Mobile Elements in Ray-Finned Fish Genomes
Abstract
:1. Introduction
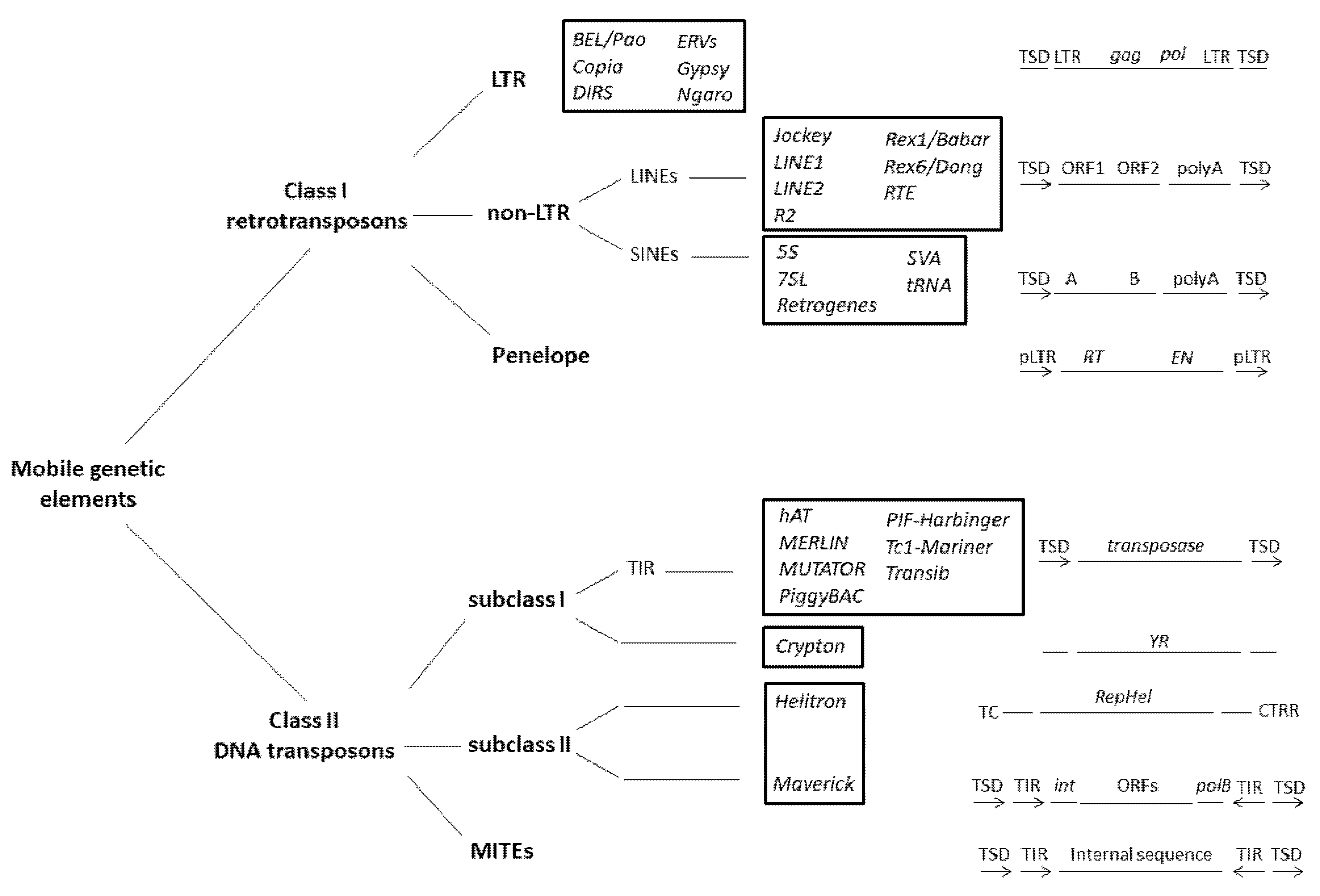
2. Mobile Elements
3. Transposable Elements in Actinopterygians
4. Rex Retroelements
5. Endogenous Retroviruses
6. TEs and Sex Chromosomes
7. Fish Transposons and the Environment
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Froese, R.; Pauly, D. FishBase. World Wide Web Electronic Publication. Version (12/2019). Available online: www.fishbase.org (accessed on 5 June 2020).
- Noack, K.; Zardoya, R.; Meyer, A. The complete mitochondrial DNA sequence of the bichir (Polypterus ornatipinnis), a basal ray-finned fish: Ancient establishment of the consensus vertebrate gene order. Genetics 1996, 144, 1165–1180. [Google Scholar]
- Venkatesh, B.; Erdmann, M.V.; Brenner, S. Molecular synapomorphies resolve evolutionary relationships of extant jawed vertebrates. Proc. Natl. Acad. Sci. USA 2001, 98, 11382–11387. [Google Scholar]
- Inoue, J.G.; Miya, M.; Tsukamoto, K.; Nishida, M. Basal actinopterygian relationships: A mitogenomic perspective on the phylogeny of the “ancient fish”. Mol. Phylogenet. Evol. 2003, 26, 110–120. [Google Scholar]
- Kikugawa, K.; Katoh, K.; Kuraku, S.; Sakurai, H.; Ishida, O.; Iwabe, N.; Miyata, T. Basal jawed vertebrate phylogeny inferred from multiple nuclear DNA-coded genes. BMC Evol. Biol. 2004, 2, 3. [Google Scholar]
- Gregory, T.R. Animal Genome Size Database. Version (12/2019). Available online: http://www.genomesize.com (accessed on 5 June 2020).
- Betancur-R, R.; Wiley, E.O.; Arratia, G.; Acero, A.; Bailly, N.; Miya, M.; Lecointre, G.; Ortí, G. Phylogenetic classification of bony fishes. BMC Evol. Biol. 2017, 17, 162. [Google Scholar]
- Bian, C.; Huang, Y.; Li, J.; You, X.; Yi, Y.; Ge, W.; Shi, Q. Divergence, evolution and adaptation in ray-finned fish genomes. Sci. China Life Sci. 2019, 62, 1003–1018. [Google Scholar]
- Dehal, P.; Boore, J.L. Two rounds of whole genome duplication in the ancestral vertebrate. PLoS Biol. 2005, 3, e314. [Google Scholar]
- Postlethwait, J.H.; Woods, I.G.; Ngo-Hazelett, P.; Yan, Y.L.; Kelly, P.D.; Chu, F.; Huang, H.; Hill-Force, A.; Talbot, W.S. Zebrafish comparative genomics and the origins of vertebrate chromosomes. Genome Res. 2000, 10, 1890–1902. [Google Scholar]
- Vandepoele, K.; de Vos, W.; Taylor, J.S.; Meyer, A.; Van de Peer, Y. Major events in the genome evolution of vertebrates: Paranome age and size differ considerably between ray-finned fishes and land vertebrates. Proc. Natl. Acad. Sci. USA 2004, 101, 1638–1643. [Google Scholar]
- Christoffels, A.; Koh, E.G.L.; Chia, J.M.; Brenner, S.; Aparicio, S.; Venkatesh, B. Fugu genome analysis provides evidence for a whole-genome duplication early during the evolution of ray-finned fishes. Mol. Biol. Evol. 2004, 21, 1146–1151. [Google Scholar]
- Meyer, A.; Van de Peer, Y. From 2R to 3R: Evidence for a fish-specific genome duplication (FSGD). Bioessays 2005, 27, 937–945. [Google Scholar]
- Marburger, S.; Alexandrou, M.A.; Taggart, J.B.; Creer, S.; Carvalho, G.; Oliveira, C.; Taylor, M.I. Whole genome duplication and transposable element proliferation drive genome expansion in Corydoradinae catfishes. Proc. R. Soc. B Biol. Sci. 2018, 285, 20172732. [Google Scholar]
- Alexandrou, M.A.; Swartz, B.A.; Matzke, N.J.; Oakley, T.H. Genome duplication and multiple evolutionary origins of complex migratory behavior in Salmonidae. Mol. Phylogenet. Evol. 2013, 69, 514–523. [Google Scholar]
- Macqueen, D.J.; Johnston, I.A. A well-constrained estimate for the timing of the salmonid whole genome duplication reveals major decoupling from species diversification. Proc. R. Soc. B 2014, 281, 2013–2881. [Google Scholar]
- Wang, J.T.; Li, J.T.; Zhang, X.F.; Sun, X.W. Transcriptome analysis reveals the time of the fourth round of genome duplication in common carp (Cyprinus carpio). BMC Genom. 2012, 13, 96. [Google Scholar]
- Costantini, M.; Cammarano, R.; Bernardi, G. The evolution of isochore patterns in vertebrate genomes. BMC Genom. 2009, 10, 146. [Google Scholar]
- Symonová, R.; Suh, A. Nucleotide composition of transposable elements likely contributes to AT/GC compositional homogeneity of teleost fish genomes. Mob. DNA 2019, 10, 49. [Google Scholar]
- Symonová, R.; Majtánová, Z.; Arias-Rodriguez, L.; Morkovský, L.; Korìnková, T.; Cavin, L.; Johnson Pokorná, M.; Dolezálková, M.; Flajshans, M.; Normandeau, E.; et al. Genome compositional organization in gars shows more similarities to mammals than to other ray-finned fish. J. Exp. Zool. Mol. Dev. Evol. 2016, 328, 607–619. [Google Scholar]
- Canapa, A.; Barucca, M.; Biscotti, M.A.; Forconi, M.; Olmo, E. Transposons, genome size, and evolutionary insights in animals. Cytogenet. Genome Res. 2015, 147, 217–239. [Google Scholar]
- Carducci, F.; Biscotti, M.A.; Barucca, M.; Canapa, A. Transposable elements in vertebrates: Species evolution and environmental adaptation. Eur. Zool. J. 2019, 86, 497–503. [Google Scholar]
- Warren, I.A.; Naville, M.; Chalopin, D.; Levin, P.; Berger, C.S.; Galiana, D.; Volff, J.N. Evolutionary impact of transposable elements on genomic diversity and lineage-specific innovation in vertebrates. Chrom. Res. 2015, 23, 505–531. [Google Scholar]
- Pujolar, J.M.; Astolfi, L.; Boscari, E.; Vidotto, E.; Barbisan, F.; Bruson, A.; Congiu, L. Tana1, a new putatively active Tc1-like transposable element in the genome of sturgeons. Mol. Phylogenet. Evol. 2013, 66, 223–232. [Google Scholar]
- Cheng, P.; Huang, Y.; Du, H.; Li, C.; Lv, Y.; Ruan, R.; Ye, H.; Bian, C.; You, X.; Xu, J.; et al. Draft genome and complete Hox-cluster characterization of the sterlet (Acipenser ruthenus). Front. Genet. 2019, 10, 776. [Google Scholar]
- Braasch, I.; Gehrke, A.R.; Smith, J.J.; Kawasaki, K.; Manousaki, T.; Pasquier, J.; Amores, A.; Desvignes, T.; Batze, P.; Catchen, J.; et al. The spotted gar genome illuminates vertebrate evolution and facilitates human-teleost comparisons. Nat. Genet. 2016, 48, 4. [Google Scholar]
- Hao, S.; Han, K.; Meng, L.; Huang, X.; Shi, C.; Zhang, M.; Wang, Y.; Liu, Q.; Zhang, Y.; Seim, I.; et al. Three genomes of Osteoglossidae shed light on ancient teleost evolution. BioRxiv 2020. [Google Scholar] [CrossRef]
- Jiang, J.; Yuan, H.; Zheng, X.; Wang, Q.; Kuang, T.; Li, J.; Liu, J.; Song, S.; Wang, W.; Cheng, F.; et al. Gene markers for exon capture and phylogenomics in ray-finned fishes. Ecol. Evol. 2019, 9, 3973–3983. [Google Scholar]
- Rodriguez, F.; Arkhipova, I.R. Transposable elements and polyploid evolution in animals. Curr. Opin. Genet. Dev. 2018, 49, 115–123. [Google Scholar]
- Symonová, R.; Howell, W.M. Vertebrate genome evolution in the light of fish cytogenomics and rDNAomics. Genes 2018, 9, 96. [Google Scholar]
