Using Nanobodies to Study Protein Function in Developing Organisms
Abstract
:1. Introduction
2. From Cultured Cells to Developing Organisms
3. Use of Functionalized Nanobodies in Multicellular Animals in the Context of Developmental Biology Studies
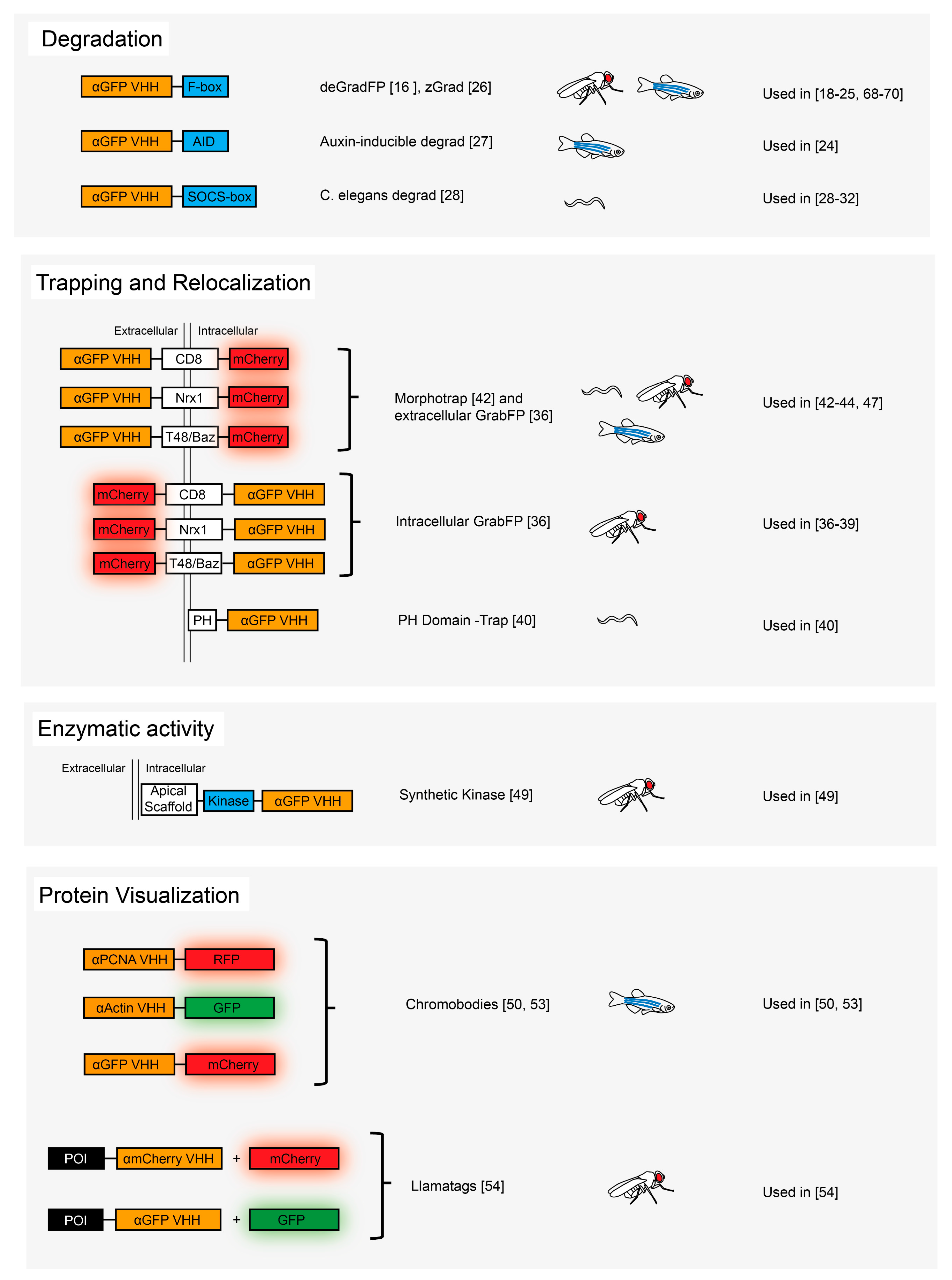
3.1. Protein Degradation
3.2. Protein Relocalization and Trapping
3.3. Protein Post-Translational Modification
3.4. Protein Visualization
3.5. Protein Scaffolding and Cell–Cell Contact Reporters
4. A look Into the Future
Author Contributions
Funding
Conflicts of Interest
References
- Hamers-Casterman, C.; Atarhouch, T.; Muyldermans, S.; Robinson, G.; Hamers, C.; Songa, E.B.; Bendahman, N.; Hamers, R. Naturally occurring antibodies devoid of light chains. Nature 1993, 363, 446–448. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; La Russa, M.; Qi, L.S. CRISPR/Cas9 in Genome Editing and Beyond. Annu. Rev. Biochem. 2016, 85, 227–264. [Google Scholar] [CrossRef] [PubMed]
- Helma, J.; Cardoso, M.C.; Muyldermans, S.; Leonhardt, H. Nanobodies and recombinant binders in cell biology. J. Cell. Biol. 2015, 209, 633–644. [Google Scholar] [CrossRef] [PubMed]
- Schumacher, D.; Helma, J.; Schneider, A.F.L.; Leonhardt, H.; Hackenberger, C.P.R. Nanobodies: Chemical Functionalization Strategies and Intracellular Applications. Angew. Chem. Int. Ed. Engl. 2018, 57, 2314–2333. [Google Scholar] [CrossRef] [PubMed]
- Beghein, E.; Gettemans, J. Nanobody Technology: A Versatile Toolkit for Microscopic Imaging, Protein-Protein Interaction Analysis, and Protein Function Exploration. Front Immunol. 2017, 8, 771. [Google Scholar] [CrossRef] [PubMed]
- Kaiser, P.D.; Maier, J.; Traenkle, B.; Emele, F.; Rothbauer, U. Recent progress in generating intracellular functional antibody fragments to target and trace cellular components in living cells. Biochim. Biophys. Acta. 2014, 1844, 1933–1942. [Google Scholar] [CrossRef] [PubMed]
- Binz, H.K.; Stumpp, M.T.; Forrer, P.; Amstutz, P.; Plückthun, A. Designing Repeat Proteins: Well-expressed, Soluble and Stable Proteins from Combinatorial Libraries of Consensus Ankyrin Repeat Proteins. J. Mol. Biol. 2003, 332, 489–503. [Google Scholar] [CrossRef]
- Koide, A.; Bailey, C.W.; Huang, X.; Koide, S. The fibronectin type III domain as a scaffold for novel binding proteins11Edited by J. Wells. J. Mol. Biol. 1998, 284, 1141–1151. [Google Scholar] [CrossRef] [PubMed]
- Bieli, D.; Alborelli, I.; Harmansa, S.; Matsuda, S.; Caussinus, E.; Affolter, M. Development and Application of Functionalized Protein Binders in Multicellular Organisms. Int. Rev. Cell Mol. Biol. 2016, 325, 181–213. [Google Scholar] [CrossRef] [PubMed]
- Harmansa, S.; Affolter, M. Protein binders and their applications in developmental biology. Development 2018, 145. [Google Scholar] [CrossRef] [PubMed]
- Boldicke, T. Single domain antibodies for the knockdown of cytosolic and nuclear proteins. Protein Sci. 2017, 26, 925–945. [Google Scholar] [CrossRef] [PubMed]
