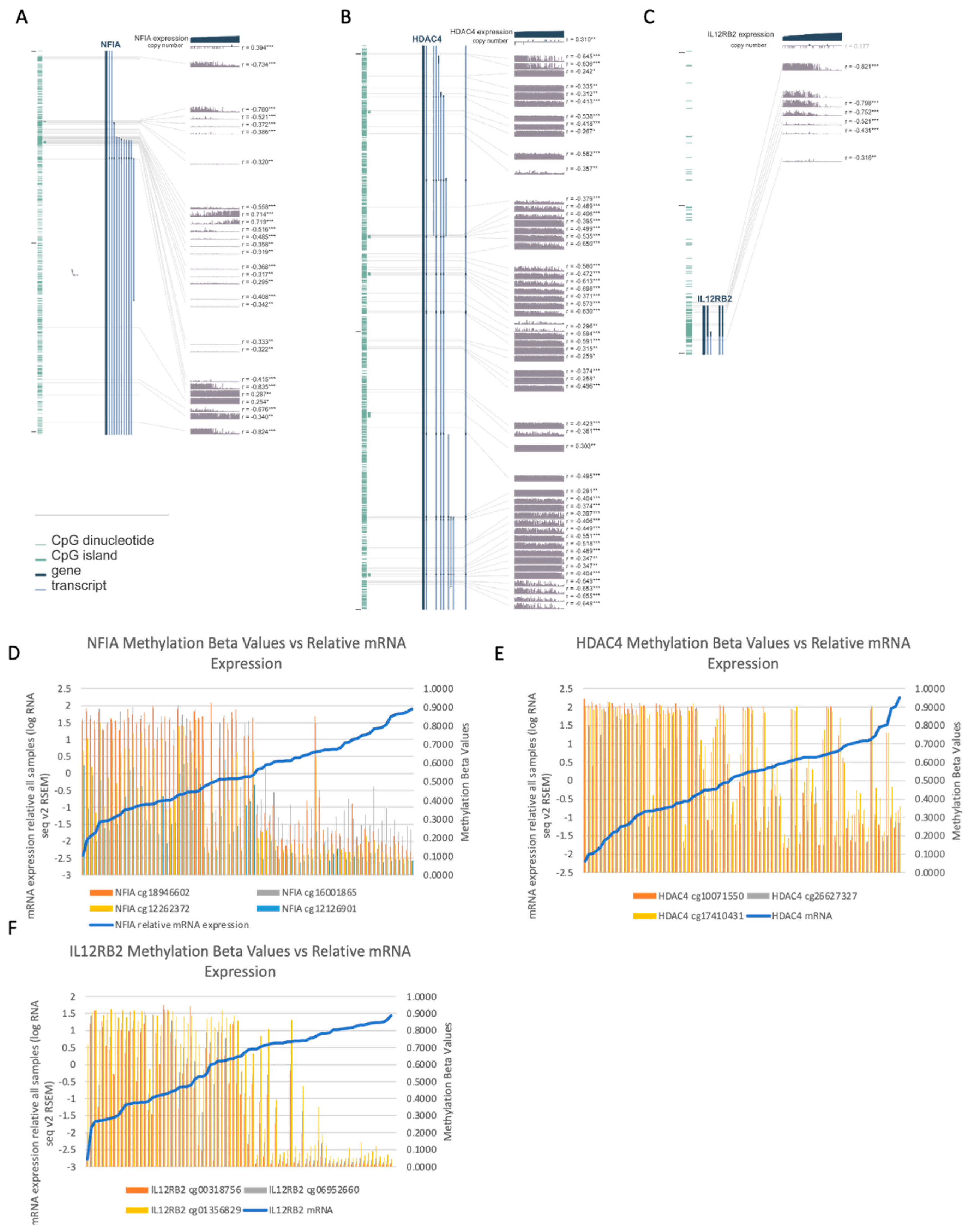
3.7. Hypermethylation of Tumor Suppressor Genes and Transcriptional and Epigenetic Regulators Are Seen in the High-Risk Group
3.8. Changes in Methylation Are Found in Genes with a Role in Epigenetic Modifications
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Singh, A.D.; Mary, E.T.; Allan, K.T. Uveal Melanoma: Trends in Incidence, Treatment, and Survival. Ophthalmology 2011, 118, 1881–1885. [Google Scholar] [CrossRef] [PubMed]
- Carvajal, R.D.; Gary, K.S.; Tongalp, T.; Brian, M.; Jasmine, H.F.; Paul, D.N. Metastatic Disease from Uveal Melanoma: Treatment Options and Future Prospects. Br. J. Ophthalmol. 2017, 101, 38–44. [Google Scholar] [CrossRef] [PubMed]
- Barker, C.A.; April, K.S. New Nccn Guidelines for Uveal Melanoma and Treatment of Recurrent or Progressive Distant Metastatic Melanoma. J. Natl. Compr. Cancer Netw. 2018, 16, 646–650. [Google Scholar] [CrossRef] [PubMed]
- Mosci, C.; Francesco, B.L.; Annalisa, B.; Sofia, M.; Joël, H.; Luca, A.; Mauro, T. Comparison of Clinical Outcomes for Patients with Large Choroidal Melanoma after Primary Treatment with Enucleation or Proton Beam Radiotherapy. Ophthalmologica 2012, 227, 190–196. [Google Scholar] [PubMed]
- Singh, A.D.; Shields, C.L.; Shields, J.A. Prognostic Factors in Uveal Melanoma. Melanoma Res. 2001, 11, 255–263. [Google Scholar]
- Ewens, K.G.; Peter, A.K.; Jennifer, R.-Y.; Saad, A.-D.; Maria, C.D.L.; Carlos, G.B.; Carol, L.S.; Arupa, G. Genomic Profile of 320 Uveal Melanoma Cases: Chromosome 8p-Loss and Metastatic Outcome. Investig. Ophthalmol. Vis. Sci. 2013, 54, 5721–5729. [Google Scholar] [CrossRef] [PubMed]
- Kilic, E.; van Walter, G.; Elisabeth, L.; Berna, H.B.; Marjan, E.T.; Cornelia, M.M.; Dion, P.; de Annelies, K.; Gregorius, P.M.L. Clinical and Cytogenetic Analyses in Uveal Melanoma. Investig. Ophthalmol. Vis. Sci. 2006, 47, 3703–3707. [Google Scholar]
- Koopmans, A.E.; Jolanda, V.; Dion, P.; Nicole, C.N.; Emine, K.; de Annelies, K. Patient Survival in Uveal Melanoma is Not Affected by Oncogenic Mutations in Gnaq and Gna11. Br. J. Cancer 2013, 109, 493–496. [Google Scholar] [CrossRef]
- Yavuzyigitoglu, S.; Anna, E.K.; Robert, M.V.; Jolanda, V.; Bert, E.; Alice, V.B.; Dion, P.; Emine, K.; de Annelies, K.; Rotterdam Ocular Melanoma Study Group. Uveal Melanomas with Sf3b1 Mutations: A Distinct Subclass Associated with Late-Onset Metastases. Ophthalmology 2016, 123, 1118–1128. [Google Scholar] [CrossRef]
