Effect of Plant Harvesting on the Performance of Constructed Wetlands during Summer
Abstract
:1. Introduction
2. Materials and Methods
2.1. Microcosm Wetland System Setup
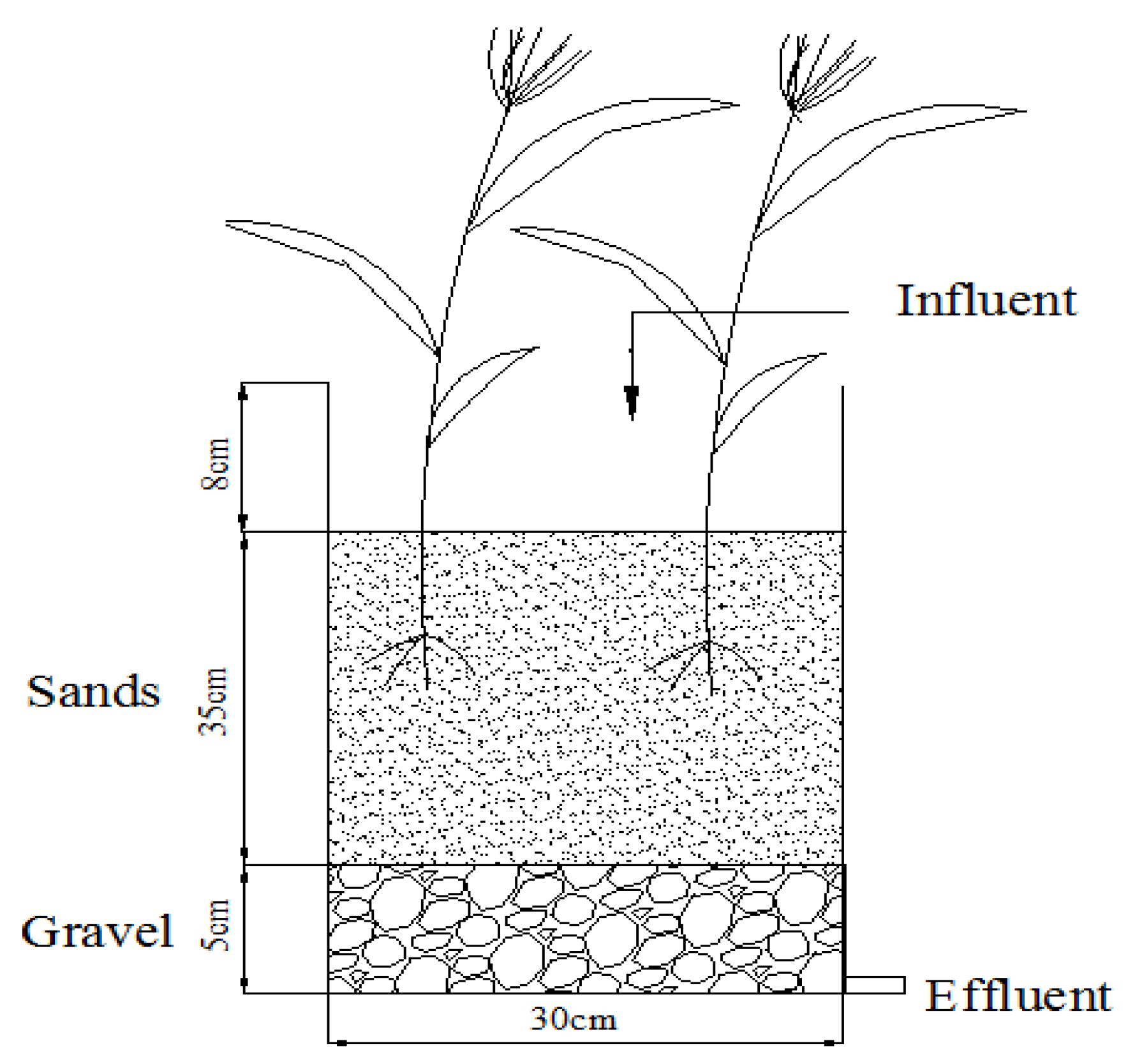
2.2. Sampling
2.3. Analysis
2.3.1. Physical and Chemical Analysis
2.3.2. Nitrogen Analysis
2.3.3. Microbial Analysis
2.3.4. Pyrosequencing and Data Analysis
2.3.5. Statistical Analyses
3. Results and Discussion
3.1. Effluent Water Parameters
| Group | Parameter | |||||
|---|---|---|---|---|---|---|
| COD (mg/L) | NH4+-N (mg/L) | Removal Efficiency of COD (%) | Removal Efficiency of NH4+-N (%) | T (°C) | DO (mg/L) | |
| Unharvested | 22.75 ± 2.91 | 1.17 ± 0.55 | 92.64 ± 3.35 | 63.23 ± 4.13 | 24.36 ± 3.87 | 1.63 ± 0.56 |
| Harvested | 18.35 ± 5.23 | 1.00 ± 0.44 | 93.70 ± 2.73 | 70.65 ± 7.47 | 24.54 ± 4.03 | 2.00 ± 0.38 |
| Cleared | 26.14 ± 4.43 | 1.97 ± 0.52 | 87.62 ± 0.26 | 57.75 ± 6.40 | 25.41 ± 4.84 | 1.63 ± 0.38 |
3.2. Nitrogen Mass Balance
| Group | Plant Dry Mass (g) | Plant Biomass (g) | Input Load (mg N/m2/day) | Output Load (mg N/m2/day) | |||
|---|---|---|---|---|---|---|---|
| Influent | Effluent | Sediment | Plant Uptake | Other Losses | |||
| Cleared | 0 | 0 | 101.21 | 13.59 | 38.97 | 0 | 48.65 |
| Harvested | 13.18 | 16.48 | 101.21 | 6.97 | 40.06 | 41.12 | 13.06 |
| Unharvested | 20.34 | 27.12 | 101.21 | 8.15 | 30.54 | 33.21 | 29.31 |
3.3. Microbial Abundance and Community
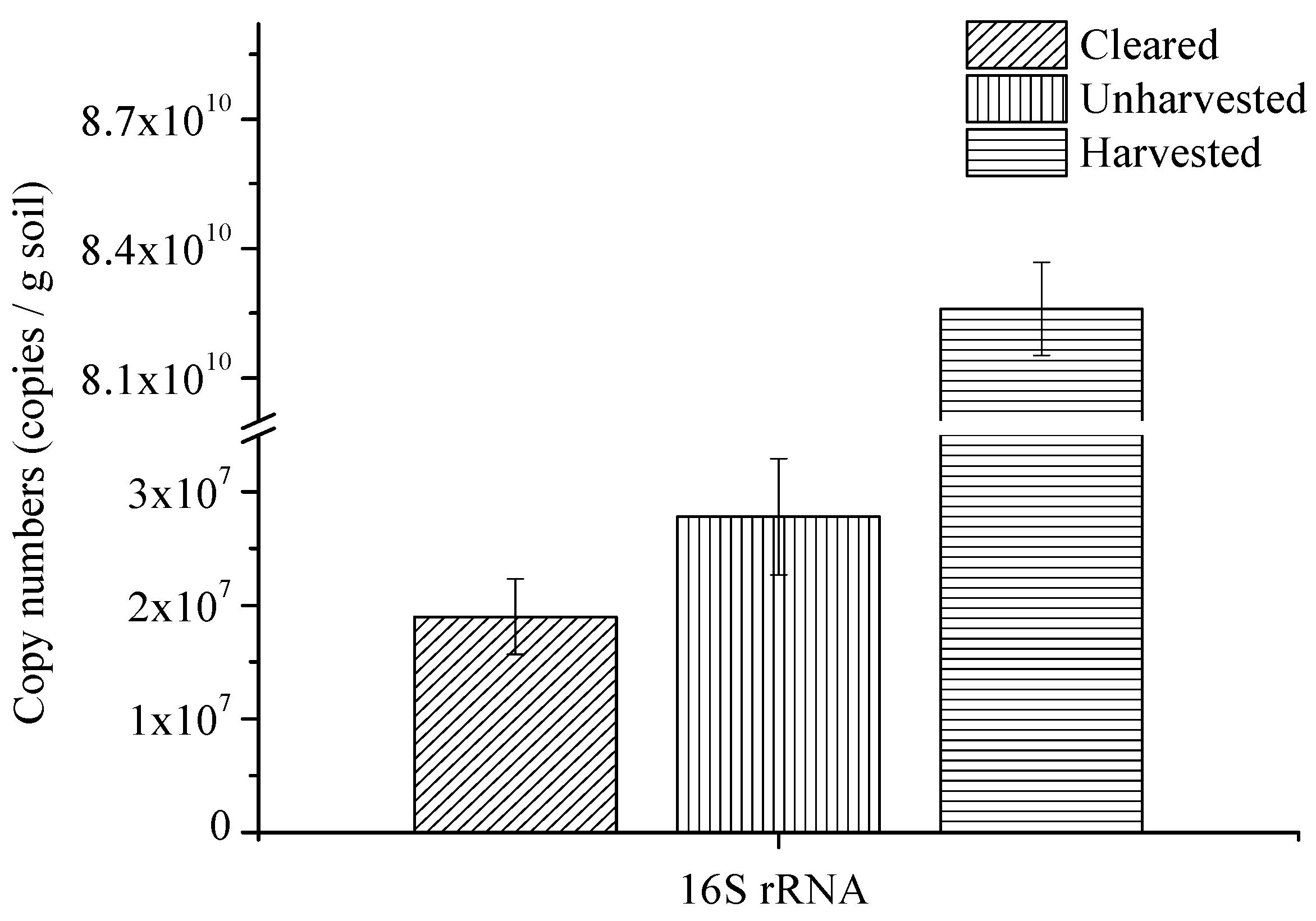
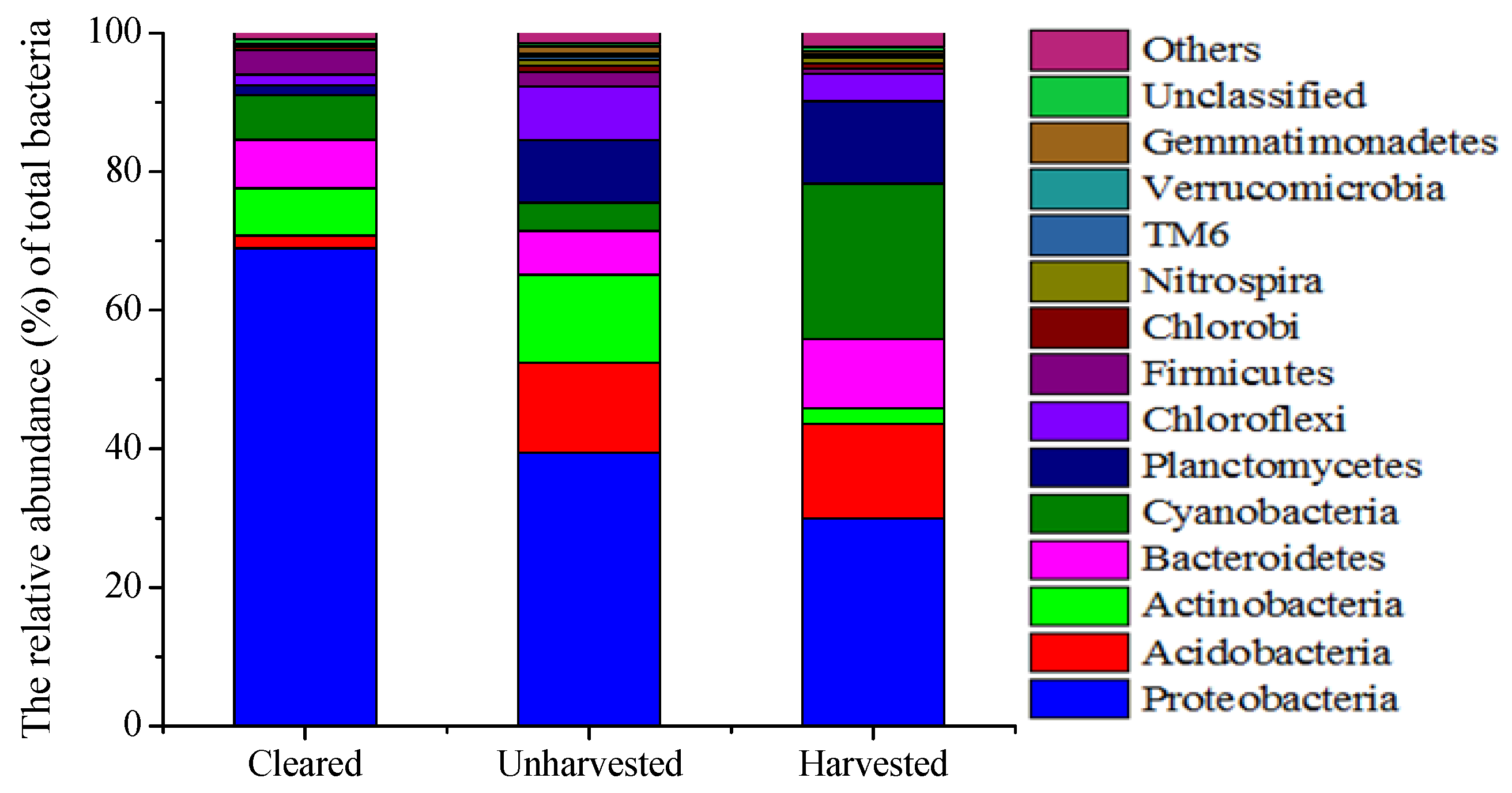
| Class | Unharvested | Harvested | Cleared |
|---|---|---|---|
| Alpha | 15.30 | 9.03 | 4.63 |
| Beta | 11.21 | 9.26 | 16.06 |
| Delta | 6.18 | 3.86 | 2.78 |
| Gamma | 6.57 | 7.60 | 45.40 |
| Unclassified | 0.01 | 0.03 | 0.06 |
| Total | 39.41 | 29.92 | 68.92 |