- Ingram, J.R.; Schmidt, F.I.; Ploegh, H.L. Exploiting Nanobodies’ Singular Traits. Annu. Rev. Immunol. 2018, 36, 695–715. [Google Scholar] [CrossRef] [PubMed]
- Housden, B.E.; Muhar, M.; Gemberling, M.; Gersbach, C.A.; Stainier, D.Y.; Seydoux, G.; Mohr, S.E.; Zuber, J.; Perrimon, N. Loss-of-function genetic tools for animal models: Cross-species and cross-platform differences. Nat. Rev. Genet. 2017, 18, 24–40. [Google Scholar] [CrossRef] [PubMed]
- Kuo, C.L.; Oyler, G.A.; Shoemaker, C.B. Accelerated neuronal cell recovery from Botulinum neurotoxin intoxication by targeted ubiquitination. PLoS ONE 2011, 6, e20352. [Google Scholar] [CrossRef] [PubMed]
- Caussinus, E.; Affolter, M. deGradFP: A System to Knockdown GFP-Tagged Proteins. Methods Mol. Biol. 2016, 1478, 177–187. [Google Scholar] [CrossRef] [PubMed]
- Caussinus, E.; Kanca, O.; Affolter, M. Fluorescent fusion protein knockout mediated by anti-GFP nanobody. Nat. Struct. Mol. Biol. 2011, 19, 117–121. [Google Scholar] [CrossRef] [PubMed]
- Rothbauer, U.; Zolghadr, K.; Tillib, S.; Nowak, D.; Schermelleh, L.; Gahl, A.; Backmann, N.; Conrath, K.; Muyldermans, S.; Cardoso, M.C.; et al. Targeting and tracing antigens in live cells with fluorescent nanobodies. Nat. Methods 2006, 3, 887–889. [Google Scholar] [CrossRef] [PubMed]
- Ochoa-Espinosa, A.; Harmansa, S.; Caussinus, E.; Affolter, M. Myosin II is not required for Drosophila tracheal branch elongation and cell intercalation. Development 2017, 144, 2961–2968. [Google Scholar] [CrossRef] [PubMed]
- Pasakarnis, L.; Frei, E.; Caussinus, E.; Affolter, M.; Brunner, D. Amnioserosa cell constriction but not epidermal actin cable tension autonomously drives dorsal closure. Nat. Cell Biol. 2016, 18, 1161–1172. [Google Scholar] [CrossRef] [PubMed]
- Córdoba, S.; Estella, C. The transcription factor Dysfusion promotes fold and joint morphogenesis through regulation of Rho1. PLoS Genetics 2018, 14, e1007584. [Google Scholar] [CrossRef] [PubMed]
- Urbano, J.M.; Naylor, H.W.; Scarpa, E.; Muresan, L.; Sanson, B. Suppression of epithelial folding at actomyosin-enriched compartment boundaries downstream of Wingless signalling in Drosophila. Development 2018, 145. [Google Scholar] [CrossRef] [PubMed]
- Lee, P.T.; Lin, G.; Lin, W.W.; Diao, F.; White, B.H.; Bellen, H.J. A kinase-dependent feedforward loop affects CREBB stability and long term memory formation. Elife 2018, 7. [Google Scholar] [CrossRef] [PubMed]
- Batz, T.; Forster, D.; Luschnig, S. The transmembrane protein Macroglobulin complement-related is essential for septate junction formation and epithelial barrier function in Drosophila. Development 2014, 141, 899–908. [Google Scholar] [CrossRef] [PubMed]
- Domsch, K.; Carnesecchi, J.; Disela, V.; Friedrich, J.; Trost, N.; Ermakova, O.; Polychronidou, M.; Lohmann, I. The Hox Transcription Factor Ubx stabilizes Lineage Commitment by Suppressing Cellular Plasticity. bioRxiv 2018. [Google Scholar] [CrossRef]