- Baylin, S.B. DNA Methylation and Gene Silencing in Cancer. Nat. Rev. Clin. Oncol. 2005, 2, S4. [Google Scholar] [CrossRef]
- Witte, T.; Christoph, P.; Clarissa, G. Pan-Cancer Patterns of DNA Methylation. Genome Med. 2014, 6, 66. [Google Scholar] [CrossRef] [PubMed]
- Ehrlich, M. DNA Methylation in Cancer: Too Much, but Also Too Little. Oncogene 2002, 21, 5400–5413. [Google Scholar] [CrossRef] [PubMed]
- Kulis, M.; Manel, E. DNA Methylation and Cancer. In Advances in Genetics; Elsevier: Amsterdam, The Netherlands, 2010; pp. 27–56. [Google Scholar]
- Esteller, M. Aberrant DNA Methylation as a Cancer-Inducing Mechanism. Annu. Rev. Pharmacol. Toxicol. 2005, 45, 629–656. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.A.; Stephen, B.B. The Epigenomics of Cancer. Cell 2007, 128, 683–692. [Google Scholar] [CrossRef] [PubMed]
- Merhavi, E.; Yoram, C.; Bat, C.R.A.; Shahar, F.; Itay, C.; Jacob, P.; Nitza, G.-C. Promoter Methylation Status of Multiple Genes in Uveal Melanoma. Investig. Ophthalmol. Vis. Sci. 2007, 48, 4403–4406. [Google Scholar] [CrossRef][Green Version]
- Robertson, A.G.; Juliann, S.; Christina, Y.; Ewan, A.G.; Junna, O.; Karen, L.M.; Julian, M.H.; Vladislav, U.; Vonn, W.; Ludmila, D. Integrative Analysis Identifies Four Molecular and Clinical Subsets in Uveal Melanoma. Cancer Cell 2017, 32, 204–220.e15. [Google Scholar] [CrossRef]
- Field, M.G.; Jeffim, N.K.; Parker, L.B.; Louie, Z.C.; Karam, A.A.; Christina, L.D.; Stefan, K.; William, J.H. Bap1 Loss is Associated with DNA Methylomic Repatterning in Highly Aggressive Class 2 Uveal Melanomas. Clin. Cancer Res. 2019, 25, 5663–5673. [Google Scholar] [CrossRef]
- Cheng, Y.; Cai, H.; Manni, W.; Xuelei, M.; Fei, M.; Shengyong, Y.; Junhong, H.; Xiawei, W. Targeting Epigenetic Regulators for Cancer Therapy: Mechanisms and Advances in Clinical Trials. Signal Transduct. Target. Ther. 2019, 4, 1–39. [Google Scholar] [CrossRef]
- Fardi, M.; Saeed, S.; Majid, F.H. Epigenetic Mechanisms as a New Approach in Cancer Treatment: An Updated Review. Genes Dis. 2018, 5, 304–311. [Google Scholar] [CrossRef]
- Aryee, M.J.; Andrew, E.J.; Hector, C.-B.; Christine, L.-A.; Andrew, P.F.; Kasper, D.H.; Rafael, A.I. Minfi: A Flexible and Comprehensive Bioconductor Package for the Analysis of Infinium DNA Methylation Microarrays. Bioinformatics 2014, 30, 1363–1369. [Google Scholar] [CrossRef]
