2.1. Selections on Whole Virus Passively Adsorbed to Polystyrene
Four llamas were immunized with mixtures of purified influenza viruses representing all (avian) H and N subtypes, and VHH phage display libraries were generated. The viruses used for immunization were preferably recent isolates from the Netherlands. Here, we only describe the isolation of NA binding VHHs. For the evaluation of selections, we performed parallel phage ELISA and phage display selection. Thus, phages were incubated with immobilized antigen at identical conditions in two wells, one of which was used for phage ELISA and the other for phage elution and transduction of
Escherichia coli. After screening of binding of soluble VHH to antigens in ELISA, we identified 188 unique VHHs binding to the nine different N subtypes. The VHHs were classified according to subtype specificity. For each subtype, they were further classified into different complementarity determining region 3 (CDR3) groups based on their CDR3 sequence. VHHs of the same CDR3 group often contain similar VH coding regions, that belong to the same VHH subfamily [
20]. They presumably arise from the same B-cell recombination process and most likely bind the same antigenic site. For further work, we preferentially selected clones from different CDR3 groups. We will only describe these 64 VHHs.
Initially, we aimed to enrich for NA binding phage by counter-selection of a HA binding phage using two sequential panning rounds on two viruses that have different H subtypes. Such a method was earlier successfully used to prevent enrichment of a phage that specifically binds to contaminating proteins present in antigen preparations [
21]. However, a selection on H5N2 virus and, subsequently, on H6N2 or H9N2 virus yielded clones that bound to H5N2 virus, but not rN2 (
Table 1). Note that this rN2 was derived from the same H5N2 virus later used for successful selection and screening of N2 binding clones (
Table 4). This makes it unlikely that the binders selected are directed against NA. They are more likely directed against the immunologically conserved NP protein [
22] or conserved epitopes of HA, possibly the stem region [
23,
24,
25]. These binders may be more readily selected, because NP and HA are more immunogenic than NA [
7,
9].
Table 1.
Phage display selection of N2 binding llama single-domain antibody fragments (VHHs) by alternating selection on N2 subtype strains. The second selection round was performed using a phage selected on 0.02 µg/mL of H5N2 virus. Viruses were always directly adsorbed to polystyrene plates. Clones were screened for binding to the indicated antigens by incubation with E. coli-produced soluble VHH.
Table 1.
Phage display selection of N2 binding llama single-domain antibody fragments (VHHs) by alternating selection on N2 subtype strains. The second selection round was performed using a phage selected on 0.02 µg/mL of H5N2 virus. Viruses were always directly adsorbed to polystyrene plates. Clones were screened for binding to the indicated antigens by incubation with E. coli-produced soluble VHH.
| | | | No. of positive clones/ No. clones screened |
|---|
| | First round selection | Second round selection | 1 µg/mL H5N2 | 1.5 µg/mL rN2 |
|---|
| Llama | Antigen | A450 nm | Antigen | A450 nm | A450 nm > 1 a | A450 nm > 0.2 a |
|---|
| 3049 | 2 µg/mL H5N2 | 0.827 | | | | |
| 3049 | 0.2 µg/mL H5N2 | 0.851 | | | | |
| 3049 | 0.02 µg/mL H5N2 | 0.775 | 0.1 µg/mL H6N2 | 1.828 | 7/13 | 0/13 |
| | | | 0.01 µg/mL H6N2 | 1.112 | 7/16 | 0/16 |
| | | | Without antigen | 0.074 | | |
| | | | 0.1 µg/mL H9N2 | 1.819 | 6/16 | 0/16 |
| | | | 0.01 µg/mL H9N2 | 1.892 | 6/16 | 0/16 |
| 3049 | Without antigen | 0.051 | | | | |
We next tried to enrich for NA binding phages during panning on low concentrations of immobilized virus by competition with a high concentration of soluble virus of the same H subtype, but a different N subtype. This should result in reduced recovery of the phage displaying HA and NP binding VHHs. Thus, selections on H7N1 were competed with H7N7 virus and
vice versa. As compared to control selections without competition that were performed in parallel, selections with competition resulted in 3–4-fold lower absorbance values in both the first and second selection round, while absorbance values remained above the background without antigen (
Table 2). This suggests that competition was at least partially successful. However, when individual clones were screened for binding of soluble VHH to H7N1 or H7N7 virus, all binders gave comparable absorbance values on both viruses (
Table 2), suggesting binding to H7 or NP. Thus, competition with soluble influenza virus did not stimulate enrichment of NA binding VHHs. These results are consistent with other reports that have indicated that pre-clearing during phage display selection is often of limited usefulness [
21,
26].
Table 2.
Phage display selection of N1 or N7 binding VHHs by competition with other viruses. Phage libraries were first incubated with or without a high concentration of soluble H7N7 or H7N1 virus and, subsequently, selected using low concentrations of immobilized, H7N1 or H7N7 virus, respectively. Clones were screened for binding to immobilized H7N1 and H7N7 virus by incubation with E. coli-produced soluble VHH.
Table 2.
Phage display selection of N1 or N7 binding VHHs by competition with other viruses. Phage libraries were first incubated with or without a high concentration of soluble H7N7 or H7N1 virus and, subsequently, selected using low concentrations of immobilized, H7N1 or H7N7 virus, respectively. Clones were screened for binding to immobilized H7N1 and H7N7 virus by incubation with E. coli-produced soluble VHH.
| | First round selection | Second round selection | No. of clones with A450 nm > 1 / No. clones screened a |
|---|
| | | A450 nm with or without competing virus | | A450 nm with or without competing virus |
|---|
| 5 µg/mL H7N1 | 5 µg/mL H7N7 | H7N1 and H7N7 |
|---|
| Llama | Antigen | without | with | Antigen | without | with |
|---|
| 3049 | 2 µg/mL H7N1 | 0.662 | 0.313 | | | | | | |
| 3049 | 0.2 µg/mL H7N1 | 0.701 | 0.278 | | | | | | |
| 3049 | 0.02 µg/mL | 0.490 | 0.152 | 0.1 µg/mL | 0.426 | 0.306 | 6/16 | 6/16 | 6/16 |
| | H7N1 | | | H7N1 | | | | | |
| | | | | 0.01 µg/mL H7N1 | 0.362 | 0.219 | 6/16 | 5/16 | 5/16 |
| | | | | Without antigen | 0.088 | 0.077 | | | |
| 3049 | 2 µg/mL H7N7 | 0.516 | 0.142 | | | | | | |
| 3049 | 0.2 µg/mL H7N7 | 0.674 | 0.161 | 0.1 µg/mL H7N7 | 0.450 | 0.141 | 7/13 | 6/13 | 6/13 |
| | | | | 0.01 µg/mL H7N7 | 0.294 | 0.091 | 6/16 | 5/16 | 5/16 |
| | | | | Without antigen | 0.099 | 0.082 | | | |
| 3049 | 0.02 µg/mL H7N7 | 0.428 | 0.094 | | | | | | |
| 3049 | Without antigen | 0.052 | 0.057 | | | | | | |
2.2. Selections on Recombinant NA
For use in phage display selection, we prepared recombinant NA (rNA) consisting of the soluble head domain of different N subtypes by expression in HEK293 cells (
Table 3). Some rNA preparations were derived from different virus strains than those used for llama immunization. Expression of rN6 was not successful, but rNA of the further eight subtypes could be produced and purified. The specific enzymatic activity of the different preparations varied considerably. We could not detect enzymatic activity of rN1, rN4 and rN5 (
Table 3), suggesting that these rNAs were produced in a misfolded form, which could be due to either an incorrect tertiary structure or lack of tetramerization.
The eight rNA preparations were used in phage display selections after direct passive adsorption to polystyrene. The three inactive rNA preparations gave the lowest (<0.171) absorbance value in phage ELISA after the first selection round (
Table 3). This could be due to misfolding of these inactive rNA preparations, since most antigenic sites on NA are conformational [
9]. A correlation between enzymatic activity and antigenicity of virus-derived NA was earlier reported [
27]. In the case of rN1 and rN5, but not of rN4, it could also be due to the different viruses that the rNA was derived from and the antigen used for llama immunization. The five enzymatically active rNA preparations gave higher absorbance values in phage ELISA (0.359 to 1.619), suggesting that selections were more efficient. Eventually, we performed two rounds of phage display selection using rNA in round one and whole virus or rNA in the second round. Soluble VHH of individual clones was screened for binding to rNA and viruses with different N subtypes to determine their subtype specificity. Especially after selection on enzymatically inactive rNA, clones that bound whole virus often did not react with rNA. This suggests that these clones are NA specific, but do not bind inactive rNA, because of its presumed misfolding. VHHs against rN1, rN4 and rN5 were, therefore, often not analyzed for binding to rNA in later experiments (
Table 4).
Such selections on rN2, rN3, rN4, rN7, rN8 and rN9 were highly efficient and resulted in diverse panels of binders, representing at least three CDR3 groups that specifically bound NA (
Table 4). Selection on rN1 similarly yielded binders representing three CDR3 groups. However, selection on rN5 yielded only one binder (IV224), possibly due to the misfolding or heterologous nature of rN5. Furthermore, selections on rN6 could not be done, because it was not available.
Table 3.
Recombinant neuraminidase (NA) proteins used.
Table 3.
