Karyotypes and COI Gene Sequences of Chironomus annularius Sensu Lato (Diptera, Chironomidae) from Russia, Mongolia, and Armenia
Abstract
:1. Introduction
2. Materials and Methods
3. Results
3.1. Morphological Characters of C. “annularius”
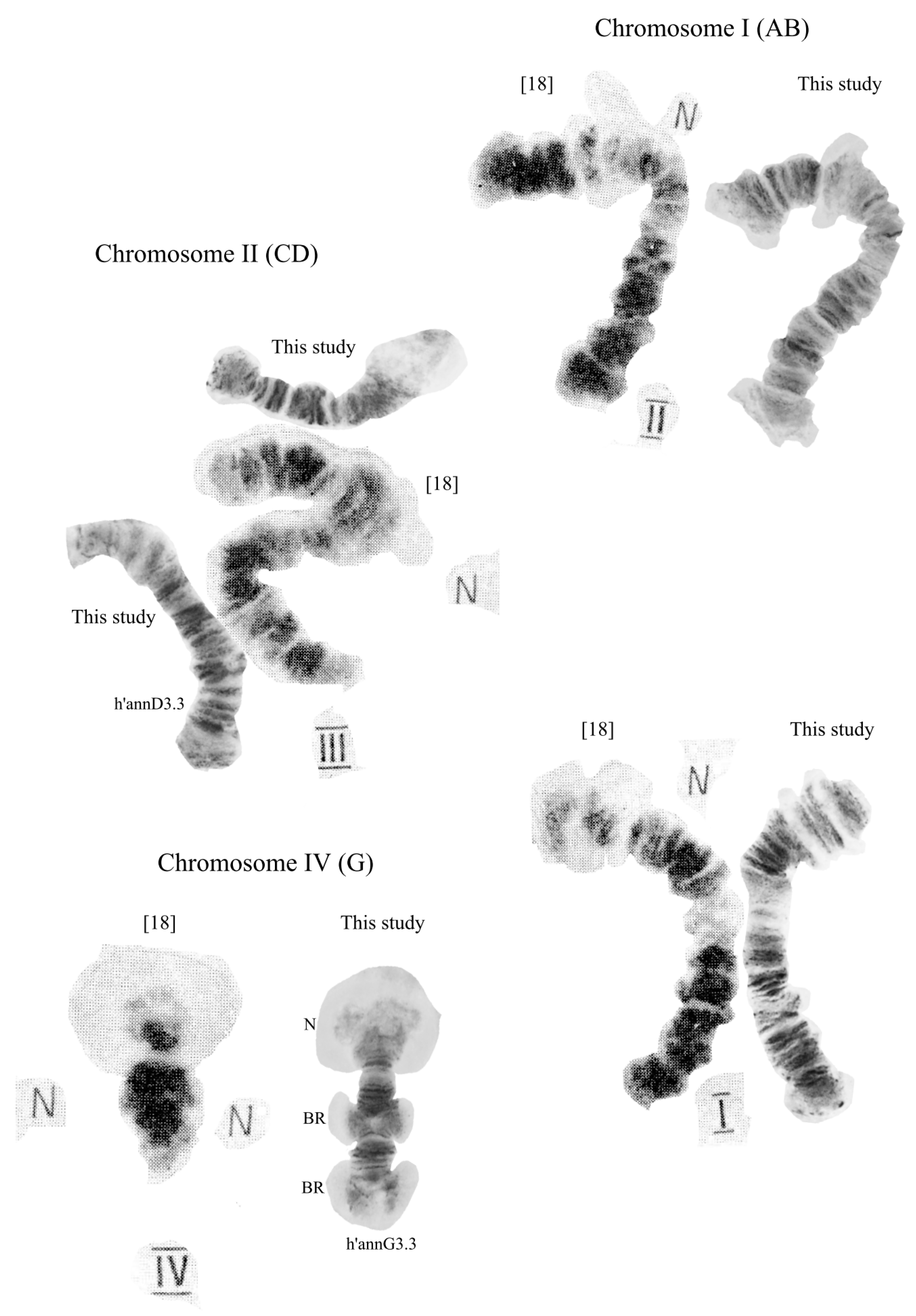
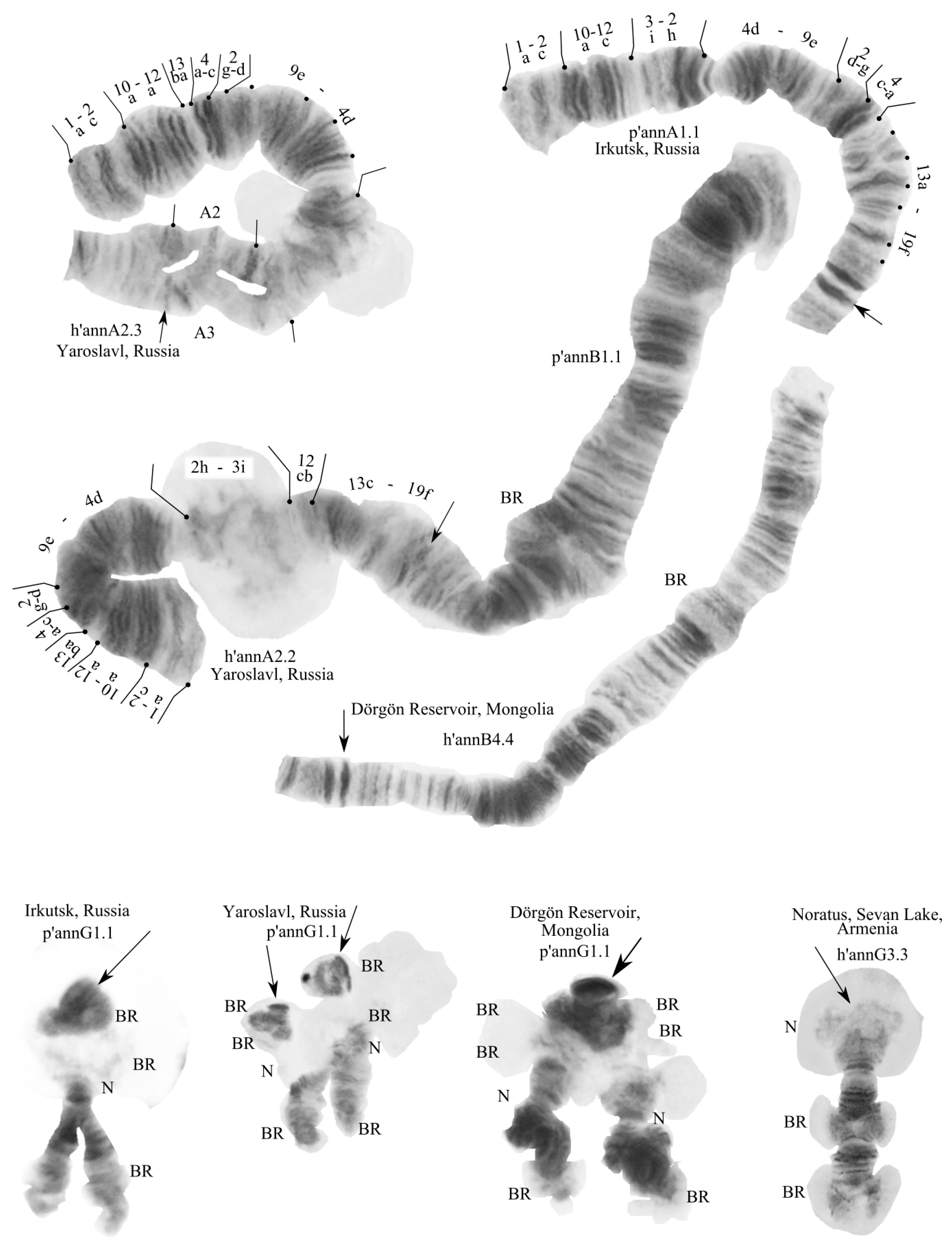
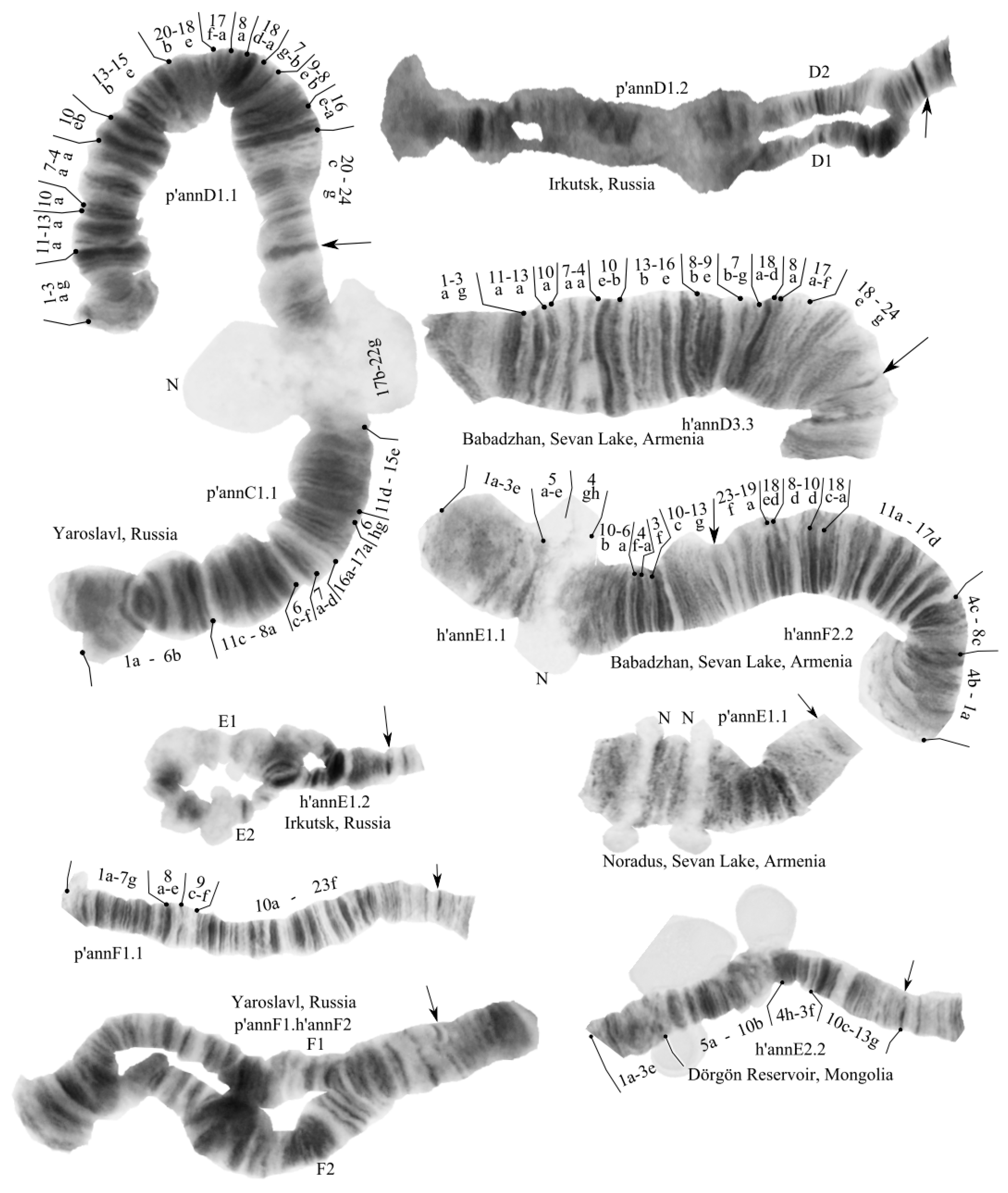
3.2. Karyotype of C. “annularius” from All Studied Populations
3.3. COI Gene Sequences and Phylogenetic Analysis of C. “annularius” from All Studied Populations
3.3.1. Genetic Distances between Chironomus “annularius” Obtained with K2P
3.3.2. Analysis of the Phylogenetic Tree
3.3.3. Analysis of the Haplotype Network
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A

Appendix B
References
- Spies, M.; Sæther, O.A. Notes and Recommendations on Taxonomy and Nomenclature of Chironomidae (Diptera). Zootaxa 2004, 752, 1. [Google Scholar] [CrossRef]
- Strenzke, K. Revision Der Gattung Chironomus Meig. i. Die Imagines von 15 Norddeutschen Arten Und Unterarten. Arch. Hydrobiol. 1959, 56, 1–42. [Google Scholar]
- Keyl, H.-G.; Keyl, I. Die Cytologische Diagnostik Der Chironomiden. I. Bestimmungstabelle Fur Die Gattung Chironomus Auf Grund Der Speicheldrusen-Chromosomen. Arch. Hydrobiol. 1959, 56, 43–57. [Google Scholar]
- Keyl, H.G. Chromosomenevolution Bei Chironomus. II. Chromosomenumbauten Und Phylogenetische Beziehungen Der Arten. Chromosoma 1962, 13, 464–514. [Google Scholar] [CrossRef]
- Shobanov, N.A. The Genus Chironomus Meigen (Diptera, Chironomidae) (Systematics, Biology, Evolution). Ph.D. Thesis, Zoological Institute of Russian Academy of Sciences, Saint-Petersburg, Russia, 2000. [Google Scholar]
- Karmokov, M.K. Karyotype Characteristics and Chromosomal Polymorphism of Chironomus “annularius” Sensu Strenzke (1959) (Diptera, Chironomidae) from the Caucasus Region. Comp. Cytogenet. 2018, 12, 267–284. [Google Scholar] [CrossRef]
- Kiknadze, I.I.; Istomina, A.G.; Golygina, V.V.; Gunderina, L.I. Karyotypes of Palearctic and Holarctic Species of the Genus Chironomus; Kiknadze, I.I., Ed.; Russian Academy of Sciences, Siberian Branch, Federal Research Center Institute of Cytology and Genetics, Academic Publishing House “GEO”: Novosibirsk, Russia, 2016; ISBN 9785990885325. [Google Scholar]
- Kiknadze, I.I.; Shilova, A.I.; Shobanov, N.A.; Zelentsov, N.I.; Grebenyuk, L.P.; Istomina, A.G.; Pratasov, V.A. Karyoypes and Larval Morphology in Tribe Chironomini. Atlas.; Nauka Publish: Novosibirsk, Russia, 1991; pp. 1–115. [Google Scholar]
