Feature Selection Methods for Extreme Learning Machines
Abstract
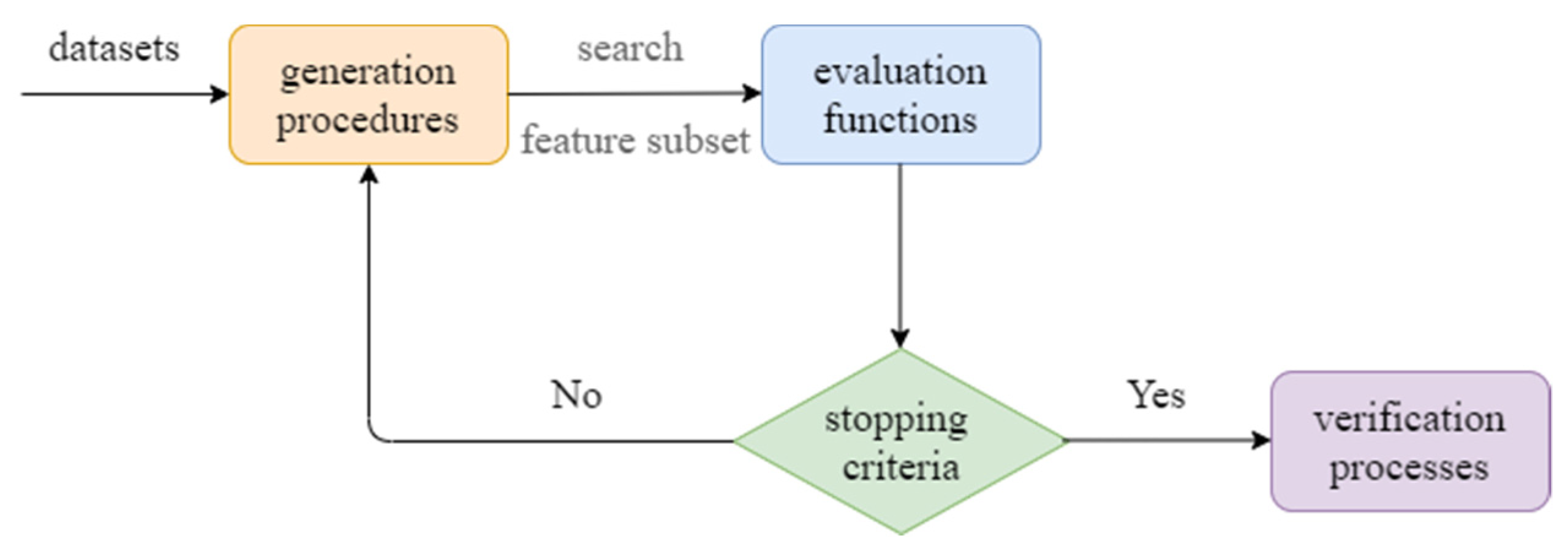
:1. Introduction
- A wrapper feature selection method is proposed for the ELM, called FELM. In the FELM, the corresponding objective function and hyperplane are introduced by adding a feature selection matrix, a diagonal matrix with element either 1 or 0 to the objective functions of the ELM. The FELM can effectively reduce the dimensionality of the input space and remove redundant features.
- The FELM is extended to the nonlinear case (called FKELM) by incorporating a kernel function that combines generalization and memorization radial basis function (RBF) kernels into the FELM. The FKELM can obtain high classification accuracy and extensive generalization by fully applying the property of memorization of training data.
- A feature ranking strategy is proposed, which can evaluate features based on their contributions to the objective functions of the FELM and FKELM. After obtaining the best matrix E, the resulting methods significantly improve the classification accuracy, generalization performance, and learning speed.
