Sequential Anaerobic/Aerobic Microbial Transformation of Chlorinated Ethenes: Use of Sustainable Approaches for Aquifer Decontamination
Abstract
:1. Introduction
2. Chlorinated Ethenes
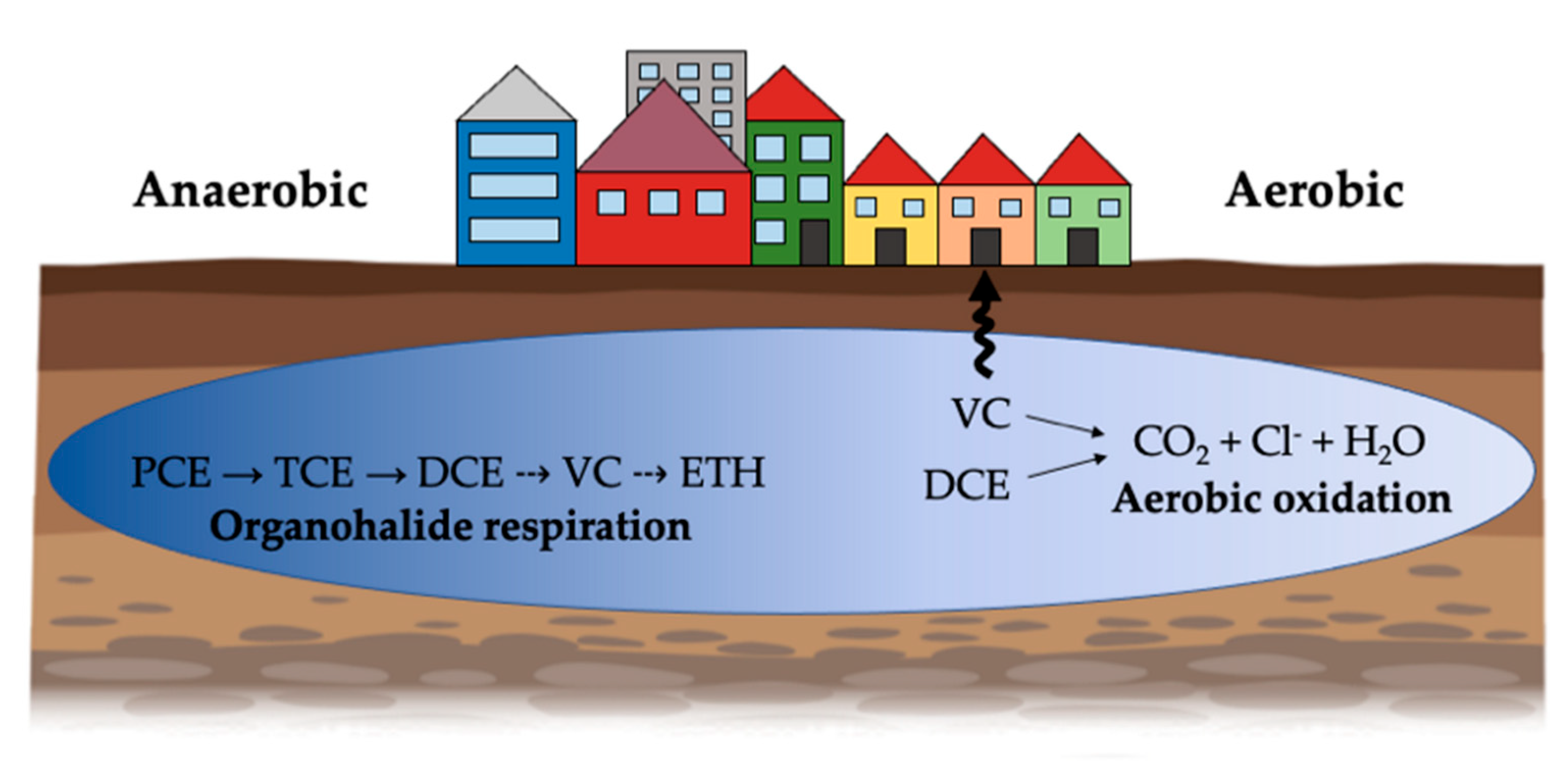
3. Microbial Transformation of Chlorinated Ethenes
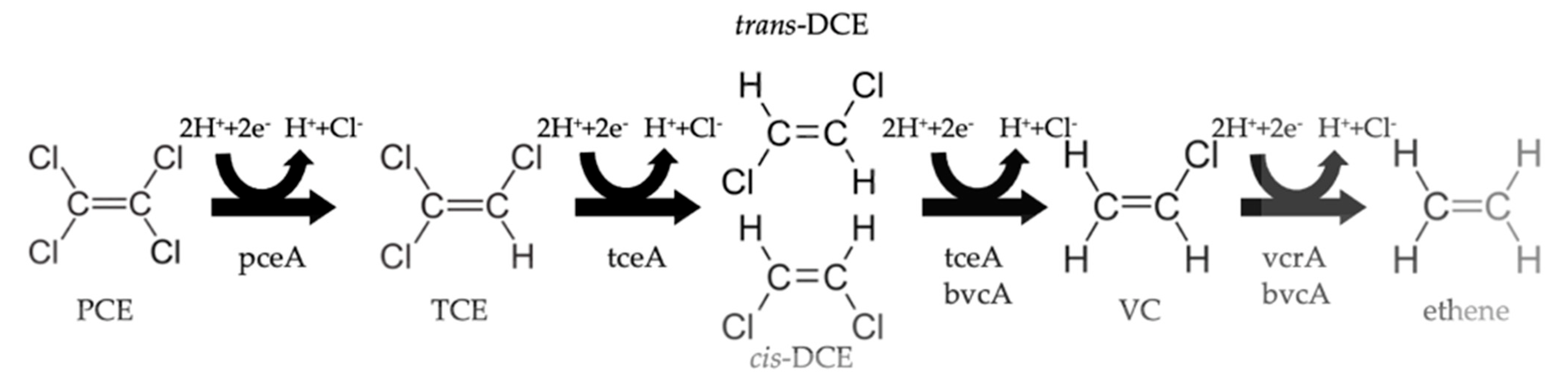
4. Organohalide Respiration
5. Aerobic Pathways of Lower Chlorinated Ethenes Biodegradation
5.1. DCE and VC Metabolic Degradation
5.2. Co-Metabolic Aerobic Degradation
6. Bio-Based Substrates for Stimulation of OHR Bacteria
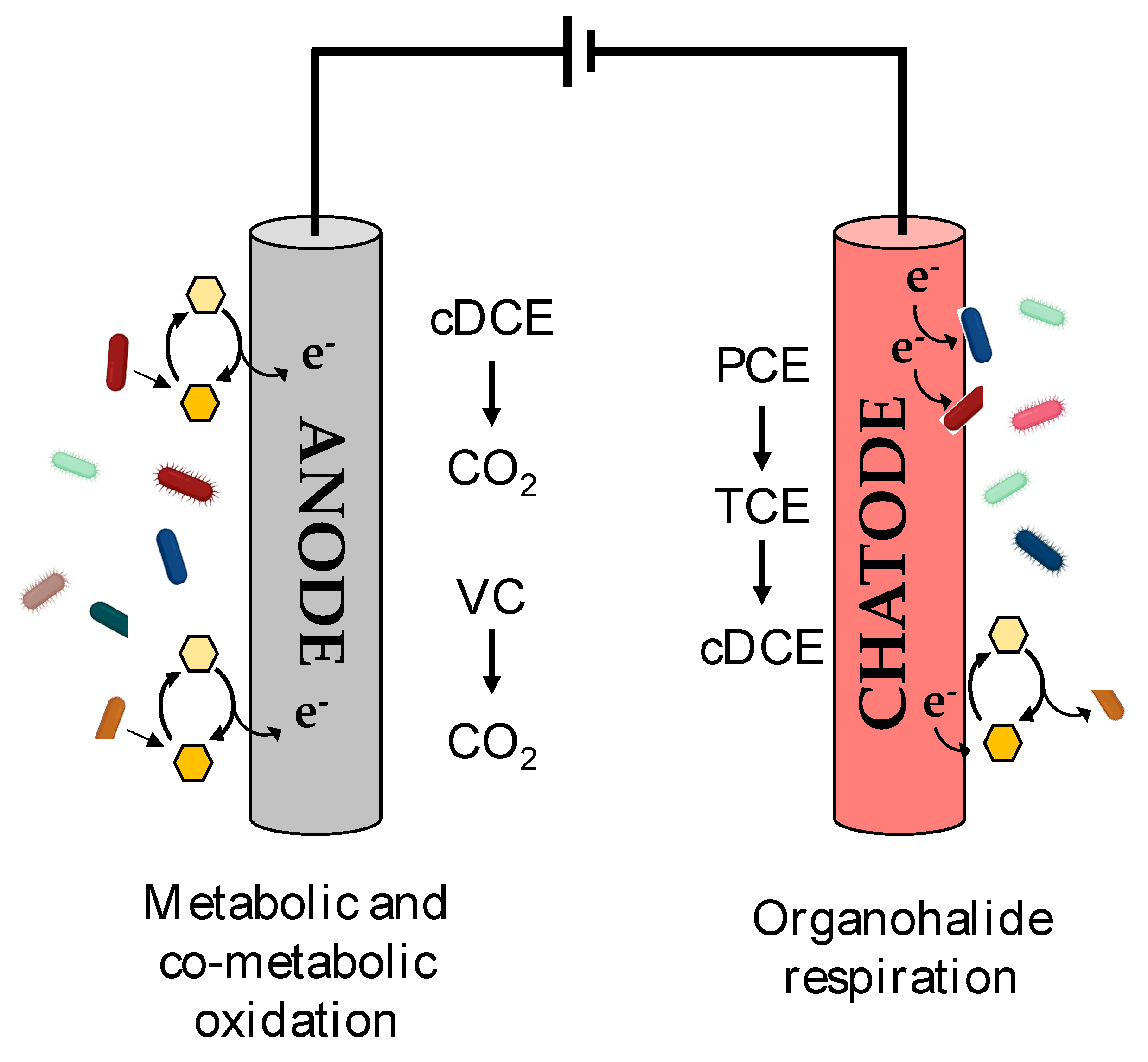
7. Bioelectrochemical Systems
8. Environmental Halogenomics: Detection and Distribution of Halo-Bacteria
9. Conclusions
Funding
Conflicts of Interest
Abbreviations
| 1,1-DCE | 1,1-trans-dichloroethene |
| 1,2-DCE | 1,2-trans-dichloroethene |
| AkMO | alkene monooxygenase |
| BES | bioelectrochemical systems |
| bvcA | vinyl chloride reductase |
| CEs | chloroethenes |
| cis-DCE | cis-dichloroethene |
| D-NAPL | dense non-aqueous phase liquid |
| EaCoMT | epoxyalkane:coenzyme M transferase |
| GSH | glutathione |
| GST | glutathione S-transferase |
| MMO | methanol monooxygenase |
| nZVI | nano zero valent iron |
| OHR | organohalide respiration |
| OHRB | Organohalide respiring bacteria |
| ORP | oxidative redox potential |
| PCE | tetrachloroethene |
| pceA | tetrachloroethene reductive dehalogenase |
| pMMO | particulate MMO |
| RDases | reductive dehalogenase homologous genes |
| sMMO | soluble MMO |
| TCE | trichloroethene |
| tceA | trichloroethene reductive dehalogenase |
| VC | vinyl chloride |
| vcrA | vinyl chloride reductase |
References
- Available online: https://www.fao.org/sustainable-development-goals/en/ (accessed on 20 September 2022).
- Hasan, M.M. Ground Water Making the Invisible Visible. Legal Lock J. 2021, 1, 69. [Google Scholar]
- Foster, S.; Gathu, J.; Eichholz, M.; Hirata, R. Climate Change: The utility groundwater role in supply security. Source 2020, 18, 50–54. [Google Scholar]
- Margat, J.; Van der Gun, J. Groundwater around the World: A Geographic Synopsis; CRC Press: Boca Raton, FL, USA, 2013. [Google Scholar]
- UNESCO. The United Nations World Water Development Report 2015; UNESCO: Paris, France, 2015. [Google Scholar]
- Available online: https://www.eea.europa.eu/data-and-maps/indicators/progress-in-management-of-contaminated-sites-3/assessment (accessed on 20 September 2022).
- Available online: https://www.eea.europa.eu/data-and-maps/data/wise-wfd-4 (accessed on 20 September 2022).