- Kiknadze, I.I.; Istomina, A.G.; Gunderina, L.I.; Aimanova, K.G.; Savvinov, D.D. Banding Sequence Pools of Chironomid of Yakutian Permafrost. Tribe Chironomini; Nauka: Novosibirsk, Russia, 1996; pp. 1–166. [Google Scholar]
- Kiknadze, I.I.; Istomina, A.G.; Golygina, V.V. The Karyotype and Chromosome Polymorphism of the Holarctic Species Chironomus «annularius» Sensu Strenzke, 1959 (Diptera, Chironomidae). Euroasian Entomol. J. 2012, 11, 95–114. [Google Scholar]
- Belyanina, S.I. A Comparative Karyotypical Characteristics of Chironomus annularius (Diptera, Chironomidae) from Different Geographic Zones. Zool. Zhurnal 1981, 60, 1030–1039. [Google Scholar]
- Michailova, P.V. The Polytene Chromosomes and Their Significance to the Family Chironomidae, Diptera. Acta Zool. Fenn. 1989, 186, 1–107. [Google Scholar]
- Petrova, N.A.; Michailova, P.V. The Population-Karyological Studies of Some Chironomidae Species (Diptera, Chironomidae). Tsitologiia 1986, 28, 727–734. [Google Scholar]
- Butler, M.G.; Kiknadze, I.I.; Cooper, J.K.; Siirin, M.T. Cytologically Identified Chironomus Species from Lakes in North Dakota and Minnesota, USA. In Chironomids: From Genes to Ecosystems; Cranston, P.S., Ed.; CSIRO Publications: Canberra, Australia, 1995; pp. 31–38. [Google Scholar]
- Kiknadze, I.I.; Gunderina, L.I.; Butler, M.G.; Wuelker, W.F.; Martin, J. Chromosomes and Continents. In Biosphere Origin and Evolution; Springer: Boston, MA, USA, 2008; pp. 349–369. [Google Scholar] [CrossRef]
- Kiknadze, I.I.; Butler, M.G.; Gunderina, L.I.; Istomina, A.G.; Gusev, V.D.; Nemytikova, L.A. Chromosomal Evolution of Nearctic and Palearctic Chironomus Species (Diptera: Chironomidae). In Proceedings of the XV International Symposium on Chironomidae, Saint Paul, MA, USA, 12–15 August 2003; pp. 203–221. [Google Scholar]
- Petrova, N.A.; Zhirov, S.V.; Yerbayeva, E.A. Description of Three Species of Chironomids (Diptera, Chironomidae) from Lake Khubsugul, Mongolia (Morphological and Karyological Aspects). EUROASIAN Entomol. J. 2014, 13, 445–450. [Google Scholar]
- Pankratova, V.Y.; Chubareva, L.A.; Petrova, N.A. On the Taxonomy of Chironomus Species (Chironomidae) from Lake Sevan. Tr. Zool. Inst. Akad. Nauk. SSSR 1980, 95, 50–55. [Google Scholar]
- Chimeno, C.; Rulik, B.; Manfrin, A.; Kalinkat, G.; Hölker, F.; Baranov, V. Facing the Infinity: Tackling Large Samples of Challenging Chironomidae (Diptera) with an Integrative Approach. PeerJ 2023, 11, e15336. [Google Scholar] [CrossRef] [PubMed]
