Comparative Study of Microbial Diversity in Different Coastal Aquifers: Determining Factors
Abstract
:1. Introduction
2. Materials and Methods
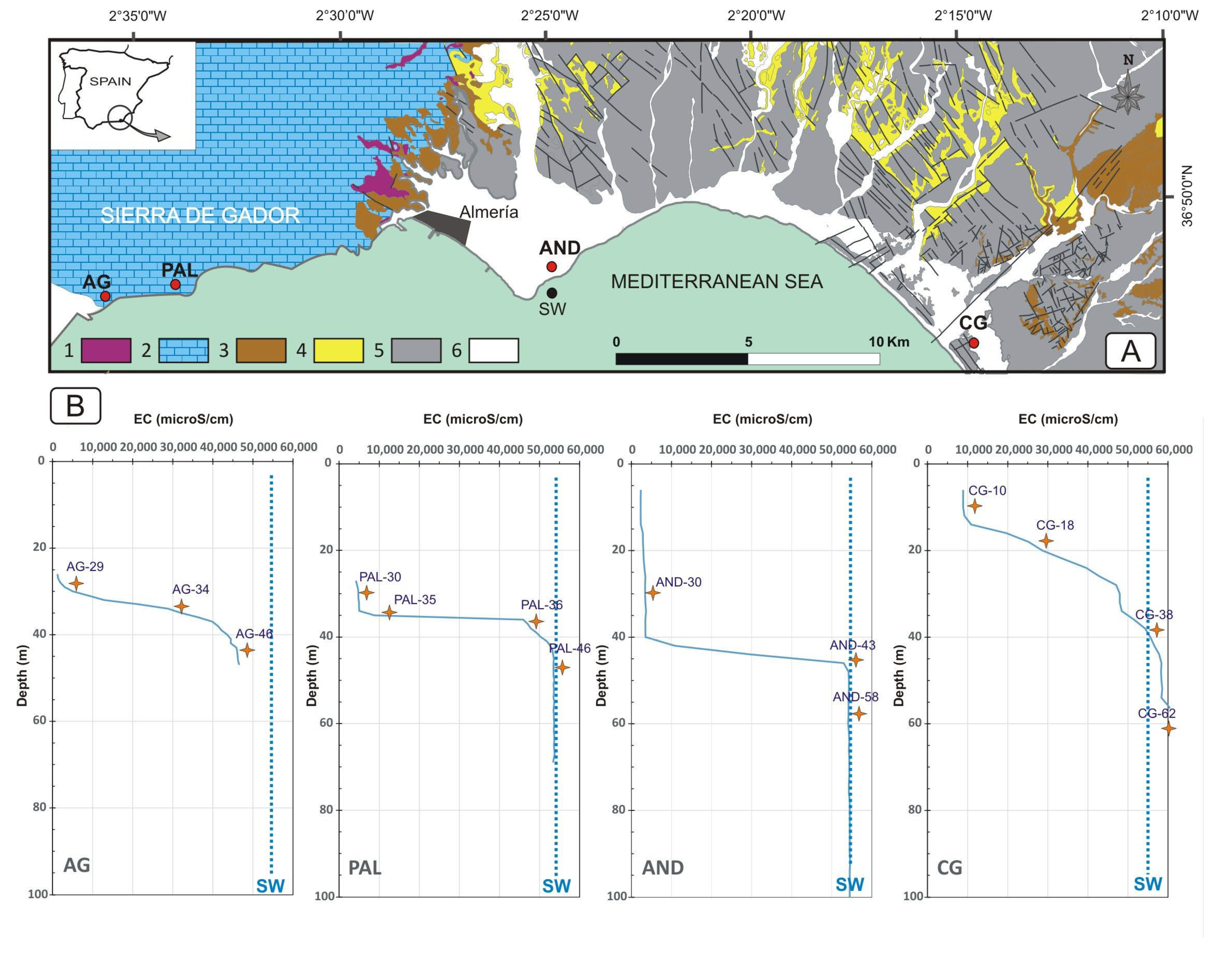
2.1. Study Sites, Sampling, and Physico-Chemical Parameters
2.2. DNA Extraction and Sequencing Analysis
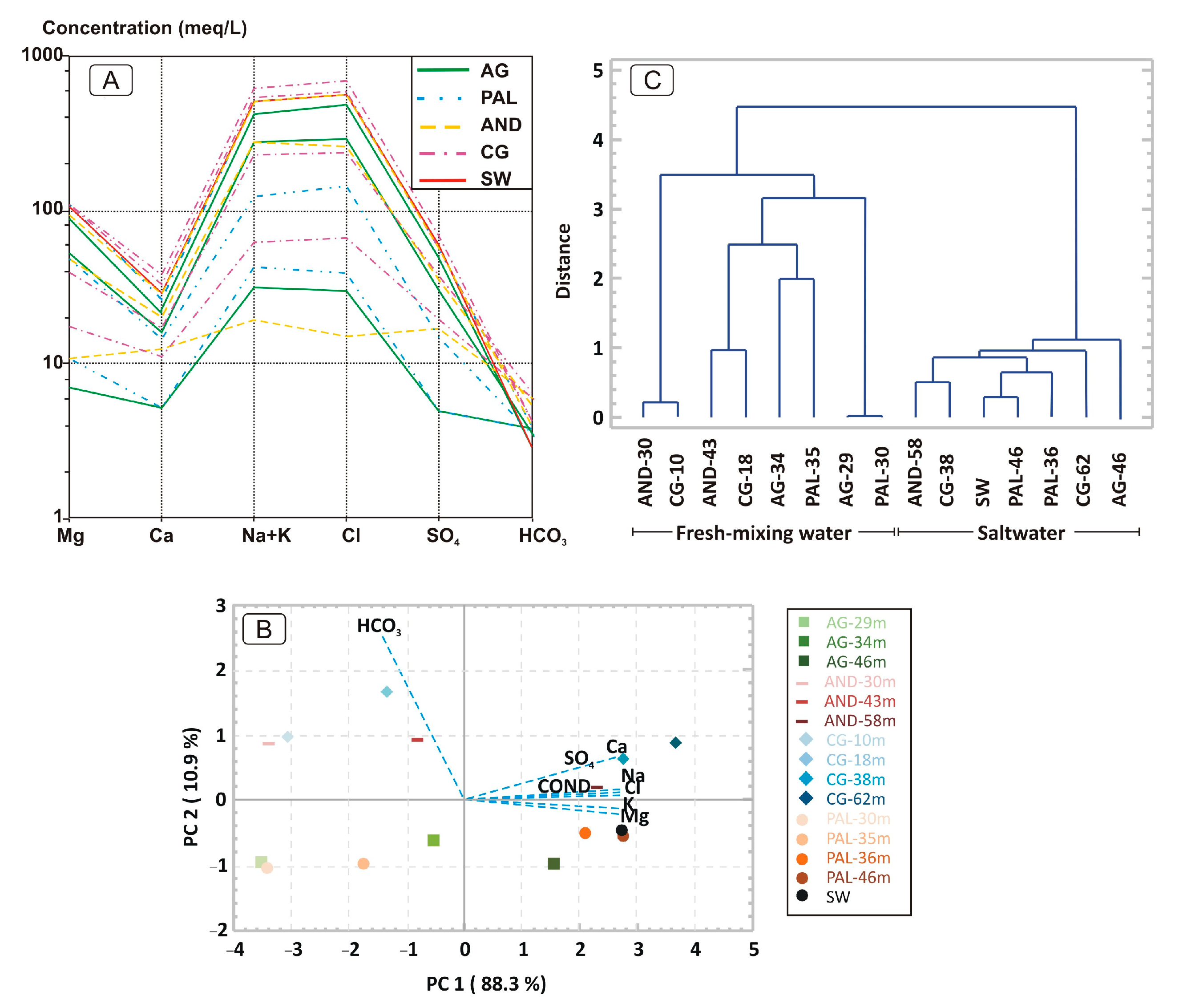
2.3. Statistical Analysis
3. Results
3.1. Hydrochemical Characterization
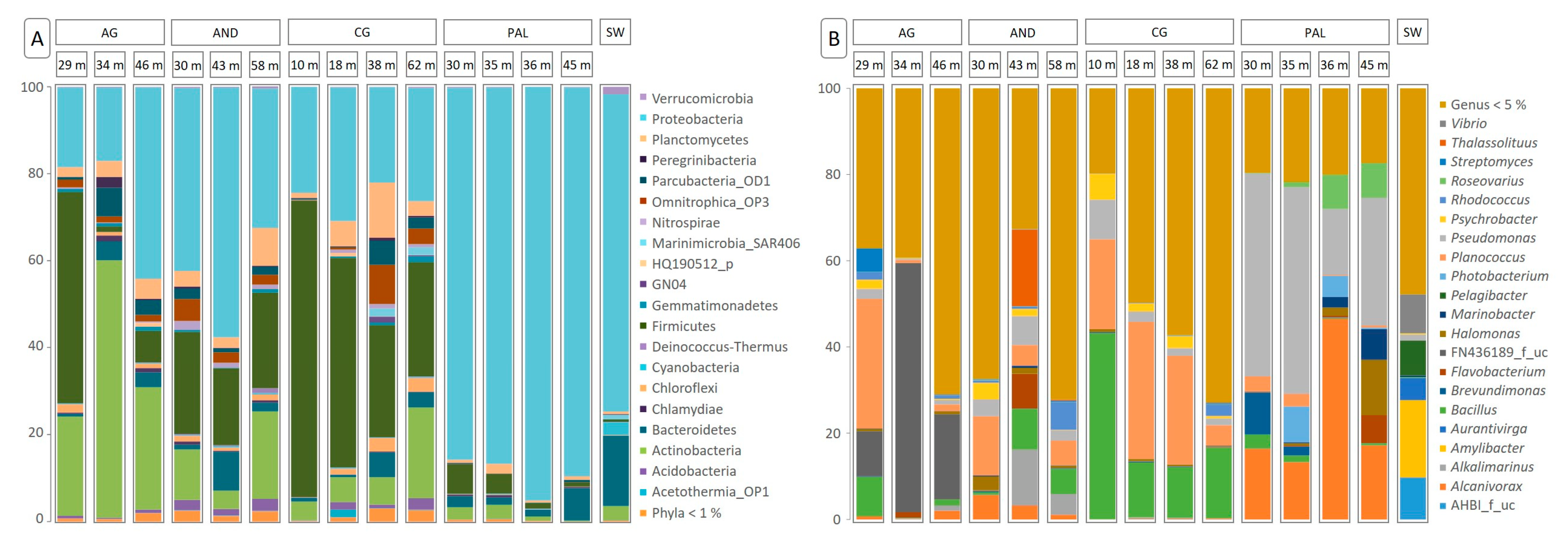
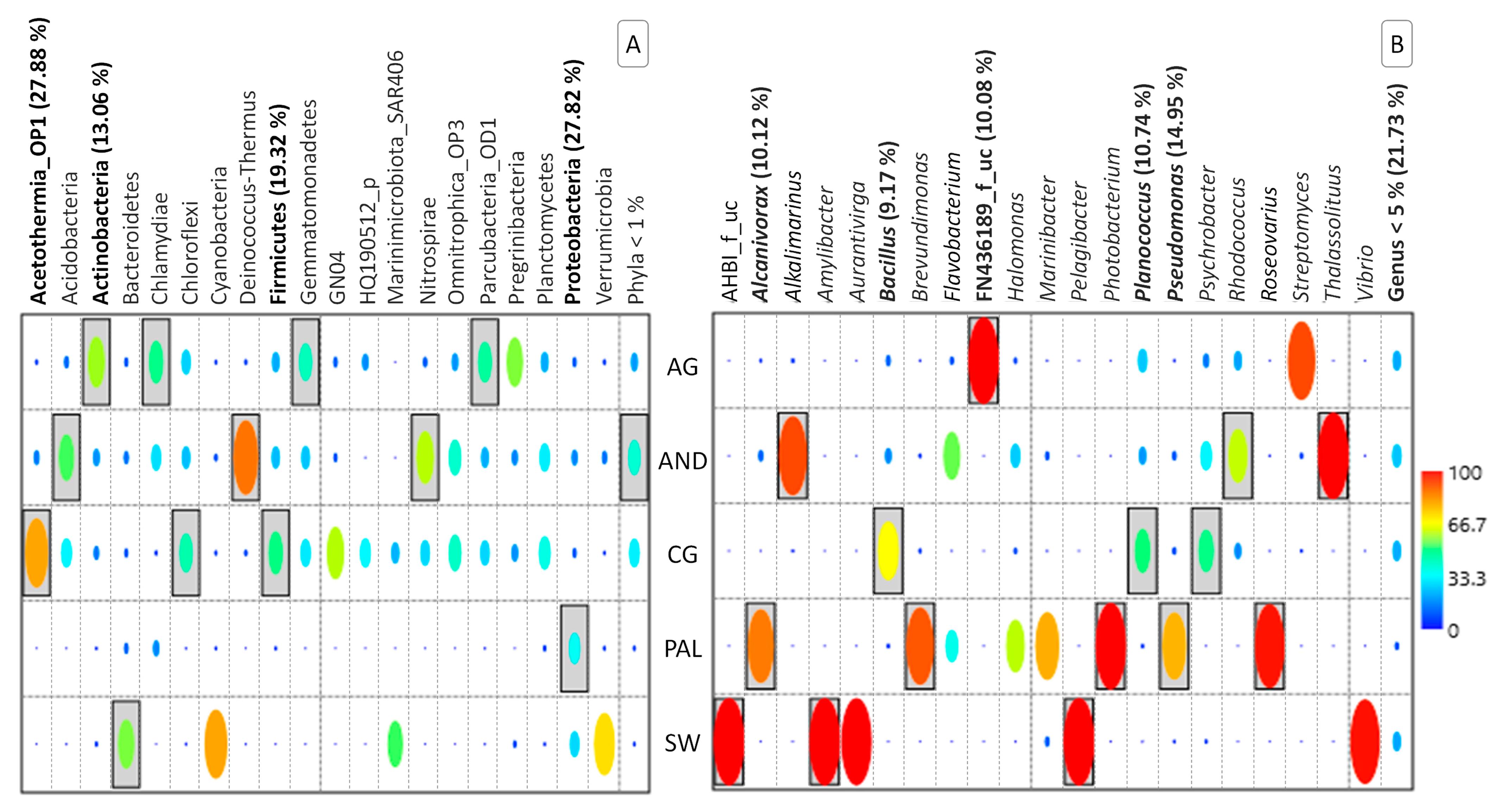
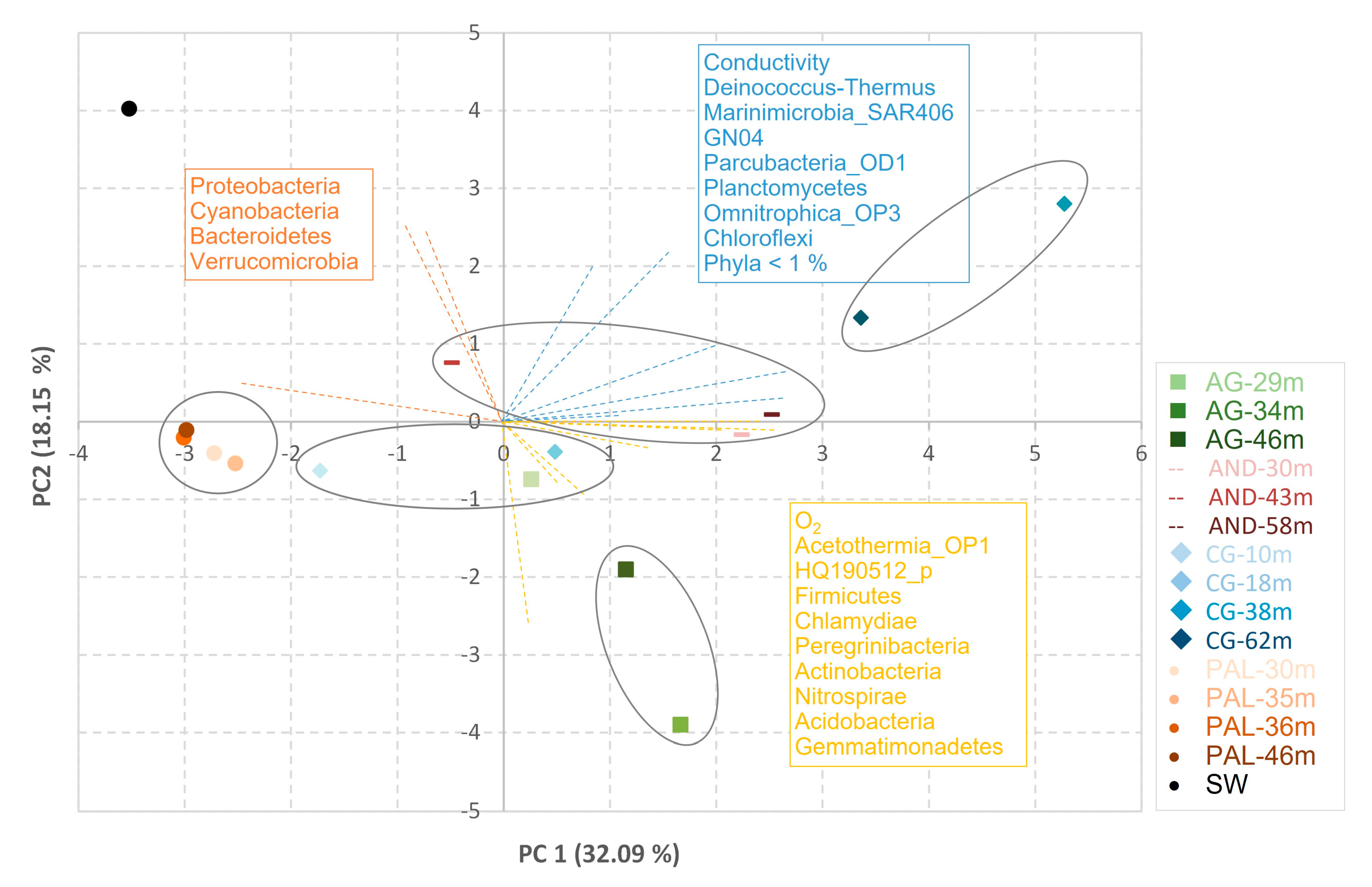
3.2. Composition of the Microbial Community
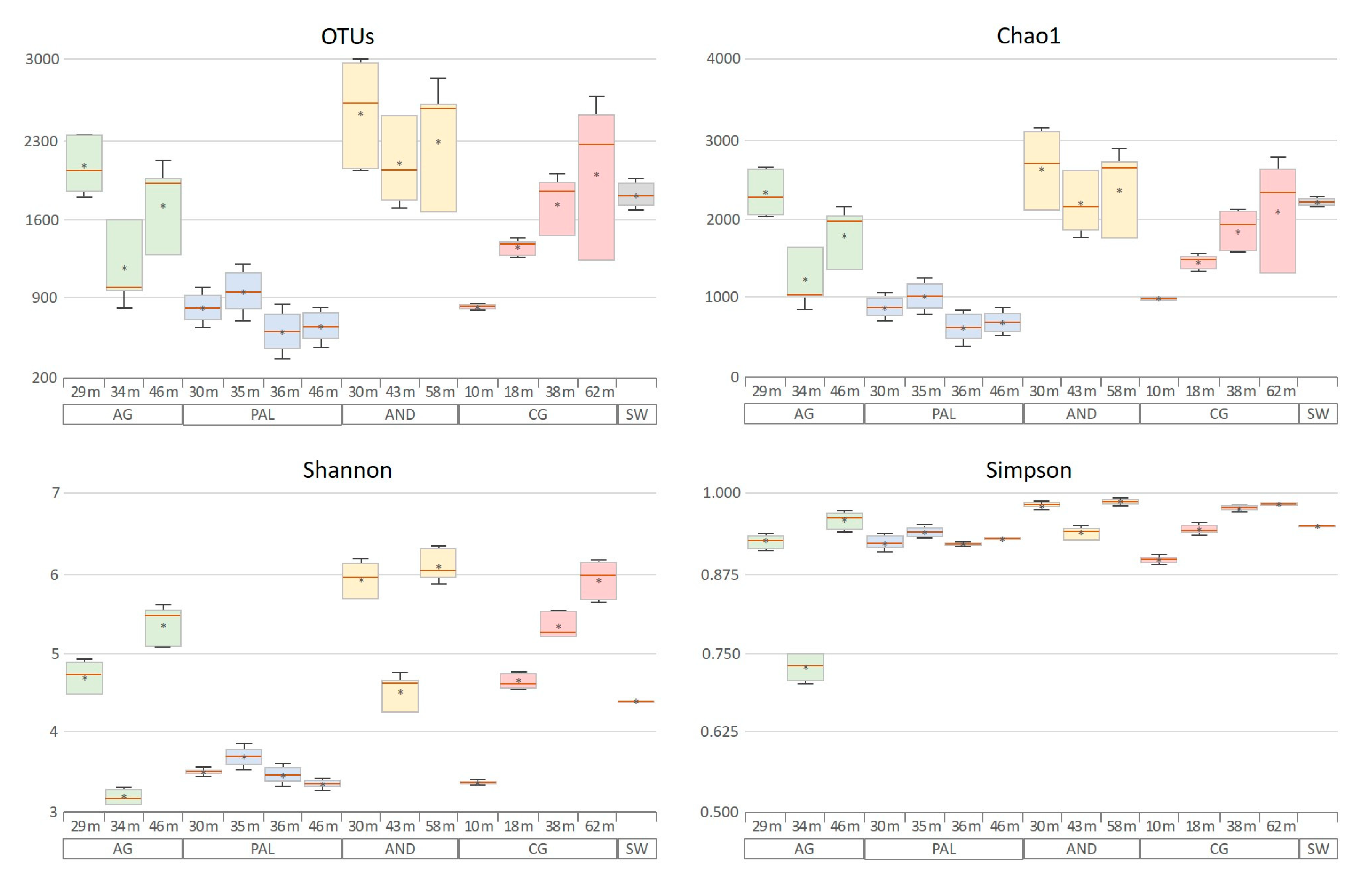
3.3. Microbial Diversity
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Taniguchi, M.; Burnett, W.C.; Cable, J.E.; Turner, J.V. Investigation of submarine groundwater discharge. Hydrol. Process. 2002, 16, 2115–2129. [Google Scholar] [CrossRef]
- Moore, W.S.; Sarmiento, J.L.; Key, R.M. Submarine groundwater discharge revealed by 228Ra distribution in the upper Atlantic Ocean. Nat. Geosci. 2008, 1, 309–311. [Google Scholar] [CrossRef]
- Moore, W.S. The Effect of Submarine Groundwater Discharge on the Ocean. Annu. Rev. Mar. Sci. 2010, 2, 59–88. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kwon, E.Y.; Kim, G.; Primeau, F.; Moore, W.S.; Cho, H.M.; DeVries, T.; Sarmiento, J.L.; Charette, M.A.; Cho, Y.K. Global estimate of submarine groundwater discharge based on an observationally constrained radium isotope model. Geophys. Res. Lett. 2014, 41, 8438–8444. [Google Scholar] [CrossRef]
- Santos, I.R.; Chen, X.; Lecher, A.L.; Sawyer, A.H.; Moosdorf, N.; Rodellas, V.; Tamborski, J.J.; Cho, H.-M.; Dimova, N.; Sugimoto, R.; et al. Submarine groundwater discharge impacts on coastal nutrient biogeochemistry. Nat. Rev. Earth Environ. 2021, 2, 307–323. [Google Scholar] [CrossRef]
- Werner, A.D.; Bakker, M.; Post, V.E.A.; Vandenbohede, A.; Lu, C.; Ataie-Ashtiani, B.; Simmons, C.T.; Barry, D.A. Seawater intrusion processes, investigation and management: Recent advances and future challenges. Adv. Water Resour. 2013, 51, 3–26. [Google Scholar] [CrossRef]
- Lu, C.; Chen, Y.; Zhang, C.; Luo, J. Steady-state freshwater–seawater mixing zone in stratified coastal aquifers. J. Hydrol. 2013, 505, 24–34. [Google Scholar] [CrossRef]
- Nielsen, P. Tidal dynamics of the water table in beaches. Water Resour. Res. 1990, 26, 2127–2134. [Google Scholar] [CrossRef] [Green Version]
- Kim, Y.; Lee, K.S.; Koh, D.C.; Lee, D.H.; Lee, S.G.; Park, W.B.; Koh, G.W.; Woo, N.C. Hydrogeochemical and isotopic evidence of groundwater salinization in a coastal aquifer: A case study in Jeju volcanic island, Korea. J. Hydrol. 2003, 270, 282–294. [Google Scholar] [CrossRef]
- Michael, H.A.; Mulligan, A.E.; Harvey, C.F. Seasonal oscillations in water exchange between aquifers and the coastal ocean. Nature 2005, 436, 1145–1148. [Google Scholar] [CrossRef]
- Chapelle, F.H.; Lovely, D.R. Rates of microbial metabolism in deep coastal plain aquifers. Appl. Environ. Microbiol. 1990, 56, 1865–1874. [Google Scholar] [CrossRef] [Green Version]
- Santoro, A.E. Microbial nitrogen cycling at the saltwater–freshwater interface. Hydrogeol. J. 2010, 18, 187–202. [Google Scholar] [CrossRef]
- McAllister, S.M.; Barnett, J.M.; Heiss, J.W.; Findlay, A.J.; Macdonald, D.J.; Dow, C.L.; Luther, G.W., III; Michael, H.A.; Chan, C.S. Dynamic hydrologic and biogeochemical processes drive microbially enhanced iron and sulfur cycling within the intertidal mixing zone of a beach aquifer. Limnol. Oceanogr. 2015, 60, 329–345. [Google Scholar] [CrossRef] [Green Version]