Recombinant neuraminidase (NA) proteins used.
| NA | Strain origin | Subtype | EMBL accession no. | Amino acids | Specific NA enzyme activity (µmol/mg/min) a | Phage display selection on 1 µg/mL rNA |
|---|
| Llamac | A450 nm |
|---|
| rN1 | A/turkey/Turkey/1/2005 | H5N1 | EF619973 | 55–449 | < 0.08 | 3049 | 0.171 |
| rN2 | A/mallard/Neth/2/2005 | H5N2 | CY041252 | 75–469 | 1.45 | 3049 | 0.474 |
| rN3 | A/ostrich/Neth/03006814/2003 | H2N3 | CY122198 | 75–469 | 4.21 | 3050 | 1.568 |
| rN4 | A/mallard/Neth/06026212-002/2006 | H8N4 | CY043850 | 74–470 | < 0.08 | 3050 | 0.094 |
| rN5 | A/turkey/Neth/07014290/2007 | H1N5 | None | 55–449 | < 0.08 | 3050 | 0.076 |
| rN6 | A/mallard/Norway/10_1334/2006 | H4N6 | FM179763 | 75–470 | ND b | ND | ND |
| rN7 | A/chicken/Neth/03010132/2003 | H7N7 | EF015553 | 74–471 | 0.32 | 3049 | 1.619 |
| rN8 | A/Bewick's swan/Neth/2/2005 | H6N8 | DQ822200 | 73–470 | 0.18 | 3050 | 0.928 |
| rN9 | A/shoveler/Neth/19/1999 | H11N9 | CY005853 | 76–470 | 0.7 | 3050 | 0.359 |
2.3. Selections on Viral NA by Immunocapture with VHHs
The VHHs isolated thus far were used as tools for specific capture of virus NA of particular strains in novel selections. For this purpose, the
E. coli-produced VHHs (suffix Y) were coated onto polystyrene plates and then incubated with virus that was treated with Triton X-100. Such a detergent treatment solubilizes the viral membrane, resulting in the release of the viral surface proteins into solution. Due to the use of a specific VHH as the capture antibody, NA is immobilized on the plate used for phage display selection, whereas the other viral proteins, especially HA and NP, are washed away. This enables to use the same strain for llama immunization and selection. Thus, we used the N5 binder IV224Y for the capture of H1N5 virus NA to select novel N5 binders from two llamas immunized with either H1N5 and H12N5 virus or with H6N5 virus. Similarly, the N1 binder, IV176Y, was used for the capture of H7N1 virus NA to select novel N1 binders (
Table 4).
Some isolated VHHs were found to cross-react with several other NAs. For example, the N7 binder IV237Y cross-reacts with N6 subtype virus (results not shown for IV237Y; however, see
Table 5,
Table 6 for yeast-produced IV237F). IV237Y is a genuine NA binder, since it binds to rN7, although with a low absorbance value (
Table 4). We used IV237Y for the capture of H14N6 or H13N6 virus to select N6 binding VHHs. This resulted in five binders of different CDR3 groups. Similarly, IV457F, resulting from selections on rN4, cross-reacted with other N subtypes with the highest absorbance value on the H14N6 virus (
Table 5) and was therefore classified as an N6 binder. Most clones isolated thus far had significantly higher absorbance values on H14N6 than on H13N6 virus. Yeast-produced N6 binders IV457F and IV461F were therefore used for the capture of H13N6 virus in novel rounds of phage display selection, resulting in five novel binders representing three novel CDR3 groups. Four of these novel binders bound well to H13N6 virus.
Another two clones, IV235 and IV456, within the same CDR3 group (
Table 4), were initially selected using N6 or N7 subtype virus, but were later found to be specific for N3 viruses. IV456F was then used for the capture of solubilized H2N3 virus and selection of a single VHH (IV540) from a llama immunized with H16N3 virus. This VHH bound inefficiently (A
450 nm = 0.098;
Table 4) to directly coated H2N3 virus, but much better (A
450 nm = 0.521) to Triton X-100 treated H2N3 virus captured with IV456F. This suggests that this clone recognizes an antigenic site that is better accessible, due to either surfactant disruption of virions or the capture of the NA in solution with a VHH.
We also selected three novel N9 binding VHHs representing three novel CDR3 groups using solubilized H15N9 virus captured with N9 binder, IV295F.
2.5. NA Specificity of Yeast-Produced VHHs
All 64 VHHs were analyzed in ELISA for binding to influenza viruses representing all N subtypes (
Table 5) to assess their cross-reaction with different N subtypes. A subset of 41 of these VHHs was biotinylated and again analyzed for binding to different influenza viruses in ELISA (
Table 6). The ELISA is much more sensitive when using biotinylated VHH and a streptavidin-peroxidase conjugate than when using non-biotinylated VHH that is detected based on its his6 tag. Some clones, such as IV417F (N1), IV105F (N2) and IV75F (N2), yield high absorbance values when biotinylated, but show no absorbance value above the background using non-biotinylated VHH.
Table 4.
Phage display selection history, antigen binding in ELISA and sequence characteristics of VHH clones selected for yeast expression.
Table 4.
Phage display selection history, antigen binding in ELISA and sequence characteristics of VHH clones selected for yeast expression.
| | | | ELISA signal with influenza viruses b | | Potential | | | | |
|---|
| N | | | | Influenza | Influenza | Homo- | N-glyco- | VHH | | Yeast |
|---|
| Sub- | VHH | | Phage Display Selection a | Virus | Virus | logous | sylation | Sub- | CDR3 | Expres- |
|---|
| type | Clone | Llama | (round 1/ round 2) | Subtype | A450 nm | Subtype | A450 nm | rNA | Sitec | Family d | Sequence | Group e | sion f |
|---|
| 1 | IV169 | 3049 | rN1/rN1 | H1N1 | 0.069 | H7N1 | 0.550 | ND | None | C | ILGGRGIMDPRERQ | A | MIRY |
| 1 | IV403 | 3049 | rN1/rN1 | H1N1 | 0.308 | H7N1 | 0.680 | ND | None | C | RIGGREIMDSRERQ | A | MIRY |
| 1 | IV417 | 3049 | rN1/IV176Y H7N1 | H1N1 | 0.079 | H7N1 | 0.583 | ND | None | 1 | AASSLSGARSRPGDYDY | B | MIRY |
| 1 | IV171 | 3049 | rN1/rN1 | H1N1 | 0.060 | H7N1 | 0.358 | ND | None | 1 | AADRYWNPVSWDY | C | MIRY |
| 1 | IV176 | 3049 | rN1/H1N1 | H1N1 | 0.086 | H7N1 | 0.673 | ND | None | 3 | AASKYGSCNGGTNENDY | D | Not done |
| 1 | IV414 | 3049 | rN1/IV176Y H7N1 | H1N1 | 0.130 | H7N1 | 0.476 | ND | None | 1 | AASERALLRLATSQWDYNY | E | MIRY |
| 1 | IV426 | 3049 | IV176Y H7N1/rN1ST | H1N1 | 0.081 | H7N1 | 0.571 | ND | None | 1 | AASQRSLPLNFLSQYDYDY | F | Episomal |
| 1 | IV422 | 3049 | rN1ST/IV176Y H7N1 | H1N1 | 0.077 | H7N1 | 0.751 | ND | None | 3 | ATDWYCSDSAELSDFTS | G | MIRY |
| 2 | IV63 | 3049 | rN2/rN2 | H5N2 | 0.195 | ND | | 1.904 | None | 1 | AADPNGRGYYY | A | Episomal |
