Are Indicator Microorganisms Predictive of Pathogens in Water?
Abstract
:1. Introduction
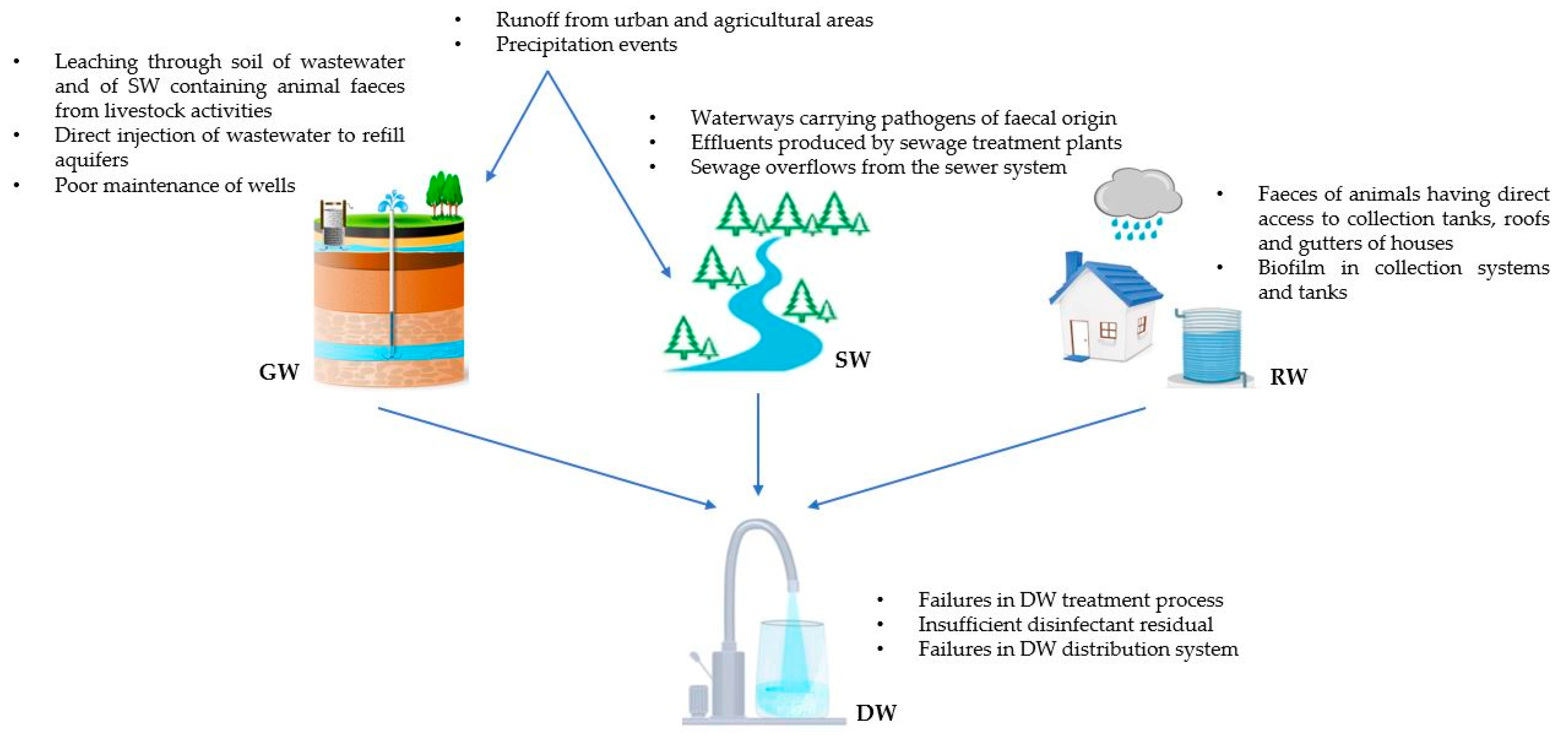
2. Origin of the Microbiological Contamination of DWSs and DW
3. Role of the IMs for the Evaluation of DW Safety
3.1. Total Coliforms
3.2. Thermotolerant Coliforms
3.3. Escherichia coli
3.4. Intestinal Enterococci
3.5. Heterotrophic Plate Count (HPC)
3.6. Clostridium perfringens and Its Spores
3.7. Bacteroides spp.
3.8. Coliphages
3.9. Methanobrevibacter smithii
3.10. Pepper Mild Mottle Virus
4. Analysis of the Relationship between IMs and Pathogens in DWSs and in DW
4.1. Groundwater
4.2. Surface Water
4.3. Rainwater
4.4. Drinking Water
5. Discussion
- The intrinsic characteristics of microorganisms, which, although similar, are not identical and therefore may present physiological differences, different decay rates, and different survival and multiplication capacities in the water environment [40];
- Seasonal variations, which affect the sources of water contamination and thus the presence and concentration of both IMs and pathogenic microorganisms (e.g., extreme weather events, urban discharges in tourist cities) [71];
- The high temporal and spatial fluctuation characterising pathogenic microorganisms, in contrast to IMs, in water environments, due to biotic and environmental factors, intrinsic characteristics of the pathogen, and its circulation in the population. This transient nature of pathogen occurrence in water means that correlation with IMs may exist at one point in time and not at another, even at the same geographic site [32]. In such a situation, the frequency of sampling becomes paramount to capture the moment of contamination [61];
- The fact that, especially in waters that are usually not heavily contaminated such as GW or DW, pathogens may not actually be present or may be present below the detection limit of the analytical method (e.g., human enteric viruses), which makes it difficult to assess the relationship with IMs [32];
- The number of samples on which the association is evaluated, as numerically large samples allow for smaller errors and thus greater reliability of the results of applied statistics. Several studies considered in this review have evaluated a small number of samples (n ≤ 22) [52,53,55,58,67]. The correlations found in these studies clearly have a different weight than those assessed on larger datasets (e.g., n = 1656 [50]; n = 964 [48]; n = 415 [69]), so it would be necessary to expand the sample size to validate the indicator–pathogen correlation;
- The volume of water sampled, as microorganisms in water environments are not uniformly distributed. This means that taking a few large-volume samples or many small-volume samples may lead to different results. Typically, the detection of pathogens requires large volumes of water (10–1000 L), because of their transient presence and generally lower concentrations than those of IMs. In contrast, the detection of IMs is normally carried out on 100–250 mL [66,74];
- The use of different detection methods (culture or molecular), characterised by different efficiency, sensitivity, and specificity, for the detection of IMs and pathogens may compromise correlation evaluation. Typically, indicator detection, especially of traditional indicators but not only, is based on the use of simple, rapid, and cost-effective culture techniques. Conversely, for most pathogens, the available culture techniques are expensive, difficult, time-consuming, and have low efficiency. For these reasons, pathogenic microorganisms are usually identified by molecular methods (PCR, qPCR, RT-qPCR). In several studies described in this work, a better ability of IMs to point out pathogen presence/absence was found when both targets were searched using the same detection method (i.e., molecular method) [59,60,62,65].