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gikas, P.; Ranieri, E.; Tchobanoglous, G. Removal of iron, chromium and lead from waste water by horizontal subsurface flow constructed wetlands. J. Chem. Technol. Biotechnol. 2013, 88, 1906–1912. [Google Scholar]
- Vymazal, J. Constructed Wetlands for Wastewater Treatment: Five Decades of Experience. Environ. Sci. Technol. 2010, 45, 61–69. [Google Scholar] [PubMed]
- Dickopp, J.; Kazda, M.; Čížková, H. Differences in rhizome aeration of Phragmites australis in a constructed wetland. Ecol. Eng. 2011, 37, 1647–1653. [Google Scholar] [CrossRef]
- Jinadasa, K.; Tanaka, N.; Sasikala, S.; Werellagama, D.; Mowjood, M.; Ng, W. Impact of harvesting on constructed wetlands performance—A comparison between Scirpus grossus and Typha angustifolia. J. Environ. Sci. Health 2008, 43, 664–671. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Xie, H.; Zhang, J.; Liang, S.; Ngo, H.H.; Guo, W.; Liu, C.; Zhao, C.; Li, H. Effect of plant harvesting on the performance of constructed wetlands during winter: Radial oxygen loss and microbial characteristics. Environ. Sci. Pollut. Res. 2014, 22, 7476–7484. [Google Scholar] [CrossRef] [PubMed]
- Ranieri, E.; Gikas, P. Effects of plants for reduction and removal of hexavalent chromium from a contaminated soil. Water Air Soil Pollut. 2014, 225, 1–9. [Google Scholar] [CrossRef]
- Gearheart, R.A. Use of constructed wetlands to treat domestic wastewater, City of Arcata, California. Water Sci. Technol. 1992, 26, 1625–1637. [Google Scholar]
- Bald, M.L. Plant Harvesting from a Constructed Wetland: Nutrient Removal and Plant Attributes; Department of Environmental Biology, University of Adelaide: Adelaide, Australia, 2001. [Google Scholar]
