Quantitative Microbial Risk Assessment for Private Wells in Flood-Impacted Areas
Abstract
:1. Introduction
2. Materials and Methods
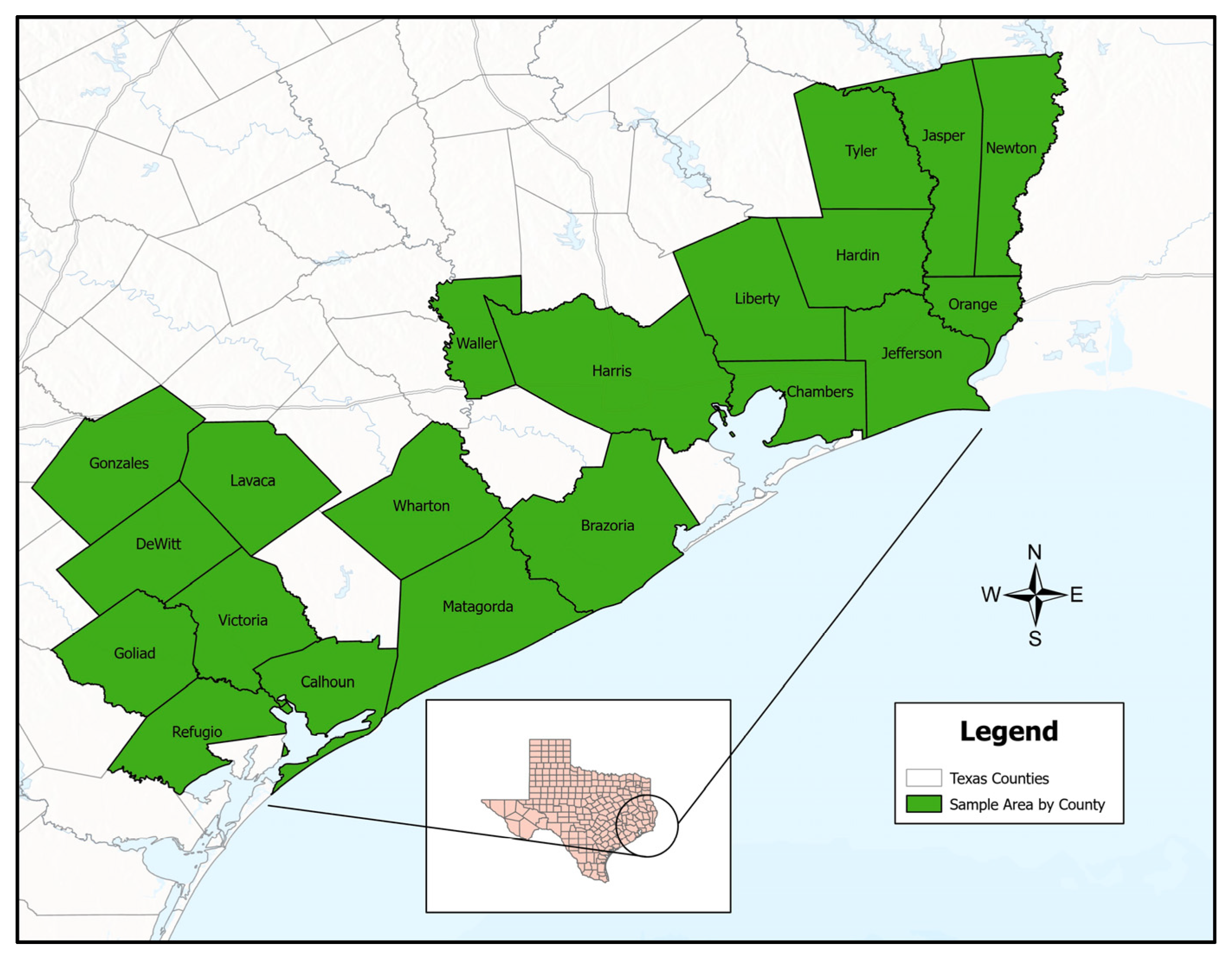
2.1. Sample Collection and Microbial Analysis
2.2. QMRA Methodology
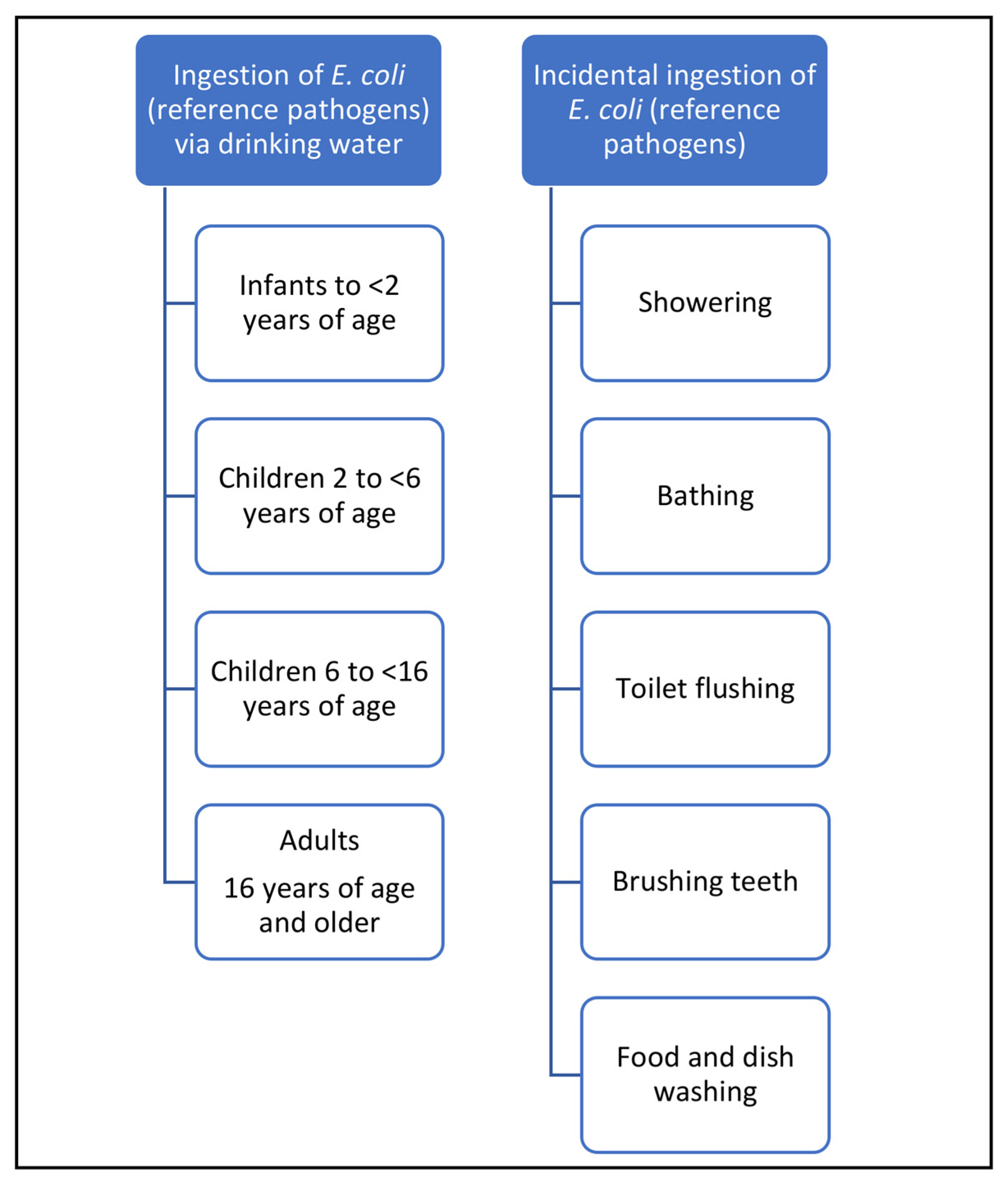
2.2.1. Exposure Models
2.2.2. Dose Calculation
| Parameters | Unit | Concentration | Source |
|---|---|---|---|
| E. coli concentration in well water | log MPN/100 mL | −4.835, 3.824 a | Environmental data |
| E. coli concentration in raw wastewater | log10 CFU/L | 6.7, 8.0 b | [46] |
| Norovirus concentration in raw wastewater | log10 copy/L | 4.7, 1.5 c | [28] |
| Cryptosporidium concentration in raw wastewater | log10 oocysts/L | −0.52, 4.7 b | [33,34,37,42,47] |