- Li, S.Y.; Zhao, Y.M.; Guo, B.X.; Li, C.H.; Sun, B.J.; Lin, X.L. Comparative Analysis of Mitogenomes of Chironomus (Diptera: Chironomidae). Insects 2022, 13, 1164. [Google Scholar] [CrossRef]
- Gadawski, P.; Montagna, M.; Rossaro, B.; Giłka, W.; Pešić, V.; Grabowski, M.; Magoga, G. DNA Barcoding of Chironomidae from the Lake Skadar Region: Reference Library and a Comparative Analysis of the European Fauna. Divers. Distrib. 2022, 28, 2838–2857. [Google Scholar] [CrossRef]
- Gadawski, P.; Rossaro, B.; Giłka, W.; Montagna, M.; Zawal, A.; Grabowski, M. First Insights into the Diversity and Ecology of Non-Biting Midges (Diptera: Chironomidae) of the Unique Ancient Skadar Lake Basin (Montenegro/Albania). J. Great Lakes Res. 2022, 48, 538–550. [Google Scholar] [CrossRef]
- Martin, J. Chromosomes as Tools in Taxonomy and Phylogeny of Chironomidae (Diptera). Entomol. Scand. 1979, 10, 67–74. [Google Scholar]
- Keyl, H.-G. Chromosomenevolution Bei Chironomus I. Strukturabwandlungen an Speicheldrüsen-Chromosomen. Chromosoma 1961, 12, 26–47. [Google Scholar] [CrossRef]
- Zamani, A.; Fric, Z.F.; Gante, H.F.; Hopkins, T.; Orfinger, A.B.; Scherz, M.D.; Bartoňová, A.S.; Pos, D.D. DNA Barcodes on Their Own Are Not Enough to Describe a Species. Syst. Entomol. 2022, 47, 385–389. [Google Scholar] [CrossRef]
- DeSalle, R.; Egan, M.G.; Siddall, M. The Unholy Trinity: Taxonomy, Species Delimitation and DNA Barcoding. Philos. Trans. R. Soc. B Biol. Sci. 2005, 360, 1905–1916. [Google Scholar] [CrossRef]
- Bolshakov, V.V.; Movergoz, E.A. Karyotype and COI Gene Sequences of Chironomus melanotus Keyl, 1961 from the Yaroslavl Region, Russia, and the Difficulties with Its Identification Using GenBank and BOLD Systems. Comp. Cytogenet. 2022, 16, 161–172. [Google Scholar] [CrossRef]
- Ilyinskaya, N.B. The Development of 4th Instar Larvae and Diapause. In Chironomus plumosus L. (Diptera, Chironomidae). Systematics, Morphology, Ecology, Production; Sokolova, N.Y., Ed.; Nauka Publishers: Moscow, Russia, 1983; pp. 167–188. [Google Scholar]
- Sæther, O. Glossary of Chironomid Morphology Terminology (Diptera: Chironomidae). Entomol. Scand. Suppl. 1980, 14, 1–51. [Google Scholar]
- Dyomin, S.Y. Variability of the Degree of Condensation of Polytene Chromosomes in the Cells of Different Organs of Chironomus plumosus Larvae from Nature. Ph.D. Thesis, Institute of Cytology of the USSR Academy of Sciences, Leningrad, Russia, 1989. [Google Scholar]