6. DW Guidelines and Quality Standards
6.1. WHO Guidelines
6.2. US EPA Regulations
- Maximum Contaminant Level Goal (MCLG) is the level of a contaminant in DW below which there is no known or expected health risk (also including risk for most sensitive people, such as infants, children, pregnant women, the elderly, and immunocompromised individuals). This goal allows for a margin of safety and is non-enforceable;
- Maximum Contaminant Level (MCL) is the highest level of a contaminant that is allowed in DW. MCLs are set as close as possible to MCLGs using the best available treatment technology and considering costs. These are enforceable standards;
- Treatment Technique (TT) is a required process intended to reduce the level of a contaminant in DW.
6.3. Canadian Guidelines
6.4. Australian Guidelines
6.5. Directive (EU) 2020/2184
7. A Focus on Quantitative Microbial Risk Assessment (QMRA)
8. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- World Health Organization. Drinking-Water. Available online: https://www.who.int/news-room/fact-sheets/detail/drinking-water (accessed on 9 August 2023).
- FAO; UNEP; WHO; WOAH. One Health Joint Plan of Action (2022–2026). Working Together for the Health of Humans, Animals, Plants and the Environment; FAO: Rome, Italy, 2022. [Google Scholar] [CrossRef]
- Zainurin, S.N.; Wan Ismail, W.Z.; Mahamud, S.N.I.; Ismail, I.; Jamaludin, J.; Ariffin, K.N.Z.; Wan Ahmad Kamil, W.M. Advancements in monitoring water quality based on various sensing methods: A systematic review. Int. J. Environ. Res. Public Health 2022, 19, 14080. [Google Scholar] [CrossRef] [PubMed]
- United Nations. The United Nations World Water Development Report 2023: Partnerships and Cooperation for Water; UNESCO: Paris, France, 2023. [Google Scholar]
- WASH. Compendium of WHO and Other UN Guidance on Health and Environment; World Health Organization: Geneva, Switzerland, 2021; (WHO/HEP/ECH/EHD/21.02). [Google Scholar]
- WHO. Water, Sanitation, Hygiene and Health: A Primer for Health Professionals; World Health Organization: Geneva, Switzerland, 2019; (WHO/CED/PHE/WSH/19.149). [Google Scholar]
- European Food Safety Authority; European Centre for Disease Prevention and Control. The European Union One Health 2021 Zoonoses Report. EFSA J. 2022, 20, 7666. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention—CDC. National Outbreak Reporting System (NORS). Available online: https://wwwn.cdc.gov/norsdashboard/ (accessed on 9 August 2023).
- UNESCO. UN-Water, 2020: United Nations World Water Development Report 2020: Water and Climate Change; UNESCO: Paris, France, 2020. [Google Scholar]
- WHO. Guidelines for Drinking-Water Quality: Fourth Edition Incorporating the First and Second Addenda; World Health Organization: Geneva, Switzerland, 2022. [Google Scholar]
- Figueras, M.J.; Borrego, J.J. New perspectives in monitoring drinking water microbial quality. Int. J. Environ. Res. Public Health 2010, 7, 4179–4202. [Google Scholar] [CrossRef] [PubMed]
- Gerba, C.P. Chapter 23—Indicator Microorganisms. In Environmental Microbiology, 3rd ed.; Pepper, I.L., Gerba, C.P., Gentry, T.J., Eds.; Academic Press: Cambridge, MA, USA, 2015; pp. 551–564. [Google Scholar]
- Saxena, G.; Bharagava, R.N.; Kaithwas, G.; Raj, A. Microbial indicators, pathogens and methods for their monitoring in water environment. J. Water Health 2015, 13, 319–339. [Google Scholar] [CrossRef] [PubMed]
- Fout, G.S.; Borchardt, M.A.; Kieke, B.A., Jr.; Karim, M.R. Human virus and microbial indicator occurrence in public-supply groundwater systems: Meta-analysis of 12 international studies. Hydrogeol. J. 2017, 25, 903–919. [Google Scholar] [CrossRef]
- United Nations. The United Nations World Water Development Report 2022: Groundwater: Making the Invisible Visible; UNESCO: Paris, France, 2022. [Google Scholar]
- Kumar, A.; Ranjan, A.; Gulati, K.; Thakur, S.; Jindal, T. Assessment of chemical and microbial contamination in groundwater through leaching of sewage waste in Delhi, India. Environ. Earth Sci. 2016, 75, 275. [Google Scholar] [CrossRef]
- Kumar, A.; Nirpen, L.; Ranjan, A.; Gulati, K.; Thakur, S.; Jindal, T. Microbial groundwater contamination and effective monitoring system. Asian J. Environ. Sci. 2014, 9, 37–48. [Google Scholar]
- World Health Organization—WHO. Protecting Surface Water for Health. Identifying, Assessing and Managing Drinking-Water Quality Risks in Surface-Water Catchments; World Health Organization: Geneva, Switzerland, 2016; ISBN 978 92 4 151612 9. [Google Scholar]
- Mackowiak, M.; Leifels, M.; Hamza, I.A.; Jurzik, L.; Wingender, J. Distribution of Escherichia coli, coliphages and enteric viruses in water, epilithic biofilms and sediments of an urban river in Germany. Sci. Total Environ. 2018, 626, 650–659. [Google Scholar] [CrossRef]
- Pandey, P.K.; Kass, P.H.; Soupir, M.L.; Biswas, S.; Singh, V.P. Contamination of water resources by pathogenic bacteria. AMB Express 2014, 4, 51. [Google Scholar] [CrossRef]
- Dean, J.; Hunter, P.R. Risk of gastrointestinal illness associated with the consumption of rainwater: A systematic review. Environ. Sci. Technol. 2012, 46, 2501–2507. [Google Scholar] [CrossRef]
