Temporal and Spatial Variations of the Bacterial Diversity in a Deep Alkaline Lake
Abstract
:1. Introduction
2. Materials and Methods
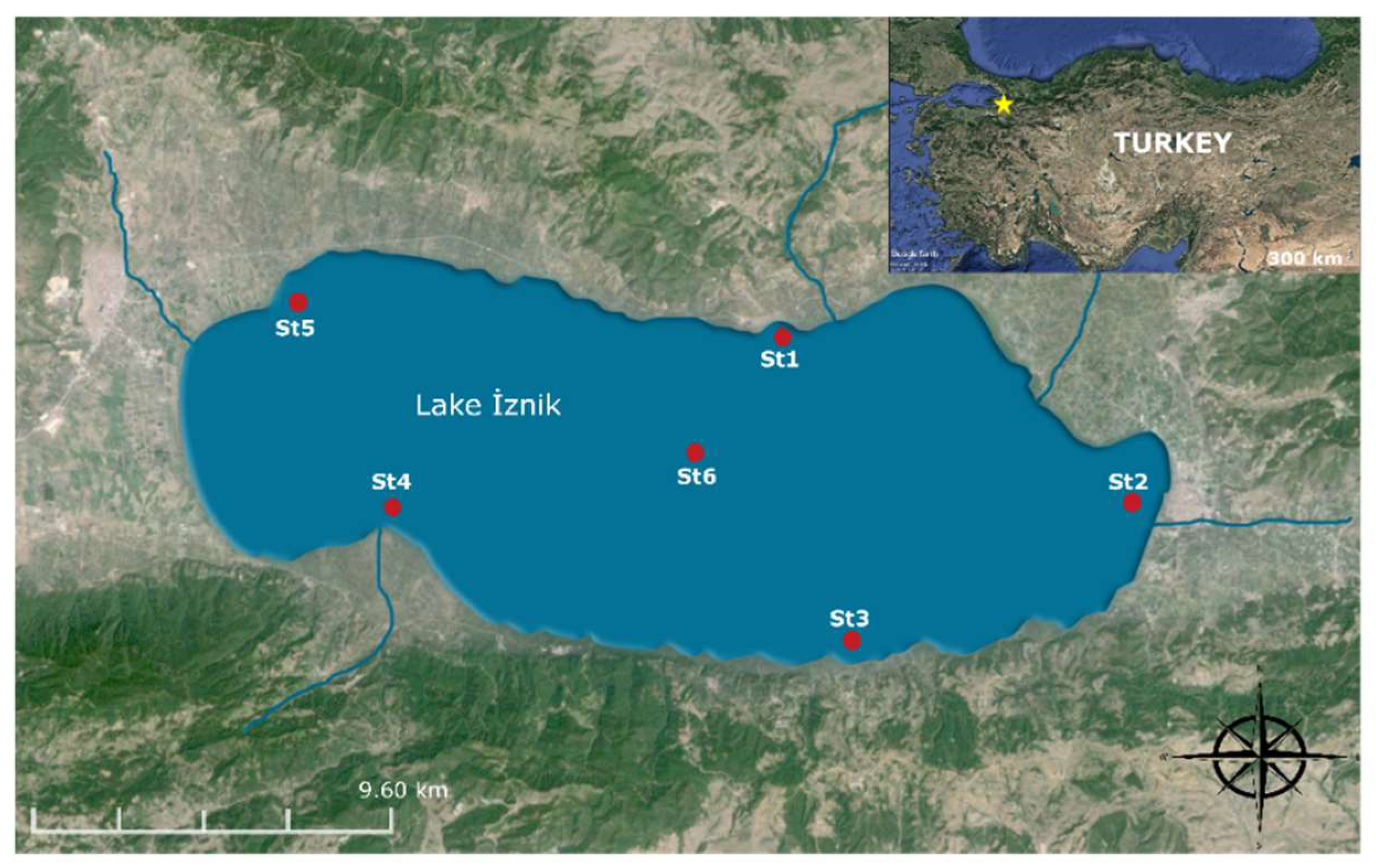
2.1. Site Description
2.2. Sample Collection and Physicochemical Analysis
2.3. DNA Extraction and Amplicon Sequencing