- Gryseels, M. Nature management experiments in a derelict reed marsh. I: Effects of winter cutting. Biol. Conserv. 1989, 47, 171–193. [Google Scholar] [CrossRef]
- Asaeda, T.; Karunaratne, S. Dynamic modeling of the growth of Phragmites australis: Model description. Aquat. Bot. 2000, 67, 301–318. [Google Scholar] [CrossRef]
- Haberl, R.; Perfler, R. Seven years of research work and experience with wastewater treatment by a reed bed system, Constructed wetlands in water pollution control. In Proceedings of the International Conference on the Use of Constructed Wetlands in Water Pollution Control, Cambridge, UK, 24–28 September 1990; p. 205.
- Jensen, S.I.; Kühl, M.; Priemé, A. Different bacterial communities associated with the roots and bulk sediment of the seagrass Zostera marina. FEMS Microbiol. Ecol. 2007, 62, 108–117. [Google Scholar] [CrossRef] [PubMed]
- Stottmeister, U.; Wießner, A.; Kuschk, P.; Kappelmeyer, U.; Kästner, M.; Bederski, O.; Müller, R.; Moormann, H. Effects of plants and microorganisms in constructed wetlands for wastewater treatment. Biotechnol. Adv. 2003, 22, 93–117. [Google Scholar] [CrossRef] [PubMed]
- Białowiec, A.; Davies, L.; Albuquerque, A.; Randerson, P.F. The influence of plants on nitrogen removal from landfill leachate in discontinuous batch shallow constructed wetland with recirculating subsurface horizontal flow. Ecol. Eng. 2012, 40, 44–52. [Google Scholar] [CrossRef]
- Ueckert, J.; Hurek, T.; Fendrik, I.; Niemann, E.-G. Radial gas diffusion from roots of rice (Oryza sativa L.) and Kallar grass (Leptochloa fusca L. Kunth), and effects of inoculation with Azospirillum brasilense Cd. Plant Soil 1990, 122, 59–65. [Google Scholar] [CrossRef]
- Wu, H.; Zhang, J.; Li, P.; Zhang, J.; Xie, H.; Zhang, B. Nutrient removal in constructed microcosm wetlands for treating polluted river water in northern China. Ecol. Eng. 2011, 37, 560–568. [Google Scholar] [CrossRef]