- Ruiz-González, C.; Rodellas, V.; Garcia-Orellana, J. The microbial dimension of submarine groundwater discharge: Current challenges and future directions. FEMS Microbiol. Rev. 2021, 45, fuab010. [Google Scholar] [CrossRef]
- Sivan, O.; Yechieli, Y.; Herut, B.; Lazar, B. Geochemical evolution and timescale of seawater intrusion into the coastal aquifer of Israel. Geochim. Cosmochim. Acta 2005, 69, 579–592. [Google Scholar] [CrossRef]
- Andersen, M.; Nyvang, V.; Jakobsen, R.; Postma, D. Geochemical processes and solute transport at the seawater/freshwater interface of a sandy aquifer. Geochim. Cosmochim. Acta 2005, 69, 3979–3994. [Google Scholar] [CrossRef]
- Liu, Y.; Jiao, J.J.; Liang, W. Tidal fluctuation influenced physicochemical parameter dynamics in coastal groundwater mixing zone. Estuaries Coasts 2018, 41, 988–1001. [Google Scholar] [CrossRef]
- Héry, C.; Volant, A.; Garing, C.; Luquot, L.; Poulichet, F.E.; Gouze, P. Diversity and geochemical structuring of bacterial communities along a salinity gradient in a carbonate aquifer subject to seawater intrusion. FEMS Microbiol. Ecol. 2014, 90, 922–934. [Google Scholar] [CrossRef] [Green Version]
- Ye, Q.; Liu, J.; Du, J.; Zhang, J. Bacterial Diversity in Submarine Groundwater along the Coasts of the Yellow Sea. Front. Microbiol. 2016, 6, 1519. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davis, M.C.; Garey, J.R. Microbial function and hydrochemistry within a stratified anchialine sinkhole: A window into coastal aquifer interactions. Water 2018, 10, 972. [Google Scholar] [CrossRef] [Green Version]
- Calvo-Martin, E.; Teira, E.; Alvarez-Salgado, X.A.; Rocha, C.; Jiang, S.; Justel-Díez, M.; Pino Ibañez, J.S. On the hidden diversity and niche specialization of the microbial realm of subterranean estuaries. Environ. Microbiol. 2022, 24, 5859–5881. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-González, C.; Rodríguez-Pie, L.; Maister, O.; Rodellas, V.; Alorda-Keinglass, A.; Diego-Feliu, M.; Folch, A.; Garcia-Orellana, J.; Gasol, J.M. High spatial heterogeneity and low connectivity of bacterial communities along a Mediterranean subterranean estuary. Mol. Ecol. 2022, 31, 5745–5764. [Google Scholar] [CrossRef]
- Sola, F.; Vargas-García, M.C.; Vallejos, A. Interrelation prokaryotic community-aquifer in a carbonate coastal environment. Aquat. Sci. 2020, 82, 13. [Google Scholar] [CrossRef]
- Unno, T.; Kim, J.; Kim, Y.; Nguyen, S.G.; Guevarra, R.B.; Kim, G.P.; Lee, J.H.; Sadowsky, M.J. Influence of seawater intrusion on microbial communities in groundwater. Sci. Total Environ. 2015, 532, 337–343. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Hu, B.X.; Dai, H.; Zhang, X.; Xia, Z.; Zhang, J. Characterizing microbial diversity and community composition of groundwater in a salt-freshwater transition zone. Sci. Total Environ. 2019, 678, 574–584. [Google Scholar] [CrossRef]
- Chen, L.; Zhang, J.; Dai, H.; Hu, B.X.; Tong, J.; Gui, D.; Zhang, X.; Xia, C. Comparison of the groundwater microbial community in a salt-freshwater mixing zone during the dry and wet seasons. J. Environ. Manag. 2020, 271, 110969. [Google Scholar] [CrossRef]
- Archana, A.; Francis, C.A.; Boehm, A.B. The beach aquifer microbiome: Research gaps and data needs. Front. Environ. Sci. 2021, 9, 653568. [Google Scholar] [CrossRef]
- Suárez-Moo, P.; Remes-Rodríguez, C.A.; Márquez-Velázquez, N.A.; Falcón, L.I.; García-Maldonado, J.Q.; Prieto-Davó, A. Changes in the sediment microbial community structure of coastal and inland sinkholes of a karst ecosystem from the Yucatan peninsula. Sci. Rep. 2022, 12, 1110. [Google Scholar] [CrossRef]