| Giardia concentration in raw wastewater | log10 cysts/L | 0.51, 4.2 b | [32,33,47] |
| Salmonella spp. concentration in raw wastewater | log10 CFU/L | 0.5, 5 b | [40,41,47] |
| E. coli O157:H7 concentration in raw wastewater | log10 CFU/L | −1, 3.3 b | [38,47] |
| Campylobacter concentration in raw wastewater | log10 MPN/L | 2.9, 4.6 b | [39,47] |
| Volume of water ingested (L) | |||
| Infants to < 2 | 0.82 d,e | [43] | |
| Children 2 to < 6 | 0.76 d,e | ||
| Children 6 to < 16 | 1.3 d,e | ||
| Adult | 2.5 d,e | ||
| Indirect ingestion (mL) | Showering | 0.058, 1.9 f,g | [48] |
| Bathing | 0.81, 63 h,i | [49] | |
| Brushing Teeth | 1.5 f,j | [50] | |
| Toilet Flushing | 0.01, 0.3 f,k | [51,52,53] | |
| Food and dish washing | 0.007, 0.008, 0.071 l | [54] |
3. Results
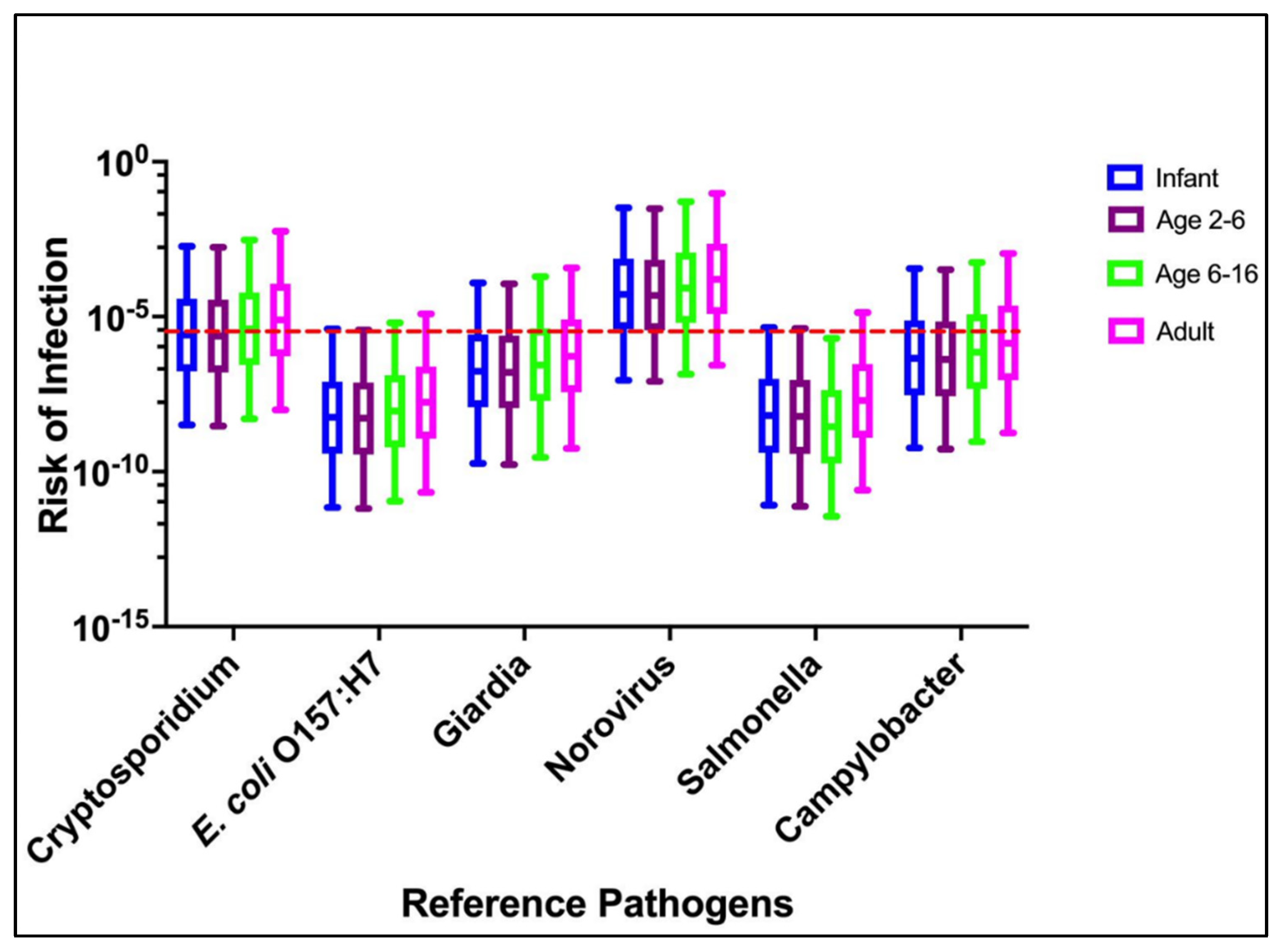
3.1. Scenario 1: Drinking Water
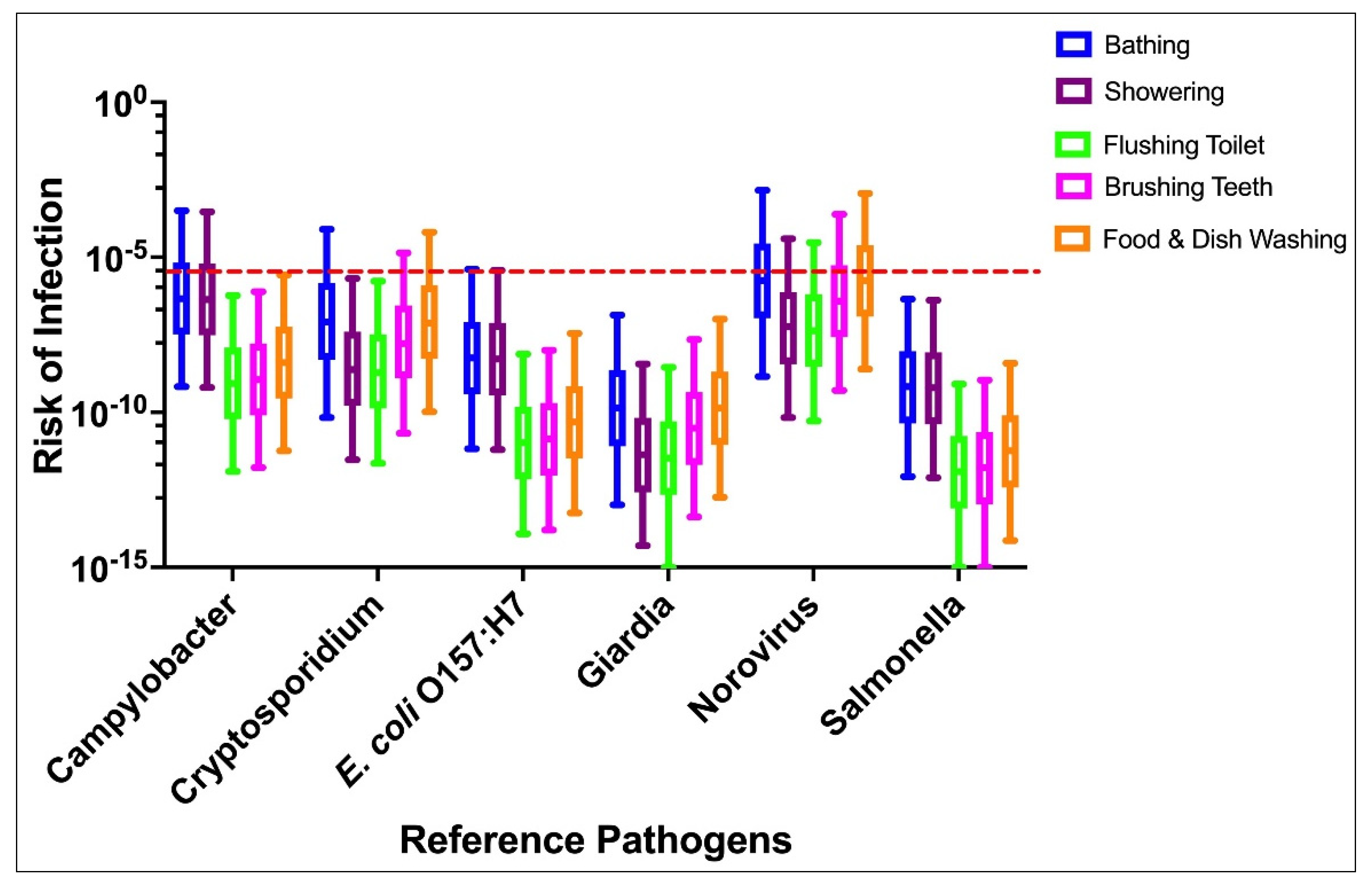
3.2. Scenario 2: Indirect Ingestion
3.3. Sensitivity Analysis
4. Discussion
4.1. Well Water Health Risks for Enteric Pathogens
4.2. Indicators for Evaluating Health Risks in Well Water
4.3. Barriers to Testing
4.4. Challenges
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- US EPA, Climate Change Indicators: Coastal Flooding. Available online: https://www.epa.gov/climate-indicators/climate-change-indicators-coastal-flooding (accessed on 1 November 2022).