| 2 | IV82 | 3049 | rN2/rN2 | H5N2 | 0.113 | ND | | 1.277 | None | 1 | AADPNGRGYYY | A | Episomal |
| 2 | IV104 | 3049 | rN2/rN2 | H5N2 | 0.180 | ND | | 1.801 | None | 1 | AADPNGRGYYY | A | Episomal |
| 2 | IV65 | 3049 | rN2/rN2 | H5N2 | 0.068 | ND | | 1.019 | None | 1 | AADRSSLQLYASRVGYDY | B | Episomal |
| 2 | IV77 | 3049 | rN2/rN2 | H5N2 | 0.168 | ND | | 1.651 | None | 1 | AASYFWSDTPFALNYRY | C | Episomal |
| 2 | IV105 | 3049 | rN2/rN2 | H5N2 | 0.060 | ND | | 1.91 | None | 1 | AGDRDGRGYVY | D | Episomal |
| 2 | IV75 | 3049 | rN2/rN2 | H5N2 | 0.059 | ND | | 1.826 | None | X | AAAVGSRWYEL | E | Episomal |
| 2 | IV78 | 3049 | rN2/rN2 | H5N2 | 0.088 | ND | | 1.306 | None | 3 | AATPIADASNYCSDYQDFMHY | F | Episomal |
| 2 | IV95 | 3049 | rN2/rN2 | H5N2 | 0.202 | ND | | 1.796 | None | 1 | AAEILYCSGYGCRDPRLYNY | G | Episomal |
| 2 | IV97 | 3049 | rN2/rN2 | H5N2 | 0.120 | ND | | 1.885 | None | 1 | SASTLLPTNDY | H | Episomal |
| 2 | IV107 | 3049 | rN2/rN2 | H5N2 | 0.120 | ND | | 1.759 | None | 1 | AAGFITGIHPYEYHY | I | Episomal |
| 2 | IV81 | 3049 | rN2/rN2 | H5N2 | 0.242 | ND | | 1.842 | 57 | 1 | AASQWDGRTWTRPHAYEY | J | Episomal |
| 2 | IV84 | 3049 | rN2/rN2 | H5N2 | 0.145 | ND | | 1.81 | 57, 61 | 1 | AADLGAYGGLGLSYDY | K | Episomal |
| 3 | IV185 | 3050 | rN3ST/H2N3 | H2N3 | 0.691 | H16N3 | 0.060 | 1.134 | None | 3 | ASIKMYRNSWSGCSALDYYGMDY | A | MIRY |
| 3 | IV191 | 3050 | rN3/rN3 | H2N3 | 0.461 | H16N3 | 0.062 | 1.157 | None | 3 | ASIIIHTRSWSGCSALDYYGMDY | A | MIRY |
| 3 | IV235 | 3049 | rN7/rN7 | H2N3 | 0.054 | H16N3 | 0.052 | ND | None | 2 | NAEEDPLYGGRRGRNY | B | MIRY |
| 3 | IV456 | 3050 | IV297Y H13N6/IV297Y H13N6 | H2N3 | 1.182 | H16N3 | 0.164 | ND | 59 | 2 | NADEDPYYGGRRGRNY | B | MIRY |
| 3 | IV187 | 3050 | rN3ST/H2N3 | H2N3 | 0.529 | H16N3 | 0.055 | 0.638 | None | 2 | AADTYYSRGYDF | C | MIRY |
| 3 | IV202 | 3050 | rN3/H2N3 | H2N3 | 0.554 | H16N3 | 0.061 | 0.382 | 60 | 3 | AADSNVGSMCRDMYADLDV | D | MIRY |
| 3 | IV540 | 3467 | IV456F H2N3/IV237F H2N3 | H2N3 | 0.098 | H16N3 | 0.058 | ND | None | X | KAAWSTL | E | Episomal |
| 4 | IV207 | 3049 | rN4/H8N4 | H8N4 | 0.268 | ND | | 0.135 | None | 2 | NAASFHAMGRDDY | A | MIRY |
| 4 | IV215 | 3049 | rN4/rN4 | H8N4 | 0.333 | ND | | ND | None | 2 | NAASFHAMGRDDY | A | MIRY |
| 4 | IV213 | 3049 | rN4/rN4 | H8N4 | 0.578 | ND | | ND | None | 2 | HAEGYYSDSGRGAWRDY | B | MIRY |
| 4 | IV297 | 3049 | rN4/rN4 | H8N4 | 0.215 | ND | | ND | None | 2 | HAEGYYSDSGRGAWRDY | B | MIRY |
| 4 | IV208 | 3049 | rN4/rN4 | H8N4 | 0.115 | ND | | ND | None | 2 | NAPWRPRGLD | C | MIRY |
| 5 | IV224 | 3050 | rN5/H1N5 | H1N5 | 0.111 | H6N5 | ND | ND | None | C | EKLIDSSGFYSADY | A | MIRY |
| 5 | IV451 | 3466 | IV224Y H1N5/IV224Y H1N5 | H1N5 | 0.222 | H6N5 | 0.115 | ND | None | 1 | AAHRTRYGLGVSAADFGS | B | MIRY |
| 5 | IV442 | 3466 | IV224Y H1N5/IV224Y H1N5 | H1N5 | 0.344 | H6N5 | 0.119 | ND | None | C | ERGSWRSSDW | C | MIRY |
| 5 | IV447 | 3050 | IV224Y H1N5/IV224Y H1N5 | H1N5 | 0.071 | H6N5 | 0.129 | ND | None | C | AKNTVSRLYGPYNE | D | MIRY |
| 6 | IV501 | 3467 | IV457F H13N6/IV457F H13N6 | H13N6 | 0.358 | H14N6 | 0.078 | ND | None | 1 | AARTTMPNEVAPHYGMDY | A | Episomal |
| 6 | IV457 | 3467 | rN4/rN4ST | H13N6 | 0.153 | H14N6 | 0.977 | ND | None | 1 | AVGFEAGYDY | B | MIRY |
| 6 | IV512 | 3467 | IV457F H13N6 / IV457F H13N6 | H13N6 | 0.675 | H14N6 | 0.758 | ND | None | 2 | NFRDYYYSELGEDY | C | Episomal |
| 6 | IV618 | 3467 | IV237Y H14N6/H14N6 | H13N6 | 0.220 | H14N6 | 0.300 | ND | None | 2 | NFRDYYYSELGEDY | C | Episomal |
| 6 | IV461 | 3467 | IV237Y H14N6/IV237Y H14N6 | H13N6 | 0.070 | H14N6 | 0.276 | ND | None | 1 | GVGPYGSRWHHDFGS | D | MIRY |
| 6 | IV462 | 3467 | IV237Y H14N6/IV237Y H14N6 | H13N6 | 0.066 | H14N6 | 0.191 | ND | None | 1 | ALSQYGLAKVENAYDS | E | Episomal |
| 6 | IV467 | 3467 | IV237Y H13N6/IV237Y H13N6 | H13N6 | 0.089 | H14N6 | 0.720 | ND | 20, 93 | 1 | AGKASTYLDYGLYWTANSYTN | F | Episomal |
| 6 | IV513 | 3467 | IV457F 1H13N6/IV457F H13N6 | H13N6 | 0.591 | H14N6 | 0.678 | ND | None | 3 | AAKWTDLSTWWDAGAMDY | G | Episomal |
| 6 | IV514 | 3467 | IV457F H13N6/IV457F H13N6 | H13N6 | 0.449 | H14N6 | 0.560 | ND | None | 1 | AADPGFNSYYSEVGLYDY | H | Episomal |
| 6 | IV619 | 3467 | IV237Y H14N6/H14N6 | H13N6 | 0.174 | H14N6 | 0.245 | ND | None | 1 | AADPGFNSYYNEVGLYDY | H | Episomal |
| 6 | IV516 | 3467 | IV461F 1H13N6/IV461F H13N6 | H13N6 | 0.126 | H14N6 | 0.595 | ND | None | 1 | AADAHAYYSGSYFNHGQLEGAAMDY | I | Episomal |
| 7 | IV246 | 3049 | rN7/H7N7 | H5N7 | 0.655 | H7N7 | 1.115 | 0.09 | None | 1 | ATTQTSGSMLEGYYGF | A | MIRY |
| 7 | IV252 | 3049 | H5N7 cH5N2/rN7 | H5N7 | 0.579 | H7N7 | 1.074 | 0.139 | None | 1 | ATTQSSGAKSEAYYGF | A | MIRY |
| 7 | IV242 | 3049 | rN7/H5N7 | H5N7 | 0.224 | H7N7 | 0.396 | 0.094 | None | 1 | AVHYSGTYSDPNSYGH | B | Episomal |
| 7 | IV237 | 3049 | rN7/rN7 | H5N7 | 0.287 | H7N7 | 0.479 | 0.108 | None | 3 | AADPGFNRGPCSDYDQRIDEYDY | C | MIRY |
| 7 | IV240 | 3049 | rN7/H5N7 | H5N7 | 0.453 | H7N7 | 0.916 | 0.106 | None | 1 | AAGSQGWGLTDAARYAY | D | MIRY |
| 8 | IV265 | 3050 | H10N8/rN8 | H3N8 | 0.812 | ND | | 0.232 | None | 3 | AAALYYSDYRGCSIYYGMDD | A | MIRY |
| 8 | IV280 | 3050 | rN8/rN8 | H3N8 | 0.475 | ND | | 1.042 | None | 2 | YYCAADRDERLWTYVTNMDY | B | MIRY |
| 8 | IV268 | 3050 | H10N8/rN8 | H3N8 | 0.707 | ND | | 0.172 | None | 1 | AAELDHGGAIMGVRRAQQYDY | C | MIRY |
| 8 | IV274 | 3466 | rN8/H10N8 | H3N8 | 0.474 | ND | | 0.183 | None | 2 | NAELVLGYGSAVDSYPMDY | D | MIRY |
| 8 | IV275 | 3467 | rN8/H10N8 | H3N8 | 0.134 | ND | | 0.527 | None | 2 | GLSHYLNWNVA | E | MIRY |
| 8 | IV284 | 3466 | rN8/rN8 | H3N8 | 0.119 | ND | | 0.769 | None | 1 | ASATASTYFSDGYVATAGHEYDS | F | MIRY |
| 9 | IV525 | 3050 | IV295F H15N9/IV295F H15N9 | H5N9g | 0.065 | H5N9h | 0.073 | ND | None | 2 | NAETHYRIDGDYDF | A | Episomal |
| 9 | IV295 | 3050 | rN9/H5N9h | H5N9g | ND | H5N9h | 0.204 | 0.144 | None | 2 | RADGVVGGY | B | MIRY |
| 9 | IV294 | 3050 | rN9/rN9 | H5N9g | ND | H5N9h | 0.132 | 0.51 | None | 2 | NFQGGSSVLDRATFDY | C | MIRY |
| 9 | IV300 | 3050 | H5N9g cH5N7/rN9 | H5N9g | ND | H5N9h | 0.474 | 0.108 | None | 2 | NADIWAIATMRAGSYDY | D | MIRY |
| 9 | IV530 | 3050 | IV295F H5N9g/IV295F H15N9 | H5N9g | 0.128 | H5N9h | 0.446 | ND | None | 3 | AAGLGSLDRLLYDY | E | Episomal |
| 9 | IV533 | 3050 | IV295F H5N9g/IV295F H15N9 | H5N9g | 0.089 | H5N9h | 0.140 | ND | None | 2 | NALLEYRRWGEYEY | F | Episomal |
Table 5.