2. Related Works
2.1. ELM
2.2. Memorization–Generalization Kernel
3. Proposed Methods
3.1. Mathematical Model
3.2. Solutions of FELM and FKELM
3.3. Computational Complexity Analysis
| Algorithm 1 Feature selection method for ELM |
| Input: Samples ; Appropriate parameters C,; A fixed and large integer k, which is the number of sweeps through E; Stopping tolerance tol = 1e-6; Output: Output weight ; 1. Set , solve the Equation (10) with the fixed E; 2. Compute each feature score by Equation (27), for , if , then set ; 3. Solve Equation (10) with the fixed E, and obtain ; 4. Repeat 5. For and : (1) Replace from 1 to 0 (or 0 to 1). (2) Compute before and after changing . (3) Keep the new only if decreases more than the tolerance. Else undo the change of , and go to (1) if . (4) Go to step 6 if the total decrease in is less than or equal to tol in the last n steps. 6. Solve the Equation (10) with the new E, and compute . 7. Until the objective function decrease of Equation (10) is less than tol if is changed. |
| Algorithm 2 Feature selection method for KELM |
| Input: Samples ; Appropriate parameters C,; Appropriate parameters in kernel ; A fixed and large integer k, which is the number of sweeps through E; Stopping tolerance tol=1e-6; Output: Output weight ; . 1. Set , solve the Equation (12) with the fixed E; 2. Compute each feature score by Equation (28), for , if , then set ; 3. Solve Equation (12) with the fixed E, and obtain ; 4. Repeat 5. For and : (1) Replace from 1 to 0 (or 0 to 1). (2) Compute be- fore and after changing . (3) Keep the new only if decreases more than the tolerance. Else undo the change of , and go to (1) if . (4) Go to step 6 if the total decrease in is less than or equal to tol in the last n steps. 6. Solve the Equation (12) with the new E, and compute . 7. Until the Equation (12) is less than tol if is changed. |
4. Experimental Results
4.1. Classification Performance