- Hamilton, K.; Reyneke, B.; Waso, M.; Clements, T.; Ndlovu, T.; Khan, W.; DiGiovanni, K.; Rakestraw, E.; Montalto, F.; Haas, C.N.; et al. A global review of the microbiological quality and potential health risks associated with roof-harvested rainwater tanks. npj Clean Water 2019, 2, 7. [Google Scholar] [CrossRef]
- Ahmed, W.; Gardner, T.; Toze, S. Microbiological quality of roof-harvested rainwater and health risks: A review. J. Environ. Qual. 2011, 40, 13–21. [Google Scholar] [CrossRef]
- Hemdan, B.A.; El-Taweel, G.E.; Goswami, P.; Pant, D.; Sevda, S. The role of biofilm in the development and dissemination of ubiquitous pathogens in drinking water distribution systems: An overview of surveillance, outbreaks, and prevention. World J. Microbiol. Biotechnol. 2021, 37, 36. [Google Scholar] [CrossRef] [PubMed]
- Moreira, N.A.; Bondelind, M. Safe drinking water and waterborne outbreaks. J. Water Health. 2017, 15, 83–96. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, S. Heterotrophic bacteria in drinking water distribution system: A review. Environ. Monit. Assess. 2012, 184, 6087–6137. [Google Scholar] [CrossRef]
- Lee, E.J.; Schwab, K.J. Deficiencies in drinking water distribution systems in developing countries. J. Water Health. 2005, 3, 109–127. [Google Scholar] [CrossRef]
- Deshmukh, R.A.; Joshi, K.; Bhand, S.; Roy, U. Recent developments in detection and enumeration of waterborne bacteria: A retrospective minireview. Microbiologyopen 2016, 5, 901–922. [Google Scholar] [CrossRef] [PubMed]
- Haramoto, E.; Kitajima, M.; Hata, A.; Torrey, J.R.; Masago, Y.; Sano, D.; Katayama, H. A review on recent progress in the detection methods and prevalence of human enteric viruses in water. Water Res. 2018, 135, 168–186. [Google Scholar] [CrossRef]
- Motlagh, A.M.; Yang, Z. Detection and occurrence of indicator organisms and pathogens. Water Environ. Res. 2019, 91, 1402–1408. [Google Scholar] [CrossRef]
- Ramírez-Castillo, F.Y.; Loera-Muro, A.; Jacques, M.; Garneau, P.; Avelar-González, F.J.; Harel, J.; Guerrero-Barrera, A.L. Waterborne pathogens: Detection methods and challenges. Pathogens 2015, 4, 307–334. [Google Scholar] [CrossRef]
- Payment, P.; Locas, A. Pathogens in water: Value and limits of correlation with microbial indicators. Ground Water 2011, 49, 4–11. [Google Scholar] [CrossRef] [PubMed]
- Waideman, M.A.; Teixeira, V.P.; Uemura, E.H.; Stamford, T.M.; Leal, D.A.G.; Stangarlin-Fiori, L.; Ferreira, S.M.R.; Taconeli, C.A.; Beux, M.R. Enterococci used as complementary indicator of fecal contamination to assess water quality from public schools in the city of Curitiba, Paraná, Brazil. Braz. J. Food Technol. 2020, 23, e2019155. [Google Scholar] [CrossRef]
- Lin, J.; Ganesh, A. Water quality indicators: Bacteria, coliphages, enteric viruses. Int. J. Environ. Health Res. 2013, 23, 484–506. [Google Scholar] [CrossRef] [PubMed]
- Ben Braïek, O.; Smaoui, S. Enterococci: Between emerging pathogens and potential probiotics. Biomed Res. Int. 2019, 2019, 5938210. [Google Scholar] [CrossRef]
- Gensberger, E.T.; Gössl, E.M.; Antonielli, L.; Sessitsch, A.; Kostić, T. Effect of different heterotrophic plate count methods on the estimation of the composition of the culturable microbial community. PeerJ 2015, 3, e862. [Google Scholar] [CrossRef] [PubMed]
- Allen, M.J.; Edberg, S.C.; Reasoner, D.J. Heterotrophic plate count bacteria--what is their significance in drinking water? Int. J. Food Microbiol. 2004, 92, 265–274. [Google Scholar] [CrossRef]
- Stelma, G.N., Jr. Use of bacterial spores in monitoring water quality and treatment. J. Water Health 2018, 16, 491–500. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, W.; Goonetilleke, A.; Gardner, T. Alternative indicators for detection and quantification of faecal pollution. Water J. Aust. Water Assoc. 2008, 35, 59–65. [Google Scholar]
- Harwood, V.J.; Staley, C.; Badgley, B.D.; Borges, K.; Korajkic, A. Microbial source tracking markers for detection of fecal contamination in environmental waters: Relationships between pathogens and human health outcomes. FEMS Microbiol. Rev. 2014, 38, 1–40. [Google Scholar] [CrossRef]
- Fitzmorris-Brisolara, K.; Maal-Bared, R.; Worley-Morse, T.; Danley-Thomson, A.; Sobsey, M. Monitoring coliphages to reduce waterborne infectious disease transmission in the One Water framework. Int. J. Hyg. Environ. Health 2022, 240, 113921. [Google Scholar] [CrossRef]
- Bonadonna, L.; Briancesco, R.; Suffredini, E.; Coccia, A.; Della Libera, S.; Carducci, A.; Verani, M.; Federigi, I.; Iaconelli, M.; Bonanno Ferraro, G.; et al. Enteric viruses, somatic coliphages and Vibrio species in marine bathing and non-bathing waters in Italy. Mar. Pollut. Bull. 2019, 149, 110570. [Google Scholar] [CrossRef] [PubMed]
- Jofre, J.; Lucena, F.; Blanch, A.R.; Muniesa, M. Coliphages as model organisms in the characterization and management of water resources. Water 2016, 8, 199. [Google Scholar] [CrossRef]