- Yang, F.; Liu, S.; Jia, C.; Wang, Y. Identification of groundwater microbial communities and their connection to the hydrochemical environment in southern Laizhou Bay, China. Environ. Sci. Pollut. Res. 2022, 29, 14263–14278. [Google Scholar] [CrossRef]
- Smart, P.L.; Dawans, J.M.; Whitaker, F. Carbonate dissolution in a modern mixing zone. Nature 1988, 335, 11–813. [Google Scholar] [CrossRef]
- Appelo, C.; Postma, D. Geochemistry, Groundwater and Pollution, 2nd ed.; AA Balkema: Rotterdam, The Netherlands, 2005; 649p. [Google Scholar]
- Sola, F.; Vallejos, A.; Moreno, L.; López-Geta, J.A.; Pulido-Bosch, A. Identification of hydrogeochemical process linked to marine intrusion induced by pumping of a semiconfined Mediterranean coastal aquifer International. J. Environ. Sci. Technol. 2013, 10, 63–76. [Google Scholar]
- Charette, M.A.; Sholkovitz, E.R. Oxidative precipitation of groundwater-derived ferrous iron in the subterranean estuary of a coastal bay. Geophys. Res. Lett. 2002, 29, 85-1–85-4. [Google Scholar] [CrossRef] [Green Version]
- Santoro, A.E.; Boehm, A.B.; Francis, C.A. Denitrifier community composition along a nitrate and salinity gradient in a coastal aquifer. Appl. Environ. Microbiol. 2006, 72, 2102–2109. [Google Scholar] [CrossRef] [Green Version]
- Spiteri, C.; Slomp, C.P.; Charette, M.A.; Tuncay, K.; Meile, C. Flow and nutrient dynamics in a subterranean estuary (Waquoit Bay, MA, USA): Field data and reactive transport modeling. Geochim. Cosmochim. Acta 2008, 72, 3398–3412. [Google Scholar] [CrossRef]
- Spiteri, C.; Slomp, C.; Tuncay, K.; Meile, C. Modeling biogeochemical processes in subterranean estuaries: Effect of flow dynamics and redox conditions on submarine groundwater discharge of nutrients. Water Resour. Res. 2008, 44, W04701. [Google Scholar] [CrossRef] [Green Version]
- Perera, E.D.; Jinno, K.; Hiroshiro, Y. Bacteria-mediated Reduction and Precipitation of Fe(OH)3 and FeS in the Subsurface of a Coastal Aquifer: A Numerical Investigation. Water Qual. Expo. Health 2010, 2, 15–30. [Google Scholar] [CrossRef]
- Couturier, M.; Tommi-Morin, G.; Sirois, M.; Rao, A.; Nozais, C.; Chaillou, G. Nitrogen transformations along a shallow subterranean estuary. Biogeosciences 2017, 14, 3321–3336. [Google Scholar] [CrossRef] [Green Version]
- Jiang, S.; Zhang, Y.; Jin, J.; Wu, Y.; Wei, Y.; Wang, X.; Rocha, C.; Pino Ibánhez, J.S.; Zhang, J. Organic carbon in a seepage face of a subterranean estuary: Turnover and microbial interrelations. Sci. Total Environ. 2020, 725, 138220. [Google Scholar] [CrossRef]
- Petrash, D.A.; Bialik, O.M.; Staudigel, P.T.; Konhauser, K.O.; Budd, D.A. Biogeochemical reappraisal of the freshwater–seawater mixing-zone diagenetic model. Sedimentology 2021, 68, 1797–1830. [Google Scholar] [CrossRef]
- Sar, P.; Dutta, A.; Bose, H.; Mandal, S.; Kazay, S.K. Deep biosphere: Microbiome of the deep terrestrial subsurface. In Microbial Diversity in Ecosystem Sustainability and Biotechnological Applications; Satyanarayana, T., Johri, B., Das, S., Eds.; Springer: Singapore, 2019; pp. 225–265. [Google Scholar] [CrossRef]
- Wegner, C.E.; Gaspar, M.; Geesink, P.; Hermann, M.; Marz, M.; Küsel, K. Biogeochemical regimes in shallow aquifers reflect the metabolic coupling of the elements nitrogen, sulfur and carbon. Appl. Environ. Microbiol. 2019, 85, e02346-18. [Google Scholar] [CrossRef] [Green Version]