- Yuan, Z.; Liu, S.; Zhou, T.; Tian, C.; Bao, L.; Dunham, R.; Liu, Z. Comparative genome analysis of 52 fish species suggests differential associations of repetitive elements with their living aquatic environments. BMC Genom. 2018, 19, 141. [Google Scholar]
- Chalopin, D.; Volff, J.N. Analysis of the spotted gar genome suggests absence of causative link between ancestral genome duplication and transposable element diversification in teleost fish. J. Exp. Zool. B Mol. Dev. Evol. 2017, 328, 629–637. [Google Scholar]
- Pasquier, J.; Braasch, I.; Batzel, P.; Cabau, C.; Montfort, J.; Nguyen, T.; Jouanno, E.; Berthelot, C.; Klopp, C.; Journot, L.; et al. Evolution of gene expression afterwhole-genome duplication: New insights from the spotted gar genome. J. Exp. Zool. Mol. Dev. Evol. 2017, 328, 709–721. [Google Scholar]
- Gornung, E. Twenty years of physical mapping of major ribosomal RNA genes across the teleosts: A review of research. Cytogenet. Genome Res. 2013, 141, 90–102. [Google Scholar]
- Spaink, H.P.; Jansen, H.J.; Dirks, R.P. Advances in genomics of bony fish. Brief. Funct. Genomics 2013, 13, 144–156. [Google Scholar]
- Kajikawa, M.; Okada, N. LINEs mobilize SINEs in the eel through a shared 3′ sequence. Cell 2002, 111, 433–444. [Google Scholar]
- Chalopin, D.; Naville, M.; Plard, F.; Galiana, D.; Volff, J.N. Comparative analysis of transposable elements highlights mobilome diversity and evolution in vertebrates. Genome Biol. Evol. 2015, 7, 567–580. [Google Scholar]
- Coluccia, E.; Pichiri, G.; Nieddu, M.; Coni, P.; Manconi, S.; Deiana, A.M.; Salvadori, S.; Mezzanotte, R. Identification of two new repetitive elements and chromosomal mapping of repetitive DNA sequences in the fish Gymnothorax unicolor (Anguilliformes: Muraenidae). Eur. J. Histochem. 2011, 55, e12. [Google Scholar]
- Durif, C.M.F.; Dufour, S.; Elie, P. Impact of silvering stage, age, body size and condition on reproductive potential of the European eel. Mar. Ecol. Prog. Ser. 2006, 327, 171–181. [Google Scholar]
- Frankowski, J.; Bastrop, R. Identification of Anguilla anguilla (L.) and Anguilla rostrata (LeSueur) and their hybrids based on a diagnostic single nucleotide polymorphism in nuclear 18SrDNA. Mol. Ecol. Res. 2010, 10, 173–176. [Google Scholar]
- Geeraerts, C.; Belpaire, C. The effects of contaminants in European eel: A review. Ecotoxicology 2010, 19, 239–266. [Google Scholar]
- Gong, X.L.; Li, S.F.; Cai, W.C.; Wang, C.H. Isolation and characterization of polymorphic microsatellite markers from Australia short-finned eel (Anguilla australis Richardson). Mol. Ecol. Res. 2008, 8, 777–779. [Google Scholar]
- Ishikawa, S.; Tsukamoto, K.; Nishida, M. Characterization of microsatellite loci from the Japanese eel Anguilla japonica. Mol. Ecol. Notes 2001, 1, 140–142. [Google Scholar]
- Itoi, S.; Nakaya, M.; Kaneko, G.; Kondo, H.; Sezaki, K.; Watabe, S. Rapid identification of eels Anguilla japonica and Anguilla anguilla by polymerase chain reaction with single nucleotide polymorphism-based specific probes. Fish Sci. 2005, 71, 1356–1364. [Google Scholar]
- Jansen, H.J.; Liem, M.; Jong-Raadsen, S.A.; Dufour, S.; Weltzien, F.A.; Swinkels, W.; Koelewijn, A.; Palstra, A.P.; Pelster, B.; Spaink, H.P.; et al. Rapid de novo assembly of the European eel genome from nanopore sequencing reads. Sci. Rep. 2017, 7, 7213. [Google Scholar]
- Kitaoka, M.; Okamura, N.; Ichinose, H.; Goto, M. Detection of SNPs in fish DNA: Application of the Fluorogenic Ribonuclease Protection (FRIP) assay for the authentication of food contents. J. Agric. Food Chem. 2008, 56, 6246–6251. [Google Scholar]
- Li, Z.; Chen, F.; Huang, C.; Zheng, W.; Yu, C.; Cheng, H.; Zhou, R. Genome-wide mapping and characterization of microsatellites in the swamp eel genome. Sci. Rep. 2017, 7, 3157. [Google Scholar]
- Maes, G.E.; Pujolar, J.M.; Raeymaekers, J.A.M.; Dannewitz, J.; Volckaert, F.A.M. Microsatellite conservation and bayesian individual assignment in four Anguilla species. Mar. Ecol. Prog. Ser. 2006, 319, 251–261. [Google Scholar]
- Pierron, F.; Daffe, G.; Lambert, P.; Couture, P.; Baudrimont, M. Retrotransposon methylation and activity in wild fish (A. anguilla): A matter of size. Environ. Poll. 2019, 245, 494–503. [Google Scholar]
- Pujolar, J.M.; Maes, G.E.; Volckaert, F.A.M. Genetic patchiness among recruits in the European eel Anguilla anguilla. Mar. Ecol. Prog. Ser. 2006, 307, 209–217. [Google Scholar]
- Suntronpong, A.; Singchat, W.; Kruasuwan, W.; Prakhongcheep, O.; Sillapaprayoon, S.; Muangmai, N.; Somyong, S.; Indananda, C.; Kraichak, E.; Peyachoknagul, S.; et al. Characterization of centromeric satellite DNAs (MALREP) in the Asian swamp eel (Monopterus albus) suggests the possible origin of repeats from transposable elements. Genomics 2020, 112, 3097–3107. [Google Scholar]
- Pujolar, J.M.; Maes, G.E.; van Houdt, J.K.J.; Zane, L. Isolation and characterization of expressed sequence tag-linked microsatellite loci for the European eel (Anguilla anguilla). Mol. Ecol. Res. 2009, 9, 233–235. [Google Scholar]
- Pujolar, J.M.; Jacobsen, M.W.; Als, T.D.; Frydenberg, J.; Magnussen, E.; Jonsson, B.; Jiang, X.; Cheng, L.; Bekkevold, D.; Maes, G.E.; et al. Assessing patterns of hybridization between North Atlantic eels using diagnostic single-nucleotide polymorphisms. Heredity 2014, 112, 627–637. [Google Scholar]
- Rozenfeld, C.; Blanca, J.; Gallego, V.; Garcìa-Carpintero, V.; Herranz-Jusdado, J.G.; Pérez, L.; Asturiano, J.F.; Cañizares, J.; Peñaranda, D.S. De novo European eel transcriptome provides insights into the evolutionary history of duplicated genes in teleost lineages. PLoS ONE 2019, 14, e0218085. [Google Scholar]
- Suntronpong, A.; Thapana, W.; Twilprawat, P.; Prakhongcheep, O.; Somyong, S.; Muangmai, N.; Peyachoknagul, S.; Srikulnath, K. Karyological characterization and identification of four repetitive element groups (the 18S–28S rRNA gene, telomeric sequences, microsatellite repeat motifs, Rex retroelements) of the Asian swamp eel (Monopterus albus). Comp. Cytogenet. 2017, 11, 435–462. [Google Scholar]
- Tseng, M.C.; Chen, C.A.; Kao, H.W.; Tzeng, W.N.; Lee, S.C. Polymorphisms of GA/GT microsatellite loci from Anguilla japonica. Mar. Biotechnol. 2001, 3, 275–280. [Google Scholar]
- Tu, Z.; Li, S.; Mao, C. The changing tails of a novel short interspersed element in Aedes aegypti: Genomic evidence for slippage retrotransposition and the relationship between 3′ tandem repeats and the poly(dA) tail. Genetics 2004, 168, 2037–2047. [Google Scholar]
- Utsunomia, R.; Pansonato-Alves, J.C.; Cardim Scacchetti, P.; Oliveira, C.; Foresti, F. Scattered organization of the histone multigene family and transposable elements in Synbranchus. Genet. Mol. Biol. 2014, 37, 30–36. [Google Scholar]
- Volff, J.N.; Körting, C.; Sweeney, K.; Schartl, M. The non-LTR retrotransposon Rex3 from the Fish Xiphophorus is widespread among teleosts. Mol. Biol. Evol. 1999, 16, 1427–1438. [Google Scholar]
