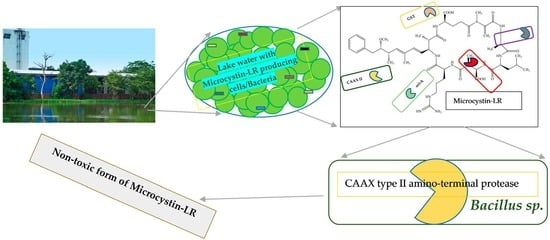
Microcystin-LR Biodegradation by Bacillus sp.: Reaction Rates and Possible Genes Involved in the Degradation
Abstract
:1. Introduction
2. Materials and Methods
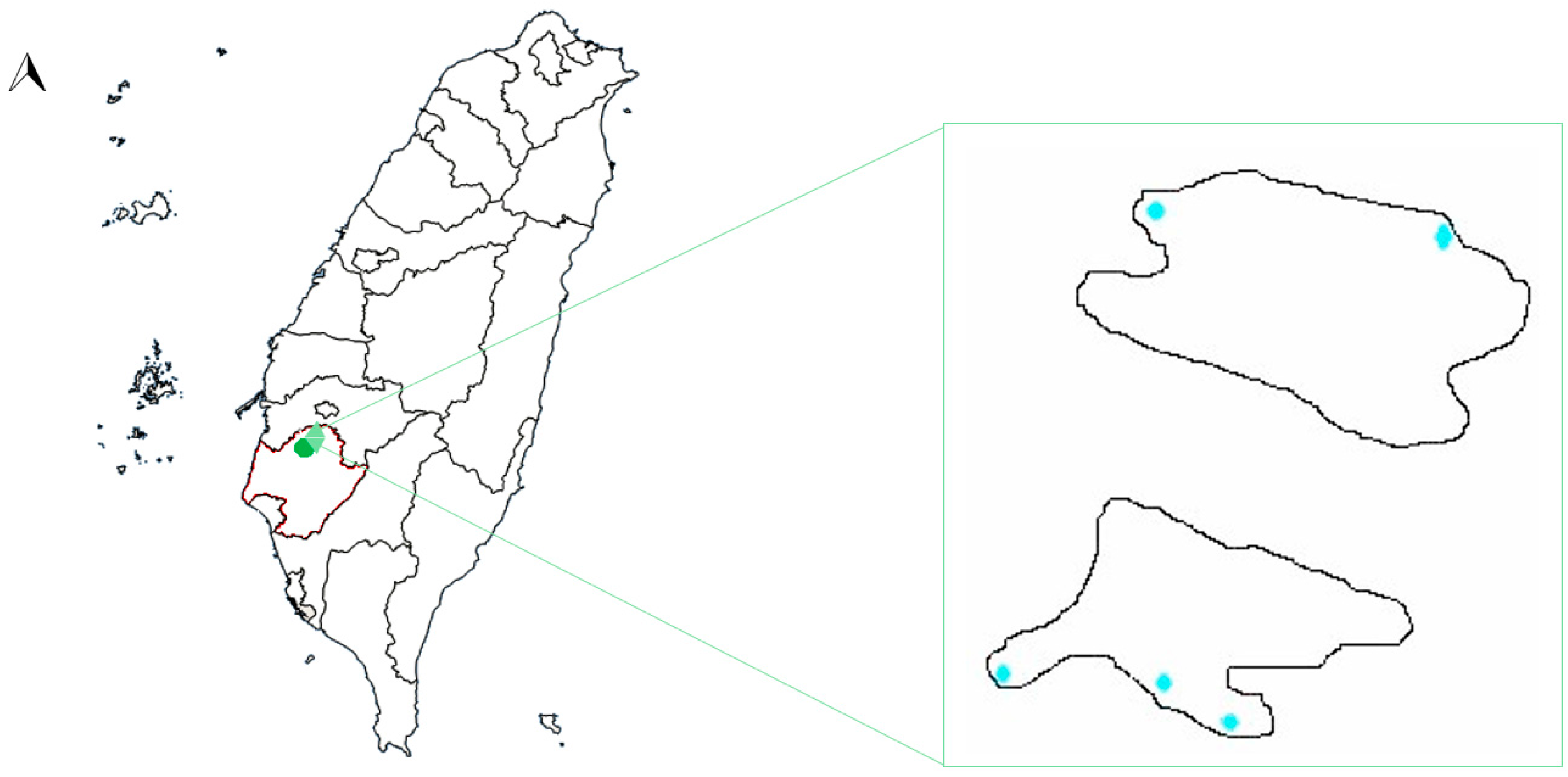
2.1. Hulupi Lake
2.2. Extraction of Crude MC-LR from M. aeruginosa Culture
2.3. MC-LR and TOC Concentration Measurements
2.4. Isolation of Bacterial Strain
2.5. MC-LR Biodegradation in HLPL Water
2.6. MC-LR Biodegradation by the Isolated Bacteria
2.7. DNA Extraction for Cyanobacteria and Bacteria
2.8. Polymerase Chain Reaction (PCR) and Quantitative PCR (q-PCR)
2.9. DNA Cloning, Sequencing, and Phylogenetic Tree Analysis for Bacteria
2.10. Statistical Analyses
3. Results and Discussion
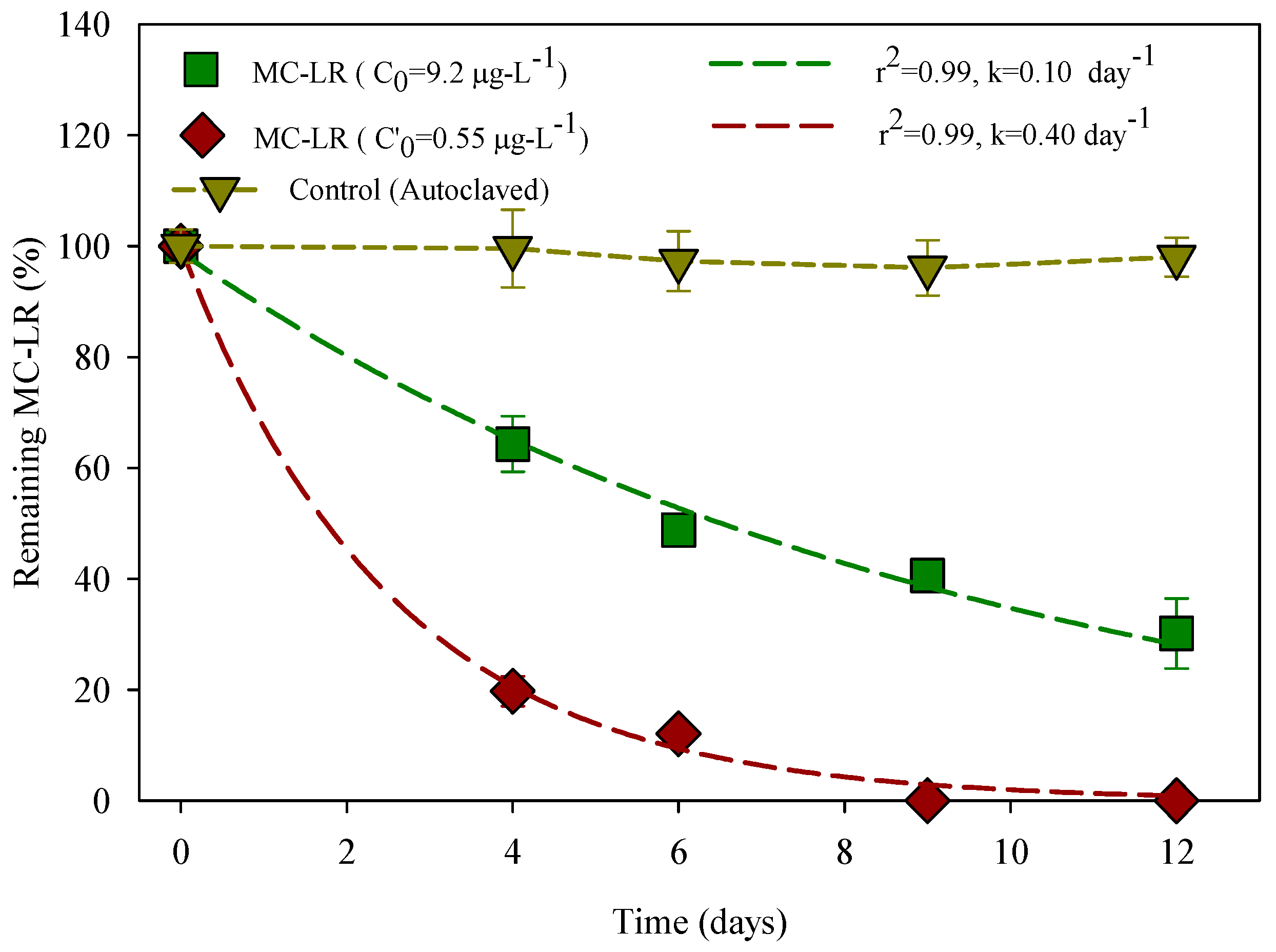
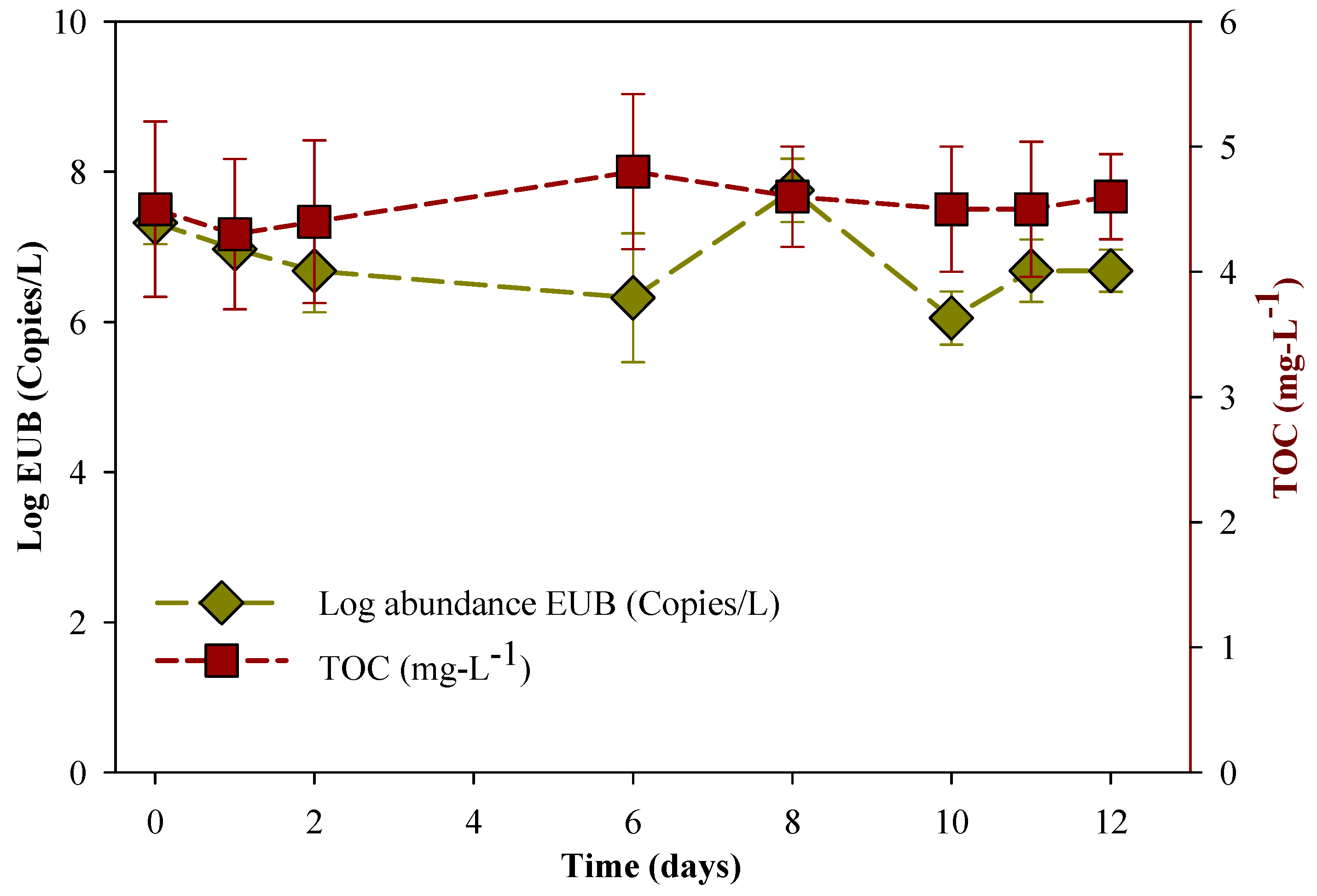
3.1. Degradation of MC-LR in HLPL Water
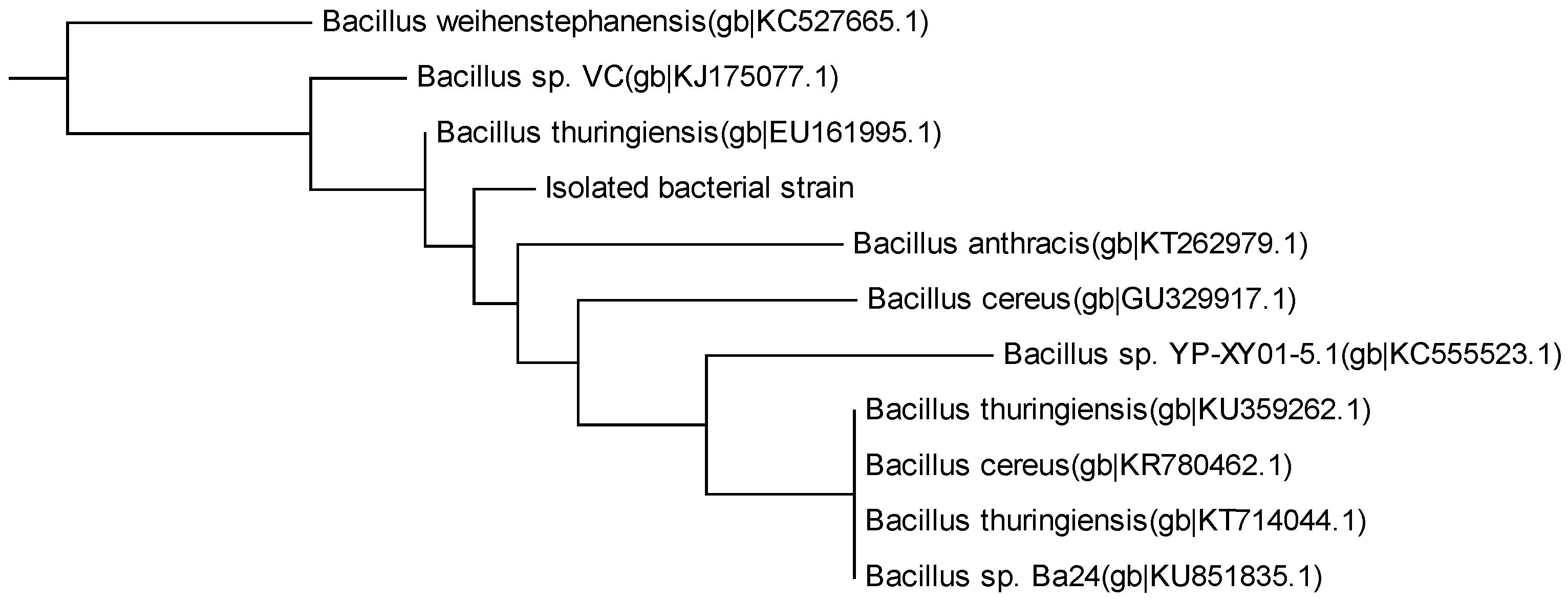
3.2. Isolated MC-LR Degrading Bacteria
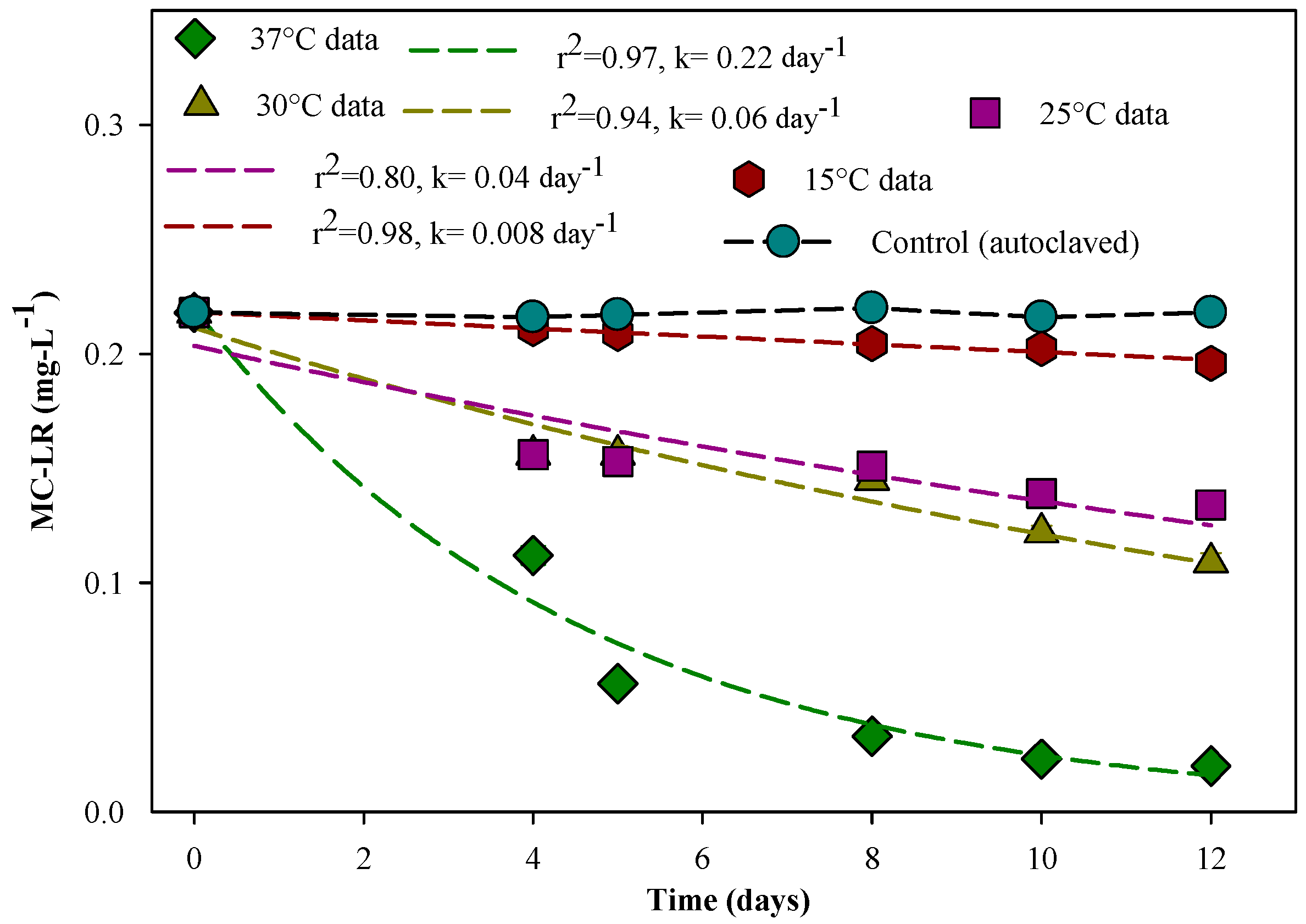
3.3. Effect of Temperature on MC-LR Degradation
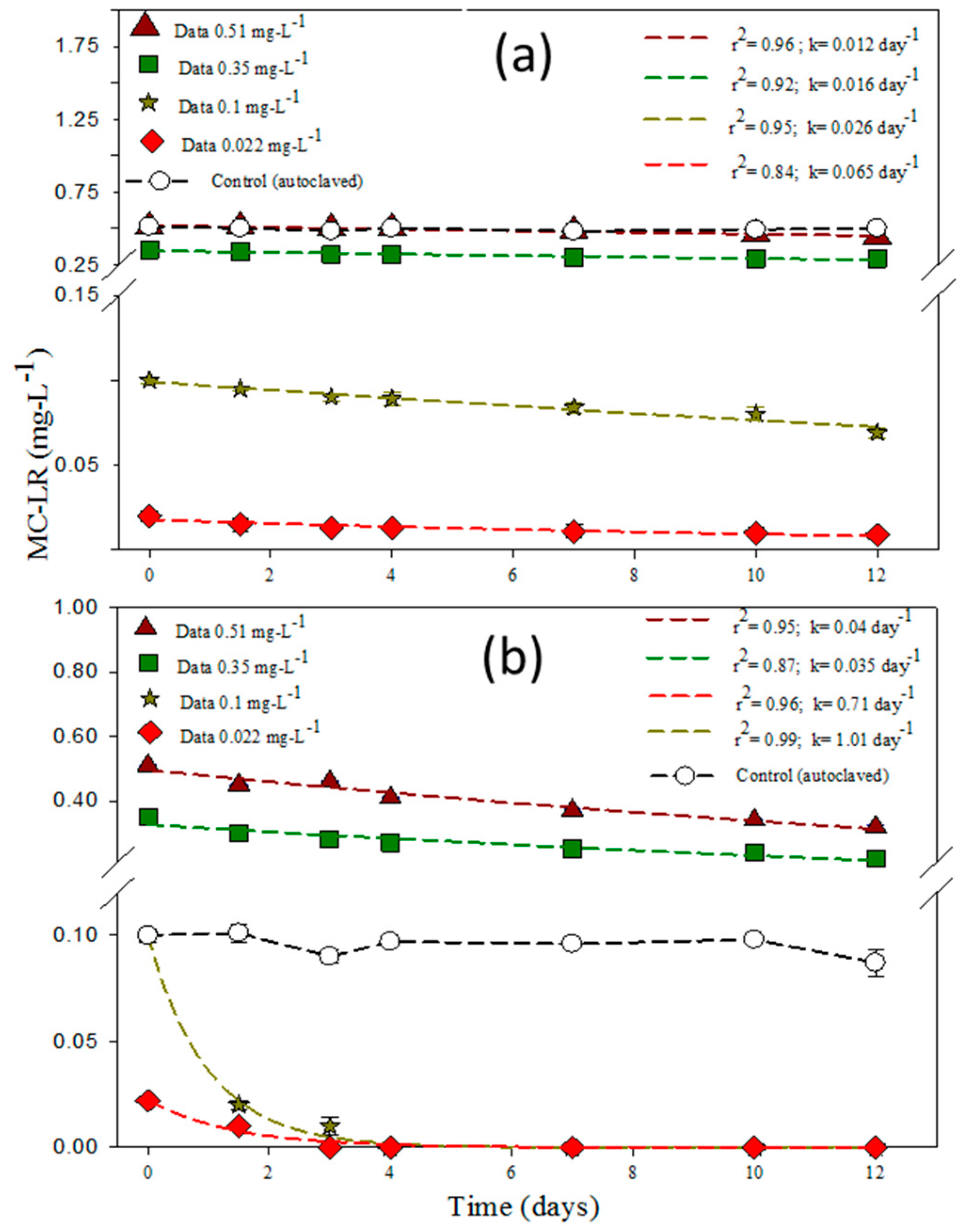
3.4. Effect of MC-LR and Bacteria Concentration on the Degradation Rates
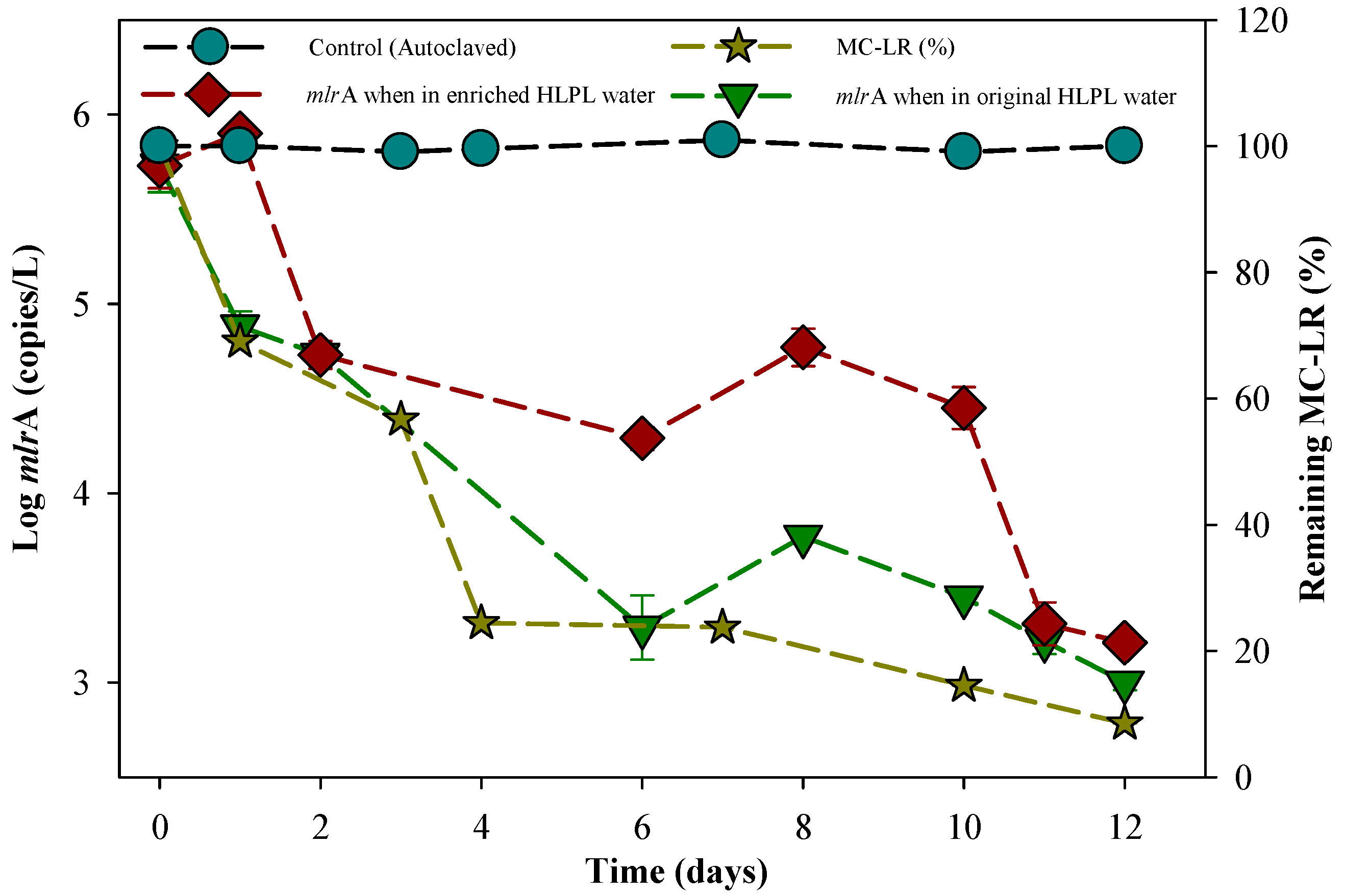
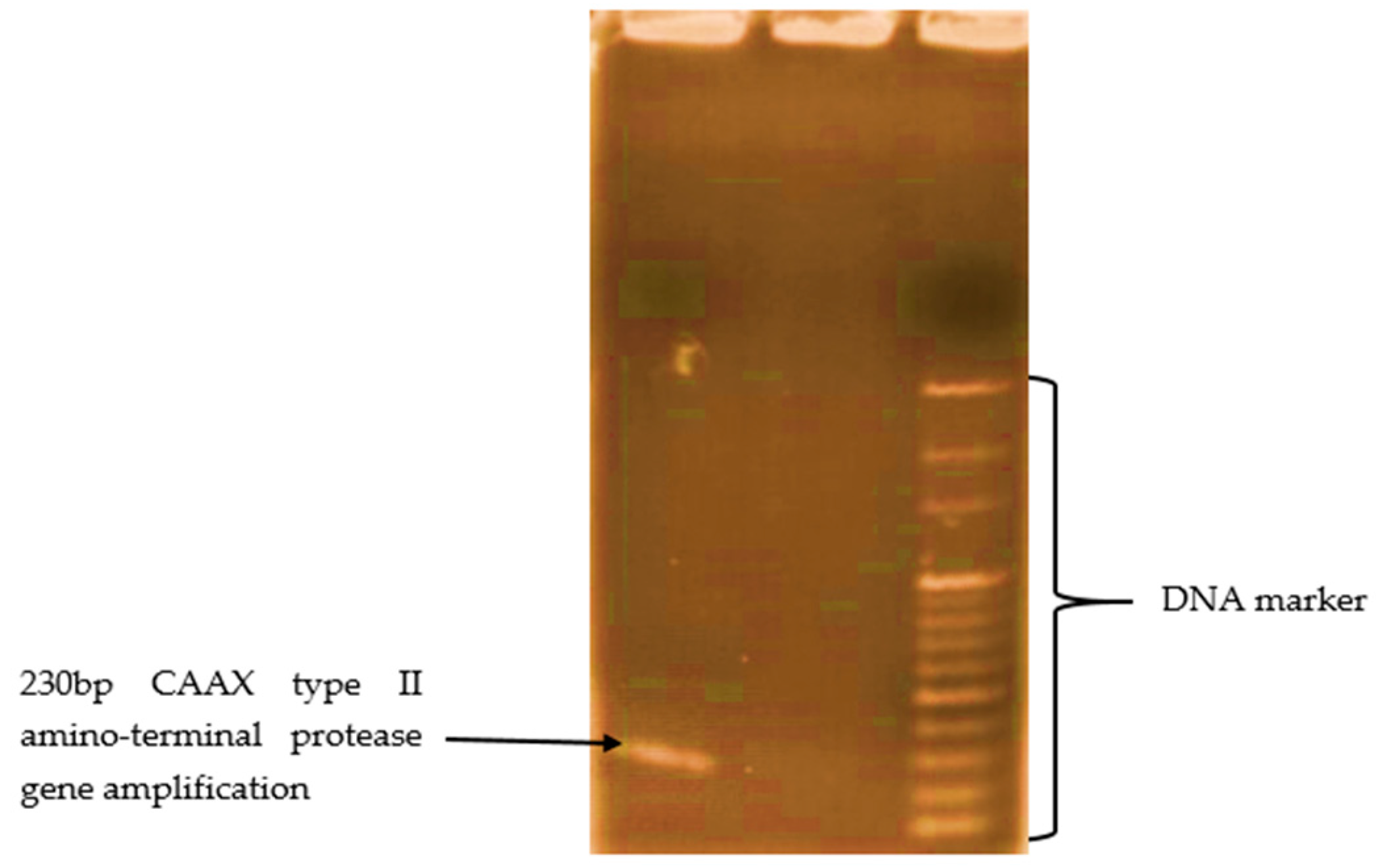
3.5. Biomolecular Aspect of MC-LR Degradation in HLPL Water
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Huo, X.; Chang, D.-W.; Tseng, J.-H.; Burch, M.D.; Lin, T.-F. Exposure of M. aeruginosa to hydrogen peroxide under light: Kinetic modeling of cell rupture and simultaneous microcystin degradation. Environ. Sci. Technol. 2015, 49, 5502–5510. [Google Scholar] [CrossRef] [PubMed]
- Antoniou, M.G.; De La Cruz, A.A.; Dionysiou, D.D. Cyanotoxins: New generation of water contaminants. J. Environ. Eng. 2005, 131, 1239–1243. [Google Scholar] [CrossRef]
- Tsao, H.-W.; Michinaka, A.; Yen, H.-K.; Giglio, S.; Hobson, P.; Monis, P.; Lin, T.-F. Monitoring of geosmin producing anabaena circinalis using quantitative PCR. Water Res. 2014, 49, 416–425. [Google Scholar] [CrossRef] [PubMed]
- Graham, J.L.; Loftin, K.A.; Meyer, M.T.; Ziegler, A.C. Cyanotoxin mixtures and taste-and-odor compounds in cyanobacterial blooms from the midwestern united states. Environ. Sci. Technol. 2010, 44, 7361–7368. [Google Scholar] [CrossRef] [PubMed]
- Chorus, I.; Bartram, J. Toxic cyanobacteria in water: A guide to public health significance. In Monitoring and Management; E&FN Spon: London, UK, 1999. [Google Scholar]