- Andrade, L.; O’Dwyer, J.; O’Neill, E.; Hynds, P. Surface Water Flooding, Groundwater Contamination, and Enteric Disease in Developed Countries: A Scoping Review of Connections and Consequences. Environ. Pollut. 2018, 236, 540–549. [Google Scholar] [CrossRef] [PubMed]
- Charrois, J.W.A. Private Drinking Water Supplies: Challenges for Public Health. CMAJ 2010, 182, 1061–1064. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beitsch, R. Few Wells Tested for Contamination After Major Flooding from Hurricanes. Available online: https://pew.org/2LgC3Su (accessed on 1 November 2022).
- Gilliland, A.E.; Pieper, K.J.; Straif-Bourgeois, S.; Rhoads, W.J.; Dai, D.; Edwards, M.; Brisolara, K.; Olexia, D.; Katner, A. Evaluation of Preparedness and Recovery Needs of Private Well Users After the Great Louisiana Flood of 2016. J. Public Health Manag. Pract. 2021, 27, 577–587. [Google Scholar] [CrossRef] [PubMed]
- US EPA. What to Do After the Flood (816-F-05-021); US EPA Office of Water: Washington, DC, USA, 2005.
- Kapoor, V.; Gupta, I.; Pasha, A.B.M.T.; Phan, D. Real-Time Quantitative PCR Measurements of Fecal Indicator Bacteria and Human-Associated Source Tracking Markers in a Texas River Following Hurricane Harvey. Environ. Sci. Technol. Lett. 2018, 5, 322–328. [Google Scholar] [CrossRef]
- Blake, E.S.; Zelinsky, D.A. Hurricane Harvey (AL092017); National Oceanic and Atmospheric Administration and National Weather Service: Washington, DC, USA, 2018; p. 77.
- Pieper, K.J.; Jones, C.N.; Rhoads, W.J.; Rome, M.; Gholson, D.M.; Katner, A.; Boellstorff, D.E.; Beighley, R.E. Microbial Contamination of Drinking Water Supplied by Private Wells after Hurricane Harvey. Environ. Sci. Technol. 2021, 55, 8382–8392. [Google Scholar] [CrossRef]
- O’Sullivan, M.B.; Garvey, P.; O’Riordan, M.; Coughlan, H.; McKeown, P.; Brennan, A.; McNamara, E. Increase in VTEC Cases in the South of Ireland: Link to Private Wells? Eurosurveillance 2008, 13, 18991. [Google Scholar] [CrossRef]
- Hrudey, S.E.; Payment, P.; Huck, P.M.; Gillham, R.W.; Hrudey, E.J. A Fatal Waterborne Disease Epidemic in Walkerton, Ontario: Comparison with Other Waterborne Outbreaks in the Developed World. Water Sci. Technol. 2003, 47, 7–14. [Google Scholar] [CrossRef]
- Eccles, K.M.; Checkley, S.; Sjogren, D.; Barkema, H.W.; Bertazzon, S. Lessons Learned from the 2013 Calgary Flood: Assessing Risk of Drinking Water Well Contamination. Appl. Geogr. 2017, 80, 78–85. [Google Scholar] [CrossRef]
- Quist, A.J.L.; Fliss, M.D.; Wade, T.J.; Delamater, P.L.; Richardson, D.B.; Engel, L.S. Hurricane Flooding and Acute Gastrointestinal Illness in North Carolina. Sci. Total Environ. 2022, 809, 151108. [Google Scholar] [CrossRef]
- Masciopinto, C.; De Giglio, O.; Scrascia, M.; Fortunato, F.; La Rosa, G.; Suffredini, E.; Pazzani, C.; Prato, R.; Montagna, M.T. Human Health Risk Assessment for the Occurrence of Enteric Viruses in Drinking Water from Wells: Role of Flood Runoff Injections. Sci. Total Environ. 2019, 666, 559–571. [Google Scholar] [CrossRef]
- Stokdyk, J.; Borchardt, M.; Firnstahl, A.; Bradbury, K.; Muldoon, M.; Kieke, B. Assessing Private Well Contamination in Grant, Iowa, and Lafayette Counties, Wisconsin: The Southwest Wisconsin Groundwater and Geology Study; USGS: Reston, VA, USA, 2022; p. 66.