- Volff, J.N.; Körting, C.; Schartl, M. Multiple lineages of the non-LTR retrotransposon Rex1 with varying success in invading fish genomes. Mol. Biol. Evol. 2000, 17, 1673–1684. [Google Scholar]
- Volff, J.N.; Körting, C.; Froschauer, A.; Sweeney, K.; Schartl, M. Non-LTR retrotransposons encoding a restriction enzyme-like endonuclease in vertebrates. J. Mol. Evol. 2001, 52, 351–360. [Google Scholar]
- Yi, M.; Chen, F.; Luo, M.; Cheng, Y.; Zhao, H.; Cheng, H.; Zhou, R. Rapid Evolution of piRNA Pathway in the Teleost Fish: Implication for an Adaptation to Transposon Diversity. Genome Biol. Evol. 2014, 6, 1393–1407. [Google Scholar]
- Ráb, P.; Yano, C.F.; Lavoué, S.; Jegede, O.I.; Bertollo, L.A.C.; Ezaz, T.; Majtánová, Z.; de Oliveira, E.A.; Cioffi, M.B. Karyotype and mapping of repetitive DNAs in the African butterfly fish Pantodon buchholzi, the sole species of the family Pantodontidae. Cytogenet. Genome Res. 2016, 149, 312–320. [Google Scholar]
- de Souza, F.H.S.; Perez, M.F.; Bertollo, L.A.C.; de Oliveira, E.A.; Lavoué, S.; Gestich, C.C.; Ráb, P.; Ezaz, T.; Liehr, T.; Viana, P.F.; et al. Interspecific Genetic Differences and Historical Demography in South American Arowanas (Osteoglossiformes, Osteoglossidae, Osteoglossum). Genes 2019, 10, 693. [Google Scholar]
- Mohindra, V.; Dangi, T.; Tripathi, R.K.; Kumar, R.; Singh, R.K.; Jena, J.K.; Mohapatra, T. Draft genome assembly of Tenualosa ilisha, Hilsa shad, provides resource for osmoregulation studies. Sci. Rep. 2019, 9, 16511. [Google Scholar]
- Machado, A.M.; Tørresen, O.K.; Kabeya, N.; Couto, A.; Petersen, B.; Felício, M.; Campos, P.F.; Fonseca, E.; Bandarra, N.; Lopes-Marques, M.; et al. “Out of the can”: A draft genome assembly, liver transcriptome, and nutrigenomics of the european sardine, Sardina pilchardus. Genes 2018, 9, 485. [Google Scholar]
- Swathi, A.; Shekhar, M.S.; Katneni, V.K.; Vijayan, K.K. Genome size estimation of brackishwater fishes and penaeid shrimps by flow cytometry. Mol. Biol. Rep. 2018, 45, 951–960. [Google Scholar]
- Gao, B.; Shen, D.; Xue, S.; Chen, C.; Cui, H.; Song, C. The contribution of transposable elements to size variations between four teleost genomes. Mob. DNA 2016, 7, 4. [Google Scholar]
- Guo, X.M.; Zhang, Q.Q.; Sun, Y.W.; Jiang, X.Y.; Zou, S.M. Tc1-like Transposase Thm3 of silver carp (Hypophthalmichthys molitrix) can mediate gene transposition in the genome of blunt snout bream (Megalobrama amblycephala). G3 2015, 5, 2601–2610. [Google Scholar]
- Micale, L.; Loviglio, M.N.; Manzoni, M.; Fusco, C.; Augello, B.; Migliavacca, E.; Cotugno, G.; Monti, E.; Borsani, G.; Reymond, A.; et al. A fish-specific transposable element shapes the repertoire of p53 target genes in zebrafish. PLoS ONE 2012, 7, e46642. [Google Scholar]
- Pereira, C.S.A.; Pazian, M.F.; Ráb, P.; Collares-Pereira, M.J. Dynamics of Rex3 in the genomes of endangered Iberian Leuciscinae (Teleostei, Cyprinidae) and their natural hybrids. Mol. Cytogenet. 2015, 8, 81. [Google Scholar]
- Ahn, S.J.; Kim, M.S.; Jang, J.H.; Lim, S.U.; Lee, H.H. MMTS, a new subfamily of Tc1-like transposons. Mol. Cells. 2008, 26, 387–395. [Google Scholar]
- Ivics, Z.; Izsvak, Z.; Minter, A.; Hackett, P.B. Identification of functional domains and evolution of Tc1-like transposable elements. Proc. Natl. Acad. Sci. USA 1996, 93, 5008–5013. [Google Scholar]
- Liu, D.; You, C.; Liu, S.; Liu, L.; Duan, W.; Chen, S.; Yan, J.; Liu, Y. Characterization of a novel Tc1-like transposon from bream (Cyprinidae, Megalobrama) and its genetic variation in the polyploidy progeny of bream-red crucian carp crosses. J. Mol. Evol. 2009, 69, 395–403. [Google Scholar]
- Ishiyama, S.; Yamazaki, K.; Kurihara, F.; Yamashita, D.; Sao, K.; Hattori, A.; Koga, A. DNA-based transposable elements with nucleotide sequence similar to Tol2 from medaka fish are prevalent in cyprinid fishes. Gene Rep. 2017, 9, 37–45. [Google Scholar]
- Xu, P.; Xu, J.; Liu, G.; Chen, L.; Zhou, Z.; Peng, W.; Jiang, Y.; Zhao, Z.; Jia, Z.; Sun, Y.; et al. The allotetraploid origin and asymmetrical genome evolution of the common carp Cyprinus carpio. Nat. Commun. 2019, 10, 4625. [Google Scholar]
- Dulz, T.A.; Lorscheider, C.A.; Nascimento, V.D.; Noleto, R.B.; Moreira-Filho, O.; Nogaroto, V.; Vicari, M.R. Comparative cytogenetics among Leporinus friderici and Leporellus vittatus populations (Characiformes, Anostomidae): Focus on repetitive DNA elements. Comp. Cytogenet. 2019, 13, 105–120. [Google Scholar]
- Barbosa, P.; de Oliveira, L.A.; Pucci, M.B.; Santos, M.E.; Moreira-Filho, O.; Vicari, M.R.; Nogaroto, V.; de Almeida, M.C.; Ferreira Artoni, R. Identification and chromosome mapping of repetitive elements in the Astyanax scabripinnis (Teleostei: Characidae) species complex. Genetica 2015, 143, 55–62. [Google Scholar]
- Daniel, S.N.; Penitente, M.; Silva, D.M.Z.A.; Hashimoto, D.T.; Ferreira, D.C.; Foresti, F.; Porto-Foresti, F. Organization and chromosomal distribution of histone genes and transposable Rex elements in the genome of Astyanax bockmanni (Teleostei, Characiformes). Cytogenet. Genome Res. 2015, 146, 311–318. [Google Scholar]
- Gao, B.; Chen, W.; Shen, D.; Wang, S.; Chen, C.; Zhang, L.; Wang, W.; Wang, X.; Song, C. Characterization of autonomous families of Tc1/Mariner transposon in neoteleost genomes. Mar. Genom. 2017, 34, 67–77. [Google Scholar]
- Martins, C.; Ferreira, I.A.; Oliveira, C.; Foresti, F.; Galetti, P.M., Jr. A tandemly repetitive centromeric DNA sequence of the fish Hoplias malabaricus (Characiformes: Erythrinidae) is derived from 5S rDNA. Genetica 2006, 127, 133–141. [Google Scholar]
- Silva, D.M.Z.A.; Pansonato-Alves, J.C.; Utsunomia, R.; Daniel, S.N.; Hashimoto, D.T.; Oliveira, C.; Porto-Foresti, F.; Foresti, F. Chromosomal organization of repetitive DNA sequences in Astyanax bockmanni (Teleostei, Characiformes): Dispersive location, association and co-localization in the genome. Genetica 2013, 141, 329–336. [Google Scholar]
- Pessenda, G.; Piscor, D.; da Silva, E.L.; Centofante, L.; Parise-Maltempi, P.P. Chromosomal organization of retrotransposon Rex1 in Astyanax species (Characiformes, Characidae). Caryologia 2018, 71, 80–84. [Google Scholar]
- Ferreira, V.A.M.; Marajo, L.; Matoso, D.A.; Ribeiro, L.B.; Feldberg, E. Chromosomal mapping of Rex retrotransposons in Tambaqui (Colossoma macropomum Cuvier, 1818) exposed to three climate change scenarios. Cytogenet. Genome Res. 2019, 159, 39–47. [Google Scholar]
- Pucci, M.B.; Nogaroto, V.; Moreira-Filho, O.; Vicari, M.R. Dispersion of transposable elements and multigene families: Microstructural variation in Characidium (Characiformes: Crenuchidae) genomes. Genet. Mol. Biol. 2018, 41, 585–592. [Google Scholar]