- Taylor, C.R.; Hook, P.B.; Stein, O.R.; Zabinski, C.A. Seasonal effects of 19 plant species on COD removal in subsurface treatment wetland microcosms. Ecol. Eng. 2011, 37, 703–710. [Google Scholar] [CrossRef]
- Calheiros, C.; Teixeira, A.; Pires, C.; Franco, A.R.; Duque, A.F.; Crispim, L.F.C.; Moura, S.C.; Castro, P.M.L. Bacterial community dynamics in horizontal flow constructed wetlands with different plants for high salinity industrial wastewater polishing. Water Res. 2010, 44, 5032–5038. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Zhang, J.; Wei, R.; Liang, S.; Li, C.; Xie, H. Nitrogen transformations and balance in constructed wetlands for slightly polluted river water treatment using different macrophytes. Environ. Sci. Pollut. Res. 2013, 20, 443–451. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Zhang, J.; Li, C.; Fan, J.; Zou, Y. Mass Balance Study on Phosphorus Removal in Constructed Wetland Microcosms Treating Polluted River Water. CLEAN Soil Air Water 2013, 41, 844–850. [Google Scholar] [CrossRef]
- Lin, Y.-F.; Jing, S.-R.; Wang, T.-W.; Lee, D.-Y. Effects of macrophytes and external carbon sources on nitrate removal from groundwater in constructed wetlands. Environ. Pollut. 2002, 119, 413–420. [Google Scholar] [CrossRef]
- Muyzer, G.; De Waal, E.C.; Uitterlinden, A.G. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl. Environ. Microbiol. 1993, 59, 695–700. [Google Scholar] [PubMed]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Xie, H.; Ngo, H.H.; Guo, W.; Zhang, J.; Liu, C.; Liang, S.; Hu, Z.; Yang, Z.; Zhao, C. Microbial abundance and community in subsurface flow constructed wetland microcosms: Role of plant presence. Environ. Sci. Pollut. Res. 2015. [Google Scholar] [CrossRef] [PubMed]