- Clark, C.G.; Price, L.; Ahmed, R.; Woodward, D.L.; Melito, P.L.; Rodgers, F.G.; Jamieson, F.; Ciebin, B.; Li, A.; Ellis, A. Characterization of Waterborne Outbreak–Associated Campylobacter Jejuni, Walkerton, Ontario. Emerg. Infect. Dis. 2003, 9, 1232–1241. [Google Scholar] [CrossRef] [Green Version]
- Haas, C.N.; Rose, J.B.; Gerba, C.P. Quantitative Microbial Risk Assessment, 2nd ed.; John Wiley & Sons: Hoboken, NJ, USA, 2014. [Google Scholar]
- Murphy, H.M.; Thomas, M.K.; Schmidt, P.J.; Medeiros, D.T.; McFadyen, S.; Pintar, K.D.M. Estimating the Burden of Acute Gastrointestinal Illness Due to Giardia, Cryptosporidium, Campylobacter, E. Coli O157 and Norovirus Associated with Private Wells and Small Water Systems in Canada. Epidemiol. Infect. 2016, 144, 1355–1370. [Google Scholar] [CrossRef] [Green Version]
- Burch, T.R.; Stokdyk, J.P.; Rice, N.; Anderson, A.C.; Walsh, J.F.; Spencer, S.K.; Firnstahl, A.D.; Borchardt, M.A. Statewide Quantitative Microbial Risk Assessment for Waterborne Viruses, Bacteria, and Protozoa in Public Water Supply Wells in Minnesota. Environ. Sci. Technol. 2022, 56, 6315–6324. [Google Scholar] [CrossRef]
- Hynds, P.D.; Gill, L.W.; Misstear, B.D. A Quantitative Risk Assessment of Verotoxigenic E. Coli (VTEC) in Private Groundwater Sources in the Republic of Ireland. Hum. Ecol. Risk Assess. Int. J. 2014, 20, 1446–1468. [Google Scholar] [CrossRef]
- Brouwer, S.; Van der Wielen, P.W.J.J.; Schriks, M.; Claassen, M.; Frijns, J. Public Participation in Science: The Future and Value of Citizen Science in the Drinking Water Research. Water 2018, 10, 284. [Google Scholar] [CrossRef] [Green Version]
- Buytaert, W.; Zulkafli, Z.; Grainger, S.; Acosta, L.; Alemie, T.C.; Bastiaensen, J.; De Bièvre, B.; Bhusal, J.; Clark, J.; Dewulf, A.; et al. Citizen Science in Hydrology and Water Resources: Opportunities for Knowledge Generation, Ecosystem Service Management, and Sustainable Development. Front. Earth Sci. 2014, 2, 26. [Google Scholar] [CrossRef] [Green Version]
- Sunger, N.; Hamilton, K.A.; Morgan, P.M.; Haas, C.N. Comparison of Pathogen-Derived ‘Total Risk’ with Indicator-Based Correlations for Recreational (Swimming) Exposure. Env. Sci. Pollut. Res. 2019, 26, 30614–30624. [Google Scholar] [CrossRef]
- Soller, J.A.; Schoen, M.E.; Bartrand, T.; Ravenscroft, J.E.; Ashbolt, N.J. Estimated Human Health Risks from Exposure to Recreational Waters Impacted by Human and Non-Human Sources of Faecal Contamination. Water Res. 2010, 44, 4674–4691. [Google Scholar] [CrossRef]
- US EPA. Quantitative Microbial Risk Assessment to Estimate Illness in Freshwater Impacted By; US EPA Office of Water: Washington, DC, USA, 2010; p. 456.
- Owens, C.E.L.; Angles, M.L.; Cox, P.T.; Byleveld, P.M.; Osborne, N.J.; Rahman, M.B. Implementation of Quantitative Microbial Risk Assessment (QMRA) for Public Drinking Water Supplies: Systematic Review. Water Res. 2020, 174, 115614. [Google Scholar] [CrossRef]
- Schoen, M.E.; Ashbolt, N.J. Assessing Pathogen Risk to Swimmers at Non-Sewage Impacted Recreational Beaches. Environ. Sci. Technol. 2010, 44, 2286–2291. [Google Scholar] [CrossRef]
- Eftim, S.E.; Hong, T.; Soller, J.; Boehm, A.; Warren, I.; Ichida, A.; Nappier, S.P. Occurrence of Norovirus in Raw Sewage—A Systematic Literature Review and Meta-Analysis. Water Res. 2017, 111, 366–374. [Google Scholar] [CrossRef] [PubMed]
- Chaban, B.; Ngeleka, M.; Hill, J.E. Detection and Quantification of 14 Campylobacter Species in Pet Dogs Reveals an Increase in Species Richness in Feces of Diarrheic Animals. BMC Microbiol 2010, 10, 73. [Google Scholar] [CrossRef] [Green Version]
- Hewitt, J.; Leonard, M.; Greening, G.E.; Lewis, G.D. Influence of Wastewater Treatment Process and the Population Size on Human Virus Profiles in Wastewater. Water Res. 2011, 45, 6267–6276. [Google Scholar] [CrossRef] [PubMed]
- Hurst, C.J.; McClellan, K.A.; Benton, W.H. Comparison of Cytopathogenicity, Immunofluorescence and In Situ DNA Hybridization as Methods for the Detection of Adenoviruses. Water Res. 1988, 22, 1547–1552. [Google Scholar] [CrossRef]
- Kitajima, M.; Haramoto, E.; Iker, B.C.; Gerba, C.P. Occurrence of Cryptosporidium, Giardia, and Cyclospora in Influent and Effluent Water at Wastewater Treatment Plants in Arizona. Sci. Total Environ. 2014, 484, 129–136. [Google Scholar] [CrossRef]