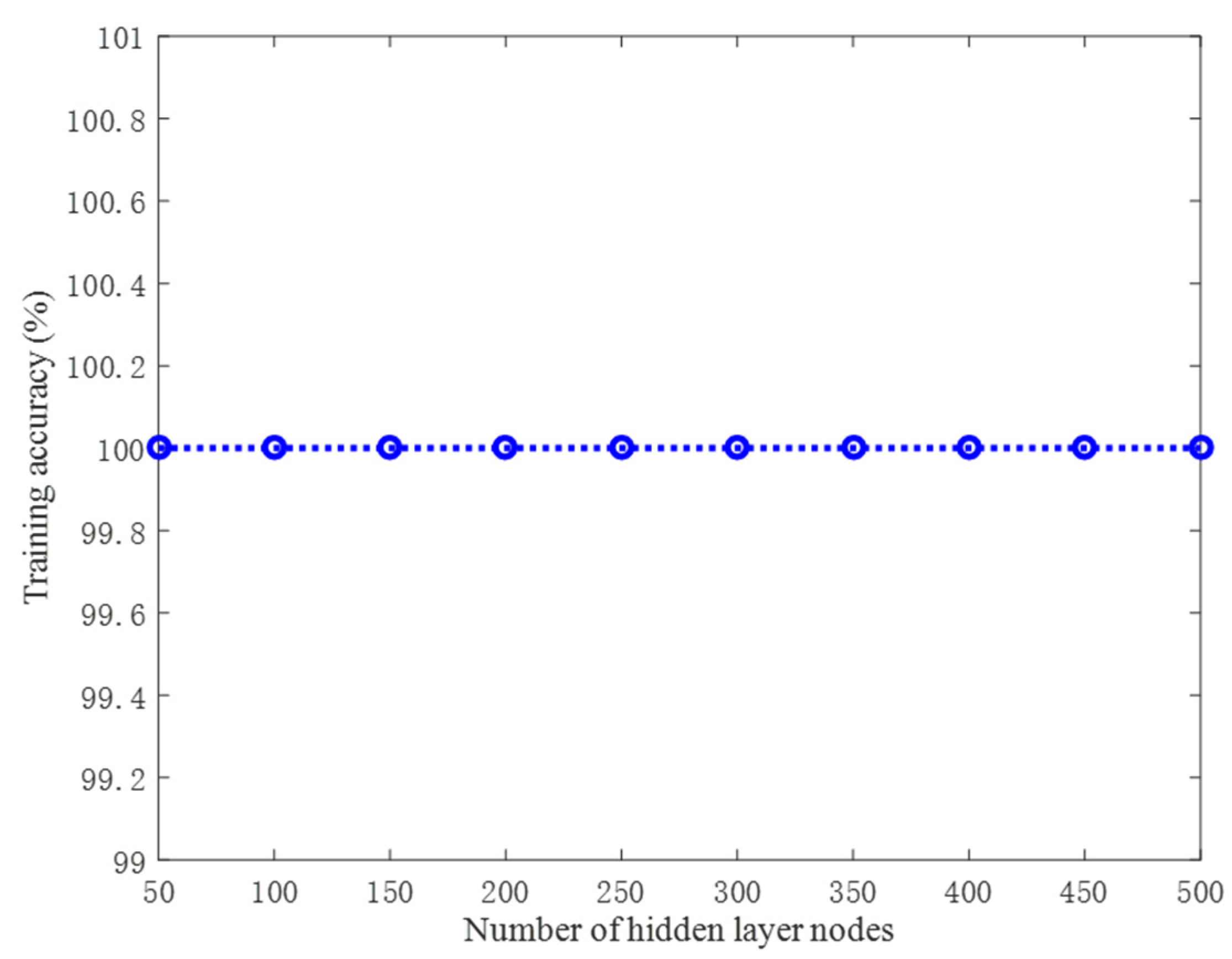
4.1.1. Performance on the Artificial Datasets
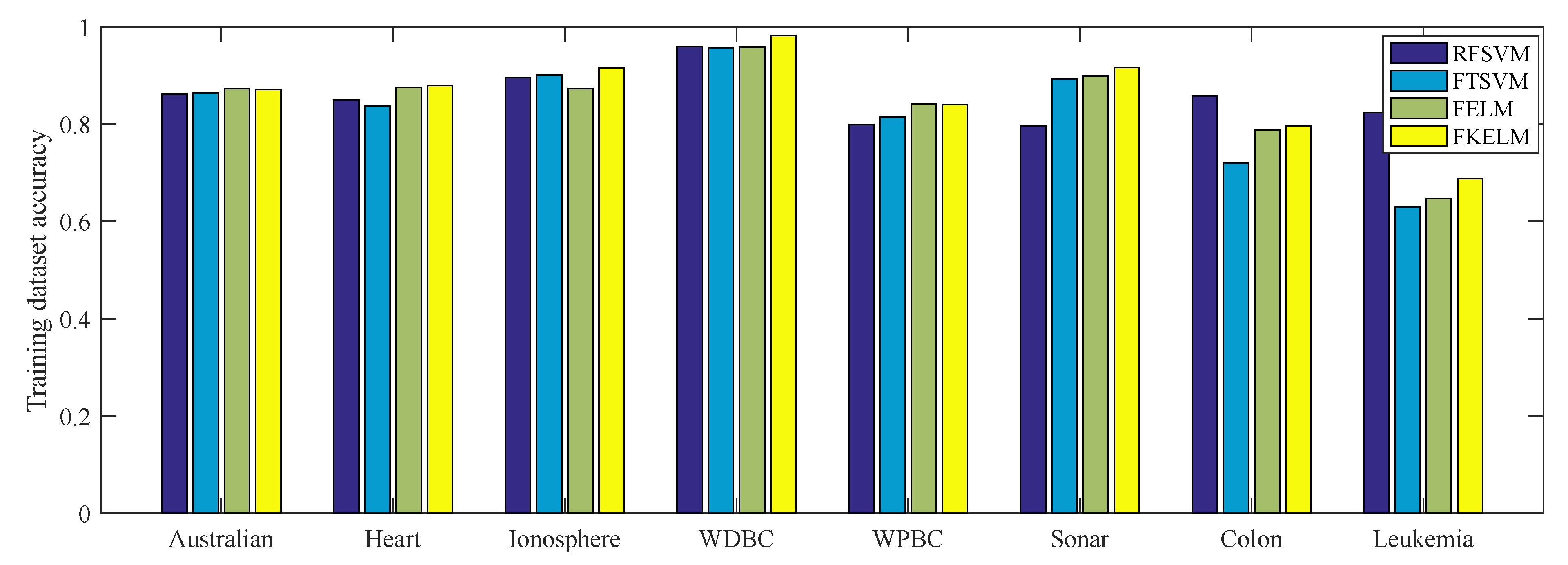
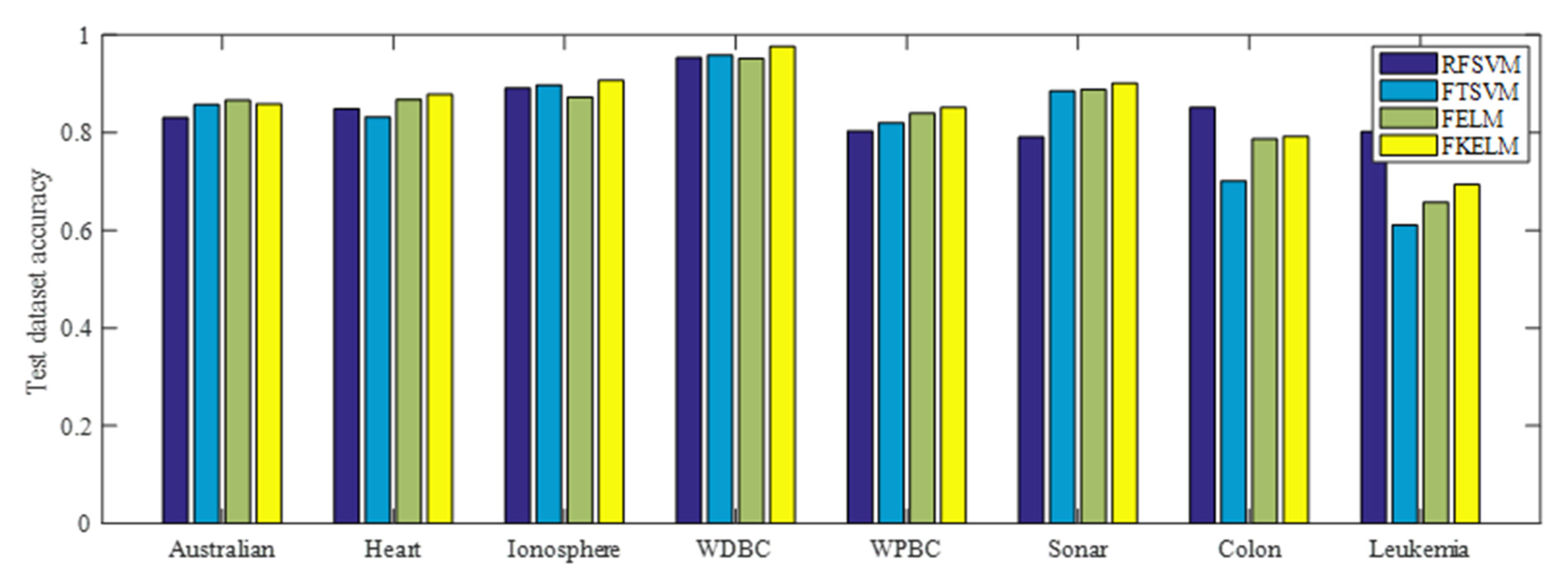
4.1.2. Performance on the Benchmark Datasets
4.2. Discussion of Results
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Dash, M.; Liu, H. Feature selection for classification. Intell. Data Anal. 1997, 1, 131–156. [Google Scholar] [CrossRef]
- Zhou, L.; Si, Y.W.; Fujita, H. Predicting the listing statuses of Chinese-listed companies using decision trees combined with an improved filter feature selection method. Knowl.-Based Syst. 2017, 128, 93–101. [Google Scholar] [CrossRef]
- Kang, M.; Rashedul, I.M.; Jaeyoung, K.; Kim, J.M.; Pecht, M. A Hybrid feature selection scheme for reducing diagnostic performance deterioration caused by outliers in Data-Driven diagnostics. IEEE Trans. Ind. Electron. 2016, 63, 3299–3310. [Google Scholar] [CrossRef]
- Zhao, J.; Chen, L.; Pedrycz, W.; Wang, W. Variational inference-based automatic relevance determination kernel for embedded feature selection of noisy industrial data. IEEE Trans. Ind. Electron. 2018, 66, 416–428. [Google Scholar] [CrossRef]
- Souza, R.; Macedo, C.; Coelho, L.; Pierezan, J.; Mariani, V.C. Binary coyote optimization algorithm for feature selection. Pattern Recognit. 2020, 107, 107470. [Google Scholar] [CrossRef]
- Sun, L.; Wang, T.X.; Ding, W.; Xu, J.; Lin, Y. Feature selection using fisher score and multilabel neighborhood rough sets for multilabel classification. Inf. Sci. 2021, 578, 887–912. [Google Scholar] [CrossRef]
- Binu, D.; Kariyappa, B.S. Rider Deep LSTM Network for Hybrid Distance Score-based Fault Prediction in Analog Circuits. IEEE Trans. Ind. Electron. 2020, 99, 10097–10106. [Google Scholar] [CrossRef]
- Jin, C.; Li, F.; Ma, S.; Wang, Y. Sampling scheme-based classification rule mining method using decision tree in big data environment. Knowl.-Based Syst. 2022, 244, 108522. [Google Scholar] [CrossRef]