- Maksimovic, J.; Lavinia, G.; Alicia, O. Swan: Subset-Quantile within Array Normalization for Illumina Infinium Humanmethylation450 Beadchips. Genome Biol. 2012, 13, R44. [Google Scholar] [CrossRef]
- Touleimat, N.; Jörg, T. Complete Pipeline for Infinium® Human Methylation 450k Beadchip Data Processing Using Subset Quantile Normalization for Accurate DNA Methylation Estimation. Epigenomics 2012, 4, 325–341. [Google Scholar] [CrossRef] [PubMed]
- Harbour, J.W.; Michael, D.O.; Elisha, D.R.; Shenghui, D.; Li, C.; Lori, A.W.; Laurin, M.C.; Katie, A.M.; Cynthia, H.; Anne, M.B. Frequent Mutation of Bap1 in Metastasizing Uveal Melanomas. Science 2010, 330, 1410–1413. [Google Scholar] [CrossRef] [PubMed]
- Landreville, S.; Olga, A.A.; Katie, A.M.; Zachary, T.K.; Michael, D.O.; Ryan, S.L.; Anne, M.B.; William, J.H. Histone Deacetylase Inhibitors Induce Growth Arrest and Differentiation in Uveal Melanoma. Clin. Cancer Res. 2012, 18, 408–416. [Google Scholar] [CrossRef] [PubMed]
- van de Nes, J.A.P.; Jasmin, N.; Stefan, K.; Claudia, H.D.M.; Thomas, H.; Dietmar, R.L.; Michael, Z. Comparing the Prognostic Value of Bap1 Mutation Pattern, Chromosome 3 Status, and Bap1 Immunohistochemistry in Uveal Melanoma. Am. J. Surg. Pathol. 2016, 40, 796–805. [Google Scholar] [CrossRef]
- El-Deiry, W.S.; Barry, T.; Joel, W.N. Tumor Evolution, Heterogeneity, and Therapy for Our Patients with Advanced Cancer: How Far Have We Come? Am. Soc. Clin. Oncol. Educ. Book 2017, 37, e8–e15. [Google Scholar] [CrossRef]
- Shen, S.Y.; Rajat, S.; Gordon, F.; Ankur, C.; Michael, H.A.R.; Dianne, C.; Philip, C.Z.; Ayelet, B.; Ting, T.W.; Tiantian, L. Sensitive Tumour Detection and Classification Using Plasma Cell-Free DNA Methylomes. Nature 2018, 563, 579. [Google Scholar] [CrossRef] [PubMed]
- Onken, M.D.; Lori, A.W.; Devron, H.C.; James, J.A.; Zelia, M.C.; Eric, N.; Thomas, M.A.; Michael, M.A., Jr.; David, S.B.; Paul, T.F. Collaborative Ocular Oncology Group Report Number 1: Prospective Validation of a Multi-Gene Prognostic Assay in Uveal Melanoma. Ophthalmology 2012, 119, 1596–1603. [Google Scholar] [CrossRef]
- Saraiva, V.S.; Caissie, A.L.; Segal, L.; Edelstein, C.; Burnier, M.N., Jr. Immunohistochemical Expression of Phospho-Akt in Uveal Melanoma. Melanoma Res. 2005, 15, 245–250. [Google Scholar] [CrossRef]