- Harwood, V.J.; Levine, A.D.; Scott, T.M.; Chivukula, V.; Lukasik, J.; Farrah, S.R.; Rose, J.B. Validity of the Indicator Organism Paradigm for Pathogen Reduction in Reclaimed Water and Public Health Protection. Appl. Environ. Microbiol. 2005, 71, 3163–3170. [Google Scholar] [CrossRef] [Green Version]
- Crockett, C.S. The Role of Wastewater Treatment in Protecting Water Supplies Against Emerging Pathogens. Water Environ. Res. 2007, 79, 221–232. [Google Scholar] [CrossRef]
- Schoen, M.E.; Ashbolt, N.J.; Jahne, M.A.; Garland, J. Risk-Based Enteric Pathogen Reduction Targets for Non-Potable and Direct Potable Use of Roof Runoff, Stormwater, and Greywater. Microb. Risk Anal. 2017, 5, 32–43. [Google Scholar] [CrossRef]
- Soller, J.A.; Schoen, M.; Steele, J.A.; Griffith, J.F.; Schiff, K.C. Incidence of Gastrointestinal Illness Following Wet Weather Recreational Exposures: Harmonization of Quantitative Microbial Risk Assessment with an Epidemiologic Investigation of Surfers. Water Res. 2017, 121, 280–289. [Google Scholar] [CrossRef]
- Nasser, A.M. Removal of Cryptosporidium by Wastewater Treatment Processes: A Review. J. Water Health 2015, 14, 1–13. [Google Scholar] [CrossRef]
- García-Aljaro, C.; Bonjoch, X.; Blanch, A.R. Combined Use of an Immunomagnetic Separation Method and Immunoblotting for the Enumeration and Isolation of Escherichia Coli O157 in Wastewaters. J. Appl. Microbiol. 2005, 98, 589–597. [Google Scholar] [CrossRef]
- Stampi, S.; Varoli, O.; Zanetti, F.; De Luca, G. Arcobacter cryaerophilus and Thermophilic campylobacters in a Sewage Treatment Plant in Italy: Two Secondary Treatments Compared. Epidemiol. Infect. 1993, 110, 633–639. [Google Scholar] [CrossRef] [Green Version]
- Lemarchand, K.; Lebaron, P. Occurrence of Salmonella Spp. and Cryptosporidium Spp. in a French Coastal Watershed: Relationship with Fecal Indicators. FEMS Microbiol. Lett. 2003, 218, 203–209. [Google Scholar] [CrossRef] [Green Version]
- Koivunen, J.; Siitonen, A.; Heinonen-Tanski, H. Elimination of Enteric Bacteria in Biological–Chemical Wastewater Treatment and Tertiary Filtration Units. Water Res. 2003, 37, 690–698. [Google Scholar] [CrossRef]
- Yang, J.; Schneider, O.D.; Jjemba, P.K.; Lechevallier, M.W. Microbial Risk Modeling for Main Breaks. J. AWWA 2015, 107, E97–E108. [Google Scholar] [CrossRef]
- MDEQ. Attachment H: Part 201 Generic Exposure Assumption Values Update; Michigan Department of Environmental Quality: Lansing, MI, USA, 2015.
- Gitter, A.; Mena, K.D.; Wagner, K.L.; Boellstorff, D.E.; Borel, K.E.; Gregory, L.F.; Gentry, T.J.; Karthikeyan, R. Human Health Risks Associated with Recreational Waters: Preliminary Approach of Integrating Quantitative Microbial Risk Assessment with Microbial Source Tracking. Water 2020, 12, 327. [Google Scholar] [CrossRef] [Green Version]
- Soller, J.A.; Eisenberg, J.N.S. An Evaluation of Parsimony for Microbial Risk Assessment Models. Environmetrics 2008, 19, 61–78. [Google Scholar] [CrossRef] [Green Version]
- Rose, J.B. Reduction of Pathogens, Indicator Bacteria, and Alternative Indicators by Wastewater Treatment and Reclamation Processes; IWA Publishing: London, UK, 2005; ISBN 978-1-84339-730-4. [Google Scholar]
- Boehm, A.B.; Graham, K.E.; Jennings, W.C. Can We Swim Yet? Systematic Review, Meta-Analysis, and Risk Assessment of Aging Sewage in Surface Waters. Environ. Sci. Technol. 2018, 52, 9634–9645. [Google Scholar] [CrossRef]
- Ahmed, W.; Vieritz, A.; Goonetilleke, A.; Gardner, T. Health Risk from the Use of Roof-Harvested Rainwater in Southeast Queensland, Australia, as Potable or Nonpotable Water, Determined Using Quantitative Microbial Risk Assessment. Appl. Environ. Microbiol. 2010, 76, 7382–7391. [Google Scholar] [CrossRef] [Green Version]
- Schets, F.M.; Schijven, J.F.; de Roda Husman, A.M. Exposure Assessment for Swimmers in Bathing Waters and Swimming Pools. Water Res. 2011, 45, 2392–2400. [Google Scholar] [CrossRef]
- Schijven, J.; Forêt, J.M.; Chardon, J.; Teunis, P.; Bouwknegt, M.; Tangena, B. Evaluation of Exposure Scenarios on Intentional Microbiological Contamination in a Drinking Water Distribution Network. Water Res. 2016, 96, 148–154. [Google Scholar] [CrossRef] [PubMed]
- National Water Quality Management Strategy. Australian Guidelines for Water Recycling: Managing Health and Environmental Risks (Phase 1); Natural Resource Management Ministerial Council; Environment Protection and Heritage Council; Australian Health Ministers’ Conference: Canberra, Australia, 2006.