- Falconer, I.R.; Burch, M.D.; Steffensen, D.A.; Choice, M.; Coverdale, O.R. Toxicity of the blue-green alga (cyanobacterium) M. Aeruginosa in drinking water to growing pigs, as an animal model for human injury and risk assessment. Environ. Toxicol. Water Qual. 1994, 9, 131–139. [Google Scholar] [CrossRef]
- Falconer, I.R. Algal toxins and human health. In Quality and Treatment of Drinking Water II; Springer: Berlin, Germany, 1998; pp. 53–82. [Google Scholar]
- Kamp, L.; Church, J.L.; Carpino, J.; Faltin-Mara, E.; Rubio, F. The effects of water sample treatment, preparation, and storage prior to cyanotoxin analysis for cylindrospermopsin, microcystin and saxitoxin. Chem. Biol. Interact. 2016, 246, 45–51. [Google Scholar] [CrossRef] [PubMed]
- Imanishi, S.; Kato, H.; Mizuno, M.; Tsuji, K.; Harada, K.-I. Bacterial degradation of microcystins and nodularin. Chem. Res. Toxicol. 2005, 18, 591–598. [Google Scholar] [CrossRef] [PubMed]
- Dietrich, D.; Hoeger, S. Guidance values for microcystins in water and cyanobacterial supplement products (blue-green algal supplements): A reasonable or misguided approach? Toxicol. Appl. Pharmacol. 2005, 203, 273–289. [Google Scholar] [CrossRef] [PubMed]
- Falconer, I.R.; Humpage, A.R. Health risk assessment of cyanobacterial (blue-green algal) toxins in drinking water. Int. J. Environ. Res. Public Health 2005, 2, 43–50. [Google Scholar] [CrossRef] [PubMed]
- De Figueiredo, D.R.; Azeiteiro, U.M.; Esteves, S.M.; Gonçalves, F.J.; Pereira, M.J. Microcystin-producing blooms—A serious global public health issue. Ecotoxicol. Environ. Saf. 2004, 59, 151–163. [Google Scholar] [CrossRef] [PubMed]
- Shen, P.; Shi, Q.; Hua, Z.; Kong, F.; Wang, Z.; Zhuang, S.; Chen, D. Analysis of microcystins in cyanobacteria blooms and surface water samples from meiliang bay, taihu lake, china. Environ. Int. 2003, 29, 641–647. [Google Scholar] [CrossRef]
- Newcombe, G.; Chorus, I.; Falconer, I.; Lin, T.-F. Cyanobacteria: Impacts of climate change on occurrence, toxicity and water quality management. Water Res. 2012, 46, 1347–1348. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Guidelines for Drinking-Water Quality. Vol. 2, Health Criteria and Other Supporting Information: Addendum; World Health Organization: Geneva, Switzerland, 1998. [Google Scholar]
- Jetoo, S.; Grover, V.I.; Krantzberg, G. The toledo drinking water advisory: Suggested application of the water safety planning approach. Sustainability 2015, 7, 9787–9808. [Google Scholar] [CrossRef]
- Carmichael, W.W. Health effects of toxin-producing cyanobacteria:“The cyanohabs”. Hum. Ecol. Risk Assess. Int. J. 2001, 7, 1393–1407. [Google Scholar] [CrossRef]