- Majone, M.; Verdini, R.; Aulenta, F.; Rossetti, S.; Tandoi, V.; Kalogerakis, N.; Agathos, S.; Puig, S.; Zanaroli, G.; Fava, F. In situ groundwater and sediment bioremediation: Barriers and perspectives at European contaminated sites. New Biotechnol. 2015, 32, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Battelle Memorial Institute. Permeable Reactive Barrier Cost and Performance Report; NAVFAC: Port Hueneme, CA, USA, 2012. [Google Scholar]
- Das, S.; Dash, H.R. Microbial bioremediation: A potential tool for restoration of contaminated areas. In Microbial Biodegradation and Bioremediation; Elsevier: Waltham, MA, USA, 2014; pp. 1–21. [Google Scholar]
- Narayanan, M.; Ali, S.S.; El-Sheekh, M. A comprehensive review on the potential of microbial enzymes in multipollutant bioremediation: Mechanisms, challenges, and future prospects. J. Environ. Manag. 2023, 334, 117532. [Google Scholar] [CrossRef] [PubMed]
- Abrahamsson, K.; Ekdahl, A.; Collen, J.; Pedersen, M. Marine algae-a source of trichloroethylene and perchloroethylene. Limnol. Oceanogr. 1995, 40, 1321–1326. [Google Scholar] [CrossRef]
- Field, J.A. Natural production of organohalide compounds in the environment. In Organohalide-Respiring Bacteria; Springer: Berlin/Heidelberg, Germany, 2016; pp. 7–29. [Google Scholar]
- Keppler, F.; Borchers, R.; Pracht, J.; Rheinberger, S.; Scholer, H.F. Natural formation of vinyl chloride in the terrestrial environment. Environ. Sci. Technol. 2002, 36, 2479–2483. [Google Scholar] [CrossRef]
- Öberg, G.M. The Biogeochemistry of Chlorine in Soil. Handb. Environ. Chem. 2004, 3, 43–62. [Google Scholar] [CrossRef]
- McCarty, P.L. Groundwater contamination by chlorinated solvents: History, remediation technologies and strategies. In In Situ Remediation of Chlorinated Solvent Plumes; Springer: New York, NY, USA, 2010; pp. 1–28. [Google Scholar]
- Moran, M.J.; Zogorski, J.S.; Squillace, P.J. Chlorinated solvents in groundwater of the United States. Environ. Sci. Technol. 2007, 41, 74–81. [Google Scholar] [CrossRef]
- Beamer, P.I.; Luik, C.E.; Abrell, L.; Camposc, S.; Martínez, M.E.; Sáez, A.E. Concentration of Trichloroethylene in Breast Milk and Household Water from Nogales, Arizona. Environ. Microbiol. 2012, 46, 9055–9061. [Google Scholar] [CrossRef] [Green Version]
- Huang, B.; Lei, C.; Wei, C.; Zeng, G. Chlorinated volatile organic compounds (Cl-VOCs) in environment—Sources, potential human health impacts, and current remediation technologies. Environ. Int. 2014, 71, 118–138. [Google Scholar] [CrossRef]
- USA EPA. Toxicity and Exposure Assessment for Children’s Health. Trichloroethylene—TEACH Chemical Summary. 2007. Available online: http://www.epa.gov/teach/chem_summ/TCE_summary.pdf. (accessed on 17 October 2022).