- Rouhani, H.; Fathabadi, A.; Baartman, J. A wrapper feature selection approach for efficient modelling of gully erosion susceptibility mapping. Prog. Phys. Geogr. 2021, 45, 580–599. [Google Scholar] [CrossRef]
- Liu, W.; Wang, J. Recursive elimination current algorithms and a distributed computing scheme to accelerate wrapper feature selection. Inf. Sci. 2022, 589, 636–654. [Google Scholar] [CrossRef]
- Pintas, J.T.; Fernandes, L.; Garcia, A. Feature selection methods for text classification: A systematic literature review. Artif. Intell. Rev. 2021, 54, 6149–6200. [Google Scholar] [CrossRef]
- Vapnik, V.N. The Nature of Statistical Learning Theory; Springer: Berlin/Heidelberg, Germany, 1995; Volume 8, p. 1564. [Google Scholar]
- Jayadeva; Khemchandani, R.; Chandra, S. Twin support vector machines for pattern classification. IEEE Trans. Ind. Electron. 2007, 29, 905–910. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.B.; Zhu, Q.Y.; Siew, C.K. Extreme learning machine: Theory and applications. Neurocomputing 2006, 70, 489–501. [Google Scholar] [CrossRef]
- Cui, D.; Huang, G.B.; Liu, T. ELM based Smile Detection using Distance Vector. Pattern Recognit. 2018, 78, 356–369. [Google Scholar] [CrossRef]
- Huang, G.B.; Zhou, H.; Ding, X.; Zhang, R. Extreme learning machine for regression and multiclass classification. IEEE Trans. Syst. Man Cybern. Part B Cybern. 2012, 42, 513–529. [Google Scholar] [CrossRef]
- Albashish, D.; Hammouri, A.I.; Braik, M.; Atwan, J.; Sahran, S. Binary biogeography-based optimization based SVM-RFE for feature selection. Appl. Soft Comput. 2020, 101, 107026. [Google Scholar] [CrossRef]
- Guyon, I.; Weston, J.; Barnhill, S. Gene selection for cancer classification using support vector machines. Mach. Learn. 2002, 46, 389–422. [Google Scholar] [CrossRef]
- Chang, Y.; Lin, C.; Guyon, I. Feature ranking using linear SVM. In Proceedings of the Workshop on the Causation and Prediction Challenge at WCCI 2008, Hong Kong, China, 3–4 June 2008; 2008; Volume 3, pp. 53–64. [Google Scholar]