- Schemberger, M.O.; Nascimento, V.D.; Coan, R.; Ramos, E.; Nogaroto, V.; Ziemniczak, K.; Valente, G.T.; Moreira-Filho, O.; Martins, C.; Vicari, M.R. DNA transposon invasion and microsatellite accumulation guide W chromosome differentiation in a Neotropical fish genome. Chromosoma 2019, 128, 547–560. [Google Scholar]
- Schemberger, M.O.; Oliveira, J.I.; Nogaroto, V.; Almeida, M.C.; Artoni, R.F.; Cestari, M.M.; Moreira-Filho, O.; Vicari, M.R. Construction and characterization of a repetitive DNA library in Parodontidae (Atinopterygii:Characiformes): A genomic and evolutionary approach to the degeneration of the W sex chromosome. Zebrafish 2014, 11, 518–527. [Google Scholar]
- Schemberger, M.O.; Nogaroto, V.; Almeida, M.C.; Ferreira Artoni, R.; Valente, G.T.; Martins, C.; Moreira-Filho, O.; Cestari, M.M.; Vicari, M.R. Sequence analyses and chromosomal distribution of the Tc1/Mariner element in Paraodontidae fish (Teleostei: Characiforme). Gene 2016, 593, 308–314. [Google Scholar]
- Terencio, M.L.; Schneider, C.H.; Gross, M.C.; Vicari, M.R.; Farias, I.P.; Passos, K.B.; Feldberg, E. Evolutionary dynamics of repetitive DNA in Semaprochilodus (Characiformes, Prochilodontidae): A fish model for sex chromosome differentiation. Sex Dev. 2013, 7, 325–333. [Google Scholar]
- Yano, C.; Bertollo, L.A.C.; Ezaz, T.; Trifonov, V.; Sember, A.; Liehr, T.; Cioffi, M.B. Highly conserved Z and molecularly diverged W chromosomes in the fish genus Triportheus (Characiformes, Triportheidae). Heredity 2017, 118, 276–283. [Google Scholar]
- Yano, C.F.; Merlo, M.A.; Portela-Bens, S.; Cioffi, M.B.; Bertollo, L.A.C.; Santos-Júnior, C.D.; Rebordinos, L. Evolutionary dynamics of multigene families in Triportheus (Characiformes, Triportheidae): A transposon mediated mechanism? Front. Mar. Sci. 2020, 7, 6. [Google Scholar]
- de Sene, V.F.; Pansonato-Alves, J.C.; Ferreira, D.C.; Utsumonia, R.; Oliveira, C.; Foresti, F. Mapping of the retrotransposable elements Rex1 and Rex3 in chromosomes of eigenmannia (Teleostei, Gymnotigformes, Sternopygidae). Cytogenet. Genome Res. 2015, 146, 319–324. [Google Scholar]
- da Silva, M.; Matoso, D.A.; Vicari, M.R.; de Almeida, M.C.; Margarido, V.P.; Artoni, R.F. Physical Mapping of 5S rDNA in two species of knifefishes: Gymnotus pantanal and Gymnotus paraguensis (Gymnotiformes). Cytogenet. Genome Res. 2011, 134, 303–307. [Google Scholar]
- da Silva, M.; Barbosa, P.; Artoni, R.F.; Feldberg, E. Evolutionary dynamics of 5S rDNA and recurrent association of transposable elements in electric fish of the family Gymnotidae (Gymnotiformes): The case of Gymnotus mamiraua. Cytogenet. Genome Res. 2016, 149, 297–303. [Google Scholar]
- Ferreira, D.C.; Oliveira, C.; Foresti, F. Chromosome mapping of retrotransposable elements Rex1 and Rex3 in three fish species in the subfamily Hypoptopomatinae (Teleostei, Siluriformes, Loricariidae). Cytogenet. Genome Res. 2011, 132, 64–70. [Google Scholar]
- Gonzalez-Gouveia, J.; Wolf, I.R.; Vilas-Boas, L.A.; Heslop-Harrison, J.S.; Schwarzacher, T.; Dias, A.L. Repetitive DNA in the catfish genome: rDNA, microsatellites, and Tc1-Mariner transposon sequences in Imparfinis species (Siluriformes, Heptapteridae). J. Hered. 2017, 108, 650–657. [Google Scholar]
- Liu, Z.; Li, P.; Kucuktas, H.; Dunham, R. Characterization of nonautonomous Tc1-like transposable elements of channel catfish (Ictalurus punctatus). Fish Physiol. Biochem. 1999, 21, 65–72. [Google Scholar]
- Primo, C.C.; Glugoski, L.; Vicari, M.R.; Nogaroto, V. Chromosome mapping and molecular characterization of the Tc1/Mariner element in Rineloricaria (Siluriformes: Loricariidae). Braz. Arch. Biol. Technol. 2018, 61, e18170623. [Google Scholar]
- Traldi, J.B.; Lui, R.L.; de Martinez, J.F.; Vicari, M.R.; Nogaroto, V.; Filho, O.M.; Blanco, D. R Chromosomal distribution of the retroelements Rex1, Rex3 and Rex6 in species of the genus Harttia and Hypostomus (Siluriformes: Loricariidae). Neotrop. Ichthyol. 2019, 17, e190010. [Google Scholar]
- da Silva, K.R.; Mariotto, S.; Centofante, L.; Parise-Maltempi, P.P. Chromosome mapping of a Tc1-like transposon in species of the catfish Ancistrus. Comp. Cytogenet. 2017, 11, 65–79. [Google Scholar]
- Hughes, L.C.; Ortí, G.; Huang, Y.; Sun, Y.; Baldwin, C.C.; Thompson, A.W.; Arcila, D.; Betancur-R, R.; Li, C.; Becker, L.; et al. Comprehensive phylogeny of ray-finned fishes (Actinopterygii) based on transcriptomic and genomic data. Proc. Natl. Acad. Sci. USA 2018, 115, 6249–6254. [Google Scholar]
- Ybazeta, G. Molecular systematics and biogeography of the Galaxiidae. Ph.D. Thesis, Univeristy of Toronto, Toronto, ON, Canada, 2012. [Google Scholar]
- Pan, Q.; Feron, R.; Yano, A.; Guyomard, R.; Jouanno, E.; Vigouroux, E.; Wen, M.; Busnel, J.M.; Bobe, J.; Concordet, J.P.; et al. Identification of the master sex determining gene in Northern pike (Esox lucius) reveals restricted sex chromosome differentiation. PLoS Genet. 2019, 15, e1008013. [Google Scholar]
- de Boer, J.G.; Yazawa, R.; Davidson, W.S.; Koop, B.F. Bursts and horizontal evolution of DNA transposon in the speciation of pseudotetraploid salmonids. BMC Genom. 2007, 8, 422. [Google Scholar]
- Faber-Hammond, J.J.; Phillips, R.B.; Brown, K.H. Comparative analysis of the shared sex-determination region (SDR) among salmonid fishes. Genome Biol. Evol. 2015, 7, 1972–1987. [Google Scholar]
- Lien, S.; Koop, B.K.; Sandve, S.R.; Miller, J.R.; Kent, M.P.; Nome, T.; Hvidsten, T.R.; Leong, J.S.; Minkley, D.R.; Zimin, A.; et al. The Atlantic salmon genome provides insights into rediploidization. Nature 2016, 533, 200–205. [Google Scholar]
- Sävilammi, T.; Primmer, C.R.; Varadharajan, S.; Guyomard, R.; Guiguen, Y.; Sandve, S.R.; Vøllestad, L.A.; Papakostas, S.; Lien, S. The chromosome-level genome assembly of european grayling reveals aspects of a unique genome evolution process within salmonids. G3 2019, 9, 1283–1294. [Google Scholar]
- Liu, K.; Xu, D.; Li, J.; Bian, C.; Duan, J.; Zhou, Y.; Zhang, M.; You, X.; You, Y.; Chen, J.; et al. Whole genome sequencing of Chinese clearhead icefish, Protosalanx hyalocranius. GigaScience 2017, 6, giw012. [Google Scholar]
- Malmstrøm, M.; Matschiner, M.; Tørresen, O.K.; Jakobsen, K.S.; Jentoft, S. Whole genome sequencing data and de novo draft assemblies for 66 teleost species. Sci. Data 2017, 4, 160132. [Google Scholar]
- Ota, K.; Tateno, Y.; Gobojori, T. Highly differentiated and conserved sex chromosome in fish species (Aulopus japonicus: Teleostei, Aulopidae). Gene 2003, 317, 187–193. [Google Scholar]
- Brynildsen, W. Transposable Elements in Teleost Fish: In Silico Exploration of TE Activity, Diversity and Abundance across 74 Teleost Fish Genomes. Master’s Thesis, University Oslo, Oslo, Norway, 2016. Available online: http://urn.nb.no/URN:NBN:no-55565 (accessed on 4 May 2020).