- IARC. Vinyl Chloride; International Agency for Research on Cancer: Ottawa, ON, Canada, 1987. [Google Scholar]
- IARC Monographs on the Identification of Carcinogenic Hazards to Humans. 2020, Volume 1–127. Available online: https://monographs.iarc.fr/list-of-classifications. (accessed on 5 September 2022).
- Lynge, E.; Anttila, A.; Hemminki, K. Organic solvents and cancer. Cancer Causes Control 1992, 55, 1353–1395. [Google Scholar]
- Bradley, P.M. History and ecology of chlororethene biodegradation: A review. Bioremediat. J. 2003, 7, 81–109. [Google Scholar] [CrossRef]
- Mattes, T.E.; Alexander, A.K.; Coleman, N.V. Aerobic biodegradation of the chloroethenes: Pathways, enzymes, ecology, and evolution. FEMS Microbiol. Rev. 2010, 34, 445–475. [Google Scholar] [CrossRef] [Green Version]
- Tiehm, A.; Schmidt, K.R. Sequential anaerobic/aerobic biodegradation of chloroethenes-aspects of field application. Curr. Opin. Biotechnol. 2011, 22, 415–421. [Google Scholar] [CrossRef]
- Anam, G.B.; Choi, J.; Ahn, Y. Reductive dechlorination of perchloroethene (PCE) and bacterial community changes in a continuous-flow, two-stage anaerobic column. Int. Biodeter. Biodegr. 2019, 138, 41–49. [Google Scholar] [CrossRef]
- Chen, S.K.; Yang, H.Y.; Huang, S.R.; Hung, J.M.; Lu, C.J.; Liu, M.H. Complete degradation of chlorinated ethenes and its intermediates through sequential anaerobic/aerobic biodegradation in simulated groundwater columns (complete degradation of chlorinated ethenes). IJEST 2020, 17, 4517–4530. [Google Scholar] [CrossRef]
- Leys, D.; Adrian, L.; Smidt, H. Organohalide respiration: Microbes breathing chlorinated molecules. Proc. R. Soc. B 2013, 368, 20120316. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Atashgahi, S.; Bosma, T.N.; Peng, P.; Smidt, H. Organohalide respiration potential in marine sediments from Aarhus Bay. FEMS Microbiol. 2022, 98, fiac073. [Google Scholar] [CrossRef]
- Maymo-Gatell, X. “Dehalococcoides Ethenogenes” Strain 195: A Novel Eubacterium that Reductively Dechlorinates Tetrachloroethene (PCE) to Ethene; Cornell University: Ithaca, NY, USA, 1997. [Google Scholar]
- Yang, Y.; Higgins, S.A.; Yan, J.; Şimşir, B.; Chourey, K.; Iyer, R.; Hettich, R.L.; Baldwin, B.; Oglen, D.M.; Löffler, F.E. Grape pomace compost harbors organohalide-respiring Dehalogenimonas species with novel reductive dehalogenase genes. ISME J. 2017, 11, 2767–2780. [Google Scholar] [CrossRef] [Green Version]
- Chen, G.; Kara Murdoch, F.; Xie, Y.; Murdoch, R.W.; Cui, Y.; Yang, Y.; Yan, Y.; Key, T.A.; Löffler, F.E. Dehalogenation of Chlorinated Ethenes to Ethene by a Novel Isolate, “Candidatus Dehalogenimonas etheniformans”. Appl. Environ. Microbiol. 2022, e00443-22. [Google Scholar] [CrossRef]
- Cichocka, D.; Nikolausz, M.; Haest, P.J.; Nijenhuis, I. Tetrachloroethene conversion to ethene by a Dehalococcoides-containing enrichment culture from Bitterfeld. FEMS Microbiol. Ecol. 2010, 72, 297–310. [Google Scholar] [CrossRef] [Green Version]
- AFCEE. Principles and Practices of Enhanced Anaerobic Bioremediation of Chlorinated Solvents; Department of Defense, Air Force Center for Environmental Excellence and the Environmental Security Technology Certification Program (ESTCP): Washington, DC, USA, 2004. [Google Scholar]
- Heimann, A.C.; Batstone, D.J.; Jakobsen, R. Methanosarcina spp. drive vinyl chloride dechlorination via interspecies hydrogen transfer. Appl. Environ. Microbiol. 2006, 72, 2942–2949. [Google Scholar] [CrossRef] [Green Version]
- Smidt, H.; de Vos, W.M. Anaerobic Microbial Dehalogenation. Annu. Rev. Microbiol. 2004, 58, 43–73. [Google Scholar] [CrossRef]
- Futagami, T.; Goto, M.; Furukawa, K. Biochemical and genetic bases of dehalorespiration. Chem. Rec. 2008, 8, 1–12. [Google Scholar] [CrossRef]
- Abe, Y.; Aravena, R.; Zopfi, J.; Parker, B.; Hunkeler, D. Evaluating the fate of chlorinated ethenes in streambed sediments by combining stable isotope, geochemical and microbial methods. J. Contam. Hydrol. 2009, 107, 10–21. [Google Scholar] [CrossRef] [Green Version]
- Hug, L.A.; Maphosa, F.; Leys, D.; Löffler, F.E.; Smidt, H.; Edwards, E.A.; Adrian, L. Overview of organohalide-respiring bacteria and a proposal for a classification system for reductive dehalogenases. Phil. Trans. R. Soc. B 2013, 368, 20120322. [Google Scholar] [CrossRef] [Green Version]
- Yan, J.; Wang, J.; Villalobos Solis, M.I.; Jin, H.; Chourey, K.; Li, X.; Yang, Y.; Yin, Y.; Hettich, R.L.; Loffler, F.E. Respiratory vinyl chloride reductive dechlorination to ethene in TceA-expressing Dehalococcoides mccartyi. Environ. Sci. Technol. 2021, 55, 4831–4841. [Google Scholar] [CrossRef]
- Men, Y.; Feil, H.; VerBerkmoes, N.C.; Shah, M.B.; Johnson, D.R.; Lee, P.K.; West, K.A.; Zinder, S.H.; Andersen, G.L.; Alvarez-Cohen, L. Sustainable syntrophic growth of Dehalococcoides ethenogenes strain 195 with Desulfovibrio vulgaris Hildenborough and Methanobacterium congolense: Global transcriptomic and proteomic analyses. ISME J. 2012, 6, 410–421. [Google Scholar] [CrossRef]
- Teng, Y.; Xu, Y.; Wang, X.; Christie, P. Function of biohydrogen metabolism and related microbial communities in environmental bioremediation. Front. Microbiol. 2019, 10, 106. [Google Scholar] [CrossRef] [Green Version]
- Ziv-El, M.; Delgado, A.G.; Yao, Y.; Kang, D.W.; Nelson, K.G.; Halden, R.U.; Krajmalnik-Brown, R. Development and characterization of DehaloR^2, a novel anaerobic microbial consortium performing rapid dechlorination of TCE to ethene. Appl. Microbiol. Biotechnol. 2011, 92, 1063–1071. [Google Scholar] [CrossRef]
- Maphosa, F.; van Passel, M.W.; de Vos, W.M.; Smidt, H. Metagenome analysis reveals yet unexplored reductive dechlorinating potential of Dehalobacter sp. E1 growing in co-culture with Sedimentibacter sp. Environ. Microbiol. Rep. 2012, 4, 604–616. [Google Scholar] [PubMed]