- Garcia, A.C.; Bargu, S.; Dash, P.; Rabalais, N.N.; Sutor, M.; Morrison, W.; Walker, N.D. Evaluating the potential risk of microcystins to blue crab (callinectes sapidus) fisheries and human health in a eutrophic estuary. Harmful Algae 2010, 9, 134–143. [Google Scholar] [CrossRef]
- Cousins, I.; Bealing, D.; James, H.; Sutton, A. Biodegradation of microcystin-lr by indigenous mixed bacterial populations. Water Res. 1996, 30, 481–485. [Google Scholar] [CrossRef]
- Chen, W.; Song, L.; Peng, L.; Wan, N.; Zhang, X.; Gan, N. Reduction in microcystin concentrations in large and shallow lakes: Water and sediment-interface contributions. Water Res. 2008, 42, 763–773. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Utsumi, M.; Gao, Y.; Li, Q.; Tian, X.; Shimizu, K.; Sugiura, N. Influences of metal ions on microcystin-lr degradation capacity and dynamics in microbial distribution of biofilm collected from water treatment plant nearby kasumigaura lake. Chemosphere 2016, 147, 230–238. [Google Scholar] [CrossRef] [PubMed]
- Edwards, C.; Graham, D.; Fowler, N.; Lawton, L.A. Biodegradation of microcystins and nodularin in freshwaters. Chemosphere 2008, 73, 1315–1321. [Google Scholar] [CrossRef] [PubMed]
- Gągała, I.; Mankiewicz-Boczek, J. The natural degradation of microcystins (cyanobacterial hepatotoxins) in fresh water-the future of modern treatment systems and water quality improvement. Pol. J. Environ. Stud. 2012, 21, 1125–1139. [Google Scholar]
- Chen, J.; Hu, L.B.; Zhou, W.; Yan, S.H.; Yang, J.D.; Xue, Y.F.; Shi, Z.Q. Degradation of microcystin-LR and RR by a Stenotrophomonas sp. strain EMS isolated from lake taihu, china. Int. J. Mol. Sci. 2010, 11, 896–911. [Google Scholar] [CrossRef] [PubMed]
- Dziga, D.; Zielinska, G.; Wladyka, B.; Bochenska, O.; Maksylewicz, A.; Strzalka, W.; Meriluoto, J. Characterization of enzymatic activity of MlrB and MlrC proteins involved in bacterial degradation of cyanotoxins microcystins. Toxins 2016, 8, 76. [Google Scholar] [CrossRef] [PubMed]
- Sumaiya, I.; Pathmalal, M.; de Silva, B.; Welgamage, A.; Edwards, C.; Lawton, L.A. Isolation and characterization of microcystin-degrading bacteria from two water bodies in sri lanka. Appl. Environ. Microbiol. 2009, 75, 6924–6928. [Google Scholar]
- Bourne, D.G.; Jones, G.J.; Blakeley, R.L.; Jones, A.; Negri, A.P.; Riddles, P. Enzymatic pathway for the bacterial degradation of the cyanobacterial cyclic peptide toxin microcystin LR. Appl. Environ. Microbiol. 1996, 62, 4086–4094. [Google Scholar] [PubMed]