- Anantharaman, K.; Brown, C.T.; Hug, L.A.; Sharon, I.; Castelle, C.J.; Probst, A.J.; Brodie, E.L. Thousands of microbial genomes shed light on interconnected biogeochemical processes in an aquifer system. Nat. Commun. 2016, 7, 13219. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Herrmann, M.; Kampe, B.; Lehmann, R.; Totsche, K.U.; Küsel, K. Environmental selection shapes the formation of near-surface groundwater microbiomes. Water Res. 2020, 170, 115341. [Google Scholar] [CrossRef] [PubMed]
- Stein, H.; Kellerman, C.; Schmidt, S.I.; Brielmann, H.; Steube, C.; Berkhoff, S.E.; Fuchs, A.; Hahn, H.J.; Thulin, B.; Griebler, C. The potential use of fauna and bacterias as ecological indicators for the assessment of groundwater quality. Environ. Monit. 2010, 12, 242–254. [Google Scholar] [CrossRef] [PubMed]
- Giménez-Forcada, E. Use of the Hydrochemical Facies Diagram (HFE-D) for the evaluation of salinization by seawater intrusion in the coastal Oropesa Plain: Comparative analysis with the coastal Vinaroz Plain, Spain. HydroResearch 2019, 2, 76–84. [Google Scholar] [CrossRef]
- Duque, C.; Olsen, J.T.; Sánchez-Úbeda, J.P.; Calvache, M.L. Groundwater salinity during 500 years of anthropogenic-driven coastline changes in the Motril-Salobreña aquifer (South East Spain). Environ. Earth Sci. 2019, 78, 471. [Google Scholar] [CrossRef]
- Vallejos, A.; Daniele, L.; Sola, F.; Molina, L.; Pulido-Bosch, A. Anthropic-induced salinization in a dolomite coastal aquifer. Hydrogeochemical processes. J. Geochem. Explor. 2020, 209, 106438. [Google Scholar] [CrossRef]
- Goyetche, T.; Luquot, L.; Carrera, J.; Martínez Pérez, L.; Folch, A. Identification and quantification of chemical reactions in a coastal aquifer to assess submarine groundwater discharge composition. Sci. Total Environ. 2022, 838, 155978. [Google Scholar] [CrossRef]
- López-Archilla, A.I.; Moreira, D.; Velasco, S.; López-García, P. Archaeal and bacterial community composition of a pristine coastal aquifer in Doñana National Park, Spain. Aquat. Microb. Ecol. 2007, 47, 123–129. [Google Scholar] [CrossRef]
- Velasco-Ayuso, S.; Acebes, P.; López-Archilla, A.I.; Montes, C.; Guerrero, M.d.C. Environmental factors controlling the spatiotemporal distribution of microbial communities in a coastal, sandy aquifer system (Doñana, southwest Spain). Hydrogeol. J. 2009, 17, 767–780. [Google Scholar] [CrossRef]
- Velasco-Ayuso, S.; Guerrero, M.C.; Montes, C.; López-Archilla, A.I. Spatiotemporal distribution of microbial communities in a coastal, sandy aquifer system (Doñana, SW Spain). Geobiology 2009, 7, 66–81. [Google Scholar] [CrossRef] [Green Version]
- Velasco-Ayuso, S.; López-Archilla, A.I.; Montes, C.; Guerrero, M.C. Microbial activities in a coastal, sandy aquifer system (Doñana natural protected area, SW Spain). Geomicrobiol. J. 2010, 27, 409–423. [Google Scholar] [CrossRef]
- Plaza-Bolaños, P.; Padilla-Sánchez, J.A.; Garrido-Frenich, A.; Romero-González, R.; Martínez-Vidal, J.L. Evaluation of soil contamination in intensive agricultural areas by pesticides and organic pollutants: South-eastern Spain as a case study. J. Environ. Monit. 2012, 14, 1182–1189. [Google Scholar] [CrossRef]
- Foster, S.; Pulido-Bosch, A.; Vallejos, A.; Molina, L.; Llop, A.; MacDonald, A.M. Impact of irrigated agriculture on groundwater-recharge salinity: A major sustainability concern in semi-arid regions. Hydrogeol. J. 2018, 26, 2781–2791. [Google Scholar] [CrossRef] [Green Version]
- Pulido-Bosch, A.; Rigol-Sánchez, J.P.; Vallejos, A.; Andreu, J.M.; Cerón, J.C.; Molina, L.; Sola, F. Impacts of agricultural irrigation on groundwater salinity. Environ. Earth Sci. 2018, 77, 197. [Google Scholar] [CrossRef] [Green Version]