Inhibition of rNA activity and binding in ELISA to influenza viruses of different subtypes of 64 yeast-produced VHHs.
Table 5.
Inhibition of rNA activity and binding in ELISA to influenza viruses of different subtypes of 64 yeast-produced VHHs.
| N Sub- type | VHH Clone | rNA enzyme inhibitiona | ELISA signal on influenza virus b |
|---|
| (IC50 in ng/mL) | H1 | H7 | H6 | H9 | H2 | H8 | H1 | H6 | H12 | H11 | H13 | H14 | H5 | H7 | H3 | H4 | H10 | H5 | H5 | H15 |
|---|
| N1 | N1 | N2 | N2 | N3 | N4 | N5 | N5 | N5 | N6 | N6 | N6 | N7 | N7 | N8 | N8 | N8 | N9c | N9d | N9 |
|---|
| 1 | IV169F | ND | 0.06 | 0.60 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.06 | 0.06 | 0.06 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 |
| 1 | IV403F | ND | 0.05 | 0.72 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.08 | 0.05 | 0.06 | 0.05 | 0.09 | 0.05 | 0.05 | 0.13 | 0.06 |
| 1 | IV417F | ND | 0.05 | 0.07 | 0.04 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.06 | 0.05 | 0.12 | 0.11 | 0.06 | 0.07 | 0.06 | 0.12 | 0.06 | 0.06 | 0.10 | 0.06 |
| 1 | IV171F | ND | 0.06 | 0.40 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.09 | 0.10 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 1 | IV414F | ND | 0.04 | 0.05 | 0.04 | 0.05 | 0.18 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.10 | 0.05 | 0.06 | 0.05 | 0.09 | 0.05 | 0.05 | 0.17 | 0.06 |
| 1 | IV426F | ND | 0.05 | 0.15 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.06 |
| 1 | IV422F | ND | 0.06 | 1.02 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.07 | 0.06 | 0.07 | 0.05 | 0.08 | 0.05 | 0.05 | 0.06 | 0.06 |
| 2 | IV63F | 15 | 0.06 | 0.06 | 0.13 | 0.12 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.07 | 0.06 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 2 | IV82F | 30 | 0.06 | 0.05 | 0.07 | 0.12 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.06 |
| 2 | IV104F | 51 | 0.05 | 0.05 | 0.04 | 0.07 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.06 | 0.05 | 0.05 | 0.06 | 0.06 |
| 2 | IV65F | 33 | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 2 | IV77F | > 1,000 | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.07 | 0.05 | 0.05 | 0.05 | 0.05 |
| 2 | IV105F | 63 | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.09 | 0.05 | 0.06 | 0.05 | 0.08 | 0.05 | 0.05 | 0.06 | 0.06 |
| 2 | IV75F | 73 | 0.06 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 2 | IV78F | 54 | 0.06 | 0.05 | 0.85 | 0.67 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.71 | 0.49 | 0.96 | 0.05 | 0.06 | 0.05 | 0.08 | 0.05 | 0.43 | 0.47 | 0.63 |
| 2 | IV95F | 55 | 0.05 | 0.05 | 0.10 | 0.07 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 2 | IV97F | 71 | 0.04 | 0.05 | 0.05 | 0.09 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.04 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 2 | IV107F | 63 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.11 | 0.06 | 0.07 | 0.05 | 0.09 | 0.06 | 0.06 | 0.07 | 0.06 |
| 2 | IV81Fe | 76 | 0.05 | 0.05 | 0.06 | 0.06 | 0.05 | 0.06 | 0.06 | 0.05 | 0.06 | 0.06 | 0.06 | 0.11 | 0.05 | 0.06 | 0.06 | 0.11 | 0.05 | 0.05 | 0.05 | 0.06 |
| 2 | IV84F | 40 | 0.06 | 0.05 | 0.56 | 0.25 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.84 | 0.86 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 3 | IV185F | 8 | 0.05 | 0.07 | 0.05 | 0.05 | 1.21 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 3 | IV191F | 5 | 0.05 | 0.05 | 0.04 | 0.05 | 0.94 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 3 | IV235F | > 1,000 | 0.05 | 0.05 | 0.04 | 0.05 | 0.79 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.07 | 0.05 | 0.06 | 0.05 | 0.06 | 0.05 | 0.05 | 0.07 | 0.06 |
| 3 | IV456Fe | > 1,000 | 0.06 | 0.05 | 0.05 | 0.05 | 1.39 | 0.05 | 0.06 | 0.05 | 0.07 | 0.06 | 0.05 | 0.08 | 0.06 | 0.06 | 0.05 | 0.07 | 0.06 | 0.06 | 0.06 | 0.06 |
| 3 | IV187F | 2 | 0.05 | 0.05 | 0.05 | 0.05 | 0.24 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 3 | IV202Fe | 13 | 0.06 | 0.06 | 0.05 | 0.06 | 0.19 | 0.05 | 0.10 | 0.06 | 0.06 | 0.06 | 0.06 | 0.10 | 0.06 | 0.07 | 0.05 | 0.09 | 0.06 | 0.06 | 0.06 | 0.06 |
| 3 | IV540F | ND | 0.04 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.06 | 0.05 | 0.05 | 0.06 | 0.05 | 0.06 | 0.07 | 0.05 | 0.05 | 0.07 |
| 4 | IV207F | ND | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.23 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.09 | 0.05 | 0.07 | 0.05 | 0.08 | 0.05 | 0.05 | 0.07 | 0.06 |
| 4 | IV215F | ND | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.31 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 4 | IV213F | ND | 0.07 | 0.10 | 0.05 | 0.05 | 0.05 | 0.43 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.07 | 0.05 | 0.07 | 0.05 | 0.06 | 0.07 | 0.08 |
| 4 | IV297F | ND | 0.05 | 0.10 | 0.04 | 0.05 | 0.05 | 0.64 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.05 | 0.06 | 0.05 | 0.06 | 0.07 | 0.07 |
| 4 | IV208F | ND | 0.09 | 0.05 | 0.05 | 0.05 | 0.05 | 0.20 | 0.06 | 0.05 | 0.06 | 0.05 | 0.05 | 0.06 | 0.06 | 0.07 | 0.05 | 0.07 | 0.06 | 0.05 | 0.05 | 0.06 |
| 5 | IV224F | ND | 0.04 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.09 | 0.08 | 0.12 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 5 | IV451F | ND | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.21 | 0.14 | 0.20 | 0.05 | 0.05 | 0.09 | 0.07 | 0.06 | 0.05 | 0.06 | 0.05 | 0.05 | 0.13 | 0.06 |