- Chetram, M.A.; Cimona, V.H. Pten Regulation of Erk1/2 Signaling in Cancer. J. Recept. Signal Transduct. 2012, 32, 190–195. [Google Scholar] [CrossRef]
- Abdel-Rahman, M.H.; Ying, Y.; Zhou, X.-P.; Elson, L.C.; Frederick, H.D.; Charis, E. High Frequency of Submicroscopic Hemizygous Deletion Is a Major Mechanism of Loss of Expression of Pten in Uveal Melanoma. J. Clin. Oncol. 2006, 24, 288–295. [Google Scholar] [CrossRef]
- Zuidervaart, W.; Van Nieuwpoort, F.; Stark, M.; Dijkman, R.; Packer, L.; Borgstein, A.M.; Pavey, S.; van der Velden, P.; Out, C.; Jager, M.J. Activation of the Mapk Pathway is a Common Event in Uveal Melanomas Although It Rarely Occurs through Mutation of Braf or Ras. Br. J. Cancer 2005, 92, 2032–2038. [Google Scholar] [CrossRef] [PubMed]
- Shoushtari, A.N.; Richard, D.C. Gnaq and Gna11 Mutations in Uveal Melanoma. Melanoma Res. 2014, 24, 525–534. [Google Scholar] [CrossRef] [PubMed]
- Krantz, B.A.; Nikita, D.; Kimberly, M.K.; Brian, P.M.; Richard, D.C. Uveal Melanoma: Epidemiology, Etiology, and Treatment of Primary Disease. Clin. Ophthalmol. 2017, 11, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, M.; Toshihiko, I.; Takahiro, N.; Rieko, K.; Akira, I.; Kenzo, H.; Yukio, N.; Takehiko, F. Aberrant Methylation of Il-12rβ2 Gene in Lung Adenocarcinoma Cells is Associated with Unfavorable Prognosis. Ann. Surg. Oncol. 2007, 14, 2636–2642. [Google Scholar] [CrossRef]
- Airoldi, I.; Emma, D.C.; Claudia, C.; Giuseppe, T.; Tommaso, D.; Emanuela, O.; Morihiro, W.; Domenico, R.; Vito, P. Endogenous Il-12 Triggers an Antiangiogenic Program in Melanoma Cells. Proc. Natl. Acad. Sci. USA 2007, 104, 3996–4001. [Google Scholar] [CrossRef] [PubMed]
- Tugues, S.; Burkhard, S.H.; Ohs, I.; Vrohlings, M.; Nussbaum, K.; Berg, J.V.; Kulig, P.; Becher, B. New Insights into Il-12-Mediated Tumor Suppression. Cell Death Differ. 2015, 22, 237. [Google Scholar] [CrossRef] [PubMed]
- Rajaii, F.; Laura, A.; Raymond, E.; Shannath, L.M.; James, T.H.; Charles, G.E. The Demethylating Agent 5-Aza Reduces the Growth, Invasiveness, and Clonogenicity of Uveal and Cutaneous Melanoma. Investig. Ophthalmol. Vis. Sci. 2014, 55, 6178–6186. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gonçalves, J.; Michael, F.E.; Fernanda, F.-F.; Andrew, E.A.; William, J.H.; Jonathan, D.L.; Márcia, R.W.; Keiran, S.M.S. Decitabine Limits Escape from Mek Inhibition in Uveal Melanoma. Pigment Cell Melanoma Res. 2020, 33, 507–514. [Google Scholar] [CrossRef] [PubMed]