- Gaci, N.; Borrel, G.; Tottey, W.; O’Toole, P.W.; Brugère, J.F. Archaea and the human gut: New beginning of an old story. World J. Gastroenterol. 2014, 20, 16062–16078. [Google Scholar] [CrossRef] [PubMed]
- Ufnar, J.A.; Wang, S.Y.; Christiansen, J.M.; Yampara-Iquise, H.; Carson, C.A.; Ellender, R.D. Detection of the nifH gene of Methanobrevibacter smithii: A potential tool to identify sewage pollution in recreational waters. J. Appl. Microbiol. 2006, 101, 44–52. [Google Scholar] [CrossRef] [PubMed]
- Maniah, K.; Nour, I.; Hanif, A.; Yassin, M.T.; Alkathiri, A.; Alharbi, Y.; Alotaibi, R.; Al-Anazi, A.E.; Eifan, S. Application of the Human Viral Surrogate Pepper Mild Mottle Virus for Wastewater Fecal Pollution Management. Water 2022, 14, 4033. [Google Scholar] [CrossRef]
- Bonanno Ferraro, G.; Suffredini, E.; Mancini, P.; Veneri, C.; Iaconelli, M.; Bonadonna, L.; Montagna, M.T.; De Giglio, O.; La Rosa, G. Pepper mild mottle virus as indicator of pollution: Assessment of prevalence and concentration in different water environments in Italy. Food. Environ. Virol. 2021, 13, 117–125. [Google Scholar] [CrossRef]
- Stokdyk, J.P.; Firnstahl, A.D.; Walsh, J.F.; Spencer, S.K.; de Lambert, J.R.; Anderson, A.C.; Rezania, L.W.; Kieke, B.A., Jr.; Borchardt, M.A. Viral, bacterial, and protozoan pathogens and fecal markers in wells supplying groundwater to public water systems in Minnesota. USA. Water Res. 2020, 178, 115814. [Google Scholar] [CrossRef]
- Richards, C.L.; Broadaway, S.C.; Eggers, M.J.; Doyle, J.; Pyle, B.H.; Camper, A.K.; Ford, T.E. Detection of pathogenic and non-pathogenic bacteria in drinking water and associated biofilms on the Crow Reservation, Montana, USA. Microb. Ecol. 2018, 76, 52–63. [Google Scholar] [CrossRef]
- De Giglio, O.; Barbuti, G.; Trerotoli, P.; Brigida, S.; Calabrese, A.; Di Vittorio, G.; Lovero, G.; Caggiano, G.; Uricchio, V.F.; Montagna, M.T. Microbiological and hydrogeological assessment of groundwater in southern Italy. Environ. Monit. Assess. 2016, 188, 638. [Google Scholar] [CrossRef]
- Ferguson, A.S.; Layton, A.C.; Mailloux, B.J.; Culligan, P.J.; Williams, D.E.; Smartt, A.E.; Sayler, G.S.; Feighery, J.; McKay, L.D.; Knappett, P.S.; et al. Comparison of fecal indicators with pathogenic bacteria and rotavirus in groundwater. Sci. Total Environ. 2012, 431, 314–322. [Google Scholar] [CrossRef]
- Haramoto, E.; Yamada, K.; Nishida, K. Prevalence of protozoa, viruses, coliphages and indicator bacteria in groundwater and river water in the Kathmandu Valley, Nepal. Trans. R. Soc. Trop. Med. Hyg. 2011, 105, 711–716. [Google Scholar] [CrossRef]
- Baker, K.H.; Hegarty, J.P. Presence of Helicobacter pylori in drinking water is associated with clinical infection. Scand. J. Infect. Dis. 2001, 33, 744–746. [Google Scholar] [CrossRef] [PubMed]
- Ashbolt, N.J. Microbial contamination of drinking water and human health from community water systems. Curr. Environ. Health Rep. 2015, 2, 95–106. [Google Scholar] [CrossRef]
- Bailey, E.S.; Hopkins, M.; Casanova, L.; Sobsey, M.D. Evaluating fecal indicator and pathogen relationships in sewage impacted surface waters to blend with reclaimed water for potable reuse in North Carolina. Pathogens 2021, 10, 1603. [Google Scholar] [CrossRef] [PubMed]
- Vadde, K.K.; McCarthy, A.J.; Rong, R.; Sekar, R. Quantification of microbial source tracking and pathogenic bacterial markers in water and sediments of Tiaoxi River (Taihu Watershed). Front. Microbiol. 2019, 10, 699. [Google Scholar] [CrossRef]
- Vesga, F.J.; Moreno, Y.; Ferrús, M.A.; Ledesma-Gaitan, L.M.; Campos, C.; Trespalacios, A.A. Correlation among fecal indicator bacteria and physicochemical parameters with the presence of Helicobacter pylori DNA in raw and drinking water from Bogotá, Colombia. Helicobacter 2019, 24, e12582. [Google Scholar] [CrossRef]
- Tandukar, S.; Sherchand, J.B.; Bhandari, D.; Sherchan, S.P.; Malla, B.; Ghaju Shrestha, R.; Haramoto, E. Presence of human enteric viruses, protozoa, and indicators of pathogens in the Bagmati River, Nepal. Pathogens 2018, 7, 38. [Google Scholar] [CrossRef]
- Liang, L.; Goh, S.G.; Vergara, G.G.; Fang, H.M.; Rezaeinejad, S.; Chang, S.Y.; Bayen, S.; Lee, W.A.; Sobsey, M.D.; Rose, J.B.; et al. Alternative fecal indicators and their empirical relationships with enteric viruses, Salmonella enterica, and Pseudomonas aeruginosa in surface waters of a tropical urban catchment. Appl. Environ. Microbiol. 2015, 81, 850–860. [Google Scholar] [CrossRef]
- Vergara, G.G.; Goh, S.G.; Rezaeinejad, S.; Chang, S.Y.; Sobsey, M.D.; Gin, K.Y. Evaluation of FRNA coliphages as indicators of human enteric viruses in a tropical urban freshwater catchment. Water Res. 2015, 79, 39–47. [Google Scholar] [CrossRef]
- Burnet, J.B.; Penny, C.; Ogorzaly, L.; Cauchie, H.M. Spatial and temporal distribution of Cryptosporidium and Giardia in a drinking water resource: Implications for monitoring and risk assessment. Sci. Total. Environ. 2014, 472, 1023–1035. [Google Scholar] [CrossRef]
- Rezaeinejad, S.; Vergara, G.G.; Woo, C.H.; Lim, T.T.; Sobsey, M.D.; Gin, K.Y. Surveillance of enteric viruses and coliphages in a tropical urban catchment. Water Res. 2014, 58, 122–131. [Google Scholar] [CrossRef]