| 5 | IV442F | ND | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.39 | 0.16 | 0.59 | 0.05 | 0.05 | 0.05 | 0.06 | 0.07 | 0.05 | 0.06 | 0.05 | 0.06 | 0.05 | 0.06 |
| 5 | IV447F | ND | 0.04 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.48 | 0.22 | 0.79 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 6 | IV501F | ND | 0.04 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.67 | 0.10 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 |
| 6 | IV457F | ND | 0.06 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.07 | 0.05 | 0.06 | 1.60 | 1.14 | 1.57 | 0.20 | 0.21 | 0.05 | 0.10 | 0.06 | 0.35 | 0.52 | 0.84 |
| 6 | IV512F | ND | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 1.58 | 1.13 | 1.51 | 0.05 | 0.06 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.06 |
| 6 | IV618F | ND | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 1.20 | 0.55 | 0.98 | 0.06 | 0.06 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.06 |
| 6 | IV461F | ND | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.07 | 0.08 | 0.13 | 0.06 | 0.07 | 0.05 | 0.08 | 0.06 | 0.06 | 0.06 | 0.06 |
| 6 | IV462F | ND | 0.05 | 0.07 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.59 | 0.05 | 0.27 | 0.06 | 0.06 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.06 |
| 6 | IV467Fe | ND | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.07 | 0.67 | 0.86 | 1.38 | 0.06 | 0.06 | 0.05 | 0.06 | 0.06 | 0.05 | 0.05 | 0.06 |
| 6 | IV513F | ND | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 1.39 | 0.99 | 1.46 | 0.07 | 0.07 | 0.05 | 0.07 | 0.05 | 0.07 | 0.08 | 0.11 |
| 6 | IV514F | ND | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.06 | 1.08 | 0.74 | 1.20 | 0.06 | 0.06 | 0.05 | 0.10 | 0.05 | 0.05 | 0.07 | 0.06 |
| 6 | IV619F | ND | 0.05 | 0.06 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.92 | 0.64 | 1.08 | 0.05 | 0.06 | 0.05 | 0.09 | 0.05 | 0.05 | 0.10 | 0.06 |
| 6 | IV516F | ND | 0.06 | 0.05 | 0.04 | 0.05 | 0.05 | 0.06 | 0.08 | 0.06 | 0.06 | 0.11 | 0.20 | 1.26 | 0.06 | 0.07 | 0.05 | 0.09 | 0.06 | 0.06 | 0.06 | 0.06 |
| 7 | IV246F | 6 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 1.09 | 1.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.05 | 0.06 |
| 7 | IV252F | 6 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.05 | 0.06 | 1.14 | 0.99 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.06 |
| 7 | IV242F | 56 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.06 | 0.22 | 0.15 | 0.05 | 0.06 | 0.06 | 0.05 | 0.05 | 0.06 |
| 7 | IV237F | 22 | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.13 | 0.09 | 0.17 | 0.17 | 0.17 | 0.05 | 0.08 | 0.05 | 0.05 | 0.09 | 0.06 |
| 7 | IV240F | 33 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.06 | 0.05 | 0.05 | 0.09 | 0.20 | 0.20 | 0.05 | 0.09 | 0.05 | 0.05 | 0.07 | 0.06 |
| 8 | IV265F | 18 | 0.04 | 0.05 | 0.04 | 0.05 | 0.07 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.13 | 0.33 | 0.10 | 0.05 | 0.05 | 0.06 |
| 8 | IV280F | 201 | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.07 | 0.05 | 0.08 | 0.10 | 0.05 | 0.07 | 0.19 | 1.44 | 0.14 | 0.06 | 0.15 | 0.06 |
| 8 | IV268F | > 1000 | 0.04 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.09 | 1.04 | 0.07 | 0.05 | 0.08 | 0.06 |
| 8 | IV274F | 124 | 0.05 | 0.92 | 0.04 | 0.05 | 0.05 | 0.69 | 0.40 | 0.14 | 0.53 | 0.05 | 0.05 | 0.08 | 0.05 | 0.06 | 0.18 | 1.64 | 0.18 | 0.05 | 0.09 | 0.06 |
| 8 | IV275F | > 1000 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.06 | 0.12 | 0.06 | 0.06 | 0.08 | 0.70 | 0.06 | 0.06 | 0.12 | 0.06 |
| 8 | IV284F | 8 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.07 | 0.06 | 0.07 | 0.06 | 0.13 | 0.06 | 0.05 | 0.05 | 0.06 |
| 9 | IV525F | > 1000 | 0.06 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.06 | 0.06 | 0.06 | 0.05 | 0.06 | 0.07 | 0.06 | 0.06 | 0.06 | 0.06 | 0.05 | 1.24 |
| 9 | IV295F | > 1000 | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.05 | 0.05 | 0.06 | 0.35 | 0.08 | 1.23 |
| 9 | IV294F | > 1000 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.11 | 0.05 | 0.05 | 0.06 | 0.06 | 0.07 | 0.11 | 0.06 | 0.07 | 0.05 | 0.08 | 0.06 | 0.07 | 0.09 | 0.10 |
| 9 | IV300F | 100 | 0.10 | 0.05 | 0.16 | 0.08 | 0.16 | 0.05 | 0.05 | 0.05 | 0.06 | 0.07 | 0.06 | 0.08 | 0.05 | 0.06 | 0.05 | 0.06 | 0.05 | 0.11 | 0.31 | 0.33 |
| 9 | IV533F | 7 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.05 | 0.05 | 0.05 | 0.24 | 0.47 | 0.70 |
| None | >1000 | 0.05 | 0.05 | 0.04 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.05 | 0.05 | 0.05 | 0.05 | 0.06 | 0.05 | 0.05 | 0.05 | 0.06 | 0.06 | 0.08 |
Figure 1.
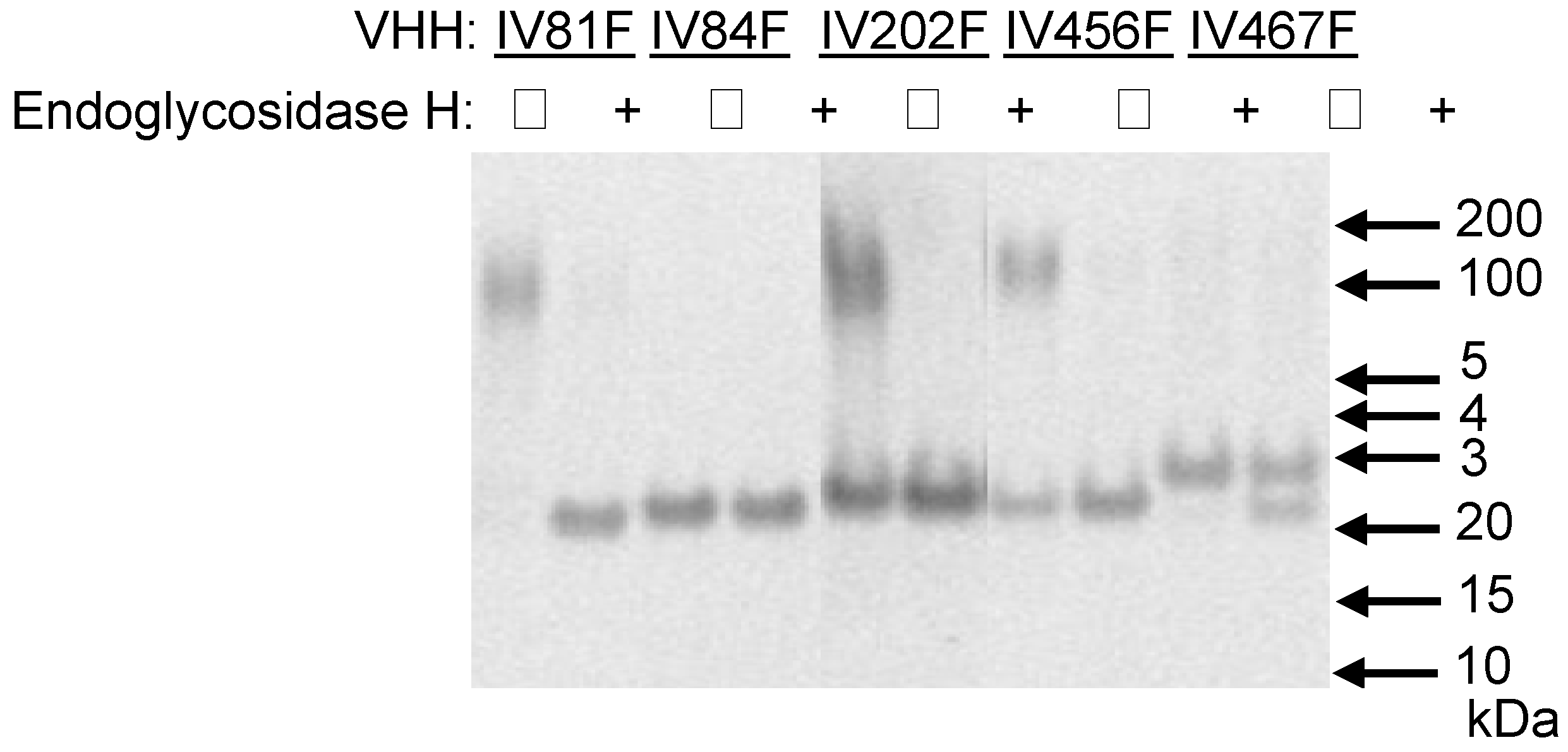
Analysis of N-glycosylation of yeast-produced VHHs. Five VHHs containing potential N-glycosylation sites were deglycosylated by endoglycosidase H treatment (+) or mock-incubated (−) prior to analysis by reducing SDS-PAGE. The position of relevant molecular mass markers (BenchmarkTM protein ladder, Invitrogen) is indicated on the right.
Figure 1.
Analysis of N-glycosylation of yeast-produced VHHs. Five VHHs containing potential N-glycosylation sites were deglycosylated by endoglycosidase H treatment (+) or mock-incubated (−) prior to analysis by reducing SDS-PAGE. The position of relevant molecular mass markers (BenchmarkTM protein ladder, Invitrogen) is indicated on the right.
Table 6.
Binding in ELISA of yeast-produced and biotinylated VHHs to influenza viruses of different subtypes.
Table 6.