- Aznar-Sánchez, J.A.; Belmonte-Ureña, L.J.; Velasco-Muñoz, J.F.; Valera, D.L. Aquifer sustainability and the use of desalinated seawater for greenhouse irrigation in the Campo de Níjar, Southeast Spain. Int. J. Environ. Res. Public Health 2019, 16, 898. [Google Scholar] [CrossRef] [Green Version]
- Tout, D. The Horticulture Industry of Almería Province, Spain. Geogr. J. 1990, 156, 304–312. [Google Scholar] [CrossRef]
- Mendoza-Fernández, A.J.; Peña-Fernández, A.; Molina, L.; Aguilera, P.A. The role of technology in greenhouse agriculture: Towards a sustainable intensification in Campo de Dalías (Almería, Spain). Agronomy 2021, 11, 101. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of general 16S ribosomal RNA genes primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Martin, M. Cut adapt removes adapter sequences from high through put sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Fu, L.; Niu, B.; Zhu, Z.; Wu, S.; Li, W. CD-HIT: Accelerated for clustering the next generation sequencing data. Bioinformatics 2012, 28, 3150–3152. [Google Scholar] [CrossRef]
- Li, W.; Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 2006, 22, 1658–1659. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef] [PubMed]
- Bokulich, N.A.; Kaehler, B.D.; Rideout, J.R.; Dillon, M.; Bolyen, E.; Knight, R.; Huttley, G.A.; Caporaso, J.G. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2′s q2-feature-classifier plugin. Microbiome 2018, 6, 90. [Google Scholar] [CrossRef] [PubMed]
- Hammer, Ø.; Harper, D.A.T.; Ryan, P.D. PAST: Paleontological Statistics software package for education and data analysis. Paleontol. Electron. 2001, 4, 9. [Google Scholar]
- Sola, F.; Vallejos, A.; Daniele, L.; Pulido-Bosch, A. Identification of a Holocene aquifer–lagoon system using hydrogeochemical data. Quat. Res. 2014, 82, 121–131. [Google Scholar] [CrossRef]
- Vallejos, A.; Sola, F.; Yechieli, Y.; Pulido-Bosch, A. Influence of the paleogeographic evolution on the groundwater salinity in a coastal aquifer. Cabo de Gata aquifer, SE Spain. J. Hydrol. 2018, 557, 55–66. [Google Scholar] [CrossRef]
- Kvålseth, T.O. Note on Biological Diversity, Evenness, and Homogeneity Measures. Oikos 1991, 62, 123–127. [Google Scholar] [CrossRef]
- Flynn, T.M.; Sandford, R.A.; Ryu, H.; Bethke, C.M.; Levne, A.D.; Ashbolt, N.J.; Santo Domingo, J.W. Functional microbial diversity explains groundwater chemistry in a pristine aquifer. BMC Microbiol. 2013, 13, 146. [Google Scholar] [CrossRef] [Green Version]
- Griebler, C.; Lueders, T. Microbial diversity in groundwater ecosystems. Freshw. Biol. 2009, 54, 649–677. [Google Scholar] [CrossRef]
- Boyd, E.S.; Cummings, D.E.; Geesey, G.G. Mineralogy influences structure and diversity of bacterial communities associated with geological substrata in a pristine aquifer. Microb. Ecol. 2007, 54, 170–182. [Google Scholar] [CrossRef]
- Kim, Y.H.; Ahn, Y. Ecology of groundwater microorganisms in aquifers. J. Life Sci. 2017, 27, 1086–1095. [Google Scholar] [CrossRef]
- Shi, Y.; Zwolinski, M.D.; Schreiber, M.E.; Bahr, J.M.; Sewell, G.W.; Hickey, W.J. Molecular analysis of microbial community structures in pristine and contaminated aquifers: Field and laboratory microcosm experiments. Appl. Environ. Microbiol. 1999, 65, 2143–2150. [Google Scholar] [CrossRef] [Green Version]
- Griebler, C.; Mindl, B.; Slezak, D.; Geiger-Kaiser, M. Distribution patterns of attached and suspended bacteria in pristine and contaminated shallow aquifers studied with an in situ sediment exposure microcosm. Aquat. Microb. Ecol. 2002, 28, 117–129. [Google Scholar] [CrossRef] [Green Version]