- Tørresen, O.K.; Star, B.; Jentoft, S.; Reinar, W.B.; Grove, H.; Miller, J.R.; Walenz, B.P.; Knight, J.; Ekholm, J.M.; Peluso, P.; et al. An improved genome assembly uncovers prolific tandem repeats in Atlantic cod. BMC Genom. 2017, 18, 95. [Google Scholar]
- Lima, L.C.B.; de Martinez, P.A.; Molina, W.F. Cytogenetic characterization of three Balistoidea fish species from the Atlantic with inferences on chromosomal evolution in the families Monacanthidae and Balistidae. Comp. Cytogenet. 2011, 5, 61–69. [Google Scholar]
- Nirchio, M.; Rossi, A.R.; Foresti, F.; Oliveira, C. Chromosome evolution in fishes: A new challenging proposal from Neotropical species. Neotrop. Ichthyol. 2014, 12, 761–770. [Google Scholar]
- Pérez Bielsa, N. Is the Small Genome Size of Pleuronectiformes Related to Small Intron Size? Master’s Thesis, Universitat de Girona, Girona, Spain, 2013. [Google Scholar]
- Tarallo, A. Genome Composition Plasticity in Marine Organisms. Ph.D. Thesis, University of Naples “Federico II”, Naples, Italy, 2016. [Google Scholar]
- Dettleff, P.; Hormazabal, E.; Aedo, J.; Fuentes, M.; Meneses, C.; Molina, A.; Valdes, J.A. Identification and evaluation of long noncoding RNAs in response to handling stress in red cusk-eel (Genypterus chilensis) via RNA-seq. Mar. Biotechnol. 2020, 22, 94–108. [Google Scholar]
- Small, C.M.; Bassham, S.; Catchen, J.; Amores, A.; Fuiten, A.M.; Brown, R.S.; Jones, A.G.; Cresko, W.A. The genome of the Gulf pipefish enables understanding of evolutionary innovations. Genome Biol. 2016, 17, 258. [Google Scholar]
- Cioffi, M.; Bertollo, L.A.; Villa, M.A.; de Oliveira, E.A.; Tanomtong, A.; Yano, C.F.; Supiwong, W.; Chaveerach, A. Genomic organization of repetitive DNA elements and its implications for the chromosomal evolution of channid fishes (Actinopterygii, Perciformes). PLoS ONE 2015, 10, e0130199. [Google Scholar]
- García, E.; Cross, I.; Portela-Bens, S.; Rodríguez, M.E.; García-Angulo, A.; Molina, B.; Cuadrado, A.; Liehr, T.; Rebordinos, L. Integrative genetic map of repetitive DNA in the sole Solea senegalensis genome shows a Rex transposon located in a proto-sex chromosome. Sci. Rep. 2019, 9, 17146. [Google Scholar]
- Roberts, B.H.; Morrongiello, J.R.; King, A.J.; Morgan, D.L.; Saunders, T.M.; Woodhead, J.; Crook, D.A. Migration to freshwater increases growth rates in a facultatively catadromous tropical fish. Oecologia 2019, 191, 253–260. [Google Scholar]
- Rodríguez, M.E.; Molina, B.; Merlo, M.A.; Arias-Pérez, A.; Portela-Bens, S.; García-Angulo, A.; Cross, I.; Liehr, T.; Rebordinos, L. Evolution of the proto sex-chromosome in Solea senegalensis. Int. J. Mol. Sci. 2019, 20, 5111. [Google Scholar]
- Bohne, A.; Zhou, Q.; Darras, A.; Schmidt, C.; Schartl, M.; Galiana-Arnoux, D.; Volff, J.N. Zisupton—A novel superfamily of DNA transposable elements recently active in fish. Mol. Biol. Evol. 2011, 29, 631–645. [Google Scholar]
- Koga, A.; Shimada, A.; Shima, A.; Sakaizumi, M.; Tachida, H.; Hori, H. Evidence for recent invasion of the medaka fish genome by the Tol2 transposable element. Genetics 2000, 155, 273–281. [Google Scholar]
- Koga, A.; Hori, H. The Tol2 transposable element of the medaka fish: An active DNA-based element naturally occurring in a vertebrate genome. Genes Genet. Syst. 2001, 76, 1–8. [Google Scholar]
- Koga, A.; Sakaizumi, M.; Hori, H. Transposable elements in medaka fish. Zool. Sci. 2002, 19, 1–6. [Google Scholar]
- Dennenmoser, S.; Sedlazeck, F.J.; Iwaszkiewicz, E.; Li, X.-Y.; Altmuller, J.; Nolte, A.W. Copy number increases of transposable elements and protein-coding genes in an invasive fish of hybrid origin. Mol. Ecol. 2017, 26, 4712–4724. [Google Scholar]
- Xiao, Y.; Xiao, Z.; Ma, D.; Liu, J.; Li, J. Genome sequence of the barred knifejaw Oplegnathus fasciatus (Temminck & Schlegel, 1844): The first chromosome-level draft genome in the family Oplegnathidae. GigaScience 2019, 8, giz013. [Google Scholar]
- Costa, G.W.W.F.; Cioffi, M.B.; Bertollo, L.A.C.; Molina, W.F. Transposable elements in fish chromosomes: A study in the marine cobia species. Cytogenet. Genome Res. 2013, 141, 126–132. [Google Scholar]
- Almeida, L.A.H.; Nunes, L.A.; Bitencourt, J.A.; Molina, W.F.; Affonso, P.R.A.M. Chromosomal evolution and cytotaxonomy in wrasses (Perciformes; Labridae). J. Hered. 2017, 108, 239–253. [Google Scholar]
- Jacobina, U.P.; Cioffi, M.B.; Souza, L.G.R.; Calado, L.L.; Tavares, M.; Manzella, J., Jr.; Bertollo, L.A.C.; Molina, W.F. Chromosome mapping of repetitive sequences in Rachycentron canadum (Perciformes: Rachycentridae): Implications for karyotypic evolution and perspectives for biotechnological uses. J. Biomed. Biotechnol. 2011, 2011, 218231. [Google Scholar]
- Gross, M.C.; Schneider, C.H.; Valente, G.T.; Porto, J.I.R.; Martins, C.; Feldeberg, E. Comparative cytogenetic analysis of the genus Symphysodon (Discus Fishes, Cichlidae): Chromosomal characteristics of retrotransposons and minor ribosomal DNA. Cytogenet. Genome Res. 2010, 127, 43–53. [Google Scholar]
- de Freitas Mourão, A.A.; Daniel, S.N.; Hashimoto, D.T.; Ferreira, C.D.; Porto-Foresti, F. Organization and distribution of repetitive DNA classes in the Cichla kelberi and Cichla piquiti Genome. Cytologia 2017, 82, 193–197. [Google Scholar]
- Fischer, C.; Bouneau, L.; Coutanceau, J.P.; Weissenbach, J.; Volff, J.N.; Ozouf-Costaz, C. Global heterochromatic colocalization of transposable elements with minisatellites in the compact genome of the pufferfish Tetraodon nigroviridis. Gene 2004, 336, 175–183. [Google Scholar]
- Santos, F.A.; Marques, D.F.; Terencio, M.L.; Feldberg, E.; Rodrigues, L.R. Cytogenetic variation of repetitive DNA elements in Hoplias malabaricus (Characiformes-Erythrinidae) from white, black and clear water rivers of the Amazon basin. Genet. Mol. Biol. 2016, 39, 40–48. [Google Scholar]
- Schneider, C.H.; Gross, M.C.; Terencio, M.L.; do Carmo, E.J.; Martins, C.; Feldberg, E. Evolutionary dynamics of retrotransposable elements Rex1, Rex3 and Rex6 in neotropical cichlid genomes. BMC Evol. Biol. 2013, 13, 152. [Google Scholar]
- Capriglione, T.; De Paolo, S.; Cocca, E. Helinoto, a Helitron2 transposon from the icefish Chionodraco hamatus, contains a region with three deubiquitinase-like domains that exhibit transcriptional activity. Comp. Biochem. Physiol. Part D Genom. Proteom. 2014, 11, 49–58. [Google Scholar]
- Capriglione, T.; Odierna, G.; Caputo, V.; Canapa, A.; Olmo, E. Characterization of a Tc1- like transposon in the Antarctic ice-fish, Chionodraco hamatus. Gene 2002, 295, 193–198. [Google Scholar]