Binding in ELISA of yeast-produced and biotinylated VHHs to influenza viruses of different subtypes.
| | ELISA signal on influenza viruses a |
|---|
| N-Subtype | VHH Clone | H1 | H7 | H6 | H9 | H2 | H8 | H1 | H6 | H12 | H11 | H13 | H14 | H5 | H7 | H3 | H4 | H10 | H5 | H5 | H15 | None |
|---|
| N1 | N1 | N2 | N2 | N3 | N4 | N5 | N5 | N5 | N6 | N6 | N6 | N7 | N7 | N8 | N8 | N8 | N9b | N9c | N9 |
|---|
| 1 | IV169F | 0.16 | 1.53 | 0.13 | 0.24 | 0.07 | 0.11 | 0.08 | 0.25 | 0.23 | 0.12 | 0.20 | 0.14 | 0.28 | 0.19 | 0.11 | 0.11 | 0.15 | 0.12 | 0.21 | 0.22 | 0.06 |
| 1 | IV403F | 0.08 | 1.43 | 0.07 | 0.12 | 0.46 | 0.06 | 0.07 | 0.12 | 0.13 | 0.11 | 0.13 | 0.09 | 0.08 | 0.10 | 0.11 | 0.08 | 0.07 | 0.13 | 0.12 | 0.07 | 0.05 |
| 1 | IV417F | 0.07 | 1.36 | 0.07 | 0.10 | 0.07 | 0.06 | 0.07 | 0.10 | 0.12 | 0.10 | 0.11 | 0.07 | 0.06 | 0.09 | 0.06 | 0.14 | 0.06 | 0.07 | 0.12 | 0.07 | 0.05 |
| 1 | IV171F | 0.08 | 1.78 | 0.07 | 0.14 | 0.07 | 0.07 | 0.08 | 0.12 | 0.13 | 0.11 | 0.14 | 0.09 | 0.46 | 0.33 | 0.07 | 0.07 | 0.07 | 0.07 | 0.08 | 0.07 | 0.05 |
| 1 | IV422F | 0.74 | 1.70 | 0.08 | 0.13 | 0.08 | 0.84 | 0.08 | 0.13 | 0.12 | 0.12 | 0.13 | 0.09 | 0.08 | 0.11 | 0.10 | 0.15 | 0.08 | 0.08 | 0.09 | 0.07 | 0.05 |
| 2 | IV82F | 0.07 | 0.07 | 0.54 | 1.32 | 0.06 | 0.06 | 0.09 | 0.10 | 0.12 | 0.09 | 0.12 | 0.07 | 0.06 | 0.07 | 0.07 | 0.06 | 0.15 | 0.07 | 0.07 | 0.06 | 0.05 |
| 2 | IV105F | 0.08 | 0.07 | 0.15 | 0.84 | 0.07 | 0.07 | 0.07 | 0.13 | 0.14 | 0.09 | 0.13 | 0.08 | 0.06 | 0.07 | 0.07 | 0.08 | 0.08 | 0.06 | 0.08 | 0.07 | 0.06 |
| 2 | IV75F | 0.10 | 0.09 | 0.09 | 1.03 | 0.08 | 0.07 | 0.10 | 0.15 | 0.14 | 0.11 | 0.15 | 0.11 | 0.08 | 0.08 | 0.08 | 0.07 | 0.09 | 0.07 | 0.10 | 0.08 | 0.06 |
| 2 | IV97F | 0.12 | 0.08 | 0.42 | 1.16 | 0.07 | 0.08 | 0.08 | 0.13 | 0.12 | 0.10 | 0.13 | 0.09 | 0.08 | 0.08 | 0.07 | 0.07 | 0.07 | 0.06 | 0.10 | 0.07 | 0.07 |
| 2 | IV81Fd | 0.08 | 0.08 | 1.29 | 1.26 | 0.07 | 0.06 | 0.08 | 0.11 | 0.13 | 0.11 | 0.15 | 0.10 | 0.07 | 0.09 | 0.12 | 0.09 | 0.07 | 0.07 | 0.08 | 0.07 | 0.05 |
| 2 | IV84F | 0.07 | 0.07 | 1.42 | 1.24 | 0.07 | 0.06 | 0.07 | 0.10 | 0.11 | 0.10 | 0.12 | 0.08 | 1.50 | 1.25 | 0.09 | 0.07 | 0.11 | 0.07 | 0.07 | 0.06 | 0.05 |
| 3 | IV191F | 0.07 | 0.08 | 0.08 | 0.13 | 1.58 | 0.06 | 0.09 | 0.11 | 0.11 | 0.10 | 0.12 | 1.21 | 0.65 | 0.14 | 0.07 | 0.07 | 0.13 | 0.06 | 0.09 | 0.07 | 0.05 |
| 3 | IV235F | 0.08 | 0.08 | 0.07 | 0.12 | 1.83 | 0.06 | 0.08 | 0.11 | 0.12 | 0.11 | 0.12 | 0.08 | 0.06 | 0.08 | 0.07 | 0.09 | 0.06 | 0.07 | 0.09 | 0.07 | 0.05 |
| 3 | IV456Fd | 0.14 | 0.11 | 0.12 | 0.14 | 1.82 | 0.07 | 0.12 | 0.16 | 0.15 | 0.13 | 0.21 | 0.18 | 0.10 | 0.11 | 0.18 | 0.10 | 0.13 | 0.10 | 0.10 | 0.09 | 0.06 |
| 3 | IV187F | 0.09 | 0.12 | 0.33 | 0.13 | 1.42 | 0.06 | 0.09 | 0.12 | 0.14 | 0.11 | 0.14 | 0.10 | 0.09 | 0.08 | 0.09 | 0.06 | 0.07 | 0.11 | 0.14 | 0.07 | 0.05 |
| 3 | IV202Fd | 0.11 | 1.42 | 0.09 | 0.14 | 1.57 | 0.06 | 0.11 | 0.15 | 0.15 | 0.15 | 0.19 | 0.14 | 0.09 | 0.10 | 0.16 | 0.54 | 0.09 | 0.08 | 0.10 | 0.08 | 0.06 |
| 3 | IV540F | 0.07 | 0.09 | 0.07 | 0.11 | 0.08 | 0.06 | 0.08 | 0.12 | 0.13 | 0.11 | 0.15 | 0.10 | 0.07 | 0.09 | 0.12 | 0.11 | 0.08 | 0.07 | 0.09 | 0.07 | 0.05 |
| 4 | IV215F | 0.08 | 0.07 | 0.06 | 0.12 | 0.06 | 1.47 | 0.07 | 0.12 | 0.12 | 0.10 | 0.13 | 0.07 | 0.07 | 0.07 | 0.07 | 0.11 | 0.08 | 0.06 | 0.08 | 0.07 | 0.06 |
| 4 | IV213F | 1.21 | 1.17 | 0.56 | 0.17 | 0.29 | 1.43 | 0.08 | 0.10 | 0.12 | 0.09 | 0.12 | 0.07 | 0.10 | 0.09 | 0.08 | 1.12 | 0.07 | 0.47 | 0.90 | 0.97 | 0.06 |
| 4 | IV208F | 0.96 | 0.08 | 0.08 | 0.12 | 0.10 | 1.21 | 0.67 | 0.10 | 0.12 | 0.10 | 0.13 | 0.09 | 0.10 | 0.07 | 0.06 | 0.06 | 0.06 | 0.07 | 0.09 | 0.07 | 0.05 |
| 5 | IV224F | 0.07 | 0.08 | 0.07 | 0.14 | 0.07 | 0.06 | 1.31 | 1.18 | 1.59 | 0.10 | 0.13 | 0.08 | 0.07 | 0.07 | 0.07 | 0.06 | 0.06 | 0.07 | 0.09 | 0.06 | 0.05 |
| 5 | IV451F | 0.07 | 0.07 | 0.07 | 0.12 | 0.08 | 0.06 | 1.29 | 1.35 | 1.55 | 0.10 | 0.12 | 0.07 | 0.07 | 0.07 | 0.06 | 0.06 | 0.06 | 0.07 | 0.08 | 0.07 | 0.05 |
| 6 | IV501F | 0.07 | 0.12 | 0.07 | 0.11 | 0.06 | 0.06 | 0.07 | 0.10 | 0.12 | 0.10 | 1.44 | 1.50 | 0.06 | 0.14 | 0.07 | 0.09 | 0.06 | 0.08 | 0.08 | 0.07 | 0.05 |
| 6 | IV512F | 0.80 | 0.13 | 0.60 | 0.35 | 0.84 | 0.06 | 0.40 | 0.19 | 0.30 | 1.49 | 1.42 | 1.41 | 0.36 | 0.35 | 0.08 | 0.65 | 0.07 | 0.08 | 0.08 | 0.07 | 0.05 |
| 6 | IV618F | 0.61 | 0.12 | 0.65 | 0.34 | 0.74 | 0.06 | 0.34 | 0.13 | 0.26 | 1.38 | 1.40 | 1.37 | 0.68 | 0.58 | 0.10 | 0.39 | 0.08 | 0.08 | 0.09 | 0.08 | 0.06 |
| 6 | IV461F | 0.07 | 0.07 | 0.08 | 0.11 | 0.07 | 0.06 | 0.09 | 0.10 | 0.13 | 1.11 | 1.19 | 1.34 | 0.06 | 0.07 | 0.12 | 0.07 | 0.09 | 0.07 | 0.08 | 0.07 | 0.06 |
| 6 | IV467Fd | 0.08 | 0.12 | 0.08 | 0.14 | 0.07 | 0.07 | 0.13 | 0.13 | 0.15 | 1.38 | 1.53 | 1.58 | 0.24 | 0.18 | 0.09 | 0.08 | 0.10 | 0.06 | 0.12 | 0.09 | 0.06 |
| 6 | IV513F | 0.08 | 0.07 | 0.07 | 0.12 | 0.06 | 0.06 | 0.08 | 0.10 | 0.12 | 1.41 | 1.32 | 1.37 | 1.24 | 1.11 | 0.07 | 0.13 | 0.06 | 0.77 | 1.14 | 1.25 | 0.05 |
| 6 | IV514F | 0.08 | 0.08 | 0.07 | 0.12 | 0.06 | 0.06 | 0.07 | 0.10 | 0.13 | 1.50 | 1.42 | 1.55 | 0.49 | 0.43 | 0.08 | 0.07 | 0.07 | 0.08 | 0.08 | 0.07 | 0.06 |