- Jansen, Y.J.L.; Gontran, V.; Kelly, S.; Van Dam, P.-J.; Teofila, S.; Mark, K.; Jean-Luc, B.V.L.; Bart, N. Phase I Clinical Trial of Decitabine (5-Aza-2′-Deoxycytidine) Administered by Hepatic Arterial Infusion in Patients with Unresectable Liver-Predominant Metastases. ESMO Open 2019, 4, e000464. [Google Scholar] [CrossRef]
- Koch, A.; Tim, D.M.; Jana, J.; Wim, V.C. Mexpress: Visualizing Expression, DNA Methylation and Clinical Tcga Data. BMC Genom. 2015, 16, 636. [Google Scholar] [CrossRef] [PubMed]
- Koch, A.; Jana, J.; Wim, V.C.; van Manon, E.; De Tim, M. Mexpress Update 2019. Nucleic Acids Res. 2019, 47, 561–565. [Google Scholar] [CrossRef]
- Fagerberg, L.; Björn, M.H.; Per, O.; Caroline, K.; Dijana, D.; Jacob, O.; Masato, H.; Simin, T.; Angelika, D.; Karolina, E. Analysis of the Human Tissue-Specific Expression by Genome-Wide Integration of Transcriptomics and Antibody-Based Proteomics. Mol. Cell. Proteom. 2014, 13, 397–406. [Google Scholar] [CrossRef] [PubMed]
- Song, H.-R.; Ignacio, G.-G.; Greg, S.S.; Deborah, L.C.; Richard, S.; Floyd, H.G.; Benjamin, D.; Anat, E.-E. Nuclear Factor Ia Is Expressed in Astrocytomas and Is Associated with Improved Survival. Neuro Oncol. 2010, 12, 122–132. [Google Scholar] [CrossRef]
- Fane, M.; Lachlan, H.; Aaron, G.S.; Michael, P. Nuclear Factor One Transcription Factors as Epigenetic Regulators in Cancer. Int. J. Cancer 2017, 140, 2634–2641. [Google Scholar] [CrossRef] [PubMed]
- Dratviman-Storobinsky, O.; Yoram, C.; Shahar, F.; Efrat, M.-S.; Shimrit, D.-B.E.; Natalia, B.; Jacob, P.; Nitza, G.-C. The Role of Rassf1a in Uveal Melanoma. Investig. Ophthalmol. Vis. Sci. 2012, 53, 2611–2619. [Google Scholar] [CrossRef][Green Version]
- Cassandri, M.; Artem, S.; Flavia, N.; Consuelo, P.; Massimiliano, A.; Michal, M.; Gerry, M.; Giuseppe, R. Zinc-Finger Proteins in Health and Disease. Cell Death Discov. 2017, 3, 17071. [Google Scholar] [CrossRef]
- Jen, J.; Wang, Y.C. Zinc Finger Proteins in Cancer Progression. J. Biomed. Sci. 2016, 23, 53. [Google Scholar] [CrossRef]
- Andrew, S.D.; Michael, J.K.; Michael, A.W.; Fiona, C.M.; Derrick, E.R.; Lois, M.M. Structural Variation in a Novel Zinc Finger Protein and Investigation of Its Role in Hirschsprung Disease. Gene Funct. Dis. 2002, 3, 69–76. [Google Scholar] [CrossRef]