- Mangasarian, O.L.; Gang, K. Feature selection for nonlinear kernel support vector machines. In Proceedings of the Seventh IEEE International Conference on Data Mining Workshops (ICDMW 2007), Omaha, NE, USA, 31 March 2008; pp. 28–31. [Google Scholar]
- Bai, L.; Wang, Z.; Shao, Y.H.; Deng, N.Y. A novel feature selection method for twin support vector machine. Knowl.-Based Syst. 2014, 59, 1–8. [Google Scholar] [CrossRef]
- Man, Z.; Huang, G.B. Special issue on extreme learning machine and deep learning networks. Neural Comput. Appl. 2020, 32, 14241–14245. [Google Scholar] [CrossRef]
- Adesina, A.F.; Jane, L.; Abdulazeez, A. Ensemble model of non-linear feature selection-based Extreme Learning Machine for improved natural gas reservoir characterization. J. Nat. Gas Sci. Eng. 2015, 26, 1561–1572. [Google Scholar]
- Vapnik, V.; Izmailov, R. Reinforced SVM method and memorization mechanisms. Pattern Recognit. 2021, 119, 108018. [Google Scholar] [CrossRef]
- Iosifidis, A.; Gabbouj, M. On the kernel Extreme Learning Machine classifier. Pattern Recognit. Lett. 2015, 54, 11–17. [Google Scholar] [CrossRef]
- UCI Machine Learning Repository. Available online: http://archive.ics.uci.edu/ml/datasets.php (accessed on 15 October 2021).
- Test, L. Gene Expression Leukemia Testing Data set from Golub. Available online: https://search.r-project.org/CRAN/refmans/SIS/html/leukemia.test.html (accessed on 15 October 2021).
- Microarray Data. Available online: https://www.sciencedirect.com/topics/computer-science/microarray-data (accessed on 15 October 2021).
- Hua, X.G.; Ni, Y.Q.; Ko, J.M.; Wong, K.Y. Modeling of temperature–frequency correlation using combined principal component analysis and support vector regression technique. J. Comput. Civ. Eng. 2007, 21, 122–135. [Google Scholar] [CrossRef]