- Edge, T.A.; Khan, I.U.; Bouchard, R.; Guo, J.; Hill, S.; Locas, A.; Moore, L.; Neumann, N.; Nowak, E.; Payment, P.; et al. Occurrence of waterborne pathogens and Escherichia coli at offshore drinking water intakes in Lake Ontario. Appl. Environ. Microbiol. 2013, 79, 5799–5813. [Google Scholar] [CrossRef] [PubMed]
- Krometis, L.A.; Characklis, G.W.; Drummey, P.N.; Sobsey, M.D. Comparison of the presence and partitioning behavior of indicator organisms and Salmonella spp. in an urban watershed. J. Water Health. 2010, 8, 44–59. [Google Scholar] [CrossRef] [PubMed]
- Lodder, W.J.; van den Berg, H.H.; Rutjes, S.A.; de Roda Husman, A.M. Presence of enteric viruses in source waters for drinking water production in The Netherlands. Appl. Environ. Microbiol. 2010, 76, 5965–5971. [Google Scholar] [CrossRef] [PubMed]
- Nieminski, E.; Durrant, G.C.; Hoyt, M.B.; Owens, M.E.; Peterson, L.; Peterson, S.; Tanner, W.D.; Rosen, J.; Clancy, J.L. Is E. Coli an appropriate surrogate for Cryptosporidium occurrence in water? J. Am. Water Works Assoc. 2010, 102, 65–78. [Google Scholar] [CrossRef]
- Cizek, A.R.; Characklis, G.W.; Krometis, L.A.; Hayes, J.A.; Simmons, O.D., III; Di Lonardo, S.; Alderisio, K.A.; Sobsey, M.D. Comparing the partitioning behavior of Giardia and Cryptosporidium with that of indicator organisms in stormwater runoff. Water Res. 2008, 42, 4421–4438. [Google Scholar] [CrossRef]
- Hachich, E.M.; Sato, M.I.; Galvani, A.T.; Menegon, J.R.; Mucci, J.L. Giardia and Cryptosporidium in source waters of São Paulo State, Brazil. Water Sci. Technol. 2004, 50, 239–245. [Google Scholar] [CrossRef]
- Payment, P.; Berte, A.; Prévost, M.; Ménard, B.; Barbeau, B. Occurrence of pathogenic microorganisms in the Saint Lawrence River (Canada) and comparison of health risks for populations using it as their source of drinking water. Can. J. Microbiol. 2000, 46, 565–576. [Google Scholar] [CrossRef]
- Ahmed, W.; Zhang, Q.; Ishii, S.; Hamilton, K.; Haas, C. Microfluidic quantification of multiple enteric and opportunistic bacterial pathogens in roof-harvested rainwater tank samples. Environ. Monit. Assess. 2018, 190, 105. [Google Scholar] [CrossRef]
- Hamilton, K.A.; Ahmed, W.; Palmer, A.; Smith, K.; Toze, S.; Haas, C.N. Seasonal assessment of opportunistic premise plumbing pathogens in roof-harvested rainwater tanks. Environ. Sci. Technol. 2017, 51, 1742–1753. [Google Scholar] [CrossRef]
- Hamilton, K.A.; Ahmed, W.; Palmer, A.; Sidhu, J.P.S.; Hodgers, L.; Toze, S.; Haas, C.N. Public health implications of Acanthamoeba and multiple potential opportunistic pathogens in roof-harvested rainwater tanks. Environ. Res. 2016, 150, 320–327. [Google Scholar] [CrossRef]
- Ahmed, W.; Brandes, H.; Gyawali, P.; Sidhu, J.P.; Toze, S. Opportunistic pathogens in roof-captured rainwater samples, determined using quantitative PCR. Water Res. 2014, 53, 361–369. [Google Scholar] [CrossRef]
- Ahmed, W.; Goonetilleke, A.; Gardner, T. Implications of faecal indicator bacteria for the microbiological assessment of roof-harvested rainwater quality in southeast Queensland, Australia. Can. J. Microbiol. 2010, 56, 471–479. [Google Scholar] [CrossRef]
- Simmons, G.; Hope, V.; Lewis, G.; Whitmore, J.; Gao, W. Contamination of potable roof-collected rainwater in Auckland, New Zealand. Water Res. 2001, 35, 1518–1524. [Google Scholar] [CrossRef] [PubMed]
- United States Environmental Protection Agency—US EPA. Understanding the Safe Drinking Water Act. EPA 816-F-04-030 June 2004. Available online: https://www.epa.gov/sdwa/overview-safe-drinking-water-act (accessed on 9 August 2023).
- United States Environmental Protection Agency—US EPA. National Primary Drinking Water Regulations. Available online: https://www.epa.gov/ground-water-and-drinking-water/national-primary-drinking-water-regulations (accessed on 9 August 2023).
- United States Environmental Protection Agency—US EPA. CCL 5 Microbial Contaminants. Available online: https://www.epa.gov/ccl/ccl-5-microbial-contaminants (accessed on 9 August 2023).
- Health Canada. Guidelines for Canadian Drinking Water Quality—Summary Tables; Water and Air Quality Bureau, Healthy Environments and Consumer Safety Branch, Health Canada: Ottawa, ON, Canada, 2022. [Google Scholar]
- NHMRC, NRMMC. Australian Drinking Water Guidelines Paper 6 National Water Quality Management Strategy; National Health and Medical Research Council, National Resource Management Ministerial Council, Commonwealth of Australia: Canberra, Australia, 2011. [Google Scholar]
- Directive (EU) 2020/2184 of the European Parliament and of the Council of 16 December 2020 on the Quality of Water Intended for Human Consumption. Available online: http://data.europa.eu/eli/dir/2020/2184/oj (accessed on 9 August 2023).
- Commission Directive (EU) 2015/1787 of 6 October 2015 amending Annexes II and III to Council Directive 98/83/EC on the Quality of Water Intended for Human Consumption. Available online: https://eur-lex.europa.eu/legal-content/EN/TXT/PDF/?uri=CELEX:01998L0083-20151027&from=EN (accessed on 9 August 2023).