- Kuznetsov, J.N.; Tristan, H.A.; Dawn, A.O.; Stefan, K.; Matthew, G.F.; Michael, A.D.; Daniel, A.R.; Mary, L.K.; William, J.H. Bap1 Regulates Epigenetic Switch from Pluripotency to Differentiation in Developmental Lineages Giving Rise to Bap1-Mutant Cancers. Sci. Adv. 2019, 5, eaax1738. [Google Scholar] [CrossRef]



| Number of Patients | 80 |
|---|---|
| Sex (M:F) | 45:35 |
| Number of Deaths | 24 |
| Age at diagnosis (average, years (range)) | 62 (22–86) |
| Follow up (average, days (range)) | 767 (4–2600) |
| Low-Risk | High-Risk | ||
|---|---|---|---|
| Number of Patients | 40 | 40 | |
| Sex (M:F) | 22:18 | 23:17 | * Χ2: 0.0508, p = 0.822 |
| Age at diagnosis (average, years (range)) | 59 (22–79) | 65 (41–86) | ** p = 0.423 |
| Follow up (average, days (range)) | 973 (6–2600) | 560 (4–1862) | |
| Metastasis | 4 | 23 | * Χ2: 20.18, p < 0.00001 |
| Death (from metastasis or unspecified) | 1 | 23 | * Χ2: 28.81, p < 0.00001 |
| Low-Risk | High-Risk | Significance | |
|---|---|---|---|
| Tumor thickness (mm) | 9.99 | 10.9 | p = 0.171 |
| Tumor diameter (mm) | 16.15 | 17.72 | p = 0.044 |
| Cell type (% spindle:epithelioid) | ~80:20 | ~50:50 | |
| Presence of closed connective loops | 13 | 30 | X2 = 14.5, p = 0.00013 |
| Extraocular extension | 2 | 5 | X2 = 1.4, p = 0.23 |
| Genetic Alteration | Low-Risk | High-Risk | Significance |
|---|---|---|---|
| GNAQ mutations | 25 | 15 | * Χ2: 5, p = 0.0253 |
| GNA11 mutations | 14 | 22 | * Χ2: 3.23, p = 0.0722 |
| BAP1 mutation | 0 | 35 | ** p < 0.00001 |
| Chromosome 3 loss | 3 | 35 | * Χ2: 51.33, p < 0.00001 |
| Chromosome 6p gain | 33 | 12 | * Χ2: 22.4, p < 0.00001 |
| Chromosome 8q gain | 23 | 37 | * Χ2: 13.06 p = 0.0003 |
| Chromosome 1 loss | 7 | 11 | * Χ2: 1.147, p = 0.284 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ferrier, S.T.; Burnier, J.V. Novel Methylation Patterns Predict Outcome in Uveal Melanoma. Life 2020, 10, 248. https://doi.org/10.3390/life10100248
Ferrier ST, Burnier JV. Novel Methylation Patterns Predict Outcome in Uveal Melanoma. Life. 2020; 10(10):248. https://doi.org/10.3390/life10100248
Chicago/Turabian StyleFerrier, Sarah Tadhg, and Julia Valdemarin Burnier. 2020. "Novel Methylation Patterns Predict Outcome in Uveal Melanoma" Life 10, no. 10: 248. https://doi.org/10.3390/life10100248
APA StyleFerrier, S. T., & Burnier, J. V. (2020). Novel Methylation Patterns Predict Outcome in Uveal Melanoma. Life, 10(10), 248. https://doi.org/10.3390/life10100248