- Ziv-El, M.; Popat, S.C.; Parameswaran, P.; Kang, D.W.; Polasko, A.; Halden, R.U.; Rittmann, B.E.; Krajmalnik-Brown, R. Using electron balances and molecular techniques to assess trichoroethene-induced shifts to a dechlorinating microbial community. Biotechnol. Bioeng. 2012, 109, 2230–2239. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Ritalahti, K.M.; Wagner, D.D.; Löffler, F.E. Unexpected specificity of interspecies cobamide transfer from Geobacter spp. to organohalide-respiring Dehalococcoides mccartyi strains. Appl. Environ. Microbiol. 2012, 78, 6630–6636. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yan, J.; Im, J.; Yang, Y.; Löffler, F.E. Guided cobalamin biosynthesis supports Dehalococcoides mccartyi reductive dechlorination activity. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2013, 368, 20120320. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Men, Y.; Lee, P.K.; Harding, K.C.; Alvarez-Cohen, L. Characterization of four TCE-dechlorinating microbial enrichments grown with different cobalamin stress and methanogenic conditions. Appl. Microbiol. Biotechnol. 2013, 97, 6439–6450. [Google Scholar] [CrossRef]
- Adrian, L.; Löffler, F.E. Organohalide-Respiring Bacteria; Springer: Berlin/Heidelberg, Germany, 2016; Volume 85. [Google Scholar]
- Richardson, R.E. Organohalide-respiring bacteria as members of microbial communities: Catabolic food webs and biochemical interactions. In Organohalide-Respiring Bacteria; Springer: Berlin/Heidelberg, Germany, 2016; pp. 309–341. [Google Scholar]
- Fennell, D.E.; Gossett, J.M.; Zinder, S.H. Comparison of butyric acid, ethanol, lactic acid, and propionic acid as hydrogen donors for the reductive dechlorination of tetrachloroethene. Environ. Sci. Technol. 1997, 31, 918–926. [Google Scholar] [CrossRef]
- Matteucci, F.; Ercole, C.; Del Gallo, M. A study of chlorinated solvent contamination of the aquifers of an industrial area in central Italy: A possibility of bioremediation. Front. Microbiol. 2015, 6, 924. [Google Scholar] [CrossRef]
- Yang, Y.; McCarty, P.L. Comparison between donor substrates for biologically enhanced tetrachloroethene DNAPL dissolution. Environ. Sci. Technol. 2002, 36, 3400–3404. [Google Scholar] [CrossRef]
- Dolfing, J. Energetic considerations in organohalide respiration. In Organohalide-Respiring Bacteria; Springer: Berlin/Heidelberg, Germany, 2016; pp. 31–48. [Google Scholar]
- Lin, W.H.; Chien, C.C.; Lu, C.W.; Hou, D.; Sheu, Y.T.; Chen, S.C.; Kao, C.M. Growth inhibition of methanogens for the enhancement of TCE dechlorination. Sci. Total Environ. 2021, 787, 147648. [Google Scholar] [CrossRef]
- Yang, Y.; McCarty, P.L. Competition for hydrogen within a chlorinated solvent dehalogenating anaerobic mixed culture. Environ. Sci. Technol. 1998, 32, 3591–3597. [Google Scholar] [CrossRef]
- Robinson, C.; Barry, D.A.; McCarty, P.L.; Gerhard, J.I.; Kouznetsova, I. pH control for enhanced reductive bioremediation of chlorinated solvent source zones. Sci. Total Environ. 2009, 407, 4560–4573. [Google Scholar] [CrossRef] [Green Version]
- Schmidt, K.R.; Gaza, S.; Voropaev, A.; Ertl, S.; Tiehm, A. Aerobic biodegradation of trichloroethene without auxiliary substrates. Water Res. 2014, 59, 112–118. [Google Scholar] [CrossRef]
- Jennings, L.K.; Chartrand, M.M.G.; Lacrampe-Couloume, G.; Lollar, B.S.; Spain, J.C.; Gossett, J.M. Proteomic and transcriptomic analyses reveal genes upregulated by cis-dichloroethene in Polaromonas sp. Strain JS666. Appl. Environ. Microbiol. 2009, 75, 3733–3744. [Google Scholar] [CrossRef] [Green Version]
- Coleman, N.V.; Mattes, T.E.; Gossett, J.M.; Spain, J.C. Biodegradation of cis-dichloroethene as the sole carbon source by a β-Proteobacterium. Appl. Environ. Microbiol. 2002, 68, 2726–2730. [Google Scholar] [CrossRef] [Green Version]
- Mattes, T.E.; Alexander, A.K.; Richardson, P.M.; Munk, A.C.; Han, C.S.; Stothard, P.; Coleman, N.V. The genome of Polaromonas sp. strain JS666: Insights into the evolution of a hydrocarbon- and xenobiotic-degrading bacterium, and features of relevance to biotechnology. Appl. Environ. Microbiol. 2008, 74, 6405–6416. [Google Scholar] [CrossRef] [Green Version]
- Coleman, N.V.; Spain, J.C. Distribution of the Coenzyme M Pathway of Epoxide Metabolism among Ethene- and Vinyl Chloride-Degrading Mycobacterium Strains. Appl. Environ. Microbiol. 2003, 69, 6041–6046. [Google Scholar] [CrossRef] [Green Version]
- Xing, Z.; Su, X.; Zhang, X.; Zhang, L.; Zhao, T. Direct aerobic oxidation (DAO) of chlorinated aliphatic hydrocarbons: A review of key DAO bacteria, biometabolic pathways and in-situ bioremediation potential. Environ. Int. 2022, 162, 107165. [Google Scholar] [CrossRef] [PubMed]
- Mattes, T.E.; Coleman, N.V.; Spain, J.C.; Gossett, J.M. Physiological and molecular genetic analyses of vinyl chloride and ethene biodegradation in Nocardioides sp. strain JS614. Arch. Microbiol. 2005, 183, 95–106. [Google Scholar] [CrossRef]
- Coleman, N.V.; Mattes, T.E.; Gossett, J.M.; Spain, J.C. Phylogenetic and kinetic diversity of aerobic vinyl chloride-assimilating bacteria from contaminated sites. Appl. Environ. Microbiol. 2002, 68, 6162–6171. [Google Scholar] [CrossRef] [Green Version]
- Jin, Y.O.; Cheung, S.; Coleman, N.V.; Mattes, T.E. Association of missense mutations in epoxyalkane coenzyme M transferase with adaptation of Mycobacterium sp. Strain JS623 to growth on vinyl chloride. Appl. Environ. Microbiol. 2010, 76, 3413–3419. [Google Scholar] [CrossRef] [Green Version]
- Liu, X.; Wu, Y.; Wilson, F.P.; Yu, K.; Lintner, C.; Cupples, A.M.; Mattes, T.E. Integrated methodological approach reveals microbial diversity and functions in aerobic groundwater microcosms adapted to vinyl chloride. FEMS Microbiol. Ecol. 2018, 94, fiy124. [Google Scholar] [CrossRef] [PubMed]
- Hartmans, S.; De Bont, J.; Tramper, J.; Luyben, K.C.A. Bacterial degradation of vinyl chloride. Biotechnol. Lett. 1985, 7, 383–388. [Google Scholar] [CrossRef]
- Malachowsky, K.; Phelps, T.; Teboli, A.; Minnikin, D.; White, D. Aerobic mineralization of trichloroethylene, vinyl chloride and aromatic compounds by Rhodococcus species. Appl. Environ. Microb. 1994, 60, 542–548. [Google Scholar] [CrossRef] [Green Version]