- Miao, H.-F.; Qin, F.; Tao, G.-J.; Tao, W.-Y.; Ruan, W.-Q. Detoxification and degradation of microcystin-LR and-RR by ozonation. Chemosphere 2010, 79, 355–361. [Google Scholar] [CrossRef] [PubMed]
- Okano, K.; Shimizu, K.; Maseda, H.; Kawauchi, Y.; Utsumi, M.; Itayama, T.; Zhang, Z.; Sugiura, N. Whole-genome sequence of the microcystin-degrading bacterium Sphingopyxis sp. Strain c-1. Genome Announc. 2015, 3. [Google Scholar] [CrossRef] [PubMed]
- Somdee, T.; Thunders, M.; Ruck, J.; Lys, I.; Allison, M.; Page, R. Degradation of [Dha7] MC-LR by a microcystin degrading bacterium isolated from lake rotoiti, new zealand. ISRN Microbiol. 2013, 2013, 596429. [Google Scholar] [CrossRef] [PubMed]
- Kormas, K.A.; Lymperopoulou, D.S. Cyanobacterial toxin degrading bacteria: Who are they? BioMed Res. Int. 2013, 2013, 877–881. [Google Scholar] [CrossRef] [PubMed]
- Bourne, D.G.; Riddles, P.; Jones, G.J.; Smith, W.; Blakeley, R.L. Characterisation of a gene cluster involved in bacterial degradation of the cyanobacterial toxin microcystin LR. Environ. Toxicol. 2001, 16, 523–534. [Google Scholar] [CrossRef] [PubMed]
- Manage, P.M.; Edwards, C.; Singh, B.K.; Lawton, L.A. Isolation and identification of novel microcystin-degrading bacteria. Appl. Environ. Microbiol. 2009, 75, 6924–6928. [Google Scholar] [CrossRef] [PubMed]
- Mankiewicz-Boczek, J.; Gagala, I.; Jurczak, T.; Jaskulska, A.; Pawelczyk, J.; Dziadek, J. Bacteria homologus to aeromonas. Capable of microcystin degradation. Open Life Sci 2015, 10, 119–129. [Google Scholar] [CrossRef]
- Saito, T.; Okano, K.; Park, H.-D.; Itayama, T.; Inamori, Y.; Neilan, B.A.; Burns, B.P.; Sugiura, N. Detection and sequencing of the microcystin lr-degrading gene, mlra, from new bacteria isolated from japanese lakes. FEMS Microbiol. Lett. 2003, 229, 271–276. [Google Scholar] [CrossRef]
- Pei, J.; Mitchell, D.A.; Dixon, J.E.; Grishin, N.V. Expansion of type ii caax proteases reveals evolutionary origin of γ-secretase subunit aph-1. J. Mol. Biol. 2011, 410, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Barrett, A.J.; Woessner, J.F.; Rawlings, N.D. Handbook of Proteolytic Enzymes; Elsevier: Amsterdam, The Netherlands, 2012. [Google Scholar]
- Mou, X.; Lu, X.; Jacob, J.; Sun, S.; Heath, R. Metagenomic identification of bacterioplankton taxa and pathways involved in microcystin degradation in lake erie. PLoS ONE 2013, 8, e61890. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Zhou, Y.; Yin, L.; Zhu, G.; Liang, G.; Pu, Y. Microcystin-degrading activity of an indigenous bacterial strain Stenotrophomonas acidaminiphila MC-LTH2 isolated from lake taihu. PLoS ONE 2014, 9, e86216. [Google Scholar] [CrossRef] [PubMed]