| 6 | IV619F | 0.07 | 0.08 | 0.07 | 0.12 | 0.07 | 0.06 | 0.07 | 0.09 | 0.15 | 1.42 | 1.38 | 1.45 | 0.08 | 0.12 | 0.12 | 0.15 | 0.08 | 0.07 | 0.07 | 0.06 | 0.05 |
| 6 | IV516F | 0.09 | 0.08 | 0.07 | 0.12 | 0.07 | 0.10 | 0.08 | 0.11 | 0.13 | 1.25 | 1.27 | 1.44 | 0.07 | 0.08 | 0.07 | 0.07 | 0.07 | 0.08 | 0.09 | 0.08 | 0.05 |
| 7 | IV242F | 0.08 | 0.08 | 0.07 | 0.26 | 0.07 | 0.07 | 0.08 | 0.11 | 0.12 | 0.11 | 0.13 | 0.09 | 1.68 | 1.57 | 0.07 | 0.08 | 0.07 | 0.08 | 0.09 | 0.07 | 0.05 |
| 7 | IV237F | 0.08 | 0.07 | 0.07 | 0.12 | 0.08 | 0.06 | 0.09 | 0.12 | 0.13 | 1.55 | 1.56 | 1.60 | 1.65 | 1.60 | 0.09 | 0.13 | 0.08 | 0.07 | 0.09 | 0.07 | 0.05 |
| 7 | IV240F | 0.07 | 0.08 | 0.07 | 0.12 | 0.07 | 0.06 | 0.07 | 0.10 | 0.12 | 0.12 | 0.14 | 0.11 | 1.53 | 1.42 | 0.08 | 0.11 | 0.06 | 0.09 | 0.09 | 0.08 | 0.05 |
| 8 | IV268F | 0.94 | 1.14 | 0.27 | 0.14 | 0.15 | 1.45 | 0.07 | 0.09 | 0.11 | 0.09 | 0.12 | 0.06 | 0.08 | 0.08 | 0.82 | 1.76 | 0.68 | 0.38 | 0.79 | 0.90 | 0.05 |
| 8 | IV275F | 0.09 | 0.09 | 0.07 | 0.14 | 0.07 | 0.07 | 0.09 | 0.13 | 0.13 | 0.15 | 0.17 | 0.15 | 0.08 | 0.08 | 0.64 | 1.61 | 0.55 | 0.14 | 0.11 | 0.31 | 0.06 |
| 8 | IV284F | 0.07 | 0.07 | 0.07 | 0.10 | 0.07 | 0.06 | 0.09 | 0.12 | 0.13 | 0.09 | 0.13 | 0.09 | 0.06 | 0.07 | 0.54 | 1.50 | 0.49 | 0.07 | 0.08 | 0.07 | 0.06 |
| 9 | IV525F | 0.07 | 0.07 | 0.07 | 0.12 | 0.07 | 0.06 | 0.07 | 0.10 | 0.13 | 0.10 | 0.12 | 0.10 | 0.07 | 0.07 | 0.16 | 0.67 | 0.16 | 0.07 | 0.08 | 1.55 | 0.06 |
| 9 | IV295F | 0.08 | 0.07 | 0.07 | 0.13 | 0.07 | 0.06 | 0.08 | 0.11 | 0.13 | 0.10 | 0.13 | 0.10 | 0.06 | 0.08 | 0.09 | 0.06 | 0.06 | 1.23 | 1.40 | 1.61 | 0.05 |
| 9 | IV530F | 0.08 | 0.10 | 0.09 | 0.12 | 0.08 | 0.06 | 0.10 | 0.12 | 0.15 | 0.12 | 0.23 | 0.15 | 0.07 | 0.08 | 0.21 | 0.08 | 0.14 | 1.19 | 1.41 | 1.55 | 0.06 |
| 9 | IV533F | 0.07 | 0.09 | 0.08 | 0.13 | 0.13 | 0.06 | 0.07 | 0.10 | 0.14 | 0.15 | 0.16 | 0.09 | 0.07 | 0.08 | 0.13 | 0.09 | 0.09 | 0.59 | 0.88 | 1.22 | 0.06 |
| | None | 0.06 | 0.07 | 0.06 | 0.10 | 0.06 | 0.06 | 0.06 | 0.10 | 0.11 | 0.08 | 0.11 | 0.06 | 0.05 | 0.06 | 0.06 | 0.05 | 0.06 | 0.06 | 0.06 | 0.06 | 0.05 |
Most VHHs bound specifically to only one N-subtype virus. Some clones, such as IV417F (N1) and IV501F (N6), even bound specifically to particular virus strains of this subtype. Such strain-specific binding is not unexpected, since HA and NA proteins of influenza virus rapidly mutate, and under selective pressure from the host immune response, novel antigenic variants are constantly selected, a process known as antigenic drift [
9]. Other VHHs, however, cross-react to influenza viruses of a different N subtype. The VHHs showing the strongest cross-reaction are summarized in
Table 7.
Table 7.
Summary of VHHs that bind to at least two different NA subtypes in ELISA.
Table 7.
Summary of VHHs that bind to at least two different NA subtypes in ELISA.
| N Sub-type | | VHH binding in ELISA to several NAs |
|---|
| VHH Clone | N1 | N2 | N3 | N4 | N5 | N6 | N7 | N8 | N9 |
|---|
| 1 | IV422F | + a | − | − | + | − | − | − | − | − |
| 4 | IV208F | + | − | − | + | + | − | − | − | − |
| 4 | IV213F | + | + | − | + | − | − | − | + | + |
| 8 | IV274F | + | − | − | + | + | − | − | + | − |
| 8 | IV268F | + | − | − | + | − | − | − | + | + |
| 9 | IV525F | − | − | − | − | − | − | − | + | + |
| 2 | IV78F | − | + | − | − | − | + | − | − | + |
| 2 | IV84F | − | + | − | − | − | − | + | − | − |
| 3 | IV202F | + | − | + | − | − | − | − | + | − |
| 3 | IV191F | − | − | + | − | − | + | + | − | − |
| 7 | IV237F | − | − | − | − | − | + | + | − | − |
| 6 | IV513F | − | − | − | − | − | + | + | − | + |
| 6 | IV512F | + | + | + | − | − | + | − | + | − |
| 6 | IV618F | + | + | + | − | − | + | + | − | − |
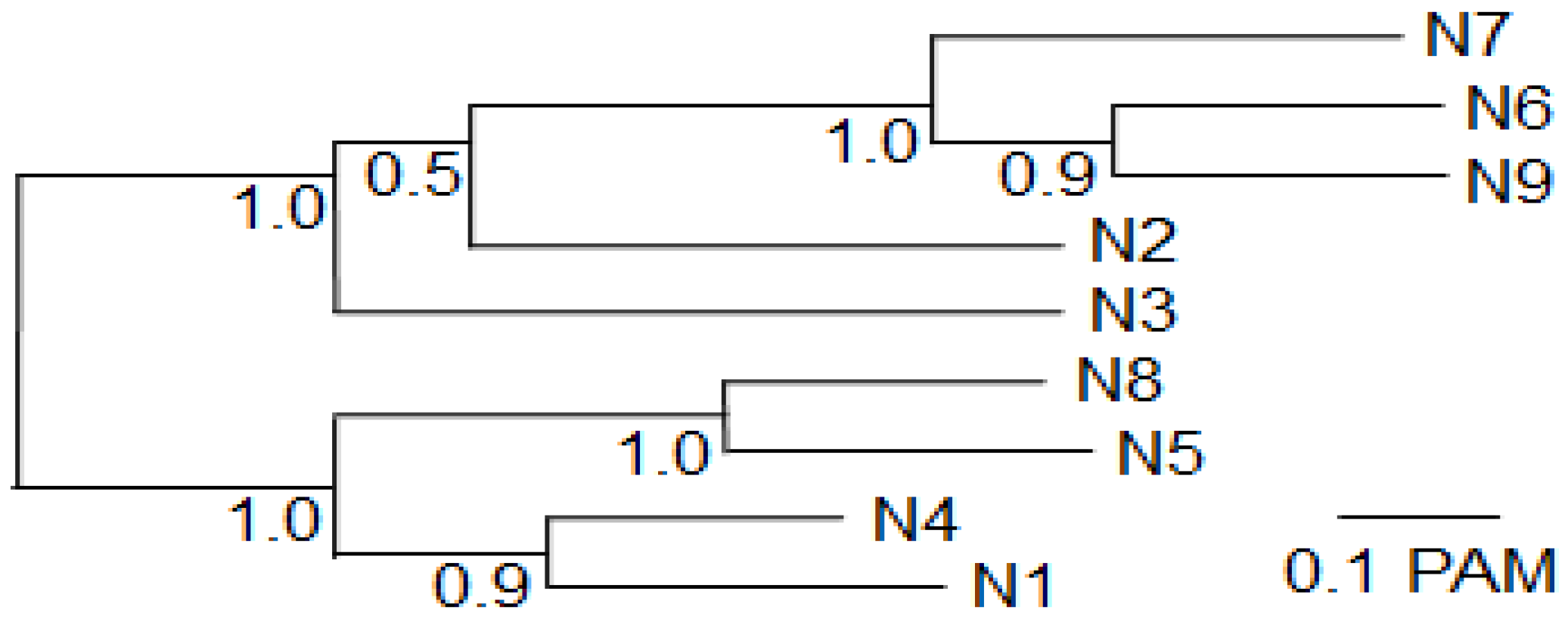
Most of the observed cross-reaction corresponds to the phylogenetic relationship between the different N subtypes. A phylogenetic tree (
Figure 2) of the NA proteins of the strains used for rNA production (
Table 3) closely resembles a previously published tree of NA proteins derived from other strains [
8], showing the generality of this tree for different strains. N1 is most related to N4 and somewhat more distantly to N5 and N8 (
Figure 2). Consistent with this, five clones bind both N1 and N4, two of which also bind N5 and three of which also bind N8 (
Table 7). The binding of four clones to both N6 and N7 (
Table 7) is also consistent with the close phylogenetic relationship between N6 and N7. However, some cross-reaction is not explained by a close phylogenetic relationship, such as the binding of IV525F to the distantly related N8 and N9. Furthermore, IV512F and IV618F, which belong to the same CDR3 group (
Table 4), bind to NAs of as much as five subtypes (
Table 7), according to the criterion used (A
450 nm > 0.5 in
Table 6). Using a less stringent criterion (A
450 nm = 0.2–0.5 in
Table 6), these VHHs also bind to other viruses, including N5 subtype viruses. Such broad cross-reaction with sometimes even lower absorbance values is also observed with N3-binder, IV456F, and N4-binder, IV213F (
Table 6).