- Verce, M.F.; Ulrich, R.L.; Freedman, D.L. Characterization of an isolate that uses vinyl chloride as a growth substrate under aerobic conditions. Appl. Environ. Microbiol. 2000, 66, 3535–3542. [Google Scholar] [CrossRef] [Green Version]
- Verce, M.F.; Ulrich, R.L.; Freedman, D.L. Transition from cometabolic to growth-linked biodegradation of vinyl chloride by a Pseudomonas sp. isolated on ethene. Environ. Sci. Technol. 2001, 35, 4242–4251. [Google Scholar] [CrossRef]
- Danko, A.S.; Luo, M.; Bagwell, C.E.; Brigmon, R.L.; Freedman, D.L. Involvement of linear plasmids in aerobic biodegradation of vinyl chloride. Appl. Environ. Microbiol. 2004, 70, 6092–6097. [Google Scholar] [CrossRef] [Green Version]
- Elango, V.K.; Liggenstoffer, A.S.; Fathepure, B.Z. Biodegradation of vinyl chloride and cis-dichloroethene by a Ralstonia sp. strain TRW-1. Appl. Microbiol. Biotechnol. 2006, 72, 1270–1275. [Google Scholar] [CrossRef]
- Paes, F.; Liu, X.; Mattes, T.E.; Cupples, A.M. Elucidating carbon uptake from vinyl chloride using stable isotope probing and Illumina sequencing. Appl. Microbiol. Biotechnol. 2015, 99, 7735–7743. [Google Scholar] [CrossRef]
- Gupta, R.S.; Lo, B.; Son, J. Phylogenomic and comparative genomic studies robustly support division of the genus Mycobacterium into an emended genus Mycobacterium and four novel genera. Front. Microbiol. 2018, 9, 67. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547. [Google Scholar] [CrossRef]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [Green Version]
- Jones, D.T.; Taylor, W.R.; Thornton, J.M. The rapid generation of mutation data matrices from protein sequences. Comput. Appl. Biosci. 1992, 8, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Richards, P.M.; Ewald, J.M.; Zhao, W.; Rectanus, H.; Fan, D.; Durant, N.; Pound, M.; Mattes, T.E. Natural biodegradation of vinyl chloride and cis-dichloroethene in aerobic and suboxic conditions. ESPR 2022, 1–14. [Google Scholar] [CrossRef]
- Fullerton, H.; Rogers, R.; Freedman, D.L.; Zinder, S.H. Isolation of an aerobic vinyl chloride oxidizer from anaerobic groundwater. Biodegradation 2014, 25, 893–901. [Google Scholar] [CrossRef]
- Zhao, H.P.; Schmidt, K.R.; Tiehm, A. Inhibition of aerobic metabolic cis-1,2-dichloroethene biodegradation by other chloroethenes. Water Res. 2010, 44, 2276–2282. [Google Scholar] [CrossRef]
- Dolinová, I.; Štrojsová, M.; Černík, M.; Němeček, J.; Macháčková, J.; Ševců, A. Microbial degradation of chloroethenes: A review. ESPR 2017, 24, 13262–13283. [Google Scholar] [CrossRef]
- Lange, C.C.; Wackett, L.P. Oxidation of aliphatic olefins by toluene dioxygenase: Enzyme rates and product identification. J. Bacteriol. 1997, 179, 3858–3865. [Google Scholar] [CrossRef] [Green Version]
- Doughty, D.M.; Sayavedra-Soto, L.A.; Arp, D.J.; Bottomley, P.J. Effects of dichloroethene isomers on the induction and activity of butane monooxygenase in the alkane-oxidizing bacterium ‘Pseudomonas butanovora’. Appl. Environ. Microbiol. 2005, 71, 6054–6059. [Google Scholar] [CrossRef] [Green Version]
- Ryoo, D.; Shim, H.; Canada, K.; Barbieri, P.; Wood, T.K. Aerobic degradation of tetrachloroethylene by toluene-o-xylene monooxygenase of Pseudomonas stutzeri OX1, Nat. Biotechnol. 2000, 18, 775–778. [Google Scholar] [CrossRef]
- Shokrollahzadeh, S.; Azizmohseni, F.; Golmohamad, F. Characterization and kinetic study of PAH–degrading Sphingopyxis ummariensis bacteria isolated from a petrochemical wastewater treatment plant. Adv. Environ. Sci. Technol. 2015, 1, 1–9. [Google Scholar] [CrossRef]
- Frascari, D.; Pinelli, D.; Nocentini, M.; Baleani, E.; Cappelletti, M.; Fedi, S. A kinetic study of chlorinated solvent cometabolic biodegradation by propane-grown Rhodococcus sp. PB1. Biochem. Eng. J. 2008, 42, 139–147. [Google Scholar] [CrossRef]
- Zalesak, M.; Ruzicka, J.; Vicha, R.; Dvorackova, M. Cometabolic degradation of dichloroethenes by Comamonas testosteroni RF2. Chemosphere 2017, 186, 919–927. [Google Scholar] [CrossRef] [PubMed]
- Bowman, J.P.; Jiménez, L.; Rosario, I.; Hazen, T.C.; Sayler, G.S. Characterization of the methanotrophic bacterial community present in a trichloroethylene-contaminated subsurface groundwater site. Appl. Environ. Microbiol. 1993, 59, 2380–2387. [Google Scholar] [CrossRef] [Green Version]
- Yoon, S.; Im, J.; Bandow, N.; Dispirito, A.A.; Semrau, J.D. Constitutive expression of pMMO by Methylocystis strain SB2 when grown on multi-carbon substrates: Implications for biodegradation of chlorinated ethenes. Environ. Microbiol. Rep. 2011, 3, 182–188. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.W.; Keeney, D.R.; Lim, D.H.; Dispirito, A.A.; Semrau, J.D. Mixed pollutant degradation by Methylosinus trichosporium OB3b expressing either soluble or particulate methane monooxygenase: Can the tortoise beat the hare. Appl. Environ. Microbiol. 2006, 72, 7503–7509. [Google Scholar] [CrossRef] [Green Version]
- Dedysh, S.N.; Knief, C.; Dunfield, P.F. Methylocella species are facultatively methanotrophic. J. Bacteriol. 2005, 187, 4665–4670. [Google Scholar] [CrossRef] [Green Version]
- Dunfield, P.F.; Belova, S.E.; Vorob’ev, A.V.; Cornish, S.L.; Dedysh, S.N. Methylocapsa aurea sp. nov., a facultatively methanotrophic bacterium possessing a particulate methane monooxygenase. Int. J. Syst. Evol. Microbiol. 2010, 60, 2659–2664. [Google Scholar] [CrossRef] [Green Version]
- Im, J.; Lee, S.W.; Yoon, S.; DiSpirito, A.A.; Semrau, J.D. Characterization of a novel facultative Methylocystis species capable of growth on methane, ethanol, and acetate. Environ. Microbiol. Rep. 2010. [Google Scholar] [CrossRef]
- Freedman, D.L.; Danko, A.S.; Verce, M.F. Substrate interactions during aerobic biodegradation of methane, ethene, vinyl chloride and 1,2-dichloroethenes. Water Sci. Technol. 2001, 43, 333–340. [Google Scholar] [CrossRef] [Green Version]
- Findlay, M.; Smoler, D.F.; Fogel, S.; Mattes, T.E. Aerobic vinyl chloride metabolism in groundwater microcosms by methanotrophic and etheneotrophic bacteria. Environ. Sci. Technol. 2016, 50, 3617–3625. [Google Scholar] [CrossRef]