- Cocca, E.; De Iorio, S.; Capriglione, T. Identification of a novel helitron transposon in the genome of Antarctic fish. Mol. Phylogenet. Evol. 2011, 58, 439–446. [Google Scholar]
- Hendry, T.A.; Freed, L.L.; Fader, D.; Fenolio, D.; Sutton, T.T.; Lopex, J.V. Ongoing transposon-mediated genome reduction in the luminous bacterial symbionts of deep-sea ceratioid anglerfishes. Am. Soc. Microbiol. mBio 2018, 9. [Google Scholar] [CrossRef] [Green Version]
- Xu, D.; Molina, W.F.; Yano, C.F.; Zhang, Y.; de Oliveira, E.A.; Lou, B.; Cioffi, M.B. Comparative cytogenetics in three Sciaenid species (Teleostei, Perciformes): Evidence of interspecific chromosomal diversification. Mol. Cytogenet. 2017, 10, 37. [Google Scholar]
- Dasilva, C.; Hadji, H.; Ozouf-Costaz, C.; Nicaud, S.; Jaillon, O.; Weissenbach, J.; Crollius, H.R. Remarkable compartmentalization of transposable elements and pseudogenes in the heterochromatin of the Tetraodon nigroviridis genome. Proc. Natl. Acad. Sci. USA 2002, 99, 13636–13641. [Google Scholar]
- Guo, M.; Wang, S.; Su, Y.; Zhou, Y.; Liu, M.; Wang, J. Molecular cytogenetic analyses of Epinephelus bruneus and Epinephelus moara (Perciformes, Epinephelidae). PeerJ 2014, 2, e412. [Google Scholar]
- Zong, W.; Gao, B.; Diaby, M.; Shen, D.; Wang, S.; Wang, Y.; Sang, Y.; Chen, C.; Wang, X.; Song, C. Traveler, a new DD35E family of Tc1/Mariner transposons, invaded vertebrates very recently. Genome Biol. Evol. 2020, 12, 66–76. [Google Scholar]
- Costa, G.W.; Cioffi, M.B.; Bertollo, L.A.; Molina, W.F. The evolutionary dynamics of ribosomal genes, histone H3, and transposable Rex elements in the genome of Atlantic snappers. J. Hered. 2016, 107, 173–180. [Google Scholar]
- Dalziel, A.C.; Tirbhowan, S.; Drapeau, H.F.; Power, C.; Jonah, L.S.; Gbotsyo, Y.A.; Dion-Côté, A.M. Using asexual vertebrates to study genome evolution and animal physiology: Banded (Fundulus diaphanus) x common killifish (F. heteroclitus) hybrid lineages as a model system. Evol. Appl. 2020, 13, 1214–1239. [Google Scholar]
- Collares-Pereira, M.J.; Matos, I.; Morgado-Santos, M.; Coelho, M.M. Natural pathways towards polyploidy in animals: The Squalius alburnoides fish complex as a model system to study genome size and genome reorganization in polyploids. Cytogenet. Genome Res. 2013, 140, 97–116. [Google Scholar]
- Auvinet, J.; Graça, P.; Belkadi, L.; Petit, L.; Bonnivard, E.; Dettai, A.; Detrich, W.H.; Ozouf-Costaz, C.; Higuet, D. Mobilization of retrotransposons as a cause of chromosomal diversification and rapid speciation: The case for the Antarctic teleost genus Trematomus. BMC Genom. 2018, 19, 339. [Google Scholar]
- Cioffi, M.B.; Martins, C.; Bertollo, L.A.C. Chromosome spreading of associated transposable elements and ribosomal DNA in the fish Erythrinus erythrinus. Implications for genome change and karyoevolution in fish. BMC Evol. Biol. 2010, 10, 271. [Google Scholar]
- Motta-Neto, C.C.; Cioffi, M.D.; Costa, G.W.; Amorim, K.D.; Bertollo, L.A.; Artoni, R.F.; Molina, W.F. Overview on karyotype stasis in atlantic grunts (Eupercaria, Haemulidae) and the evolutionary extensions for other marine fish groups. Front. Mar. Sci. 2019, 6, 628. [Google Scholar]
- Auvinet, J.; Graça, P.; Ghigliotti, L.; Pisano, E.; Dettaï, A.; Ozouf-Costaz, C.; Higuet, D. Insertion hot spots of DIRS1 retrotransposon and chromosomal diversifications among the Antarctic teleosts Nototheniidae. Int. J. Mol. Sci. 2019, 20, 701. [Google Scholar]
- Volff, J.N.; Bouneau, L.; Ozouf-Costaz, C.; Fischer, C. Diversity of retrotransposable elements in compact pufferfish genomes. Trends Genet. 2003, 19, 12. [Google Scholar]
- Wicker, T.; Sabot, F.; Hua-Van, A.; Bennetzen, J.L.; Capy, P.; Chalhoub, B.; Flavell, A.; Leroy, P.; Morgante, M.; Panaud, O.; et al. A unified classification system for eukaryotic transposable elements. Nat. Rev. Genet. 2007, 8, 973–982. [Google Scholar]
- Sun, C.; Shepard, D.B.; Chong, R.A.; López Arriaza, J.; Hall, K.; Castoe, T.A.; Feschotte, C.; Pollock, D.D.; Mueller, R.L. LTR retrotransposons contribute to genomic gigantism in plethodontid salamanders. Genome Biol. Evol. 2012, 40, 168–183. [Google Scholar]
- Makałowski, W.; Gotea, V.; Pande, A.; Makałowska, I. Transposable elements: Classification, identification, and their use as a tool for comparative genomics. In Evolutionary Genomics: STATISTICAL and Computational Methods, Methods in Molecular Biology; Anisimova, M., Ed.; Springer: New York, NY, USA, 2019; Volume 1910. [Google Scholar] [CrossRef] [Green Version]
- Kazazian, H.H., Jr. Mobile elements: Drivers of genome evolution. Science 2004, 303, 1626–1632. [Google Scholar]
- Kramerov, D.A.; Vassetzky, N.S. Origin and evolution of SINEs in eukaryotic genomes. Heredity 2011, 107, 487–495. [Google Scholar]
- Evgen’ev, M.B.; Arkhipova, I.R. Penelope-like elements–A new class of retroelements: Distribution, function and possible evolutionary significance. Cytogenet. Genome Res. 2005, 110, 510–521. [Google Scholar]
- Arkhipova, I.R.; Yushenova, I.A.; Rodriguez, F. Endonuclease-containing Penelope retrotransposons in the bdelloid rotifer Adineta vaga exhibit unusual structural features and play a role in expansion of host gene families. Mob. DNA 2013, 4, 19. [Google Scholar]
- Bourque, G.; Burns, K.H.; Gehring, M.; Goburnovam, V.; Scluanov, A.; Hammell, M.; Imbeault, M.; Izsvák, Z.; Levin, H.L.; Macfarlan, T.S.; et al. Ten things you should know about transposable elements. Genome Biol. 2018, 19, 199. [Google Scholar]
- Goerner-Potvin, P.; Bourque, G. Computational tools to unmask transposable elements. Nat. Rev. Genet. 2018, 19, 688–704. [Google Scholar]
- Grabundzija, I.; Messing, S.A.; Thomas, J.; Cosby, R.L.; Bilic, I.; Miskey, C.; Gogol-Döring, A.; Kapitonov, V.; Diem, T.; Dalda, A.; et al. Helitron transposon reconstructed from bats reveals a novel mechanism of genome shuffling in eukaryotes. Nat. Commun. 2016, 7, 10716. [Google Scholar]
- Kapitonov, V.V.; Jurka, J. Self-synthesizing DNA transposons in eukaryotes. Proc. Natl. Acad. Sci. USA 2006, 103, 4540–4545. [Google Scholar]
- Feschotte, C.; Swamy, L.; Wessler, S.R. Genome-wide analysis of Mariner-like transposable elements in rice reveals complex relationships with stowaway miniature inverted repeat transposable elements (MITEs). Genetics 2003, 163, 747–758. [Google Scholar]
- Shao, F.; Han, M.; Peng, Z. Evolution and diversity of transposable elements in fish genomes. Sci. Rep. 2019, 9, 15399. [Google Scholar]