- Demšar, J. Statistical comparisons of classifiers over multiple data sets. J. Mach. Learn. Res. 2006, 7, 1–30. [Google Scholar]
- Iman, L.; Davenport, J.M. Approximations of the critical region of the Friedman statistic. Commun. Stat. A 1998, A9, 571–595. [Google Scholar]
- Zheng, F.; Webb, G.I.; Suraweera, P.; Zhu, L. Subsumption resolution: An efficient and effective technique for semi–naive Bayesian learning. Mach. Learn. 2012, 87, 93–125. [Google Scholar] [CrossRef]






| Dataset | Function Definition |
|---|---|
| Self-defining function | mean value (0,0) and (0.8,0.8) |
| Two-moon |
| Dataset | Number of Test Samples | Number of Training Samples | Range of Independent Variables |
|---|---|---|---|
| Self-defining function | 50 | 350 | |
| Two-moon | 100 | 501 |
| Dataset | Algorithm | Training Accuracy | Test Accuracy | Time (s) | Parameters |
|---|---|---|---|---|---|
| Two-moon | RFSVM | 0.9948 | 0.9600 | 0.8632 | 2–8, –, –, 22, 2–3, – |
| FTSVM | 1.0000 | 0.9819 | 0.6929 | –, 2–3, 22, 21, 2–3, – | |
| FELM | 1.0000 | 1.0000 | 0.3208 | 2–6, –, –, 22, 2–1, – | |
| FKELM | 1.0000 | 1.0000 | 0.3524 | 2–6, –, –, 22, 2–1, 0.3 | |
| Self-defining function | RFSVM | 0.8977 | 0.9012 | 0.5007 | 2–7, –, –, 20, 2–3, – |
| FTSVM | 0.9293 | 0.9110 | 0.2410 | –, 20, 23, 21, 2–4, – | |
| FELM | 0.9922 | 0.9879 | 0.1375 | 2–5, –, –, 22, 2–2, – | |
| FKELM | 0.9972 | 0.9913 | 0.1398 | 2–5, –, –, 22, 2–2, 0.4 |
| Dataset | Number of Test Samples | Number of Training Samples | Number of Features |
|---|---|---|---|
| Australian | 207 | 483 | 14 |
| Heart | 81 | 189 | 13 |
| Ionosphere | 105 | 246 | 33 |
| WDBC | 171 | 398 | 30 |
| WPBC | 59 | 139 | 34 |
| Sonar | 62 | 146 | 60 |
| Colon | 18 | 44 | 2000 |
| Leukaemia | 21 | 51 | 7129 |
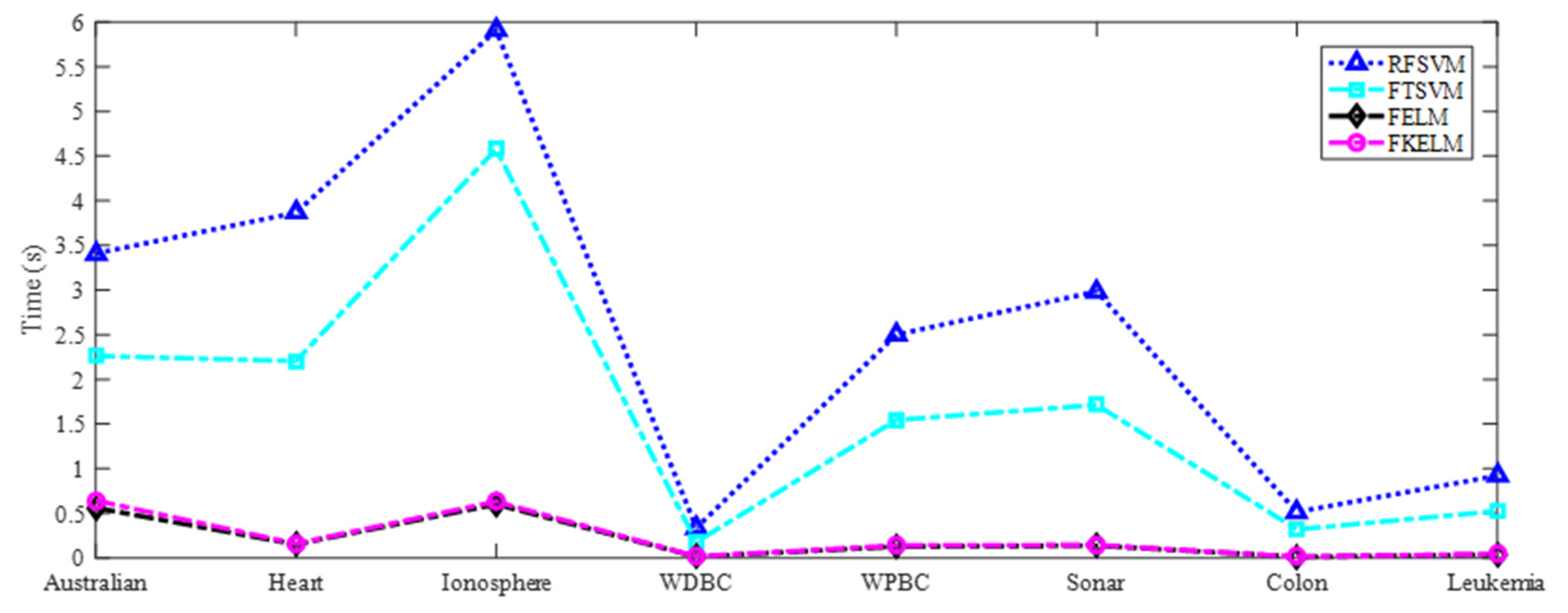
| Dataset | Algorithm | Training accuracy | Test accuracy | Time (s) | Parameter (C,C1,C2,Ψ,σ,τ) |
|---|---|---|---|---|---|
| Australian | RFSVM | 0.8617 | 0.8310 | 3.4092 | 24, –, –, 22, 2–7, – |