- Brix, H. Functions of macrophytes in constructed wetlands. Water Sci. Technol. 1994, 29, 71–78. [Google Scholar]
- Huang, J.; Cai, W.; Zhong, Q.; Wang, S. Influence of temperature on micro-environment, plant eco-physiology and nitrogen removal effect in subsurface flow constructed wetland. Ecol. Eng. 2013, 60, 242–248. [Google Scholar] [CrossRef]
- Armstrong, J.; Armstrong, W.; Beckett, P.M. Phragmites australis: Venturi-and humidity-induced pressure flows enhance rhizome aeration and rhizosphere oxidation. New Phytol. 1992, 120, 197–207. [Google Scholar] [CrossRef]
- Toet, S.; Bouwman, M.; Cevaal, A.; Verhoeven, J.T.A. Nutrient removal through autumn harvest of Phragmites australis and Thypha latifolia shoots in relation to nutrient loading in a wetland system used for polishing sewage treatment plant effluent. J. Environ. Sci. Health 2005, 40, 1133–1156. [Google Scholar] [CrossRef]
- Nikolausz, M.; Kappelmeyer, U.; Székely, A.; Rusznyák, A.; Márialigeti, K.; Kästner, M. Diurnal redox fluctuation and microbial activity in the rhizosphere of wetland plants. Eur. J. Soil Biol. 2008, 44, 324–333. [Google Scholar] [CrossRef]
- Overmann, J.; van Gemerden, H. Microbial interactions involving sulfur bacteria: Implications for the ecology and evolution of bacterial communities. FEMS Microbiol. Rev. 2000, 24, 591–599. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.-Y.; Franz, E.H.; Chen, S. Oxygen fluxes and ammonia removal efficiencies in constructed treatment wetlands. Water Environ. Res. 2001, 73, 661–666. [Google Scholar] [CrossRef] [PubMed]
- Breen, P.F. A mass balance method for assessing the potential of artificial wetlands for wastewater treatment. Water Res. 1990, 24, 689–697. [Google Scholar] [CrossRef]