- Dziga, D.; Wasylewski, M.; Wladyka, B.; Nybom, S.; Meriluoto, J. Microbial degradation of microcystins. Chem. Res. Toxicol. 2013, 26, 841–852. [Google Scholar] [CrossRef] [PubMed]
- Sadler, T.; von Elert, E. Physiological interaction of daphnia and microcystis with regard to cyanobacterial secondary metabolites. Aquat. Toxicol. 2014, 156, 96–105. [Google Scholar] [CrossRef] [PubMed]
- Bury, N.; Flik, G.; Eddy, F.; Codd, G. The effects of cyanobacteria and the cyanobacterial toxin microcystin-LR on Ca2+ transport and Na+/K+-atpase in tilapia gills. J. Exp. Biol. 1996, 199, 1319–1326. [Google Scholar] [PubMed]
- Hu, L.B.; Yang, J.D.; Zhou, W.; Yin, Y.F.; Chen, J.; Shi, Z.Q. Isolation of a Methylobacillus sp. That degrades microcystin toxins associated with cyanobacteria. New Biotechnol. 2009, 26, 205–211. [Google Scholar] [CrossRef] [PubMed]
- Nhmrc, N. Australian Drinking Water Guidelines. 2004. Available online: https://www.nhmrc.gov.au/guidelines-publications/eh34 (accessed on 20 August 2016). [Google Scholar]
- Fischer, W.J.; Garthwaite, I.; Miles, C.O.; Ross, K.M.; Aggen, J.B.; Chamberlin, A.R.; Towers, N.R.; Dietrich, D.R. Congener-independent immunoassay for microcystins and nodularins. Environ. Sci. Technol. 2001, 35, 4849–4856. [Google Scholar] [CrossRef] [PubMed]
- Sepahi, A.A.; Golpasha, I.D.; Emami, M.; Nakhoda, A. Isolation and characterization of crude oil degrading bacillus spp. J. Environ. Health Sci. Eng. 2008, 5, 149–154. [Google Scholar]
- Bartram, J.; Cotruvo, J.; Exner, M.; Fricker, C.; Glasmacher, A. Heterotrophic Plate Counts and Drinking-Water Safety: The Significance of Hpcs for Water Quality and Human Health; IWA Publishing: London, UK, 2003. [Google Scholar]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Hoefel, D.; Adriansen, C.M.; Bouyssou, M.A.; Saint, C.P.; Newcombe, G.; Ho, L. Development of an mlrA gene-directed TaqMan PCR assay for quantitative assessment of microcystin-degrading bacteria within water treatment plant sand filter biofilms. Appl. Environ. Microbiol. 2009, 75, 5167–5169. [Google Scholar] [CrossRef] [PubMed]
- Muyzer, G.; De Waal, E.C.; Uitterlinden, A.G. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl. Environ. Microbiol. 1993, 59, 695–700. [Google Scholar] [PubMed]
- Lloyd-Jones, G.; Lau, P. Glutathione s-transferase-encoding gene as a potential probe for environmental bacterial isolates capable of degrading polycyclic aromatic hydrocarbons. Appl. Environ. Microbiol. 1997, 63, 3286–3290. [Google Scholar] [PubMed]