| FTSVM | 0.8638 | 0.8573 | 2.2620 | –, 23, 25, 22, 2–8, – | |
| FELM | 0.8732 | 0.8662 | 0.5612 | 2–5, –, –, 21, 2–1, – | |
| FKELM | 0.8707 | 0.8595 | 0.6425 | 2–5, –, –, 21, 2–1, 0.6 | |
| Heart | RFSVM | 0.8496 | 0.8485 | 3.8659 | 21, –, –, 21, 2–7, – |
| FTSVM | 0.8370 | 0.8321 | 2.2051 | –, 22, 22, 21, 2–9, – | |
| FELM | 0.8759 | 0.8679 | 0.1528 | 2–2, –, –, 20, 2–2, – | |
| FKELM | 0.8796 | 0.8785 | 0.1607 | 2–2, –, –, 20, 2–2, 0.4 | |
| Ionosphere | RFSVM | 0.8957 | 0.8912 | 5.9061 | 2–1, –, –, 20, 2–3, – |
| FTSVM | 0.9007 | 0.8976 | 4.5770 | –, 20, 23, 22, 2–2, – | |
| FELM | 0.8729 | 0.8728 | 0.6012 | 2–5, –, –, 22, 2–2, – | |
| FKELM | 0.9157 | 0.9072 | 0.6314 | 2–5, –, –, 22, 2–2, 0.4 | |
| WDBC | RFSVM | 0.9597 | 0.9537 | 0.3372 | 2–2, –, –, 21, 2–3, – |
| FTSVM | 0.9573 | 0.9586 | 0.1798 | –, 20, 20, 21, 2–4, – | |
| FELM | 0.9592 | 0.9518 | 0.0146 | 23, –, –, 20, 2–5, – | |
| FKELM | 0.9823 | 0.9762 | 0.0177 | 25, –, –, 20, 2–5, 0.5 | |
| WPBC | RFSVM | 0.7992 | 0.8034 | 2.5007 | 27, –, –, 21, 2–5, – |
| FTSVM | 0.8141 | 0.8202 | 1.5451 | –, 25, 26, 21, 2–5, – | |
| FELM | 0.8422 | 0.8397 | 0.1279 | 23, –, –, 21, 2–8, – | |
| FKELM | 0.8409 | 0.8513 | 0.1398 | 23, –, –, 21, 2–8, 0.3 | |
| Sonar | RFSVM | 0.7972 | 0.7911 | 2.9832 | 22, –, –, 20, 2–3, – |
| FTSVM | 0.8933 | 0.8849 | 1.7132 | –, 21, 25, 21, 2–2, – | |
| FELM | 0.8995 | 0.8879 | 0.1375 | 23, –, –, 22, 2–4, – | |
| FKELM | 0.9172 | 0.9013 | 0.1452 | 24, –, –, 22, 2–4, 0.5 | |
| Colon | RFSVM | 0.8583 | 0.8512 | 0.5160 | 22, –, –, 20, 21, – |
| FTSVM | 0.7208 | 0.7010 | 0.3197 | –, 20, 23, 21, 20, – | |
| FELM | 0.7882 | 0.7871 | 0.0133 | 2–1, –, –, 22, 22, – | |
| FKELM | 0.7971 | 0.7922 | 0.0139 | 2–1, –, –, 22, 20, 0.6 | |
| Leukaemia | RFSVM | 0.8235 | 0.8017 | 0.9209 | 27, –, –, 23, 2–5, – |
| FTSVM | 0.6293 | 0.6110 | 0.5244 | –, 25, 26, 23, 2–5, – | |
| FELM | 0.6471 | 0.6572 | 0.0399 | 24, –, –, 24, 2–6, – | |