- Schoen, M.E.; Xue, X.; Hawkins, T.R.; Ashbolt, N.J. Comparative Human Health Risk Analysis of Coastal Community Water and Waste Service Options. Environ. Sci. Technol. 2014, 48, 9728–9736. [Google Scholar] [CrossRef] [PubMed]
- Mayer, P.W.; DeOreo, W.B.; Opitz, E.M.; Kiefer, J.C.; Davis, W.Y.; Dziegielewski, B.; Nelson, J.O. Residential End Uses of Water; AWWA Research Foundation; American Water Works Association: Denver, CO, USA, 1999. [Google Scholar]
- An, W.; Zhang, D.; Xiao, S.; Yu, J.; Yang, M. Quantitative Health Risk Assessment of Cryptosporidium in Rivers of Southern China Based on Continuous Monitoring. Environ. Sci. Technol. 2011, 45, 4951–4958. [Google Scholar] [CrossRef] [PubMed]
- Teunis, P.F.M.; Nagelkerke, N.J.D.; Haas, C.N. Dose Response Models for Infectious Gastroenteritis. Risk Anal. 1999, 19, 1251–1260. [Google Scholar] [CrossRef]
- Teunis, P.F.M.; Ogden, I.D.; Strachan, N.J.C. Hierarchical Dose Response of E. Coli O157:H7 from Human Outbreaks Incorporating Heterogeneity in Exposure. Epidemiol. Infect. 2008, 136, 761–770. [Google Scholar] [CrossRef]
- Medema, G.J.; Teunis, P.F.M.; Havelaar, A.H.; Haas, C.N. Assessment of the Dose-Response Relationship of Campylobacter Jejuni. Int. J. Food Microbiol. 1996, 30, 101–111. [Google Scholar] [CrossRef]
- Rose, J.B.; Gerba, C.P. Use of Risk Assessment for Development of Microbial Standards. Water Sci. Technol. 1991, 24, 29–34. [Google Scholar] [CrossRef]
- Eisenberg, J.N.; Seto, E.Y.W.; Olivieri, A.W.; Spear, R.C. Quantifying Water Pathogen Risk in an Epidemiological Framework. Risk Anal. 1996, 16, 549–563. [Google Scholar] [CrossRef]
- U.S. EPA. National Primary Drinking Water Regulations: Long Term 2 Enhanced Surface Water Treatment Rule. Final Rule. Fed. Regist. 2006, 67, 1811–1844. [Google Scholar]
- Messner, M.J.; Berger, P.; Nappier, S.P. Fractional Poisson—A Simple Dose-Response Model for Human Norovirus. Risk Anal. 2014, 34, 1820–1829. [Google Scholar] [CrossRef]
- Haas, C.N.; Rose, J.B.; Gerba, C.P. Quantitative Microbial Risk Assessment, 1st ed.; John Wiley & Sons, Ltd.: New York, NY, USA, 1999. [Google Scholar]
- Van Abel, N.; Schoen, M.; Kissel, J.; Meschke, J. Comparison of Risk Predicted by Multiple Norovirus Dose-Response Models and Implications for Quantitative Microbial Risk Assessment. Risk Anal. Off. Publ. Soc. Risk Anal. 2017, 37, 245–264. [Google Scholar] [CrossRef] [PubMed]
- Vergara, G.G.R.V.; Rose, J.B.; Gin, K.Y.H. Risk Assessment of Noroviruses and Human Adenoviruses in Recreational Surface Waters. Water Res. 2016, 103, 276–282. [Google Scholar] [CrossRef] [PubMed]
- Boehm, A.B.; Soller, J.A.; Shanks, O.C. Human-Associated Fecal Quantitative Polymerase Chain Reaction Measurements and Simulated Risk of Gastrointestinal Illness in Recreational Waters Contaminated with Raw Sewage. Environ. Sci. Technol. Lett. 2015, 2, 270–275. [Google Scholar] [CrossRef]
- Hamilton, K.A.; Ahmed, W.; Toze, S.; Haas, C.N. Human Health Risks for Legionella and Mycobacterium Avium Complex (MAC) from Potable and Non-Potable Uses of Roof-Harvested Rainwater. Water Res. 2017, 119, 288–303. [Google Scholar] [CrossRef] [PubMed]
- Regli, S.; Rose, J.B.; Haas, C.N.; Gerba, C.P. Modeling the Risk From Giardia and Viruses in Drinking Water. J. AWWA 1991, 83, 76–84. [Google Scholar] [CrossRef]
- Signor, R.S.; Ashbolt, N.J. Comparing Probabilistic Microbial Risk Assessments for Drinking Water against Daily Rather than Annualised Infection Probability Targets. J. Water Health 2009, 7, 535–543. [Google Scholar] [CrossRef] [Green Version]
- Messner, M.J.; Berger, P. Cryptosporidium Infection Risk: Results of New Dose-Response Modeling. Risk Anal. 2016, 36, 1969–1982. [Google Scholar] [CrossRef]
- John, D.E.; Rose, J.B. Review of Factors Affecting Microbial Survival in Groundwater. Environ. Sci. Technol. 2005, 39, 7345–7356. [Google Scholar] [CrossRef] [Green Version]