- Takeda, M.; Sami, M.M.; Wang, Y.C. A homeostatic apical microtubule network shortens cells for epithelial folding via a basal polarity shift. Nat. Cell Biol. 2018, 20, 36–45. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, N.; Colak-Champollion, T.; Knaut, H. zGrad: A nanobody-based degron system to inactivate proteins in zebrafish. bioRxiv 2019, 518944. [Google Scholar] [CrossRef] [PubMed]
- Daniel, K.; Icha, J.; Horenburg, C.; Muller, D.; Norden, C.; Mansfeld, J. Conditional control of fluorescent protein degradation by an auxin-dependent nanobody. Nat. Commun. 2018, 9, 3297. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Tang, N.H.; Lara-Gonzalez, P.; Zhao, Z.; Cheerambathur, D.K.; Prevo, B.; Chisholm, A.D.; Desai, A.; Oegema, K. A toolkit for GFP-mediated tissue-specific protein degradation in C. elegans. Development 2017, 144, 2694–2701. [Google Scholar] [CrossRef] [PubMed]
- Sallee, M.D.; Zonka, J.C.; Skokan, T.D.; Raftrey, B.C.; Feldman, J.L. Tissue-specific degradation of essential centrosome components reveals distinct microtubule populations at microtubule organizing centers. PLoS Biology 2018, 16, e2005189. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.W.; Tang, N.H.; Andrusiak, M.G.; Wu, Z.; Chisholm, A.D.; Jin, Y. A Neuronal piRNA Pathway Inhibits Axon Regeneration in C. elegans. Neuron 2018, 97, 511–519.e6. [Google Scholar] [CrossRef] [PubMed]
- Kurup, N.; Li, Y.; Goncharov, A.; Jin, Y. Intermediate filament accumulation can stabilize microtubules in Caenorhabditis elegans motor neurons. Proc. Natl. Acad. Sci. USA 2018, 115, 3114–3119. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Wu, D.; Quintin, S.; Green, R.A.; Cheerambathur, D.K.; Ochoa, S.D.; Desai, A.; Oegema, K. NOCA-1 functions with gamma-tubulin and in parallel to Patronin to assemble non-centrosomal microtubule arrays in C. elegans. Elife 2015, 4, e08649. [Google Scholar] [CrossRef] [PubMed]
- Shin, Y.J.; Park, S.K.; Jung, Y.J.; Kim, Y.N.; Kim, K.S.; Park, O.K.; Kwon, S.H.; Jeon, S.H.; Trinh le, A.; Fraser, S.E.; et al. Nanobody-targeted E3-ubiquitin ligase complex degrades nuclear proteins. Sci. Rep. 2015, 5, 14269. [Google Scholar] [CrossRef] [PubMed]
- Perez-Mockus, G.; Mazouni, K.; Roca, V.; Corradi, G.; Conte, V.; Schweisguth, F. Spatial regulation of contractility by Neuralized and Bearded during furrow invagination in Drosophila. Nat. Commun. 2017, 8, 1594. [Google Scholar] [CrossRef] [PubMed]
- Urban, E.; Nagarkar-Jaiswal, S.; Lehner, C.F.; Heidmann, S.K. The Cohesin Subunit Rad21 Is Required for Synaptonemal Complex Maintenance, but Not Sister Chromatid Cohesion, during Drosophila Female Meiosis. PLoS Genetics 2014, 10, e1004540. [Google Scholar] [CrossRef] [PubMed]
- Harmansa, S.; Alborelli, I.; Bieli, D.; Caussinus, E.; Affolter, M. A nanobody-based toolset to investigate the role of protein localization and dispersal in Drosophila. Elife 2017, 6. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Vanderzalm, P.J.; Ludwig, M.; Su, T.; Tokamov, S.A.; Fehon, R.G. Yorkie Functions at the Cell Cortex to Promote Myosin Activation in a Non-transcriptional Manner. Dev. Cell 2018, 46, 271–284.e5. [Google Scholar] [CrossRef] [PubMed]