| FKELM | 0.6882 | 0.6938 | 0.0451 | 24, –, –, 24, 2–5, 0.6 |
| Dataset | RFSVM | FTSVM | FELM | FKELM | |
|---|---|---|---|---|---|
| Australian | Training | 4 | 3 | 1 | 2 |
| Test | 4 | 3 | 1 | 2 | |
| Heart | Training | 3 | 4 | 2 | 1 |
| Test | 3 | 4 | 2 | 1 | |
| Ionosphere | Training | 3 | 2 | 4 | 1 |
| Test | 3 | 2 | 4 | 1 | |
| WDBC | Training | 2 | 4 | 3 | 1 |
| Test | 3 | 2 | 4 | 1 | |
| WPBC | Training Test | 4 4 | 3 3 | 1 1 | 2 2 |
| Sonar | Training | 4 | 3 | 2 | 1 |
| Test | 4 | 3 | 2 | 1 | |
| Colon | Training | 1 | 4 | 3 | 2 |
| Test | 1 | 4 | 3 | 2 | |
| Leukaemia | Training | 1 | 4 | 3 | 2 |
| Test | 1 | 4 | 3 | 2 | |
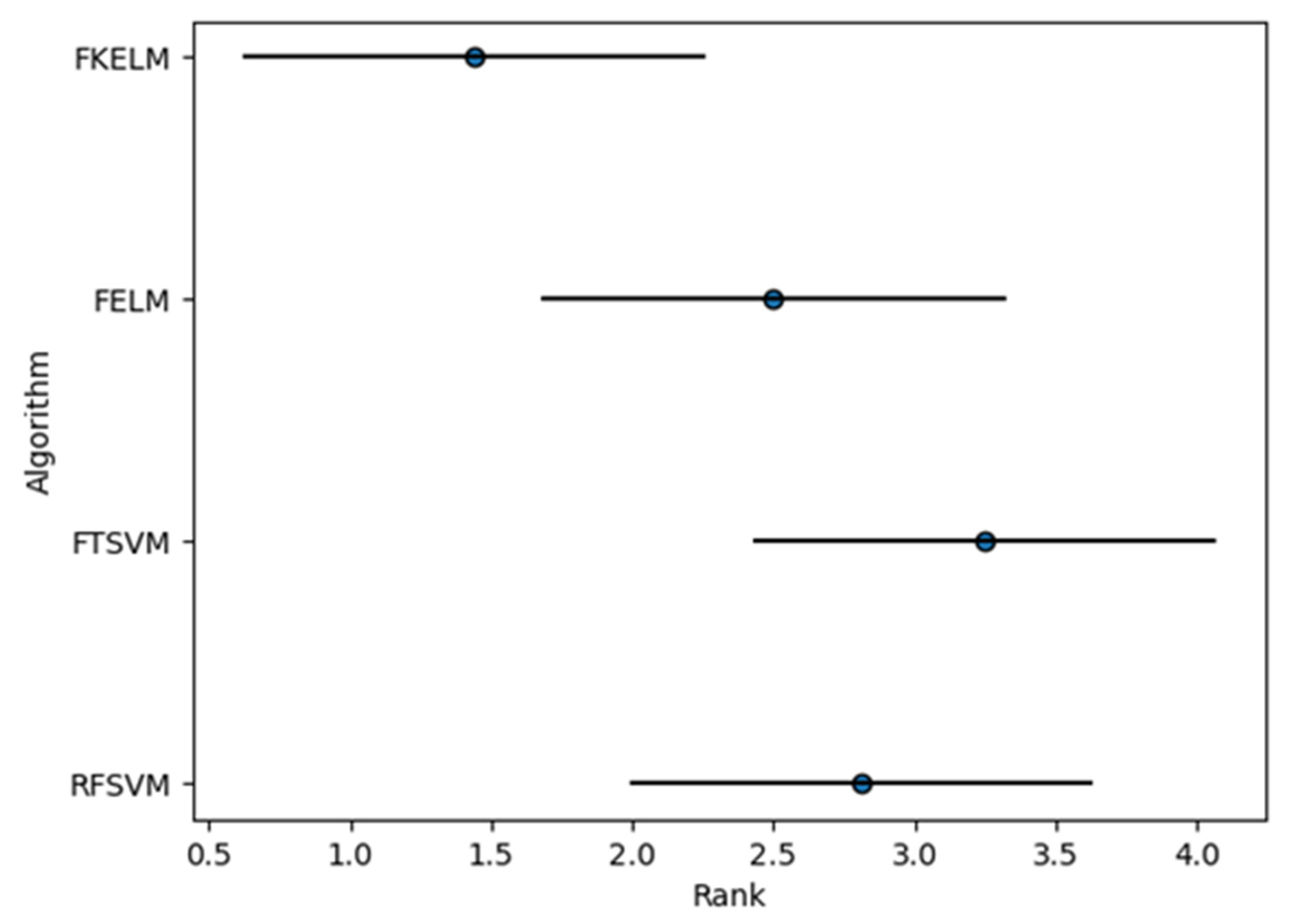
| Average rank | 2.8125 | 3.25 | 2.5 | 1.4375 | |
| Algorithm | Size of Dataset | Time | Classification Performance |
|---|---|---|---|
| RFSVM | Small and medium samples | Much | Poor |
| Large samples | Much | Sightly good | |
| FTSVM | Small and medium samples | Medium | Poor |
| Large samples | Medium | Poor | |
| FELM | Small and medium samples | Little | Significantly good |
| Large samples | Little | Good | |
| FKELM | Small and medium samples | Little | Significantly good |
| Large samples | Little | Good |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fu, Y.; Wu, Q.; Liu, K.; Gao, H. Feature Selection Methods for Extreme Learning Machines. Axioms 2022, 11, 444. https://doi.org/10.3390/axioms11090444
Fu Y, Wu Q, Liu K, Gao H. Feature Selection Methods for Extreme Learning Machines. Axioms. 2022; 11(9):444. https://doi.org/10.3390/axioms11090444
Chicago/Turabian StyleFu, Yanlin, Qing Wu, Ke Liu, and Haotian Gao. 2022. "Feature Selection Methods for Extreme Learning Machines" Axioms 11, no. 9: 444. https://doi.org/10.3390/axioms11090444
APA StyleFu, Y., Wu, Q., Liu, K., & Gao, H. (2022). Feature Selection Methods for Extreme Learning Machines. Axioms, 11(9), 444. https://doi.org/10.3390/axioms11090444