- Rasheduzzaman, M.; Singh, R.; Haas, C.N.; Tolofari, D.; Yassaghi, H.; Hamilton, K.A.; Yang, Z.; Gurian, P.L. Reverse QMRA as a decision support tool: Setting acceptable concentration limits for Pseudomonas aeruginosa and Naegleria fowleri. Water 2019, 11, 1850. [Google Scholar] [CrossRef]
- Hamouda, M.A.; Jin, X.; Xu, H.; Chen, F. Quantitative microbial risk assessment and its applications in small water systems: A review. Sci. Total Environ. 2018, 645, 993–1002. [Google Scholar] [CrossRef] [PubMed]
- Amoueyan, E.; Ahmad, S.; Eisenberg, J.N.S.; Gerrity, D. Equivalency of indirect and direct potable reuse paradigms based on a quantitative microbial risk assessment framework. Microb. Risk Anal. 2019, 12, 60–75. [Google Scholar] [CrossRef]
- World Health Organization—WHO. Quantitative Microbial Risk Assessment: Application for Water Safety Management; World Health Organization: Geneva, Switzerland, 2016; ISBN 978 92 4 156537 0. [Google Scholar]
- Howard, G.; Pedley, S.; Tibatemwa, S. Quantitative microbial risk assessment to estimate health risks attributable to water supply: Can the technique be applied in developing countries with limited data? J. Water Health 2006, 4, 49–65. [Google Scholar] [CrossRef] [PubMed]
- Machdar, E.; van der Steen, N.P.; Raschid-Sally, L.; Lens, P.N. Application of Quantitative Microbial Risk Assessment to analyze the public health risk from poor drinking water quality in a low income area in Accra, Ghana. Sci. Total Environ. 2013, 449, 134–142. [Google Scholar] [CrossRef]
- Petterson, S.R.; Stenström, T.A.; Ottoson, J. A theoretical approach to using faecal indicator data to model norovirus concentration in surface water for QMRA: Glomma River, Norway. Water Res. 2016, 91, 31–37. [Google Scholar] [CrossRef]
- Petterson, S.R.; Mitchell, V.G.; Davies, C.M.; O’Connor, J.; Kaucner, C.; Roser, D.; Ashbolt, N. Evaluation of three full-scale stormwater treatment systems with respect to water yield, pathogen removal efficacy and human health risk from faecal pathogens. Sci. Total Environ. 2016, 543, 691–702. [Google Scholar] [CrossRef] [PubMed]
- Petterson, S.; Grøndahl-Rosado, R.; Nilsen, V.; Myrmel, M.; Robertson, L.J. Variability in the recovery of a virus concentration procedure in water: Implications for QMRA. Water Res. 2015, 87, 79–86. [Google Scholar] [CrossRef] [PubMed]
- van Lieverloo, J.H.; Blokker, E.J.; Medema, G. Quantitative microbial risk assessment of distributed drinking water using faecal indicator incidence and concentrations. J. Water Health 2007, 5, 131–149. [Google Scholar] [CrossRef] [PubMed]
- Ryan, M.; Hamilton, K.; Hamilton, M.; Haas, C.N. Evaluating the potential for a Helicobacter pylori drinking water guideline. Risk Anal. 2014, 34, 1651–1662. [Google Scholar] [CrossRef] [PubMed]
- Emelko, M.B.; Schmidt, P.J.; Borchardt, M.A. Confirming the need for virus disinfection in municipal subsurface drinking water supplies. Water Res. 2019, 157, 356–364. [Google Scholar] [CrossRef] [PubMed]
| IM | Significance | Surrogate for Pathogens |
|---|---|---|
| Total coliforms | Process indicators | - |
| Thermotolerant coliforms | Indicators of faecal contamination by humans and warm-blooded animals | - |
| Escherichia coli | Indicator of faecal contamination by humans and warm-blooded animals | Surrogate for bacterial enteric pathogens: pathogenic E. coli, Salmonella, Shigella |
| Intestinal enterococci | Indicators of faecal contamination by humans and warm-blooded animals | - |
| Heterotrophic plate count (HPC) | Process and water quality indicator | - |
| Clostridium perfringens and its spores | Indicator of remote faecal contamination by humans and warm-blooded animals | Surrogate for protozoan pathogens: Cryptosporidium, Giardia |
| Bacteroides spp. | Indicators of faecal contamination by humans and warm-blooded animals, they can be used to identify the source of faecal pollution | - |
| Coliphages | Indicators of faecal contamination by humans and warm-blooded animals | Surrogate for enteric viruses |
| Methanobrevibacter smithii | Indicator of human faecal contamination | - |
| Pepper mild mottle virus (PMMoV) | Indicator of human faecal contamination | Possible surrogate for enteric viruses |
| Reference and Publication Year | Country | GW Type | Number of Samples | IMs and Detection Method | Pathogens and Detection Method | Correlation |
|---|---|---|---|---|---|---|
| [48], 2020 | Minnesota, USA | Disinfected and undisinfected GW from public water supply wells 1 | 964 | Culture method: total coliforms, E. coli Molecular method: PMMoV, human Bacteroides HF183, Bacteroidales-like HumM2 | Molecular method: C. jejuni, enteropathogenic E. coli, Shiga-toxin-1- and 2-producing bacteria, Salmonella, adenovirus, human enterovirus, human polyomavirus, norovirus GI and GII, rotavirus group A, hepatitis A virus, Giardia, Cryptosporidium | E. coli (detected in only 3 samples) not considered for correlation. Hepatitis A virus, norovirus GI, enteropathogenic E. coli, and Shiga-toxin-1-producing bacteria never detected. Positive significant: for all IMs except PMMoV on total samples (logistic regression, p < 0.01). Not significant: for any IMs at each well (p > 0.05). |
| [49], 2018 | Montana, USA | GW used as DW | 57 2 | Culture method: total coliforms, E. coli, HPC | Culture method: Mycobacterium, Helicobacter Molecular method: Mycobacterium, Helicobacter | Positive significant: between HPC and Mycobacterium (logistic regression, p = 0.05). |
| [50], 2016 | Apulia region, Italy | GW from wells for emergency use | 1656 | Culture method: E. coli, total coliforms, enterococci, HPC 22 °C and 37 °C | Culture method: Salmonella spp. | Positive significant: between Salmonella spp. and E. coli, total coliforms and enterococci (Spearman correlation coefficient, p < 0.0001). No indication on correlation for HPC. |
| [51], 2012 | Bangladesh | GW from shallow tube wells as a primary source of DW | 50 | Culture method: total coliforms, E. coli, F+ RNA coliphages Molecular method: E. coli | Molecular method: Shigella, Vibrio, pathogenic E. coli, rotavirus | Positive significant: between pathogenic E. coli and culturable E. coli, total coliforms and F+ RNA coliphages; between molecular E. coli and Shigella (Spearman rank correlations, p < 0.05). |