- Du, K.; Stöck, M.; Kneitz, S.; Klopp, C.; Woltering, J.M.; Adolfi, M.C.; Feron, R.; Prokopov, D.; Makunin, A.; Kichigin, I.; et al. The sterlet sturgeon genome sequence and the mechanisms of segmental rediploidization. Nat. Ecol. Evol. 2020, 4, 841–852. [Google Scholar]
- Dion-Cote, A.M.; Renaut, S.; Normandeau, E.; Bernatchez, L. RNA-seq reveals transcriptomic shock involving transposable elements reactivation in hybrids of young lake whitefish species. Mol. Biol. Evol. 2014, 31, 1188–1199. [Google Scholar]
- Oliver, K.R.; Greene, W.K. Mobile DNA and the TE-Thrust hypothesis: Supporting evidence from the primates. Mob. DNA 2011, 2, 8. [Google Scholar]
- Grahn, R.A.; Rinehart, T.A.; Cantrell, M.A.; Wichman, H.A. Extinction of LINE-1 activity coincident with a major mammalian radiation in rodents. Cytogenet. Genome Res. 2005, 110, 407–415. [Google Scholar]
- Platt, R.N.; Vandewege, M.W.; Kern, C.; Schmidt, C.J.; Hoffmann, F.G.; Ray, D.A. Large numbers of novel miRNAs originate from DNA transposons and are coincident with a large species radiation in bats. Mol. Biol. Evol. 2014, 31, 1536–1545. [Google Scholar]
- Burke, W.D.; Muller, F.; Eickbush, T.H. R4, a non-LTR retrotransposon specific to the large subunit rRNA genes of nematodes. Nucleic Acid Res. 1995, 23, 4628–4634. [Google Scholar]
- Yang, J.; Malik, H.S.; Eickbush, T.H. Identification of the endonuclease domain encoded by R2 and other site-specific, non-long terminal repeat retrotransposable elements. Proc. Natl. Acad. Sci. USA 1999, 96, 7847–7852. [Google Scholar]
- Yano, C.F.; Poltronieri, J.; Bertollo, L.A.C.; Ferreira Artoni, R.; Liehr, T.; Cioffi, M.B. Chromosomal mapping of repetitive DNAs in Triportheus trifurcatus (Characidae, Characiformes): Insights into the differentiation of the Z and W chromosomes. PLoS ONE 2014, 9, e90946. [Google Scholar]
- Pansonato-Alves, J.C.; Serrano, E.A.; Utsumomia, R.; Scacchetti, P.C.; Oliveira, C.; Foresti, F. Mapping five repetitive DNA classes in sympatric species of Hypostomus (Teleostei: Siluriformes: Loricariidae): Analysis of chromosomal variability. Rev. Fish. Biol. Fisher. 2013, 4, 447–489. [Google Scholar]
- Splendore de Borba, R.; Lourenço da Silva, E.; Parise-Maltempi, P.P. Chromosome mapping of retrotransposable elements Rex1 and Rex3 in Leporinus Spix, 1829 species (Characiformes: Anostomidae) and its relationships among heterochromatic segments and W sex chromosome. Mob. Genet. Elem. 2013, 3, e27460. [Google Scholar]
- Fantinatti, B.E.A.; Mazzuchelli, J.; Valente, G.T.; Cabral-de-Mello, D.C.; Martins, C. Genomic content and new insights on the origin of the B chromosome of the cichlid fish Astatotilapia latifasciata. Genetica 2011, 139, 1273–1282. [Google Scholar]
- Valente, G.T.; Mazzuchelli, J.; Ferreira, I.A.; Poletto, A.B. Cytogenetic mapping of the retroelements Rex1, Rex3 and Rex6 among cichlid fish: New insights on the chromosomal distribution of transposable elements. Cytogenet. Genome Res. 2011, 133, 34–42. [Google Scholar]
- Mazzuchelli, J.; Martins, C. Genomic organization of repetitive DNAs in the cichlid fish Astronotus ocellatus. Genetica 2009, 136, 461–469. [Google Scholar]
- Teixeira, W.G.; Ferreira, I.A.; Cabral-de-Mello, D.C.; Mazzuchelli, J.; Valente, G.T.; Pinhal, D.; Poletto, A.B.; Venere, P.C.; Martins, C. Organization of repeated DNA elements in the genome of the cichlid fish Cichla kelberi and its contributions to the knowledge of fish genomes. Cytogenet. Genome Res. 2009, 125, 224–234. [Google Scholar]
- Griffin, D.K.; Harvey, S.C.; Campos-Ramos, R.; Ayling, L.J.; Bromage, N.R.; Masabanda, J.S.; Penman, D.J. Early origins of the X and Y chromosomes: Lessons from tilapia. Cytogenet. Genome Res. 2002, 99, 157–163. [Google Scholar]
- Ozouf-Costaz, C.; Brandt, J.; Korting, C.; Pisano, E.; Bonillo, C.; Coutanceau, J.P.; Volff, J.N. Genome dynamics and chromosomal localization of the non-LTR retrotransposons Rex1 and Rex3 in Antarctic fish. Antarct. Sci. 2004, 16, 51–57. [Google Scholar]
- Supiwong, W.; Liehr, T.; Cioffi, M.B.; Chaveerach, A.; Kosyakova, N.; Pinthong, K.; Tanee, T.; Tanomtong, A. Chromosomal evolution in naked catfishes (Bagridae, Siluriformes): A comparative chromosome mapping study. Zool. Anz. 2014, 253, 316–320. [Google Scholar]
- Favarato, R.M.; Ribeiro, L.B.; Feldberg, E.; Matoso, D.A. Chromosomal mapping of transposable elements of the Rex family in the bristlenose catfish, ancistrus (Siluriformes, Loricariidae), from the amazonian region. J. Hered. 2017, 108, 254–261. [Google Scholar]
- Ferreira, D.C.; Porto-Foresti, F.; Oliveira, C.; Foresti, F. Transposable elements as a potential source for understanding the fish genome. Mob. Genet. Elements 2011, 1, 112–117. [Google Scholar]
- Bouneau, L.; Fischer, C.; Ozouf-Costaz, C.; Froschauer, A.; Jaillon, O.; Coutanceau, J.P.; Korting, C.; Weissenbach, J.; Bernot, A.; Volff, J.N. An active non-LTR retrotransposon with tandem structure in the compact genome of pufferfish Tetraodon nigroviridis. Genome Res. 2003, 13, 1686–1695. [Google Scholar]
- Carducci, F.; Barucca, M.; Canapa, A.; Biscotti, M.A. Rex retroelements and teleost genomes: An overview. Int. J. Mol. Sci. 2018, 19, 3653. [Google Scholar]
- Malik, H.S.; Henikoff, S.; Eickbush, T.H. Poised for contagion: Evolutionary origins of the infectious abilities of invertebrate retroviruses. Genome Res. 2000, 10, 1307–1318. [Google Scholar]
- Feschotte, C.; Gilbert, C. Endogenous viruses: Insights into viral evolution and impact on host biology. Nat. Rev. Genet. 2012, 13, 283–296. [Google Scholar]
- Hayward, A.; Cornwallis, C.K.; Jern, P. Pan-vertebrate comparative genomics unmasks retrovirus macroevolution. Proc. Natl. Acad. Sci. USA 2015, 112, 464–469. [Google Scholar]
- Xu, X.; Zhao, H.; Gong, Z.; Han, G.-Z. Endogenous retroviruses of non-avian/mammalian vertebrates illuminate diversity and deep history of retroviruses. Plos Pathog. 2018, 14, e1007072. [Google Scholar]
- Naville, M.; Volff, J.N. Endogenous retroviruses in fish genomes: From relics of past infections to evolutionary innovations? Front. Microbiol. 2016, 7, 1197. [Google Scholar]
- Basta, H.A.; Cleveland, S.B.; Clinton, R.A.; Dimitrov, A.G.; McClure, M.A. Evolution of teleost fish retroviruses: Characterization of new retroviruses with cellular genes. J. Virol. 2009, 83, 10152–10162. [Google Scholar]
- Shen, C.H.; Steiner, L.A. Genome structure and thymic expression of an endogenous retrovirus in zebrafish. J. Virol. 2004, 78, 899–911. [Google Scholar]
- Llorens, C.; Muñoz-Pomer, A.; Bernad, L.; Botella, H.; Moya, A. Network dynamics of eukaryotic LTR retroelements beyond phylogenetic trees. Biol. Direct. 2009, 4, 41. [Google Scholar]