- Ressurreicao, M.; Warrington, S.; Strutt, D. Rapid Disruption of Dishevelled Activity Uncovers an Intercellular Role in Maintenance of Prickle in Core Planar Polarity Protein Complexes. Cell Rep. 2018, 25, 1415–1424.e6. [Google Scholar] [CrossRef] [PubMed]
- Amcheslavsky, A.; Wang, S.; Fogarty, C.E.; Lindblad, J.L.; Fan, Y.; Bergmann, A. Plasma Membrane Localization of Apoptotic Caspases for Non-apoptotic Functions. Dev. Cell 2018, 45, 450–464.e3. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, J.; Peglion, F.; Martin, J.; Hubatsch, L.; Reich, J.; Hirani, N.; Gubieda, A.G.; Roffey, J.; Fernandes, A.R.; St Johnston, D.; et al. aPKC Cycles between Functionally Distinct PAR Protein Assemblies to Drive Cell Polarity. Dev. Cell 2017, 42, 400–415.e9. [Google Scholar] [CrossRef] [PubMed]
- Rogers, K.W.; Schier, A.F. Morphogen Gradients: From Generation to Interpretation. Annu. Rev. Cell Dev. Biol. 2011, 27, 377–407. [Google Scholar] [CrossRef] [PubMed]
- Harmansa, S.; Hamaratoglu, F.; Affolter, M.; Caussinus, E. Dpp spreading is required for medial but not for lateral wing disc growth. Nature 2015, 527, 317–322. [Google Scholar] [CrossRef] [PubMed]
- Pani, A.M.; Goldstein, B. Direct visualization of a native Wnt in vivo reveals that a long-range Wnt gradient forms by extracellular dispersal. Elife 2018, 7. [Google Scholar] [CrossRef] [PubMed]
- Almuedo-Castillo, M.; Blassle, A.; Morsdorf, D.; Marcon, L.; Soh, G.H.; Rogers, K.W.; Schier, A.F.; Muller, P. Scale-invariant patterning by size-dependent inhibition of Nodal signalling. Nat. Cell Biol. 2018, 20, 1032–1042. [Google Scholar] [CrossRef] [PubMed]
- Morsdorf, D.; Muller, P. Tuning protein diffusivity with membrane tethers. Biochemistry 2018. [Google Scholar] [CrossRef] [PubMed]
- Fridy, P.C.; Li, Y.; Keegan, S.; Thompson, M.K.; Nudelman, I.; Scheid, J.F.; Oeffinger, M.; Nussenzweig, M.C.; Fenyo, D.; Chait, B.T.; et al. A robust pipeline for rapid production of versatile nanobody repertoires. Nat. Methods 2014, 11, 1253–1260. [Google Scholar] [CrossRef] [PubMed]
- Ramat, A.; Hannaford, M.; Januschke, J. Maintenance of Miranda Localization in Drosophila Neuroblasts Involves Interaction with the Cognate mRNA. Curr. Biol. 2017, 27, 2101–2111.e5. [Google Scholar] [CrossRef] [PubMed]
- Bertrand, E.; Chartrand, P.; Schaefer, M.; Shenoy, S.M.; Singer, R.H.; Long, R.M. Localization of ASH1 mRNA particles in living yeast. Mol. Cell 1998, 2, 437–445. [Google Scholar] [CrossRef]
- Roubinet, C.; Tsankova, A.; Pham, T.T.; Monnard, A.; Caussinus, E.; Affolter, M.; Cabernard, C. Spatio-temporally separated cortical flows and spindle geometry establish physical asymmetry in fly neural stem cells. Nat. Commun. 2017, 8, 1383. [Google Scholar] [CrossRef] [PubMed]
- Panza, P.; Maier, J.; Schmees, C.; Rothbauer, U.; Sollner, C. Live imaging of endogenous protein dynamics in zebrafish using chromobodies. Development 2015, 142, 1879–1884. [Google Scholar] [CrossRef] [PubMed]