| [52], 2011 | Katmandu Valley, Nepal | GW from shallow wells for daily water supply | 9 | Culture method: E. coli, total coliforms Molecular method: F-RNA coliphages | Immunofluorescent microscopy: Cryptosporidium, Giardia Molecular method: human adenoviruses, noroviruses GI and GII | Positive significant: 3 samples negative for E. coli and target pathogens; 5/6 E. coli-positive samples also positive for at least one of target pathogens (chi-square test, p < 0.05). No indication on correlation of pathogens with total coliforms and F-RNA coliphages. |
| [53], 2001 | Pennsylvania, USA | Private untreated well water consumed as DW | 22 | Culture method: E. coli | Culture method: H. pylori | Positive significant: between E. coli and H. pylori (Fisher’s exact test, p = 0.0011). |
| Reference and Publication Year | Country | SW Type | Number of Samples | Indicators and Detection Method | Pathogens and Detection Method | Correlation |
|---|---|---|---|---|---|---|
| [55], 2021 | North Carolina, USA | River water and sewage-impacted reservoirs, both sources for DW treatment plants | 22 | Culture method: total coliforms, E. coli, Enterococcus spp., C. perfringens, somatic and F+ coliphages | Immunofluorescent microscopy: Cryptosporidium, Giardia Culture method: Salmonella spp. Molecular method: adenovirus, norovirus | Positive significant:
Not significant: for any IMs by treatment plant (n = 5) (Spearman’s rank correlation, p > 0.05). Norovirus never detected. |
| [56], 2019 | China | Tiaoxi River, an important source of water supply | 45 | Culture method: faecal coliforms Molecular method: Bacteroidales (total, human-, swine-, and avian-associated) | Molecular method: C. jejuni, pathogenic Leptospira spp., Shigella spp., Shiga-toxin-producing E. coli (STEC), E. coli O157:H7 | Positive significant: between STEC and total and human Bacteroidales (Spearman’s coefficient correlation, p < 0.05). |
| [57], 2019 | Bogotá, Colombia | Catchment of raw superficial waters | 155 | Culture method: total coliforms, E. coli, spores of sulphite-reducing clostridia | Molecular method: H. pylori | Not significant: for any IMs (Spearman correlation coefficient and Tau-b Kendall correlation coefficient, p > 0.05). |
| [58], 2018 | Nepal | Bagmati River water used untreated for domestic use | 18 | Culture method: total coliforms, E. coli Molecular method: E. coli, Enterococcus spp., human Bacteroidales HF183, PMMoV | Immunofluorescent microscopy: Cryptosporidium, Giardia Molecular method: Aichi virus 1, enteroviruses, human cosaviruses, human adenoviruses, noroviruses GI and GII, group A rotaviruses, saliviruses | Positive significant: between all IMs and total human enteric viruses (bivariate correlation with Pearson coefficients, p < 0.05) |
| [59], 2015 | Singapore | Tributaries and reservoir used for DW | 148 | Culture method: E. coli, enterococci, somatic and F-coliphages Molecular method: E. coli, enterococci, Bacteroides thetaiotaomicron, M. smithii | Culture method: Salmonella spp. Molecular method: rotavirus, astrovirus, norovirus GI and GII, adenovirus | Positive significant: between Salmonella spp. and E. coli, enterococci, somatic and F-coliphages, and B. thetaiotaomicron (Spearman’s rank correlation, p < 0.05); between adenovirus and norovirus GII and molecular E. coli (p < 0.05) and molecular enterococci (p < 0.01); between norovirus GII and M. smithii (p < 0.05) and B. thetaiotaomicron (p < 0.01); between norovirus GI and M. smithii (p < 0.01). |
| [60], 2015 | Singapore | Marina Reservoir, an important source of DW | 75 | Culture method: somatic and F-coliphages Molecular method: F-coliphages | Molecular method: adenovirus, norovirus GI and GII, astrovirus, rotavirus | Positive significant: between cultural F-coliphages and norovirus GII and rotavirus; between molecular human-specific GII coliphages and all human enteric viruses (Kendall Tau-b correlation, p < 0.05). |
| [61], 2014 | Luxembourg | Raw SW (sub-catchments, river, reservoir, and inlet of a DW treatment plant) | 121 | Culture method: E. coli | Immunofluorescent microscopy: Cryptosporidium, Giardia | Positive significant: between E. coli and Giardia in the river and both protozoa in the reservoir and at the inlet of the DW treatment plant (nonparametric Spearman rank correlation test, p < 0.05). |
| [62], 2014 | Singapore | Reservoir for water supply | 65 | Culture method: somatic and male-specific coliphages | Molecular method: adenovirus, norovirus GI and GII, astrovirus, rotavirus | Positive significant: between male-specific coliphages and norovirus GI and GII (Kendall’s Tau-b correlation, p < 0.05). |
| [63], 2013 | Canada | Raw lake water entering three DW treatment plants (DWTP1, DWTP2, DWTP3) | 298 | Culture method: total coliforms, E. coli, enterococci, clostridia Molecular method: human Bacteroidales HF183 | Culture method: thermophilic Campylobacter, cultivable enteric viruses Immunofluorescent microscopy: Cryptosporidium, Giardia Molecular method: Cryptosporidium, Giardia | Positive significant: between enterococci and enteric viruses in DWTP1 and DWTP3 influents (Spearman rank correlation test, p < 0.05); between clostridia and enteric viruses in DWTP1 (p < 0.05); between clostridia and Cryptosporidium in DWTP2 influent (p < 0.05). |
| [64], 2010 | North Carolina, USA | Creek and pond tributaries of Jordan Lake, a DWS | 83 | Culture method: faecal coliforms, E. coli, enterococci, C. perfringens spores, somatic and male-specific coliphages | Culture method: Salmonella spp. | Positive significant: between Salmonella spp. and faecal coliforms, E. coli, enterococci, C. perfringens spores, and somatic coliphages (Spearman rank correlation, p < 0.05). |
| [65], 2010 | Netherlands | Source water for DW production (intake area or upstream of a source water intake area) | 75 | Culture method: somatic and F-specific coliphages | Culture method: enteroviruses, reoviruses Molecular method: noroviruses, rotaviruses | Positive significant: between the two coliphages and enteroviruses (p < 0.0005). |