- Schartl, M.; Walter, R.B.; Shen, Y.; Garcia, T.; Catchen, J.; Amores, A.; Braasch, I.; Chalopin, D.; Volff, J.N.; Lesch, K.P.; et al. The genome of the platyfish, Xiphophorus maculatus, provides insights into evolutionary adaptation and several complex traits. Nat. Genet. 2013, 45, 567–572. [Google Scholar]
- Naville, M.; Warren, I.A.; Haftek-Terreau, Z.; Chalopin, D.; Brunet, F.; Levin, P.; Galiana, D.; Volff, J.N. Not so bad after all: Retroviruses and long terminal repeat retrotransposons as a source of new genes in vertebrates. Clin. Microbiol. Infect. 2016, 22, 312–323. [Google Scholar]
- Feschotte, C. Transposable elements and the evolution of regulatory networks. Nat. Rev. Genet. 2008, 9, 397–405. [Google Scholar]
- Chuong, E.B.; Elde, N.C.; Feschotte, C. Regulatory evolution of innate immunity through co-option of endogenous retroviruses. Science 2016, 351, 1083–1087. [Google Scholar]
- Dechaud, C.; Volff, J.; Schartl, M.; Naville, M. Sex and the TEs: Transposable elements in sexual development and function in animals. Mob. DNA 2019, 10, 42. [Google Scholar]
- Ezaz, T.; Deakin, J.E. Repetitive sequence and sex chromosome evolution in vertebrates. Adv. Evol. Biol. 2014, 2014, 104683. [Google Scholar]
- Herpin, A.; Braasch, I.; Kraeussling, M.; Schmidt, C.; Thoma, E.C.; Nakamura, S.; Tanaka, M.; Schartl., M. Transcriptional rewiring of the sex determining dmrt1 gene duplicate by transposable elements. PLoS Genet. 2010, 6, e1000844. [Google Scholar]
- Schartl, M.; Schories, S.; Wakamatsu, Y.; Nagao, Y.; Hashimoto, H.; Bertin, C.; Mourot, B.; Schmidt, C.; Wilhelm, D.; Centanin, L.; et al. Sox5 is involved in germ-cell regulation and sex determination in medaka following co-option of nested transposable elements. BMC Biol. 2018, 16, 16. [Google Scholar]
- Tomaszkiewicz, M.; Chalopin, D.; Schartl, M.; Galiana, D.; Volff, J.N. A multicopy Y chromosomal SGNH hydrolase gene expressed in the testis of the platyfish has been captured and mobilized by a Helitron transposon. BMC Genet. 2014, 15, 44. [Google Scholar]
- Lubieniecki, K.P.; Lin, S.; Cabana, E.I.; Li, J.; Lai, Y.Y.Y.; Davidson, W.S. Genomic instability of the sex-determining locus in Atlantic salmon (Salmo salar). G3 2015, 5, 2513–2522. [Google Scholar]
- Sotero-Caio, C.G.; Platt, R.N., II; Suh, A.; Ray, D.A. Evolution and diversity of transposable elements in vertebrate genomes. Genome Biol. Evol. 2017, 9, 161–177. [Google Scholar]
- Casacuberta, E.; González, J. The impact of transposable elements in environmental adaptation. Mol. Ecol. 2013, 6, 1503–1517. [Google Scholar]
- Carducci, F.; Biscotti, M.A.; Forconi, M.; Barucca, M.; Canapa, A. An intriguing relationship between teleost Rex3 retroelement and environmental temperature. Biol. Lett. 2019, 15, 20190279. [Google Scholar]
- Schmidt, A.L.; Anderson, L.M. Repetitive DNA elements as mediators of genomic change in response to environmental cues. Biol. Rev. Camb. Philos. Soc. 2006, 81, 531–543. [Google Scholar]
- Fablet, M.; Vieira, C. Evolvability, epigenetics and transposable elements. Biomol. Concepts 2011, 2, 331–341. [Google Scholar]
- Canapa, A.; Biscotti, M.A.; Barucca, M.; Carducci, F.; Carotti, E.; Olmo, E. Shedding light upon the complex net of genome size, genome composition and environment in chordates. Eur. Zool. J. 2020, 87, 192–202. [Google Scholar]
- Biscotti, M.A.; Barucca, M.; Canapa, A. New insights into the genome repetitive fraction of the Antarctic bivalve Adamussium colbecki. PLoS ONE 2018, 13, e0194502. [Google Scholar]
- Chu, C.G.; Tan, C.T.; Yu, G.T.; Zhong, S.; Xu, S.S.; Yan, L. A novel retrotransposon inserted in the dominant Vrn-B1 allele confers spring growth habit in tetraploid wheat (Triticum turgidum L.). G3 2011, 1, 637–645. [Google Scholar]
- Fujino, K.; Hashida, S.N.; Ogawa, T.; Natsume, T.; Uchiyama, T.; Mikami, T.; Kishima, Y. Temperature controls nuclear import of Tam3 transposase in Antirrhinum. Plant J. 2011, 65, 146–155. [Google Scholar]
- Makarevitch, I.; Waters, A.J.; West, P.T.; Stitzer, M.; Hirsch, C.N.; Ross-Ibarra, J.; Springer, N.M. Transposable elements contribute to activation of maize genes in response to abiotic stress. PLoS Genet. 2015, 11, e1004915. [Google Scholar]

| Teleost Orders | References |
|---|---|
| Polypteriformes | |
| Acipenseriformes | [24,25] |
| Amiiformes | |
| Lepisosteiformes | [19,20,26,27,28,29,30,31,32,33,34,35] |
| Elopiformes | [36] |
| Albuliformes | |
| Notacanthiformes | |
| Anguilliformes | [24,32,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62] |
| Osteoglossomorpha | [63,64] |
| Clupeiformes | [65,66] |
| Alepocephaliformes | |
| Gonorynchiformes | [67] |
| Cypriniformes | [14,68,69,70,71,72,73,74,75,76] |
| Characiformes | [77,78,79,80,81,82,83,84,85,86,87,88,89,90,91] |
| Gymnotiformes | [92,93,94] |
| Siluriformes | [14,31,95,96,97,98,99,100] |
| Lepidogalaxiiformes | |
| Argentiniformes | [101] |
| Galaxiiformes | [102] |
| Esociformes | [103] |
| Salmoniformes | [104,105,106,107] |
| Osmeriformes | [108,109] |
| Stomiatiformes | [109] |
| Ateleopodiformes | [109] |
| Aulopiformes | [109,110] |
| Myctiophiformes | [109] |
| Lampriformes | [109] |
| Percopsiformes | [109] |
| Zeiformes | [109] |
| Stylephoriformes | [109] |
| Gadiformes | [111,112,109] |
| Polymixiiformes | [109] |
| Trachicthtyiformes | |
| Beryciformes | [34,109,113,114,115,116] |
| Holocentriformes | [109] |
| Ophidiaria | [109,111,117] |
| Batrachoidiaria | [109] |
| Pelagiaria | [109] |
| Synghatharia | [118] |
| Gobiaria | [109] |
| Anabantaria | [109,119] |
| Carangaria | [109,120,121,122] |
| Ovalentaria | [109,123,124,125,126] |
| Eupercaria | [30,31,109,119,127,128,129,130,131,132,133,134,135,136,137,138,139,140,141,142,143,144,145,146,147,148,149,150,151,152] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carducci, F.; Barucca, M.; Canapa, A.; Carotti, E.; Biscotti, M.A. Mobile Elements in Ray-Finned Fish Genomes. Life 2020, 10, 221. https://doi.org/10.3390/life10100221
Carducci F, Barucca M, Canapa A, Carotti E, Biscotti MA. Mobile Elements in Ray-Finned Fish Genomes. Life. 2020; 10(10):221. https://doi.org/10.3390/life10100221
Chicago/Turabian StyleCarducci, Federica, Marco Barucca, Adriana Canapa, Elisa Carotti, and Maria Assunta Biscotti. 2020. "Mobile Elements in Ray-Finned Fish Genomes" Life 10, no. 10: 221. https://doi.org/10.3390/life10100221
APA StyleCarducci, F., Barucca, M., Canapa, A., Carotti, E., & Biscotti, M. A. (2020). Mobile Elements in Ray-Finned Fish Genomes. Life, 10(10), 221. https://doi.org/10.3390/life10100221