- Chen, W.Y.; Wu, J.H. Microbiome composition resulting from different substrates influences trichloroethene dechlorination performance. J. Environ. Manag. 2022, 303, 114145. [Google Scholar] [CrossRef]
- Conrad, M.E.; Brodie, E.L.; Radtke, C.W.; Bill, M.; Delwiche, M.E.; Lee, M.H.; Swift, D.L.; Colwell, F.S. Field evidence for co-metabolism of trichloroethene stimulated by addition of electron donor to groundwater. Environ. Sci. Technol. 2010, 44, 4697–4704. [Google Scholar] [CrossRef] [Green Version]
- Steffan, R.J.; Schaefer, C.E. Current and future bioremediation applications: Bioremediation from a practical and regulatory perspective. In Organohalide-Respiring Bacteria; Springer: Berlin/Heidelberg, Germany, 2016; pp. 517–540. [Google Scholar]
- Blazquez-Palli, N.; Rosell, M.; Varias, J.; Bosch, M.; Soler, A.; Vicent, T.; Marco-Urrea, E. Multi-method assessment of the intrinsic biodegradation potential of an aquifer contaminated with chlorinated ethenes at an industrial area in Barcelona (Spain). Environ. Pollut. 2019, 244, 165–173. [Google Scholar] [CrossRef]
- Li, J.; Hu, A.; Bai, S.; Yang, X.; Sun, Q.; Liao, X.; Yu, C.P. Characterization and performance of lactate-feeding consortia for reductive dechlorination of trichloroethene. Microorganisms 2021, 9, 751. [Google Scholar] [CrossRef]
- Tomita, R.; Yoshida, N.; Meng, L. Formate: A promising electron donor to enhance trichloroethene-to-ethene dechlorination in Dehalococcoides-augmented groundwater ecosystems with minimal bacterial growth. Chemosphere 2022, 307, 136080. [Google Scholar] [CrossRef]
- Sheu, Y.T.; Tsang, D.C.; Dong, C.D.; Chen, C.W.; Luo, S.G.; Kao, C.M. Enhanced bioremediation of TCE-contaminated groundwater using gamma poly-glutamic acid as the primary substrate. J. Clean. Prod. 2018, 178, 108–118. [Google Scholar] [CrossRef]
- Fu, F.; Dionysiou, D.D.; Liu, H. The use of zero-valent iron for groundwater remediation and wastewater treatment: A review. J. Hazard Mater. 2014, 267, 194–205. [Google Scholar] [CrossRef]
- Summer, D.; Schöftner, P.; Wimmer, B.; Pastar, M.; Kostic, T.; Sessitsch, A.; Gerzabeck, M.H.; Reichenauer, T.G. Synergistic effects of microbial anaerobic dechlorination of perchloroethene and nano zero-valent iron (nZVI)–A lysimeter experiment. New Biotechnol. 2020, 57, 34–44. [Google Scholar] [CrossRef]
- Matturro, B.; Pierro, L.; Frascadore, E.; Petrangeli Papini, M.; Rossetti, S. Microbial community changes in a chlorinated solvents polluted aquifer over the field scale treatment with poly-3-hydroxybutyrate as amendment. Front. Microbiol. 2018, 9, 1664. [Google Scholar] [CrossRef] [Green Version]
- Bertolini, M.; Zecchin, S.; Beretta, G.P.; De Nisi, P.; Ferrari, L.; Cavalca, L. Effectiveness of permeable reactive bio-barriers for bioremediation of an organohalide-polluted aquifer by natural-occurring microbial community. Water 2021, 13, 2442. [Google Scholar] [CrossRef]
- Masut, E.; Battaglia, A.; Ferioli, L.; Legnani, A.; Cruz Viggi, C.; Tucci, M.; Resitano, M.; Milani, A.; de Laurentiis, C.; Matturro, B.; et al. A microcosm treatability study for evaluating wood mulch-based amendments as electron donors for trichloroethene (TCE) reductive dechlorination. Water 2021, 13, 1949. [Google Scholar] [CrossRef]
- Öztürk, Z.; Tansel, B.; Katsenovich, Y.; Sukop, M.; Laha, S. Highly organic natural media as permeable reactive barriers: TCE partitioning and anaerobic degradation profile in eucalyptus mulch and compost. Chemosphere 2012, 89, 665–671. [Google Scholar] [CrossRef] [PubMed]
- McLean, J.E.; Ervin, J.; Zhou, J.; Sorensen, D.L.; Dupont, R.R. Biostimulation and bioaugmentation to enhance reductive dechlorination of tce in a long-term flow through column study. Ground Water Monit. Remediat. 2015, 35, 76–88. [Google Scholar] [CrossRef]
- Mora, R.H.; Macbeth, T.W.; MacHarg, T.; Gundarlahalli, J.; Holbrook, H.; Schiff, P. Enhanced bioremediation using whey powder for a trichloroethene plume in a high-sulfate, fractured granitic aquifer. Remediat. J. J. Environ. Cleanup Costs Technol. Tech. 2008, 18, 7–30. [Google Scholar] [CrossRef]
- Robles, A.; Yellowman, T.L.; Joshi, S.; Mohana Rangan, S.; Delgado, A.G. Microbial chain elongation and subsequent fermentation of elongated carboxylates as H2-producing processes for sustained reductive dechlorination of chlorinated ethenes. Environ. Sci. Technol. 2021, 55, 10398–10410. [Google Scholar] [CrossRef]
- De Groof, V.; Coma, M.; Arnot, T.; Leak, D.J.; Lanham, A.B. Medium chain carboxylic acids from complex organic feedstocks by mixed culture fermentation. Molecules 2019, 24, 398. [Google Scholar] [CrossRef] [Green Version]
- Aulenta, F.; Catervi, A.; Majone, M.; Panero, S.; Reale, P.; Rossetti, S. Electron transfer from a solid-state electrode assisted by methyl viologen sustains efficient microbial reductive dechlorination of TCE. Environ. Sci. Technol. 2007, 41, 2554–2559. [Google Scholar] [CrossRef]
- Lohner, S.T.; Becker, D.; Mangold, K.M.; Tiehm, A. Sequential reductive and oxidative biodegradation of chloroethenes stimulated in a coupled bioelectro-process. Environ. Sci. Technol. 2011, 45, 6491–6497. [Google Scholar] [CrossRef]
- Verdini, R.; Aulenta, F.; De Tora, F.; Lai, A.; Majone, M. Relative contribution of set cathode potential and external mass transport on TCE dechlorination in a continuous-flow bioelectrochemical reactor. Chemosphere 2015, 136, 72–78. [Google Scholar] [CrossRef]
- Wang, X.; Aulenta, F.; Puig, S.; Esteve-Núnez, A.; He, Y.; Mu, Y.; Rabaey, K. Microbial electrochemistry for bioremediation. ESE 2020, 1, 100013. [Google Scholar] [CrossRef]
- Lovley, D.R. Happy together: Microbial communities that hook up to swap electrons. ISME J. 2017, 11, 327–336. [Google Scholar] [CrossRef] [Green Version]
- Yang, Y.; Xu, M.; Guo, J.; Sun, G. Bacterial extracellular electron transfer in bioelectrochemical systems. Process Biochem. 2021, 47, 1707–1714. [Google Scholar] [CrossRef]
- Zheng, T.; Li, J.; Ji, Y.; Zhang, W.; Fang, Y.; Xin, F.; Dong, W.; Wei, P.; Ma, J.; Jiang, M. Progress and prospects of bioelectrochemical systems: Electron transfer and its applications in the microbial metabolism. Front. Bioeng. Biotechnol. 2020, 8, 10. [Google Scholar] [CrossRef] [Green Version]