- Lloyd-Jones, G.; Laurie, A.D.; Hunter, D.W.; Fraser, R. Analysis of catabolic genes for naphthalene and phenanthrene degradation in contaminated new zealand soils. FEMS Microbiol. Ecol. 1999, 29, 69–79. [Google Scholar] [CrossRef]
- Engelbrektson, A.; Kunin, V.; Wrighton, K.C.; Zvenigorodsky, N.; Chen, F.; Ochman, H.; Hugenholtz, P. Experimental factors affecting pcr-based estimates of microbial species richness and evenness. ISME J. 2010, 4, 642–647. [Google Scholar] [CrossRef] [PubMed]
- Ahn, J.-H.; Kim, M.-C.; Shin, H.-C.; Choi, M.-K.; Yoon, S.-S.; Kim, T.; Song, H.-G.; Lee, G.H.; Ka, J.-O. Improvement of PCR amplification bias for community structure analysis of soil bacteria by denaturing gradient gel electrophoresis. J. Microbiol. Biotechnol. 2006, 16, 1561–1569. [Google Scholar]
- Schreier, P.H.; Cortese, R. A fast and simple method for sequencing DNA cloned in the single-stranded bacteriophage M13. J. Mol. Biol. 1979, 129, 169–172. [Google Scholar] [CrossRef]
- Knoche, K.; Kephart, D. Cloning blunt-end Pfu DNA polymerase-generated PCR fragments into pGEM®-T vector systems. Promega Notes 1999, 71, 10. [Google Scholar]
- Ncbi, R.C. Database resources of the national center for biotechnology information. Nucleic Acids Res. 2016, 44, D7. [Google Scholar]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Jordaan, K.; Bezuidenhout, C. Bacterial community composition of an urban river in the north west province, south africa, in relation to physico-chemical water quality. Environ. Sci. Pollut. Res. 2016, 23, 5868–5880. [Google Scholar] [CrossRef] [PubMed]
- Himberg, K.; Keijola, A.-M.; Hiisvirta, L.; Pyysalo, H.; Sivonen, K. The effect of water treatment processes on the removal of hepatotoxins frommicrocystis andoscillatoria cyanobacteria: A laboratory study. Water Res. 1989, 23, 979–984. [Google Scholar] [CrossRef]
- Harada, K.-I.; Tsuji, K.; Watanabe, M.F.; Kondo, F. Stability of microcystins from cyanobacteria-III.* Effect of pH and temperature. Phycologia 1996, 35, 83–88. [Google Scholar] [CrossRef]
- Jones, G.J.; Orr, P.T. Release and degradation of microcystin following algicide treatment of a M. Aeruginosa bloom in a recreational lake, as determined by hplc and protein phosphatase inhibition assay. Water Res. 1994, 28, 871–876. [Google Scholar] [CrossRef]
- Farhadkhani, M.; Nikaeen, M.; Adergani, B.A.; Hatamzadeh, M.; Nabavi, B.F.; Hassanzadeh, A. Assessment of drinking water quality from bottled water coolers. Iran. J. Public Health 2014, 43, 674. [Google Scholar] [PubMed]