- Dévai, G.; Wülker, W.F.; Scholl, A. Revision Der Gattung Chironomus Meigen (Diptera). IX. C. Balatonicus Sp. n. Aus Flachsee Balaton. Acta Zool. Acad. Sci. Hung. 1983, 29, 357–374. [Google Scholar]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Kimura, M. A Simple Method for Estimating Evolutionary Rates of Base Substitutions through Comparative Studies of Nucleotide Sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian Phylogenetic Inference under Mixed Models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; van der Mark, P.; Huelsenbeck, J.P. Bayesian Phylogenetic Analysis Using MRBAYES. In The Phylogenetic Handbook; Cambridge University Press: Cambridge, UK, 2009; pp. 210–266. [Google Scholar]
- Karmokov, M.K. Karyotype Characteristics, Chromosomal Polymorphism and Gene COI Sequences of Chironomus Heteropilicornis Wülker, 1996 (Diptera, Chironomidae) from the South Caucasus. Comp. Cytogenet. 2019, 13, 339–357. [Google Scholar] [CrossRef] [PubMed]
- Bolshakov, V.V.; Prokin, A.A.; Artemenko, S. V Karyotype and COI Gene Sequence of Chironomus heteropilicornis Wülker, 1996 (Diptera, Chironomidae) from the Gydan Peninsula, Russia. Comp. Cytogenet. 2021, 15, 447–458. [Google Scholar] [CrossRef]
- Librado, P.; Rozas, J. DnaSP v5: A Software for Comprehensive Analysis of DNA Polymorphism Data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef]
- Leigh, J.W.; Bryant, D. POPART: Full-Feature Software for Haplotype Network Construction. Methods Ecol. Evol. 2015, 6, 1110–1116. [Google Scholar] [CrossRef]
- Webb, C.J.; Scholl, A. Identification of Larvae of European Species of Chironomus Meigen (Diptera: Chironomidae) by Morphological Characters. Syst. Entomol. 1985, 10, 353–372. [Google Scholar] [CrossRef]
- Shilova, A.I. A New Species of the Genus Chironomus (Diptera, Chironomidae) from Lake Sevan. Zool. Zhurnal 1983, 62, 245–251. [Google Scholar]
- Guryev, V.; Makarevitch, I.; Blinov, A.; Martin, J. Phylogeny of the Genus Chironomus (Diptera) Inferred from DNA Sequences of Mitochondrial Cytochrome b and Cytochrome Oxidase I. Mol. Phylogenet. Evol. 2001, 19, 9–21. [Google Scholar] [CrossRef] [PubMed]
- Gunderina, L.I.; Kiknadze, I.I.; Golygina, V.V. Differentiation of the Cytogenetic Structure of Natural Populations in the Plumosus Group of Sibling Species Chironomus balatonicus, Chironomus entis, Chironomus muratensis, Chironomus nudiventris (Chironomidae: Diptera). Genetica 1999, 35, 606–614. [Google Scholar]