- Rollefson, J.B.; Stephen, C.S.; Tien, M.; Bond, D.R. Identification of an extracellular polysaccharide network essential for cytochrome anchoring and biofilm formation in Geobacter sulfurreducens. J. Bacteriol. 2011, 193, 1023–1033. [Google Scholar] [CrossRef] [Green Version]
- Kouzuma, A.; Meng, X.Y.; Kimura, N.; Hashimoto, K.; Watanabe, K. Disruption of the putative cell surface polysaccharide biosynthesis gene SO3177 in Shewanella oneidensis MR-1 enhances adhesion to electrodes and current generation in microbial fuel cells. Appl. Environ. Microbiol. 2010, 76, 4151–4157. [Google Scholar] [CrossRef] [Green Version]
- Lai, A.; Aulenta, F.; Mingazzini, M.; Palumbo, M.T.; Papini, M.P.; Verdini, R.; Majone, M. Bioelectrochemical approach for reductive and oxidative dechlorination of chlorinated aliphatic hydrocarbons (CAHs). Chemosphere 2017, 169, 351–360. [Google Scholar] [CrossRef]
- Aulenta, F.; Tocca, L.; Verdini, R.; Reale, P.; Majone, M. Dechlorination of trichloroethene in a continuous-flow bioelectrochemical reactor: Effect of cathode potential on rate, selectivity, and electron transfer mechanisms. Environ. Sci. Technol. 2011, 45, 8444–8451. [Google Scholar] [CrossRef]
- Aulenta, F.; Canosa, A.; Roma, L.D.; Reale, P.; Panero, S.; Rossetti, S.; Majone, M. Influence of mediator immobilization on the electrochemically assisted microbial dechlorination of trichloroethene (TCE) and cis-dichloroethene (cis-DCE). J. Chem. Technol. Biotechnol. Int. Res. Process Environ. Clean Technol. 2009, 84, 864–870. [Google Scholar] [CrossRef]
- Zeppilli, M.; Matturro, B.; Dell’Armi, E.; Cristiani, L.; Papini, M.P.; Rossetti, S.; Majone, M. Reductive/oxidative sequential bioelectrochemical process for Perchloroethylene (PCE) removal: Effect of the applied reductive potential and microbial community characterization. J. Environ. Chem. Eng. 2021, 9, 104657. [Google Scholar] [CrossRef]
- Lohner, S.T.; Tiehm, A. Application of electrolysis to stimulate microbial reductive PCE dechlorination and oxidative VC biodegradation. Environ. Sci. Technol. 2009, 43, 7098–7104. [Google Scholar] [CrossRef]
- Aulenta, F.; Verdini, R.; Zeppilli, M.; Zanaroli, G.; Fava, F.; Rossetti, S.; Majone, M. Electrochemical stimulation of microbial cis-dichloroethene (cis-DCE) oxidation by an ethene-assimilating culture. New Biotechnol. 2013, 30, 749–755. [Google Scholar] [CrossRef] [PubMed]
- Zeppilli, M.; Dell’Armi, E.; Cristiani, L.; Petrangeli Papini, M.; Majone, M. Reductive/oxidative sequential bioelectrochemical process for perchloroethylene removal. Water 2019, 11, 2579. [Google Scholar] [CrossRef] [Green Version]
- Strycharz, S.M.; Woodard, T.L.; Johnson, J.P.; Nevin, K.P.; Sanford, R.A.; Leoffler, F.E.; Lovley, D.R. Graphite electrode as a sole electron donor for reductive dechlorination of tetrachloroethene by Geobacter lovleyi. Appl. Environ. Microbiol. 2008, 74, 5943–5947. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leitao, P.; Rossetti, S.; Nouws, H.P.A.; Danko, A.S.; Majone, M.; Aulenta, F. Bioelectrochemically-assisted reductive dechlorination of 1,2-dichloroethane by a Dehalococcoides-enriched microbial culture. Bioresour. Technol. 2015, 195, 78–82. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rowe, A.R.; Rajeev, P.; Jain, A.; Pirbadian, S.; Okamoto, A.; Gralnick, J.A.; El-Naggar, M.Y.; Nealson, K.H. Tracking electron uptake from a cathode into Shewanella cells: Implications for energy acquisition from solid-substrate electron donors. MBio 2018, 9, e02203-17. [Google Scholar] [CrossRef] [Green Version]
- Chen, F.; Liang, B.; Li, Z.L.; Yang, J.Q.; Huang, C.; Lyu, M.; Yuan, Y.; Nan, J.; Wang, A.J. Bioelectrochemical assisted dechlorination of tetrachloroethylene and 1,2-dichloroethane by acclimation of anaerobic sludge. Chemosphere 2019, 227, 514–521. [Google Scholar] [CrossRef]
- Wang, S.; He, J.; Shen, C.; Manefield, M.J. Editorial: Organohalide respiration: New findings in metabolic mechanisms and bioremediation applications. Front. Microbiol. 2019, 10, 526. [Google Scholar] [CrossRef]
- Meng, L.; Yoshida, N.; Li, Z. Soil microorganisms facilitated the electrode-driven trichloroethene dechlorination to ethene by Dehalococcoides species in a bioelectrochemical system. Environ. Res. 2022, 209, 112801. [Google Scholar] [CrossRef]
- Chen, F.; Li, Z.L.; Yang, J.Q.; Liang, B.; Lin, X.Q.; Nan, J.; Wang, A.J. Effects of different carbon substrates on performance, microbiome community structure and function for bioelectrochemical-stimulated dechlorination of tetrachloroethylene. J. Chem. Eng. 2018, 352, 730–736. [Google Scholar] [CrossRef]
- Dell’Armi, E.; Rossi, M.M.; Taverna, L.; Petrangeli Papini, M.; Zeppilli, M. Evaluation of the bioelectrochemical approach and different electron donors for biological trichloroethylene reductive dechlorination. Toxics 2022, 10, 37. [Google Scholar] [CrossRef]
- Cecconet, D.; Sabba, F.; Devecseri, M.; Callegari, A.; Capodaglio, A.G. In situ groundwater remediation with bioelectrochemical systems: A critical review and future perspectives. Environ. Int. 2020, 137, 105550. [Google Scholar] [CrossRef]
- Palma, E.; Daghio, M.; Franzetti, A.; Petrangeli Papini, M.; Aulenta, F. The bioelectric well: A novel approach for in situ treatment of hydrocarbon-contaminated groundwater. Microb. Biotechnol. 2018, 11, 112–118. [Google Scholar]
- Zanini, A.; Ghirardi, M.; Emiliani, R. A multidisciplinary approach to evaluate the effectiveness of natural attenuation at a contaminated site. Hydrology 2021, 8, 101. [Google Scholar] [CrossRef]
- Hunkeler, D.; Aravena, R.; Berry-Spark, K.; Cox, E. Assessment of degradation pathways in an aquifer with mixed chlorinated hydrocarbon contamination using stable isotope analysis. Environ. Sci. Technol. 2005, 39, 5975–5981. [Google Scholar] [CrossRef] [Green Version]