- Cabeen, M.T.; Jacobs-Wagner, C. Bacterial cell shape. Nat. Rev. Microbiol. 2005, 3, 601–610. [Google Scholar] [CrossRef] [PubMed]
- Park, H.D.; Sasaki, Y.; Maruyama, T.; Yanagisawa, E.; Hiraishi, A.; Kato, K. Degradation of the cyanobacterial hepatotoxin microcystin by a new bacterium isolated from a hypertrophic lake. Environ. Toxicol. 2001, 16, 337–343. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Shimizu, K.; Utsumi, M.; Nakamoto, T.; Sakharkar, M.K.; Zhang, Z.; Sugiura, N. Dynamics of the functional gene copy number and overall bacterial community during microcystin-LR degradation by a biological treatment facility in a drinking water treatment plant. J. Biosci. Bioeng. 2011, 111, 695–701. [Google Scholar] [CrossRef] [PubMed]
- Ho, L.; Hoefel, D.; Palazot, S.; Sawade, E.; Newcombe, G.; Saint, C.P.; Brookes, J.D. Investigations into the biodegradation of microcystin-LR in wastewaters. J. Hazard. Mater. 2010, 180, 628–633. [Google Scholar] [CrossRef] [PubMed]
- Maatouk, I.; Bouaïcha, N.; Plessis, M.J.; Périn, F. Detection by 32P-postlabelling of 8-oxo-7, 8-dihydro-2′-deoxyguanosine in DNA as biomarker of microcystin-LR-and nodularin-induced DNA damage in vitro in primary cultured rat hepatocytes and in vivo in rat liver. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2004, 564, 9–20. [Google Scholar] [CrossRef] [PubMed]
- Ding, W.X.; Shen, H.M.; Ong, C.N. Critical role of reactive oxygen species and mitochondrial permeability transition in microcystin-induced rapid apoptosis in rat hepatocytes. Hepatology 2000, 32, 547–555. [Google Scholar] [CrossRef] [PubMed]
- Gehringer, M.M.; Shephard, E.G.; Downing, T.G.; Wiegand, C.; Neilan, B.A. An investigation into the detoxification of microcystin-LR by the glutathione pathway in balb/c mice. Int. J. Biochem. Cell Biol. 2004, 36, 931–941. [Google Scholar] [CrossRef] [PubMed]
- Campos, A.; Vasconcelos, V. Molecular mechanisms of microcystin toxicity in animal cells. Int. J. Mol. Sci. 2010, 11, 268–287. [Google Scholar] [CrossRef] [PubMed]
- Dziga, D.; Wladyka, B.; Zielińska, G.; Meriluoto, J.; Wasylewski, M. Heterologous expression and characterisation of microcystinase. Toxicon 2012, 59, 578–586. [Google Scholar] [CrossRef] [PubMed]
- Read, T.D.; Salzberg, S.L.; Pop, M.; Shumway, M.; Umayam, L.; Jiang, L.; Holtzapple, E.; Busch, J.D.; Smith, K.L.; Schupp, J.M. Comparative genome sequencing for discovery of novel polymorphisms in bacillus anthracis. Science 2002, 296, 2028–2033. [Google Scholar] [CrossRef] [PubMed]
| Target Gene | Primers | Probe/Dye | Use | References |
|---|---|---|---|---|