- Ma, J.; Liu, H.; Tong, L.; Wang, Y.; Chen, R.; Liu, S.; Zhao, L.; Li, Z.; Caiet, L. Relationships between microbial communities and groundwater chemistry in two pristine confined groundwater aquifers in central China. Hydrol. Process. 2019, 33, 1993–2005. [Google Scholar] [CrossRef]
- Rizoulis, A.; Elliot, D.R.; Rolfe, S.A.; Thornton, S.F.; Banwart, S.A.; Pickup, R.W.; Scholes, J.D. Diversity of planktonic and attached bacterial communities in a phenol-contaminated sandstone aquifer. Microb. Ecol. 2013, 66, 84–95. [Google Scholar] [CrossRef]
- Retter, A.; Karwautz, C.; Griebler, C. Groundwater microbial communities in times of climate change. Curr. Issues Mol. Biol. 2021, 41, 509–538. [Google Scholar] [CrossRef]
- Li, D.; Alidina, M.; Drewes, J.E. Role of primary substrate composition on microbial structure and function and trace organic chemical attenuation in managed aquifer recharge systems. Appl. Microbiol. Biotechnol. 2014, 98, 5747–5756. [Google Scholar] [CrossRef]
- Mak, M.S.H.; Rao, P.; Lo, I.M.C. Effects of hardness and alkalinity on the removal of arsenic(V) from humic acid-deficient and humic acid-rich groundwater by zero-valent iron. Water Res. 2009, 43, 4296–4304. [Google Scholar] [CrossRef]
- Feng, Q.; Li, X.; Cheng, Y.; Meng, L.; Meng, Q. Removal of humic acid from groundwater by electrocoagulation. J. China Univ. Min. Technol. 2007, 17, 513–520. [Google Scholar] [CrossRef]
- Hofmann, R.; Uhl, J.; Hertkorn, N.; Griebler, C. Linkage between dissolved organic matter transformation, bacterial carbon production, and diversity in a shallow oligotrophic aquifer: Results from flow-through sediment microcosm experiments. Front. Microbiol. 2020, 11, 2425. [Google Scholar] [CrossRef]
- Wiegand, S.; Jogler, M.; Jogler, C. On the maverick Planctomycetes. FEMS Microbiol. Rev. 2018, 42, 739–760. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fillinger, L.; Zhou, X.; Kellermann, C.; Griebler, C. Non-random processes determine the colonization of groundwater sediments by microbial communities in a pristine porous aquifer. Environ. Microbiol. 2019, 21, 327–342. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lazar, C.S.; Lehmann, R.; Stoll, W.; Rosenberger, J.; Totsche, K.U.; Küsel, K. The endolithic bacterial diversity of shallow bedrock ecosystems. Sci. Total Environ. 2019, 679, 35–44. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Hermans, S.M.; Totsche, K.U.; Lehmann, R.; Herrmann, M.; Küsel, K. Groundwater bacterial communities evolve over time in response to recharge. Water Res. 2021, 201, 117290. [Google Scholar] [CrossRef]
- Meng, L.; Zuo, R.; Wang, J.; Yang, J.; Li, Q.; Chen, Q. The spatial variations of correlation between microbial diversity and groundwater quality derived from a riverbank filtration site, northeast China. Sci. Total Environ. 2020, 706, 135855. [Google Scholar] [CrossRef]
- Yamamoto, K.; Hackley, K.; Kelly, W.; Panno, S.; Sekiguchi, Y.; Sanford, R.A.; Liu, W.; Kamagata, Y.; Tamaki, H. Diversity and geochemical community assembly processes of the living rare biosphere in a sand-and-gravel aquifer ecosystem in the Midwestern United States. Sci. Rep. 2019, 9, 13484. [Google Scholar] [CrossRef] [Green Version]
- Rocca, J.D.; Simonin, M.; Bernhardt, E.S.; Washburne, A.D.; Wright, J.P. Rare microbial taxa emerge when communities collide: Freshwater and marine microbiome responses to experimental mixing. Ecology 2020, 101, e02956. [Google Scholar] [CrossRef]