- Wen, L.L.; Li, Y.; Zhu, L.; Zhao, H.P. Influence of non-dechlorinating microbes on trichloroethene reduction based on vitamin B12 synthesis in anaerobic cultures. Environ. Pollut. 2020, 259, 113947. [Google Scholar] [CrossRef]
- Wu, Y.J.; Liu, P.W.G.; Hsu, Y.S.; Whang, L.M.; Lin, T.F.; Hung, W.N.; Cho, K.C. Application of molecular biological tools for monitoring efficiency of trichloroethylene remediation. Chemosphere 2019, 233, 697–704. [Google Scholar] [CrossRef]
- Jin, D.; Zhang, F.; Shi, Y.; Kong, X.; Xie, Y.; Du, X.; Li, Y.; Zhang, R. Diversity of bacteria and archaea in the groundwater contaminated by chlorinated solvents undergoing natural attenuation. Environ. Res. 2020, 185, 109457. [Google Scholar] [CrossRef]
- Garza-Rubalcava, U.; Hatzinger, P.B.; Schanzle, D.; Lavorgna, G.; Hedman, P.; Jackson, W.A. Improved assessment and performance monitoring of a biowall at a chlorinated solvent site using high-resolution passive sampling. J. Contam. Hydrol. 2022, 246, 103962. [Google Scholar] [CrossRef]
- Yoshikawa, M.; Zhang, M.; Kawabe, Y.; Katayama, T. Effects of ferrous iron supplementation on reductive dechlorination of tetrachloroethene and on methanogenic microbial community. FEMS Microbiol. Ecol. 2021, 97, fiab069. [Google Scholar] [CrossRef]
- Lo, K.H.; Lu, C.W.; Chien, C.C.; Sheu, Y.T.; Lin, W.H.; Chen, S.C.; Kao, C.M. Cleanup chlorinated ethene-polluted groundwater using an innovative immobilized Clostridium butyricum column scheme: A pilot-scale study. J. Environ. Manag. 2022, 311, 114836. [Google Scholar] [CrossRef]
- Hellal, J.; Joulian, C.; Urien, C.; Ferreira, S.; Denonfoux, J.; Hermon, L.; Vuilleumier, S.; Imfeld, G. Chlorinated ethene biodegradation and associated bacterial taxa in multi-polluted groundwater: Insights from biomolecular markers and stable isotope analysis. Sci. Total Environ. 2021, 763, 142950. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.O.; Mattes, T.E. A Quantitative PCR Assay for aerobic, vinyl chloride- and ethene-assimilating microorganisms in groundwater. Environ. Sci. Technol. 2010, 44, 9036–9041. [Google Scholar] [CrossRef] [PubMed]
- Mattes, T.E.; Jin, Y.O.; Livermore, J.; Pearl, M.; Liu, X. Abundance and activity of vinyl chloride (VC)-oxidizing bacteria in a dilute groundwater VC plume biostimulated with oxygen and ethene. Appl. Microbiol. Biotechnol. 2015, 99, 9267–9276. [Google Scholar] [CrossRef] [PubMed]
- Atashgahi, S.; Maphosa, F.; Dogan, E.; Smidt, H.; Springael, D.; Dejonghe, W. Small-scale oxygen distribution determines the vinyl chloride biodegradation pathway in surficial sediments of riverbed hyporheic zones. FEMS Microbiol. Ecol. 2012, 84, 133–142. [Google Scholar] [CrossRef] [Green Version]
- Liang, Y.; Cook, L.J.; Mattes, T.E. Temporal abundance and activity trends of vinyl chloride (VC)-degrading bacteria in a dilute VC plume at Naval Air Station Oceana. Environ. Sci. Pollut. Res. 2017, 24, 13760–13774. [Google Scholar] [CrossRef]
- Richards, P.M.; Liang, Y.; Johnson, R.L.; Mattes, T.E. Cryogenic soil coring reveals coexistence of aerobic and anaerobic vinyl chloride degrading bacteria in a chlorinated ethene contaminated aquifer. Water Res. 2019, 157, 281–291. [Google Scholar] [CrossRef] [Green Version]
- Němeček, J.; Marková, K.; Špánek, R.; Antoš, V.; Kozubek, P.; Lhotský, O.; Černík, M. Hydrochemical conditions for aerobic/anaerobic biodegradation of chlorinated ethenes—A multi-site assessment. Water 2020, 12, 322. [Google Scholar] [CrossRef] [Green Version]



| Compound | Appearance | Water Solubility at 25 °C (g L−1) a | Density at 20 °C (g cm−3) a | Vapor Pressure at 20 °C (kPa) a | Autoignition Temperature (°C) a | Carcinogenicity b | Law Limits (µg L−1) Directive 2000/60/EC | |
|---|---|---|---|---|---|---|---|---|
| Vinyl chloride (VC) (C2H3Cl) | Colorless gas | Slightly soluble | 0.91 | 516.95 | 472° | Group 1 (2012) | 0.5 | |
| cis-dichloroethene (cis-DCE) (C2H2Cl2) | Colorless liquid | 1–5 | 1.28 | 26.66 | 460° | N | - | |
| trans-dichloroethene (trans-DCE) (C2H2Cl2) | 1,1-trans-DCE | Colorless liquid | 2.5 | 1.213 | 66.5 | 460° | Group 3 (1999) | 0.05 |
| 1,2-trans-DCE | Colorless liquid | <1.0 | 1.25 | 53.33 | 460° | N | 60 | |
| Trichloroethene (TCE) (C2HCl3) | Colorless liquid | 1.280 | 1.46 | 7.8 | >410° | Group 1 (2014) | 1.5 | |
| Tetrachloroethene (PCE) (C2Cl4) | Colorless liquid | 0.15 | 1.63 | 1.9 | >650° | Group 2A (2014) | 1.1 | |
| Genus | Species/Strains | Isolation Sources | References |
|---|---|---|---|
| Mycobacterium | aurum strain L1 | Contaminated soil | [69] |
| strains JS60 | Contaminated groundwater | [66] | |
| strains JS61 | Activated sludge | [66] | |
| strains JS616 | Sediment of industrial site | [66] | |
| strains JS617 | Activated carbon of pump and treatment plant | [66] | |
| Rhodococcus | rhodochrous | PCE degrading enrichment culture | [70] |
| Nocardioides | sp. strain JS614 | Soil of industrial site | [66] |
| Pseudomonas | aeruginosa strain DL1 | Activated sludge | [71] |
| aeruginosa strain MF1 | VC degrading enrichment culture | [72] | |
| putida strain AJ | Hazardous waste site | [73] | |
| Ochrobactrum | sp. strain TD | Hazardous waste site | [73] |
| Ralstonia | sp. strain TRW-1 | Chloroethene degrading enrichment culture | [74] |
| Brevundimonas | sp. | Contaminated groundwater | [75] |
| Rhodoferax | sp. | Contaminated groundwater | [75] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bertolini, M.; Zecchin, S.; Cavalca, L. Sequential Anaerobic/Aerobic Microbial Transformation of Chlorinated Ethenes: Use of Sustainable Approaches for Aquifer Decontamination. Water 2023, 15, 1406. https://doi.org/10.3390/w15071406
Bertolini M, Zecchin S, Cavalca L. Sequential Anaerobic/Aerobic Microbial Transformation of Chlorinated Ethenes: Use of Sustainable Approaches for Aquifer Decontamination. Water. 2023; 15(7):1406. https://doi.org/10.3390/w15071406
Chicago/Turabian StyleBertolini, Martina, Sarah Zecchin, and Lucia Cavalca. 2023. "Sequential Anaerobic/Aerobic Microbial Transformation of Chlorinated Ethenes: Use of Sustainable Approaches for Aquifer Decontamination" Water 15, no. 7: 1406. https://doi.org/10.3390/w15071406