- Toze, S.; Bekele, E.; Page, D.; Sidhu, J.; Shackleton, M. Use of Static Quantitative Microbial Risk Assessment to Determine Pathogen Risks in an Unconfined Carbonate Aquifer Used for Managed Aquifer Recharge. Water Res. 2010, 44, 1038–1049. [Google Scholar] [CrossRef]
- Sidhu, J.P.S.; Toze, S.; Hodgers, L.; Barry, K.; Page, D.; Li, Y.; Dillon, P. Pathogen Decay during Managed Aquifer Recharge at Four Sites with Different Geochemical Characteristics and Recharge Water Sources. J. Environ. Qual. 2015, 44, 1402–1412. [Google Scholar] [CrossRef]
- Loomer, D.B.; MacQuarrie, K.T.B.; Al, T.A.; Bragdon, I.K.; Loomer, H.A. Temporal Variability of Dissolved Methane and Inorganic Water Chemistry in Private Well Water in New Brunswick, Canada. Appl. Geochem. 2018, 94, 53–66. [Google Scholar] [CrossRef]
- Bauder, J.W.; Sinclair, K.N.; Lund, R.E. Physiographic and Land Use Characteristics Associated with Nitrate-Nitrogen in Montana Groundwater. J. Environ. Qual. 1993, 22, 255–262. [Google Scholar] [CrossRef]
- Rowles III, L.S.; Hossain, A.I.; Ramirez, I.; Durst, N.J.; Ward, P.M.; Kirisits, M.J.; Araiza, I.; Lawler, D.F.; Saleh, N.B. Seasonal Contamination of Well-Water in Flood-Prone Colonias and Other Unincorporated U.S. Communities. Sci. Total Environ. 2020, 740, 140111. [Google Scholar] [CrossRef]
- Stallard, M.A.; Mulhern, R.; Greenwood, E.; Franklin, T.; Engel, L.S.; Fisher, M.B.; Sobsey, M.D.; Zanib, H.; Noble, R.T.; Stewart, J.R.; et al. Occurrence of Male-Specific and Somatic Coliphages and Relationship with Rainfall in Privately-Owned Wells from Peri-urban and Rural Households. Water Res. X 2021, 12, 100102. [Google Scholar] [CrossRef]
- Mapili, K.; Pieper, K.J.; Dai, D.; Pruden, A.; Edwards, M.A.; Tang, M.; Rhoads, W.J. Legionella pneumophila Occurrence in Drinking Water Supplied by Private Wells. Lett. Appl. Microbiol. 2020, 70, 232–240. [Google Scholar] [CrossRef]
- Hexemer, A.; Pintar, K.; Bird, T.; Zentner, S.; Garcia, H.; Pollari, F. An Investigation of Bacteriological and Chemical Water Quality and the Barriers to Private Well Water Sampling in a Southwestern Ontario Community. J. Water Health 2008, 6, 521–525. [Google Scholar] [CrossRef]
- Wisconsin Environmental Public Health Tracking Program Reducing Barriers to Well Testing: Marquette County, Wisconsin 2019.
- Ishii, S.; Sadowsky, M.J. Escherichia Coli in the Environment: Implications for Water Quality and Human Health. Microb. Environ. 2008, 23, 101–108. [Google Scholar] [CrossRef] [Green Version]
- Dai, D.; Rhoads, W.J.; Katner, A.; Strom, L.; Edwards, M.A.; Pruden, A.; Pieper, K.J. Molecular Survey of Legionella and Naegleria Fowleri in Private Well Water and Premise Plumbing Following the 2016 Louisiana Flood. Environ. Sci. Water Res. Technol. 2019, 5, 1464–1477. [Google Scholar] [CrossRef]
| Pathogen | Probability of Infection | References |
| Salmonella spp. | 1 − (1 + dose/2884)−0.3126 | [55,62] |
| Campylobacter jejuni | 1 − (1 + (dose/7.59))−0.145 | [57] |
| E. coli O157:H7 | 1 − (1 + (dose/48.8))−0.248 | [56] |
| Cryptosporidium | 1 − exp(−0.09 × dose) | [60] |
| Giardia | 1 − exp(−0.01982 × dose) | [58,59] |
| Norovirus | 0.72 × (1 − exp(−dose/1)) | [61,63] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gitter, A.; Boellstorff, D.E.; Mena, K.D.; Gholson, D.M.; Pieper, K.J.; Chavarria, C.A.; Gentry, T.J. Quantitative Microbial Risk Assessment for Private Wells in Flood-Impacted Areas. Water 2023, 15, 469. https://doi.org/10.3390/w15030469
Gitter A, Boellstorff DE, Mena KD, Gholson DM, Pieper KJ, Chavarria CA, Gentry TJ. Quantitative Microbial Risk Assessment for Private Wells in Flood-Impacted Areas. Water. 2023; 15(3):469. https://doi.org/10.3390/w15030469
Chicago/Turabian StyleGitter, Anna, Diane E. Boellstorff, Kristina D. Mena, Drew M. Gholson, Kelsey J. Pieper, Carlos A. Chavarria, and Terry J. Gentry. 2023. "Quantitative Microbial Risk Assessment for Private Wells in Flood-Impacted Areas" Water 15, no. 3: 469. https://doi.org/10.3390/w15030469