- Shobanov, N.A.; Bolshakov, V.V. Karyofund of Chironomus plumosus (L.) (Diptera, Chironomidae): V. Terminal and Interstitial Populations. Cell Tissue Biol. 2011, 5, 196–201. [Google Scholar] [CrossRef]
- Bolshakov, V.; Movergoz, E.; Stolbov, V. Karyotypes and COI Gene Sequences of Chironomus agilis2, Ch. balatonicus, and Camptochironomus tentans (Diptera, Chironomidae) from Kurchatskoe Lake, Tyumen Region, Russia. Diversity 2022, 14, 1044. [Google Scholar] [CrossRef]
- Bolshakov, V.V.; Andreeva, A.M. Peculiarities of Structural Organization of Hemoglobin of Chironomus plumosus L. (Diptera: Chironomidae). J. Evol. Biochem. Physiol. 2012, 48, 265–271. [Google Scholar] [CrossRef]
- Osmulski, P.A.; Leyko, W. Structure, Function and Physiological Role of Chironomus Haemoglobin. Comp. Biochem. Physiol. Part B Comp. Biochem. 1986, 85, 701–722. [Google Scholar] [CrossRef]
- Bolshakov, V. V Variability of Karyofunds of Chironomus plumosus (Diptera, Chironomidae) from the Rybinsk Reservoir during the Period of Climate Warming. Inl. Water Biol. 2021, 14, 256–262. [Google Scholar] [CrossRef]
- Shilova, A.I.; Zelentsov, N.I. To the Chironomid Fauna of Lake Sevan (Diptera, Chironomidae). Biol. Inl. Waters. Inf. Bull. 1988, 79, 48–52. [Google Scholar]
- Makarevich, I.F.; Berezikov, E.V.; Guryev, V.P.; Blinov, A.G. Molecular Phylogeny of the Chironomus Genus Deduced from Nucleotide Sequences of Two Nuclear Genes, Ssp160 and the Globin 2b Gene. Mol. Biol. 2000, 34, 606–612. [Google Scholar] [CrossRef]
- Demin, A.G.; Polukonova, N. Divergence Time Estimation of Chironomids Chironomus Genus (Diptera) Using the «molecular Clock» Hypothesis. Entomol. Parasitol. Res. Volga Reg. 2008, 7, 8–13. [Google Scholar]
- Shobanov, N.A.; Zotov, S.D. Cytological Aspects of the Phylogeny of the Genus Chironomus Meigen (Diptera, Chironomidae). Entomol. Rev. 2001, 80, 180–193. [Google Scholar]
- Papusheva, E.; Gruhl, M.C.; Berezikov, E.; Groudieva, T.; Scherbik, S.V.; Martin, J.; Blinov, A.; Bergtrom, G. The Evolution of SINEs and LINEs in the Genus Chironomus (Diptera). J. Mol. Evol. 2004, 58, 269–279. [Google Scholar] [CrossRef]
- Guryev, V.P.; Blinov, A.G. Phylogenetic Relationships among Holarctic Populations of Chironomus entis and Chironomus plumosus in View of Possible Horisontal Transfer of Mitochondrial Genes. Russ. J. Genet. 2002, 38, 239–243. [Google Scholar] [CrossRef]