- Herrmann, M.; Wegner, C.E.; Taubert, M.; Geesink, P.; Lehmann, K.; Yan, L.; Lehmann, R.; Totsche, K.U.; Küselet, K. Predominance of Cand. Patescibacteria in groundwater is caused by their preferential mobilization from soils and flourishing under oligotrophic conditions. Front. Microbiol. 2019, 10, 1407. [Google Scholar] [CrossRef] [Green Version]
- Kundu, K.; Weber, N.; Griebler, C.; Elsner, M. Phenotypic heterogeneity as key factor for growth and survival under oligotrophic conditions. Environ. Microbiol. 2020, 22, 3339–3356. [Google Scholar] [CrossRef]
- Goldscheider, N.; Hunkeler, D.; Rossi, P. Review: Microbial biocenoses in pristine aquifers and an assessment of investigative methods. Hydrogeol. J. 2006, 14, 926–941. [Google Scholar] [CrossRef] [Green Version]
- Spiers, A.J.; Buckling, A.; Rainey, P.B. The causes of the Pseudomonas diversity. Microbiology 2000, 146, 2345–2350. [Google Scholar] [CrossRef] [Green Version]
- Luna, G.M.; Danovaro, R. Rapid identification of Pseudomonas spp. from aquatic samples using terminal restriction fragment length polymorphism analysis. J. Rapid Methods Autom. Microbiol. 2008, 16, 351–373. [Google Scholar] [CrossRef]
- Zhang, L.; Gao, G.; Tang, X.; Shao, K. Can the freshwater bacterial communities shift to the “marine-like” taxa? J. Basic Microbiol. 2014, 54, 1264–1272. [Google Scholar] [CrossRef]
- Jurelivicius, D.; Alvarez, V.M.; Marques, J.M.; de Sousa Lima, L.R.F.; Dias, F.D.A.; Seldin, L. Bacterial community response to petroleum hydrocarbon amendments in freshwater, marine, and hypersaline water-containing microcosms. Appl. Environ. Microbiol. 2013, 79, 5927–5935. [Google Scholar] [CrossRef] [Green Version]
- Li, P.; Jiang, D.; Li, B.; Dai, X.; Wang, Y.; Jiang, Z.; Wang, Y. Comparative survey of bacterial and archaeal communities in high arsenic shallow aquifers using 454 pyrosequencing and traditional methods. Ecotoxicology 2014, 23, 1878–1889. [Google Scholar] [CrossRef]
- Sar, P.; Paul, D.; Sarkar, A.; Bharadwaj, R.; Kazy, S.K. Microbiology of arsenic-contaminated groundwater. In Microbiology for Minerals, Metals, Materials and the Environment; Pandey, B.D., Natarajan, K.A., Eds.; CRC Press: Boca Raton, FL, USA, 2015; pp. 477–525. [Google Scholar]
- Dey, U.; Chatterjee, S.; Mondal, N.K. Isolation and characterization of arsenic-resistant bacteria and possible application in bioremediation. Biotechnol. Rep. 2016, 10, 1–7. [Google Scholar] [CrossRef] [Green Version]
- Tiquia, S.M.; Davis, D.; Hadid, H.; Kasparian, S.; Ismail, M.; Sahly, R.; Shim, J.; Singh, S.; Murray, K.S. Halophilic and halotolerant bacteria from river waters and shallow groundwater along the Rounge river of Southeastern Michigan. Environ. Technol. 2007, 28, 297–307. [Google Scholar] [CrossRef]
- Li, Y.; Huang, D.; Sun, W.; Sun, X.; Yan, G.; Gao, W.; Lin, H. Characterizing sediment bacterial community and identifying the biological indicators in a seawater-freshwater transition zone during the wet and dry seasons. Environ. Sci. Pollut. Res. 2022, 29, 41219–41230. [Google Scholar] [CrossRef]
- Welter, D.K.; Ruaud, A.; Henseler, Z.M.; De Jong, H.N.; van Coeverden de Groot, P.; Michaux, J.; Gormezano, L.; Walters, J.L.; Youngblut, N.D.; Ley, R.E. Free-living, psychrotrophic bacteria of the genus Psychrobacter are descendants of pathobions. MSystems 2021, 6, e00258-21. [Google Scholar] [CrossRef]
- Bañeras, L.; Llorens, L.; Díaz-Guerra, L.; Gispert, M.; Hernández-del Amo, E.; Massart, S.; Verdaguer, D. Resilience of microbial communities in Mediterranean soil after induced drought and manipulated UV radiation. Eur. J. Sol. Sci. 2022, 73, e13218. [Google Scholar] [CrossRef]