- Reddy, K.R.; Debusk, W.F. Nutrient storage capabilities of aquatic and wetland plants. In Aquatic Plants for Wastewater Treatment & Resource Recovery; Magnolia Public Inc.: Orlando, FL, USA, 1987; pp. 337–353. [Google Scholar]
- Sawaittayothin, V.; Polprasert, C. Nitrogen mass balance and microbial analysis of constructed wetlands treating municipal landfill leachate. Bioresour. Technol. 2007, 98, 565–570. [Google Scholar] [CrossRef] [PubMed]
- Vymazal, J. Removal of nutrients in various types of constructed wetlands. Sci. Total Environ. 2007, 380, 48–65. [Google Scholar] [CrossRef] [PubMed]
- Gagnon, V.; Chazarenc, F.; Comeau, Y.; Brisson, J. Influence of macrophyte species on microbial density and activity in constructed wetlands. Water Sci. Technol. 2007, 56, 249–254. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Mitsuya, Y.; Gharizadeh, B.; Ronaghi, M.; Shafer, R.W. Characterization of mutation spectra with ultra-deep pyrosequencing: Application to HIV-1 drug resistance. Genome Res. 2007, 17, 1195–1201. [Google Scholar] [CrossRef] [PubMed]
- Subashchandrabose, S.R.; Ramakrishnan, B.; Megharaj, M.; Venkateswarlu, K.; Naidu, R. Consortia of cyanobacteria/microalgae and bacteria: Biotechnological potential. Biotechnol. Adv. 2011, 29, 896–907. [Google Scholar] [CrossRef] [PubMed]
- Cheng, W.; Zhang, J.; Wang, Z.; Wang, M.; Xie, S. Bacterial communities in sediments of a drinking water reservoir. Ann. Microbiol. 2014, 64, 875–878. [Google Scholar] [CrossRef]
- Herrmann, M.; Saunders, A.M.; Schramm, A. Archaea dominate the ammonia-oxidizing community in the rhizosphere of the freshwater macrophyte Littorella uniflora. Appl. Environ. Microbiol. 2008, 74, 3279–3283. [Google Scholar] [CrossRef] [PubMed]
- Bodelier, P.L.; Dedysh, S.N. Microbiology of wetlands. Front. Microbiol. 2013. [Google Scholar] [CrossRef]