- Polukonova, N.V.; Demin, A.G.; Miuge, N.S.; Shaĭkevich, E.V.; Djomin, A.G.; Mugue, N.S.; Shaikevich, E.V.; Demin, A.G.; Miuge, N.S.; Shaĭkevich, E. V Comparison of Chironomus usenicus and Chironomus curabilis with Species of the Group plumosus (Diptera) Inferred from the Mitochondrial DNA Gene COI and by the Polytene Chromosomes Banding Pattern. Russ. J. Genet. 2009, 45, 1029–1035. [Google Scholar] [CrossRef]
- Bolshakov, V.; Prokin, A.; Pavlov, D.; Akkizov, A.; Movergoz, E. Karyotypes and COI Gene Sequences of Chironomus sp. Le1 (Kiknadze and Salova, 1996), Ch. laetus (Belyanina and Filinkova, 1996) and Their Hybrid from the Yamal Peninsula, Arctic Zone of Russia. Insects 2022, 13, 1112. [Google Scholar] [CrossRef]
- Proulx, I.; Martin, J.; Carew, M.; Hare, L. Using Various Lines of Evidence to Identify Chironomus Species (Diptera: Chironomidae) in Eastern Canadian Lakes. Zootaxa 2013, 3741, 401–458. [Google Scholar] [CrossRef] [PubMed]
- Ekrem, T.; Willassen, E.; Stur, E. A Comprehensive DNA Sequence Library Is Essential for Identification with DNA Barcodes. Mol. Phylogenet. Evol. 2007, 43, 530–542. [Google Scholar] [CrossRef]
- Kondo, N.I.; Ueno, R.; Ohbayashi, K.; Golygina, V.V.; Takamura, K. DNA Barcoding Supports Reclassification of Japanese Chironomus Species (Diptera: Chironomidae). Entomol. Sci. 2016, 19, 337–350. [Google Scholar] [CrossRef]
- Lin, X.; Stur, E.; Ekrem, T. Exploring Genetic Divergence in a Species-Rich Insect Genus Using 2790 DNA Barcodes. PLoS ONE 2015, 10, e0138993. [Google Scholar] [CrossRef]
- Song, C.; Lin, X.-L.; Wang, Q.; Wang, X.-H. DNA Barcodes Successfully Delimit Morphospecies in a Superdiverse Insect Genus. Zool. Scr. 2018, 47, 311–324. [Google Scholar] [CrossRef]
- Rewicz, T.; Tończyk, G.; Trębicki, Ł.; Gadawski, P.; Mamos, T.; Pešić, V.; Zawal, A.; Grabowski, M. DNA Barcode-Based Survey Documents Underestimated Diversity and Intricate Phylogeographic Patterns of Aquatic Heteroptera in an Endangered Balkan Biodiversity Hotspot: Ancient Lake Skadar Basin. Biodivers. Conserv. 2023. [Google Scholar] [CrossRef]
- Morinière, J.; Balke, M.; Doczkal, D.; Geiger, M.F.; Hardulak, L.A.; Haszprunar, G.; Hausmann, A.; Hendrich, L.; Regalado, L.; Rulik, B.; et al. A DNA Barcode Library for 5,200 German Flies and Midges (Insecta: Diptera) and Its Implications for Metabarcoding-based Biomonitoring. Mol. Ecol. Resour. 2019, 19, 900–928. [Google Scholar] [CrossRef]
- Martin, J. Personal Web Page. 2023. Available online: http://www.chironomidae.net/Martin/NACytfiles/Sp3d.htm (accessed on 20 August 2023).