| [66], 2010 | Utah, USA | Raw SW of seven DW treatment plants (reservoir and stream) | 228 | Culture method: E. coli | Immunofluorescent microscopy: Cryptosporidium | Negative significant: between E. coli and Cryptosporidium in winter for reservoir samples (nonparametric Spearman correlations). |
| [67], 2008 | New York, USA | Tributaries of the Kensico Reservoir, a primary source of DW for the city of New York | 16 | Culture method: faecal coliforms, E. coli, enterococci, C. perfringens spores, somatic and F+ coliphages | Immunofluorescent microscopy: Cryptosporidium, Giardia | Not significant: for any IMs (nonparametric Spearman rank statistical test, p > 0.5). |
| [68], 2004 | São Paulo, Brazil | SW used to supply DW | 278 | Culture method: faecal coliforms, C. perfringens | Immunofluorescent microscopy: Giardia, Cryptosporidium | Positive significant: between IMs and Giardia for total samples (Spearman rank correlation coefficient, p < 0.0001) and for only 3/10 watersheds (p < 0.05). Cryptosporidium not considered for correlation (low number of positive samples). |
| [69], 2000 | Canada | Raw river water | 415 | Culture method: total coliforms, faecal coliforms, C. perfringens | Culture method: cultivable human enteric viruses Immunofluorescent microscopy: Cryptosporidium, Giardia | Positive significant: between all IMs and all pathogens (Spearman correlation analysis, p < 0.001; logistic regression analysis, p ≤ 0.001). |
| Reference and Publication Year | Country | RW Type | Number of Samples | Indicators and Detection Method | Pathogens and Detection Method | Correlation |
|---|---|---|---|---|---|---|
| [70], 2018 | Australia | Roof-harvested RW for potable and non-potable use | 144 | Culture method: E. coli, Enterococcus spp. Molecular method: E. coli, Enterococcus spp. | Molecular method: enteropathogenic E. coli, Shiga-toxin-producing E. coli, Shigella spp., Salmonella spp., C. jejuni, C. lari, C. perfringens, Listeria monocytogenes, V. cholera, V. parahaemolyticus | Only Shigella spp. was detected. Negative significant: between IMs and Shigella spp. (odds ratio, p < 0.05). |
| [71], 2017 | Australia | Roof-harvested RW for potable and non-potable use | 144 | Culture method: E. coli, Enterococcus spp. | Molecular method: M. avium, M. intracellulare | Not significant: for any IMs (nonparametric Kendall’s Tau correlation, p > 0.05). |
| [72], 2016 | Australia | Roof-harvested RW for potable and non-potable use | 134 | Culture method: total coliforms, E. coli, Enterococcus spp. | Molecular method: M. avium, M. intracellulare | Positive significant: between M. intracellulare and E. coli and total coliforms (nonparametric Kendall’s Tau correlation, p < 0.05). Negative significant: between Enterococcus spp. and M. avium (p < 0.05). |
| [73], 2014 | Australia | Roof-captured RW for potable and non-potable use | 72 | Culture method: E. coli, Enterococcus spp. | Molecular method: A. hydrophila | Not significant: for any IMs (binary logistic regression analysis, p > 0.05). |
| [74], 2010 | Australia | Roof-harvested RW for drinking and non-potable use | 100 | Culture method: E. coli, enterococci, spore-forming C. perfringens | Molecular method: A. hydrophila, C. coli, C. jejuni, Salmonella, E. coli O157, Giardia lamblia, Cryptosporidium parvum | Not significant: for any IMs (binary logistic regression, p > 0.05). E. coli O157 and C. parvum never detected. |
| [75], 2001 | Auckland, New Zealand | Domestic roof-collected RW consumed as DW | 125 | Culture method: HPC, total coliforms, faecal coliforms, enterococci | Culture method: Salmonella spp., Campylobacter spp., Aeromonas spp. Immunofluorescent microscopy: Cryptosporidium, Giardia | Positive significant: between Aeromonas spp. and all IMs (p < 0.05). Campylobacter spp. and Giardia never detected. |
| Ever-Present Correlation | |
|---|---|
| IM | Pathogen |
| E. coli | Adenovirus Norovirus Shigella spp. |
| Bacteroides | Norovirus Salmonella spp. Shiga-toxin-2-producing bacteria (including E. coli) |
| Ever-Absent Correlation | |
| IM | Pathogen |
| E. coli | Campylobacter (coli and jejuni) Rotavirus |
| Enterococci | Campylobacter (coli and jejuni) Cryptosporidium Giardia |
| C. perfringens and its spores | Campylobacter (coli and jejuni) Helicobacter |
| PMMoV | Cryptosporidium Giardia |
| IM | Pathogen |
|---|---|
| Total coliforms | Cryptosporidium Salmonella spp. |
| E. coli | Giardia Salmonella spp. |
| Enterococci | Salmonella spp. |
| WHO Guidelines | US EPA Regulations | Canadian Guidelines | Australian Guidelines | Directive (EU) 2020/2184 | ||||
|---|---|---|---|---|---|---|---|---|
| Guideline values | MCLG 2 | MCL 2 | TT 2 | Maximum acceptable concentration | Treatment target | Non-mandatory standards | Parametric values | |
| Total coliforms | Zero | 5.0% 3 | - | 0/100 mL 7 | - | 0/100 mL 8 | ||
| Thermotolerant (faecal) coliforms | 0/100 mL | Zero | - 4 | - | 0/100 mL | |||
| E. coli | 0/100 mL 1 | Zero | - 4 | - | 0/100 mL | - | 0/100 mL | 0/100 mL 8 |
| Intestinal enterococci | 0/100 mL 8 | |||||||
| HPC | N/A | <500 colonies per millilitre | (HPC 22 °C) No abnormal change | |||||
| C. perfringens including spores | 0/100 mL 9 | |||||||
| Coliphages | 0/100 mL | (Somatic coliphages) 50 PFU/100 mL (for raw water) 10 | ||||||
| Cryptosporidium | Zero | - | 99% removal 5 | - | Minimum 3 log removal and/or inactivation of cysts and oocysts | |||
| Giardia lamblia | Zero | - | 99.9% removal or inactivation | - | Minimum 3 log removal and/or inactivation of cysts and oocysts | |||
| Legionella | Zero | - | - 6 | <1000 CFU/L 11 | ||||
| Enteric viruses | Zero | - | 99.9% removal or inactivation | - | Minimum 4 log reduction (removal and/or inactivation) | |||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Richiardi, L.; Pignata, C.; Fea, E.; Bonetta, S.; Carraro, E. Are Indicator Microorganisms Predictive of Pathogens in Water? Water 2023, 15, 2964. https://doi.org/10.3390/w15162964
Richiardi L, Pignata C, Fea E, Bonetta S, Carraro E. Are Indicator Microorganisms Predictive of Pathogens in Water? Water. 2023; 15(16):2964. https://doi.org/10.3390/w15162964
Chicago/Turabian StyleRichiardi, Lisa, Cristina Pignata, Elisabetta Fea, Silvia Bonetta, and Elisabetta Carraro. 2023. "Are Indicator Microorganisms Predictive of Pathogens in Water?" Water 15, no. 16: 2964. https://doi.org/10.3390/w15162964