| mlrA | qmlrAf (5′-AGCCCKGGCCCRCTGC-3′) | qmlrA-tm (5′-FAM-TGCCSCAGCTSCTCAAGAAGTTTG-BHQ1-3′) TaqMan® | q-PCR | [49] |
| qmlrAr (5′-ATGCCARGCCCACCACAT-3′) | - | |||
| EUB (Total bacteria) | EUB338 5′-ACTCCTACGGGAGGCAGCAG-3′ | SYBR®Green I | q-PCR | [50] |
| EUB518 5′-ATTACCGCGGCTGCTGG-3′ | - | |||
| GST | p3-24F (5′-ATGAAACTGTTCATCAGCC-3′) | - | PCR | [51,52] |
| p3-24R (5′-CCAGCATCACGTACAGATAG-3′) | - | |||
| DNA cloning (Isolated bacteria) | 1392R 5′-ACGGGCGGTGTGTRC-3′ | - | PCR | [53] |
| 338F 5′-ACTCCTACGGGAGGCAGCAG-3′ | - | [54] | ||
| Plasmid insert vector M13 (Isolated bacteria DNA sequencing) | M13 Reverse (5′-CAGGAAACAGCTATGAC-3′) | - | PCR | [55] |
| M13 Forward (5′-GTAAAACGACGGCCAG-3′) | - | |||
| CAAX type II amino-protease gene | CAAX F2 (5′-TGGACTTTTTGGCATCTTCC-3′) | - | PCR | This study |
| CAAX R2 (5′-GGGTTGGTGGAATGAGTTTG-3′) | - |
| Microorganisms | mlrA | GST | CAAX | Temp. (°C) | Degradation Rate (mg·L−1·Day−1) | k (Day−1) | References |
|---|---|---|---|---|---|---|---|
| Bacterioplankton (Hulupi Lake) | +/# | - | + | 25 | 6 × 10−5 | 0.1 | This study |
| Bacillus sp. | - | - | + | 37 | 5 × 10−2 | 0.22 | This study |
| Mixed Arthrobacter spp., Brevibacterium sp., and Rhodococcus sp | - | ∞ | ∞ | 25 | 1.67 | 0.33 * | [33] |
| Stenotrophomonas acidaminiphila MC-LTH2 | - | ∞ | ∞ | 30 | 3.0 | 0.14 * | [39] |
| Bacterioplankton microcosm (Lake Erie) | - | + | ∞ | room temp. | Qualitative degradation | Qualitative degradation | [38] |
| Aeromonas sp. | - | ∞ | ∞ | 25 | Qualitative degradation | Qualitative degradation | [34] |
| Sphingomonas sp. Y2 (AB084247) | + | ∞ | ∞ | 30 | 5.4 | 0.27 * | [65] |
| Sphingomonas isolate NV-3 | + | ∞ | ∞ | 30 | 8.3 × 10−3 | 0.33 * | [30] |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kansole, M.M.R.; Lin, T.-F. Microcystin-LR Biodegradation by Bacillus sp.: Reaction Rates and Possible Genes Involved in the Degradation. Water 2016, 8, 508. https://doi.org/10.3390/w8110508
Kansole MMR, Lin T-F. Microcystin-LR Biodegradation by Bacillus sp.: Reaction Rates and Possible Genes Involved in the Degradation. Water. 2016; 8(11):508. https://doi.org/10.3390/w8110508
Chicago/Turabian StyleKansole, Michelline M. R., and Tsair-Fuh Lin. 2016. "Microcystin-LR Biodegradation by Bacillus sp.: Reaction Rates and Possible Genes Involved in the Degradation" Water 8, no. 11: 508. https://doi.org/10.3390/w8110508
APA StyleKansole, M. M. R., & Lin, T.-F. (2016). Microcystin-LR Biodegradation by Bacillus sp.: Reaction Rates and Possible Genes Involved in the Degradation. Water, 8(11), 508. https://doi.org/10.3390/w8110508