- Weeraratne, T.C.; Surendran, S.N.; Reimer, L.J.; Wondji, C.S.; Perera, M.D.B.; Walton, C.; Parakrama Karunaratne, S.H.P. Molecular Characterization of Anopheline (Diptera: Culicidae) Mosquitoes from Eight Geographical Locations of Sri Lanka. Malar. J. 2017, 16, 234. [Google Scholar] [CrossRef] [PubMed]
- Batovska, J.; Blacket, M.J.; Brown, K.; Lynch, S.E. Molecular Identification of Mosquitoes (Diptera: Culicidae) in Southeastern Australia. Ecol. Evol. 2016, 6, 3001–3011. [Google Scholar] [CrossRef] [PubMed]
- Pfenninger, M.; Nowak, C.; Kley, C.; Steinke, D.; Streit, B. Utility of DNA Taxonomy and Barcoding for the Inference of Larval Community Structure in Morphologically Cryptic Chironomus (Diptera) Species. Mol. Ecol. 2007, 16, 1957–1968. [Google Scholar] [CrossRef]
- Panda, D.; Barik, T.K. Molecular Characterization and Genetic Divergence of Seven Culex Mosquito (Diptera: Culicidae) Species Using Mt COI Gene from Odisha State, India. J. Basic Appl. Zool. 2022, 83, 41. [Google Scholar] [CrossRef]
- Shobanov, N.A.; Kiknadze, I.I.; Butler, M.G. Palearctic and Nearctic Chironomus (Camptochironomus) tentans (Fabricius) Are Different Species (Diptera, Chironomidae). Entomol. Scand. 1999, 30, 311–322. [Google Scholar]
- Kiknadze, I.I.; Michailova, P.V.; Istomina, A.G.; Golygina, V.V.; Panis, L.I.; Krastanov, B. The Chromosomal Polymorphism and Divergence of Populations in Chironomus nuditarsis Str. (Diptera, Chironomidae). Tsitologiia 2006, 48, 595–609. [Google Scholar] [PubMed]


| Banding Sequences | Populations | |||
|---|---|---|---|---|
| Yaroslavl, Russia | Irkutsk, Russia | Dörgön Reservoir, Mongolia | Sevan Lake, Armenia | |
| A1 | * | * | ||
| A2 | * | * | ||
| A3 | * | |||
| B1 | * | * | ||
| B2 | * | |||
| B4 | * | * | ||
| C1 | * | * | * | * |
| D1 | * | * | ||
| D2 | * | * | ||
| D3 | * | |||
| E1 | * | * | * | |
| E2 | * | * | ||
| F1 | * | * | ||
| F2 | * | * | * | |
| G1 | * | * | * | |
| G3 | * | |||
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| No. | C. annularius AF192189 Novosibirsk | C. annularius OR342373 Yaroslavl Reg. | C. annularius OR342379 Irkutsk Reg. | C. annularius OR342375 Mongolia | C. annularius OR342376 Mongolia | C. annularius OR342381 Armenia | C. annularius OR342383 Armenia | C. annularius OR342387 Armenia | C. annularius ON975033 China | C. annularius XJDQD1011-18 China | C. annularius XJDQD1025-18 China | C. annularius XJDQD1026-18 China | C. annularius MZ626747 Finland | C. annularius MZ657059 Finland | C. annularius BSCHI129-17 Sweden | C. annularius BSCHI604-17 Sweden | C. annularius OP927527 Germany | C. annularius OP927683 Germany | C. annularius MT535143 Montenegro | C. annularius MT535187 Montenegro |
| 1 | ||||||||||||||||||||