- Araya, R.; Tani, K.; Takagi, T.; Yamaguchi, N.; Nasu, M. Bacterial activity and community composition in stream water and biofilm from an urban river determined by fluorescent in situ hybridization and DGGE analysis. FEMS Microbiol. Ecol. 2003, 43, 111–119. [Google Scholar] [CrossRef] [PubMed]
- Kirchman, D.L. The ecology of Cytophaga–Flavobacteria in aquatic environments. FEMS Microbiol. Ecol. 2002, 39, 91–100. [Google Scholar] [CrossRef]
- Williams, K.P.; Gillespie, J.J.; Sobral, B.W.; Nordberg, E.K.; Snyder, E.E.; Shallom, J.M.; Dickerman, A.W. Phylogeny of gammaProteobacteria. J. Bacteriol. 2010, 192, 2305–2314. [Google Scholar] [CrossRef] [PubMed]
- Ebeling, J.M.; Timmons, M.B.; Bisogni, J. Engineering analysis of the stoichiometry of photoautotrophic, autotrophic, and heterotrophic removal of ammonia–nitrogen in aquaculture systems. Aquaculture 2006, 257, 346–358. [Google Scholar] [CrossRef]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, Z.; Wang, Q.; Zhang, J.; Xie, H.; Feng, S. Effect of Plant Harvesting on the Performance of Constructed Wetlands during Summer. Water 2016, 8, 24. https://doi.org/10.3390/w8010024
Yang Z, Wang Q, Zhang J, Xie H, Feng S. Effect of Plant Harvesting on the Performance of Constructed Wetlands during Summer. Water. 2016; 8(1):24. https://doi.org/10.3390/w8010024
Chicago/Turabian StyleYang, Zhongchen, Qian Wang, Jian Zhang, Huijun Xie, and Suping Feng. 2016. "Effect of Plant Harvesting on the Performance of Constructed Wetlands during Summer" Water 8, no. 1: 24. https://doi.org/10.3390/w8010024
APA StyleYang, Z., Wang, Q., Zhang, J., Xie, H., & Feng, S. (2016). Effect of Plant Harvesting on the Performance of Constructed Wetlands during Summer. Water, 8(1), 24. https://doi.org/10.3390/w8010024