- Seki, T.; Matsumoto, A.; Omura, S.; Takahashi, Y. Distribution and isolation of strains belonging to the order Solirubrobacterales. J. Antibiot. 2015, 68, 763–766. [Google Scholar] [CrossRef] [PubMed]
- Sola, F.; Vallejos, A.; Lopez-Geta, J.A.; Pulido-Bosch, A. The role of aquifer media in improving the quality of seawater feed to desalination plants. Water Resour. Manag. 2013, 27, 1377–1392. [Google Scholar] [CrossRef]
- Stein, S.; Russak, A.; Sivan, O.; Yechieli, Y.; Rahav, E.; Oren, Y.; Kasher, R. Saline Groundwater from Coastal Aquifers as a Source for Desalination. Environ. Sci. Technol. 2016, 50, 1955–1963. [Google Scholar] [CrossRef] [PubMed]
| SAMPLE | pH | EC (µS/cm) | O2 (mg/L) | T (°C) | Ca (meq/L) | Mg (meq/L) | Na (meq/L) | K (meq/L) | Cl (meq/L) | HCO3 (meq/L) | SO4 (meq/L) | NO3 (mg/L) | DOC (mg/L) | %SW | W-TYPE |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| AND-30 | 7.14 | 3660 | 6.99 | 21.5 | 12.57 | 10.94 | 18.78 | 0.61 | 15.26 | 5.90 | 17.11 | 100.00 | 0.0004 | 2 | Na-Ca-Mg-SO4-Cl |
| AND-43 | 7.48 | 27,600 | 8.14 | 21.5 | 20.11 | 48.36 | 267.69 | 5.24 | 260.65 | 5.40 | 34.17 | 95.58 | 0.0026 | 45 | Na-Cl |
| AND-58 | 7.45 | 55,600 | 5.5 | 21.6 | 27.74 | 93.17 | 506.55 | 10.87 | 561.64 | 3.90 | 54.78 | 3.45 | 0.0005 | 98 | Na-Cl |
| AG-29 | 7.70 | 3970 | 8.16 | 21.3 | 5.14 | 7.15 | 31.88 | 0.61 | 29.90 | 3.90 | 4.93 | 30.28 | 0.0000 | 5 | Na-Cl |
| AG-34 | 7.84 | 29,800 | 6.75 | 21.2 | 16.22 | 53.21 | 274.67 | 5.65 | 293.37 | 3.60 | 29.98 | 30.88 | 0.0008 | 51 | Na-Cl |
| AG-46 | 7.73 | 47,800 | 8.16 | 20.8 | 21.31 | 86.60 | 419.65 | 9.16 | 480.96 | 2.70 | 47.60 | 4.21 | 0.0028 | 84 | Na-Cl |
| CG-10 | 7.35 | 9330 | 5.21 | 22.7 | 11.33 | 17.52 | 61.14 | 1.10 | 66.57 | 6.00 | 19.49 | 216.62 | 0.0026 | 11 | Na-Cl-SO4 |
| CG-18 | 7.47 | 25,600 | 6.68 | 22.3 | 16.92 | 39.56 | 225.76 | 4.27 | 233.85 | 6.40 | 37.16 | 21.57 | 0.0031 | 41 | Na-Cl |
| CG-38 | 7.31 | 57,400 | 6.01 | 22.7 | 32.98 | 102.22 | 537.12 | 10.90 | 592.95 | 4.20 | 60.15 | 0.94 | 0.0061 | 104 | Na-Cl |
| CG-62 | 7.05 | 65,700 | 4.58 | 23.3 | 37.72 | 109.62 | 606.99 | 12.51 | 691.11 | 4.20 | 68.71 | 0.51 | 0.0044 | 121 | Na-Cl |
| PAL-30 | 8.00 | 5040 | 7.92 | 22.4 | 5.24 | 10.77 | 42.79 | 0.66 | 38.93 | 3.80 | 4.93 | 12.33 | 0.0024 | 7 | Na-Mg-Cl |
| PAL-35 | 7.71 | 15,770 | 8.26 | 22.2 | 14.67 | 49.01 | 119.65 | 4.07 | 143.30 | 3.50 | 14.80 | 10.55 | 0.0013 | 25 | Na-Mg-Cl |
| PAL-36 | 7.40 | 48,400 | 8.61 | 22.2 | 28.99 | 110.28 | 432.31 | 10.26 | 486.32 | 3.10 | 48.84 | 1.64 | 0.0084 | 85 | Na-Cl |
| PAL-46 | 7.37 | 56,100 | 4.52 | 21.5 | 26.30 | 107.81 | 497.82 | 13.45 | 567.56 | 3.00 | 56.90 | 0.77 | 0.0014 | 99 | Na-Cl |
| SW | 7.99 | 54,500 | — | 16.4 | 28.89 | 106.33 | 510.92 | 11.33 | 572.07 | 2.90 | 58.65 | 0.00 | 0.0039 | 100 | Na-Cl |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vargas-García, M.d.C.; Sola, F.; Vallejos, Á. Comparative Study of Microbial Diversity in Different Coastal Aquifers: Determining Factors. Water 2023, 15, 1337. https://doi.org/10.3390/w15071337
Vargas-García MdC, Sola F, Vallejos Á. Comparative Study of Microbial Diversity in Different Coastal Aquifers: Determining Factors. Water. 2023; 15(7):1337. https://doi.org/10.3390/w15071337
Chicago/Turabian StyleVargas-García, María del Carmen, Fernando Sola, and Ángela Vallejos. 2023. "Comparative Study of Microbial Diversity in Different Coastal Aquifers: Determining Factors" Water 15, no. 7: 1337. https://doi.org/10.3390/w15071337