Figure 2.
Phylogenetic tree of the nine influenza A virus N subtypes used in this study. The protein sequences of the complete NA of the nine strains used for recombinant NA production (
Table 3) were aligned and used for phylogenetic analysis. The statistical support as measured by 100 cycles of bootstrap analysis is indicated at each node.
Figure 2.
Phylogenetic tree of the nine influenza A virus N subtypes used in this study. The protein sequences of the complete NA of the nine strains used for recombinant NA production (
Table 3) were aligned and used for phylogenetic analysis. The statistical support as measured by 100 cycles of bootstrap analysis is indicated at each node.
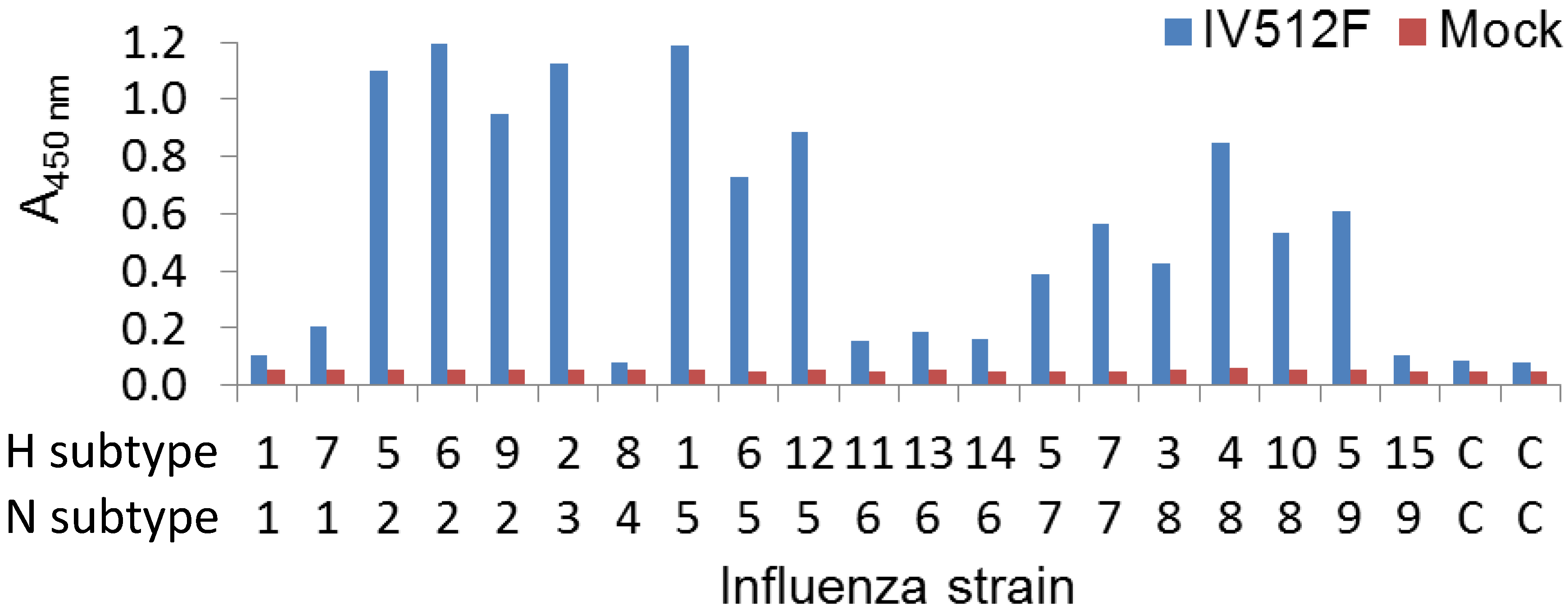
We further analyzed the cross-reaction of IV512F with many different subtypes in a sandwich ELISA setup by coating plates with IV512F and subsequent detection of bound NA of various Triton X-100 solubilized subtypes. This confirmed the binding to the various subtypes observed in the direct ELISA, including N5, although binding to N1 and N6 was inefficient (
Figure 3). Furthermore, in sandwich ELISA, IV512F bound to one of the two N9 viruses used. Thus, IV512F binds to influenza viruses off all subtypes, except N4, although not to all viruses of each subtype.
Figure 3.
Cross-reaction of IV512F to NAs of different subtypes in sandwich ELISA. Plates were coated with or without (Mock) IV512F and incubated with Triton X-100 solubilized viruses of different subtypes. C indicates control incubation without influenza virus. Bound NA was subsequently detected using different biotinylated VHHs recognizing the various NAs. The H5N9 strain used was A/chicken/Italy/22A/1998.
Figure 3.
Cross-reaction of IV512F to NAs of different subtypes in sandwich ELISA. Plates were coated with or without (Mock) IV512F and incubated with Triton X-100 solubilized viruses of different subtypes. C indicates control incubation without influenza virus. Bound NA was subsequently detected using different biotinylated VHHs recognizing the various NAs. The H5N9 strain used was A/chicken/Italy/22A/1998.
Two universally conserved epitopes in NA of all nine subtypes have been identified earlier by bioinformatic analysis and subsequent antibody generation using peptide immunization [
12]. One of these epitopes is close to the NA enzymatic site, whereas the other is located near the NA N-terminus, which forms the cytoplasmic tail. Possibly IV512F and the related VHH IV618F bind to one of these epitopes. The binding to these epitopes was improved by detergent treatment, which was critically dependent on detergent concentration [
12]. Thus, different absorbance values obtained when IV512F was analyzed by direct (
Table 6) or sandwich ELISA (
Figure 3) could be due to the use of detergent only in the latter assay.
The ability of IV512F to recognize a conserved epitope on NA could be due to the small size of a VHH. It has previously been suggested that some VHHs can recognize antigenic sites that are not recognized by the larger conventional antibody fragments, because these sites are located in cavities or clefts that can only be accessed by small antibody fragments. This is exemplified by VHHs binding to the enzyme active site of carbonic anhydrase or alpha-amylase [
28] or to cryptic epitopes on the trypanosome surface [
29]. Interestingly, this trypanosome antigenic site is conserved and lies underneath a layer of surface glycoproteins that is immunodominant and shows high antigenic variation to evade the host immune response.
We also measured the ability of VHHs to inhibit rNA enzymatic activity (
Table 5). Evidently, this could only be done for rNA preparations that showed enzymatic activity (rN2, rN3, rN7, rN8 and rN9). Most VHHs neutralized rNA, consistent with earlier observations with conventional antibodies [
9]. Several VHHs do not reach 50% inhibition at the highest VHH concentration analyzed. Clone IV77F inhibits at most 40% of the rN2, but does this at low VHH concentrations (IC
20 = 3 ng/mL). This could indicate that IV77F recognizes an epitope that is present in only 40% of the rN2 molecules. Clones, IV235F and IV456F, which belong to the same CDR3 group, also do not inhibit rN3, although they bind to H2N3 virus (
Table 5), which was also used for rN3 production. These clones possibly bind to an epitope that is not involved in enzyme activity. The absence of rNA inhibition by clones, IV268F (N8), IV275F (N8), IV294F (N9), IV295F (N9) and IV525F (N9), could be due to their strain-specific binding (
Table 5), since the rNA was derived from a different strain than used in ELISA. The cross-reaction of VHHs to viruses of different N subtype never correlates with their H subtype, making it unlikely that the cross-reaction is due to HA binding. Further proof that the selected VHHs bind to NA is based on their binding to rNA (
Table 4) or inhibition of rNA enzyme activity (
Table 5), which is, however, only available for enzymatically active rNA. The NA binding specificity of IV512F is further proven by its binding to different NAs in sandwich ELISA with other, biotinylated VHHs that unequivocally bind NA, because they bind rNA (
Table 4).