- Burgess, A.; Lorca, T.; Castro, A. Quantitative Live Imaging of Endogenous DNA Replication in Mammalian Cells. PLoS ONE 2012, 7, e45726. [Google Scholar] [CrossRef] [PubMed]
- Rocchetti, A.; Hawes, C.; Kriechbaumer, V. Fluorescent labelling of the actin cytoskeleton in plants using a cameloid antibody. Plant Methods 2014, 10, 12. [Google Scholar] [CrossRef] [PubMed]
- Kamiyama, D.; McGorty, R.; Kamiyama, R.; Kim, M.D.; Chiba, A.; Huang, B. Specification of Dendritogenesis Site in Drosophila aCC Motoneuron by Membrane Enrichment of Pak1 through Dscam1. Dev. Cell 2015, 35, 93–106. [Google Scholar] [CrossRef] [PubMed]
- Bothma, J.P.; Norstad, M.R.; Alamos, S.; Garcia, H.G. LlamaTags: A Versatile Tool to Image Transcription Factor Dynamics in Live Embryos. Cell 2018, 173, 1810–1822.e16. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.C.; Szikra, T.; Kozorovitskiy, Y.; Teixiera, M.; Sabatini, B.L.; Roska, B.; Cepko, C.L. A nanobody-based system using fluorescent proteins as scaffolds for cell-specific gene manipulation. Cell 2013, 154, 928–939. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.C.; Rudolph, S.; Dhande, O.S.; Abraira, V.E.; Choi, S.; Lapan, S.W.; Drew, I.R.; Drokhlyansky, E.; Huberman, A.D.; Regehr, W.G.; et al. Cell type-specific manipulation with GFP-dependent Cre recombinase. Nat. Neurosci. 2015, 18, 1334–1341. [Google Scholar] [CrossRef] [PubMed]
- Morsut, L.; Roybal, K.T.; Xiong, X.; Gordley, R.M.; Coyle, S.M.; Thomson, M.; Lim, W.A. Engineering Customized Cell Sensing and Response Behaviors Using Synthetic Notch Receptors. Cell 2016, 164, 780–791. [Google Scholar] [CrossRef] [PubMed]
- Rothbauer, U.; Zolghadr, K.; Muyldermans, S.; Schepers, A.; Cardoso, M.C.; Leonhardt, H. A versatile nanotrap for biochemical and functional studies with fluorescent fusion proteins. Mol. Cell Proteomics 2008, 7, 282–289. [Google Scholar] [CrossRef] [PubMed]
- Kirchhofer, A.; Helma, J.; Schmidthals, K.; Frauer, C.; Cui, S.; Karcher, A.; Pellis, M.; Muyldermans, S.; Casas-Delucchi, C.S.; Cardoso, M.C.; et al. Modulation of protein properties in living cells using nanobodies. Nat. Struct. Mol. Biol. 2010, 17, 133–138. [Google Scholar] [CrossRef] [PubMed]
- Shimomura, O.; Johnson, F.H.; Saiga, Y. Extraction, Purification and Properties of Aequorin, a Bioluminescent Protein from the Luminous Hydromedusan, Aequorea. J. Cellular Comparat. Physiol. 1962, 59, 223–239. [Google Scholar] [CrossRef]
- Prasher, D.C.; Eckenrode, V.K.; Ward, W.W.; Prendergast, F.G.; Cormier, M.J. Primary structure of the Aequorea victoria green-fluorescent protein. Gene 1992, 111, 229–233. [Google Scholar] [CrossRef]