| 2 | 4.4 | |||||||||||||||||||
| 3 | 4.4 | 1.1 | ||||||||||||||||||
| 4 | 4.4 | 1.9 | 2.4 | |||||||||||||||||
| 5 | 4.7 | 2.1 | 1.6 | 1.9 | ||||||||||||||||
| 6 | 7.0 | 5.3 | 5.0 | 5.0 | 4.7 | |||||||||||||||
| 7 | 7.0 | 5.3 | 5.0 | 5.0 | 4.7 | 0.0 | ||||||||||||||
| 8 | 7.0 | 5.3 | 5.0 | 5.0 | 4.7 | 0.0 | 0.0 | |||||||||||||
| 9 | 6.4 | 5.6 | 5.5 | 5.0 | 5.2 | 1.9 | 1.9 | 1.9 | ||||||||||||
| 10 | 4.1 | 0.8 | 0.8 | 2.2 | 1.9 | 5.2 | 5.2 | 5.2 | 5.8 | |||||||||||
| 11 | 3.8 | 0.5 | 0.5 | 1.9 | 1.6 | 5.0 | 5.0 | 5.0 | 5.5 | 0.3 | ||||||||||
| 12 | 3.8 | 0.5 | 0.5 | 1.9 | 1.6 | 5.0 | 5.0 | 5.0 | 5.5 | 0.3 | 0.0 | |||||||||
| 13 | 4.1 | 2.2 | 2.7 | 0.8 | 2.2 | 5.8 | 5.8 | 5.8 | 5.8 | 2.4 | 2.2 | 2.2 | ||||||||
| 14 | 4.1 | 2.2 | 2.7 | 0.8 | 2.2 | 5.8 | 5.8 | 5.8 | 5.8 | 2.4 | 2.2 | 2.2 | 0.0 | |||||||
| 15 | 3.8 | 0.5 | 0.5 | 1.9 | 1.6 | 5.0 | 5.0 | 5.0 | 5.5 | 0.3 | 0.0 | 0.0 | 2.2 | 2.2 | ||||||
| 16 | 4.1 | 2.2 | 2.7 | 0.8 | 2.2 | 5.8 | 5.8 | 5.8 | 5.8 | 2.4 | 2.2 | 2.2 | 0.0 | 0.0 | 2.2 | |||||
| 17 | 3.8 | 0.5 | 0.5 | 1.9 | 1.6 | 5.0 | 5.0 | 5.0 | 5.5 | 0.3 | 0.0 | 0.0 | 2.2 | 2.2 | 0.0 | 2.2 | ||||
| 18 | 3.8 | 0.5 | 0.5 | 1.9 | 1.6 | 5.0 | 5.0 | 5.0 | 5.5 | 0.3 | 0.0 | 0.0 | 2.2 | 2.2 | 0.0 | 2.2 | 0.0 | |||
| 19 | 5.8 | 5.5 | 6.1 | 6.1 | 5.8 | 6.4 | 6.4 | 6.4 | 6.1 | 5.8 | 5.5 | 5.5 | 5.2 | 5.2 | 5.5 | 5.2 | 5.5 | 5.5 | ||
| 20 | 6.1 | 5.5 | 6.1 | 6.1 | 5.5 | 6.7 | 6.7 | 6.7 | 6.4 | 5.8 | 5.5 | 5.5 | 5.2 | 5.2 | 5.5 | 5.2 | 5.5 | 5.5 | 0.3 | |
| 21 | 6.4 | 5.8 | 6.4 | 6.4 | 5.8 | 7.0 | 7.0 | 7.0 | 6.7 | 6.1 | 5.8 | 5.8 | 5.5 | 5.5 | 5.8 | 5.5 | 5.8 | 5.8 | 0.5 | 0.3 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bolshakov, V.; Prokin, A.; Mayor, T.; Hakobyan, S.; Movergoz, E. Karyotypes and COI Gene Sequences of Chironomus annularius Sensu Lato (Diptera, Chironomidae) from Russia, Mongolia, and Armenia. Water 2023, 15, 3114. https://doi.org/10.3390/w15173114
Bolshakov V, Prokin A, Mayor T, Hakobyan S, Movergoz E. Karyotypes and COI Gene Sequences of Chironomus annularius Sensu Lato (Diptera, Chironomidae) from Russia, Mongolia, and Armenia. Water. 2023; 15(17):3114. https://doi.org/10.3390/w15173114
Chicago/Turabian StyleBolshakov, Viktor, Alexander Prokin, Tatyana Mayor, Susanna Hakobyan, and Ekaterina Movergoz. 2023. "Karyotypes and COI Gene Sequences of Chironomus annularius Sensu Lato (Diptera, Chironomidae) from Russia, Mongolia, and Armenia" Water 15, no. 17: 3114. https://doi.org/10.3390/w15173114
APA StyleBolshakov, V., Prokin, A., Mayor, T., Hakobyan, S., & Movergoz, E. (2023). Karyotypes and COI Gene Sequences of Chironomus annularius Sensu Lato (Diptera, Chironomidae) from Russia, Mongolia, and Armenia. Water, 15(17), 3114. https://doi.org/10.3390/w15173114