- Moutel, S.; Bery, N.; Bernard, V.; Keller, L.; Lemesre, E.; de Marco, A.; Ligat, L.; Rain, J.C.; Favre, G.; Olichon, A.; et al. NaLi-H1: A universal synthetic library of humanized nanobodies providing highly functional antibodies and intrabodies. Elife 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Boersma, S.; Khuperkar, D.; Verhagen, B.M.P.; Sonneveld, S.; Grimm, J.B.; Lavis, L.D.; Tanenbaum, M.E. Multi-color single molecule imaging uncovers extensive heterogeneity in mRNA decoding. bioRxiv 2018. [Google Scholar] [CrossRef]
- Braun, M.B.; Traenkle, B.; Koch, P.A.; Emele, F.; Weiss, F.; Poetz, O.; Stehle, T.; Rothbauer, U. Peptides in headlock—A novel high-affinity and versatile peptide-binding nanobody for proteomics and microscopy. Sci. Rep. 2016, 6, 19211. [Google Scholar] [CrossRef] [PubMed]
- Tanenbaum, M.E.; Gilbert, L.A.; Qi, L.S.; Weissman, J.S.; Vale, R.D. A Protein-Tagging System for Signal Amplification in Gene Expression and Fluorescence Imaging. Cell 2014, 159, 635–646. [Google Scholar] [CrossRef] [PubMed]
- Zhao, N.; Kamijo, K.; Fox, P.; Oda, H.; Morisaki, T.; Sato, Y.; Kimura, H.; Stasevich, T.J. A genetically encoded probe for imaging HA-tagged protein translation, localization, and dynamics in living cells and animals. bioRxiv 2018, 474668. [Google Scholar] [CrossRef]
- Tang, J.C.; Drokhlyansky, E.; Etemad, B.; Rudolph, S.; Guo, B.; Wang, S.; Ellis, E.G.; Li, J.Z.; Cepko, C.L. Detection and manipulation of live antigen-expressing cells using conditionally stable nanobodies. Elife 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Blattner, A.C.; Chaurasia, S.; McKee, B.D.; Lehner, C.F. Separase Is Required for Homolog and Sister Disjunction during Drosophila melanogaster Male Meiosis, but Not for Biorientation of Sister Centromeres. PLoS Genet. 2016, 12, e1005996. [Google Scholar] [CrossRef] [PubMed]
- Raychaudhuri, N.; Dubruille, R.; Orsi, G.A.; Bagheri, H.C.; Loppin, B.; Lehner, C.F. Transgenerational Propagation and Quantitative Maintenance of Paternal Centromeres Depends on Cid/Cenp-A Presence in Drosophila Sperm. PLoS Biology 2012, 10, e1001434. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.-H.; Zhang, P.; Kim, H.J.; Mitrea, D.M.; Sarkar, M.; Freibaum, B.D.; Cika, J.; Coughlin, M.; Messing, J.; Molliex, A.; et al. C9orf72 Dipeptide Repeats Impair the Assembly, Dynamics, and Function of Membrane-Less Organelles. Cell 2016, 167, 774–788. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Aguilar, G.; Matsuda, S.; Vigano, M.A.; Affolter, M. Using Nanobodies to Study Protein Function in Developing Organisms. Antibodies 2019, 8, 16. https://doi.org/10.3390/antib8010016
Aguilar G, Matsuda S, Vigano MA, Affolter M. Using Nanobodies to Study Protein Function in Developing Organisms. Antibodies. 2019; 8(1):16. https://doi.org/10.3390/antib8010016
Chicago/Turabian StyleAguilar, Gustavo, Shinya Matsuda, M. Alessandra Vigano, and Markus Affolter. 2019. "Using Nanobodies to Study Protein Function in Developing Organisms" Antibodies 8, no. 1: 16. https://doi.org/10.3390/antib8010016
APA StyleAguilar, G., Matsuda, S., Vigano, M. A., & Affolter, M. (2019). Using Nanobodies to Study Protein Function in Developing Organisms. Antibodies, 8(1), 16. https://doi.org/10.3390/antib8010016