2.4. Sequence Analysis
3. Results
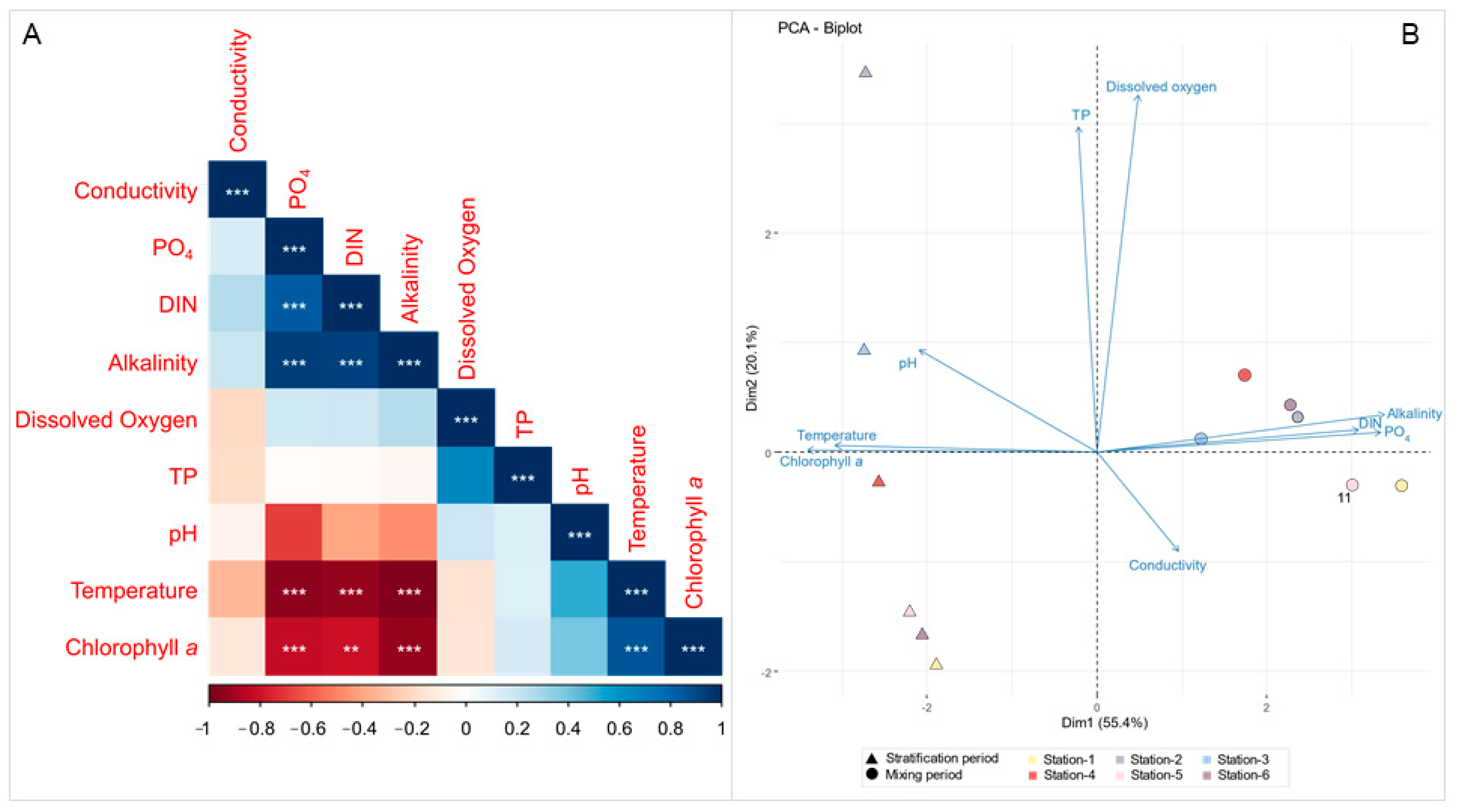
3.1. Environmental Conditions
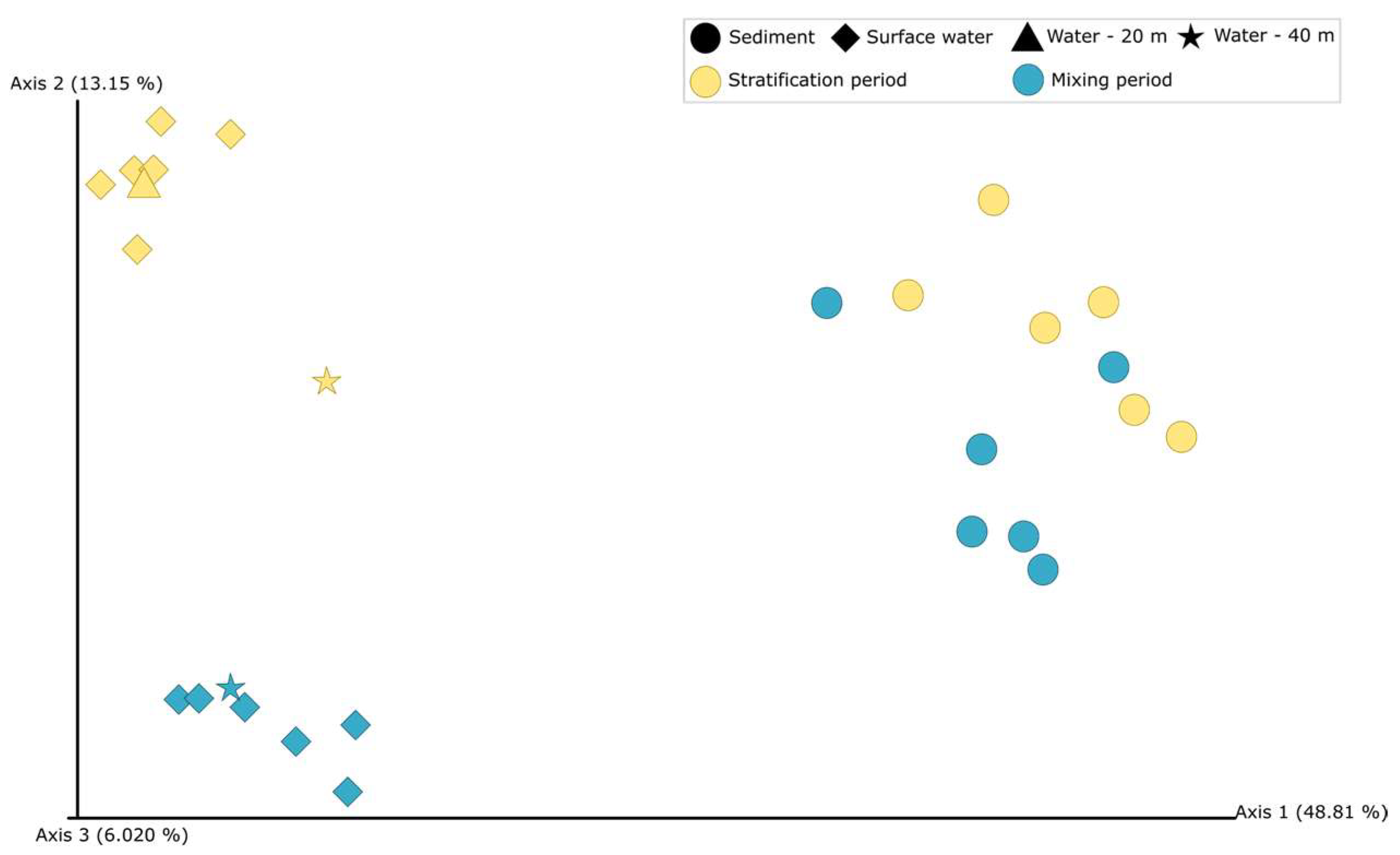
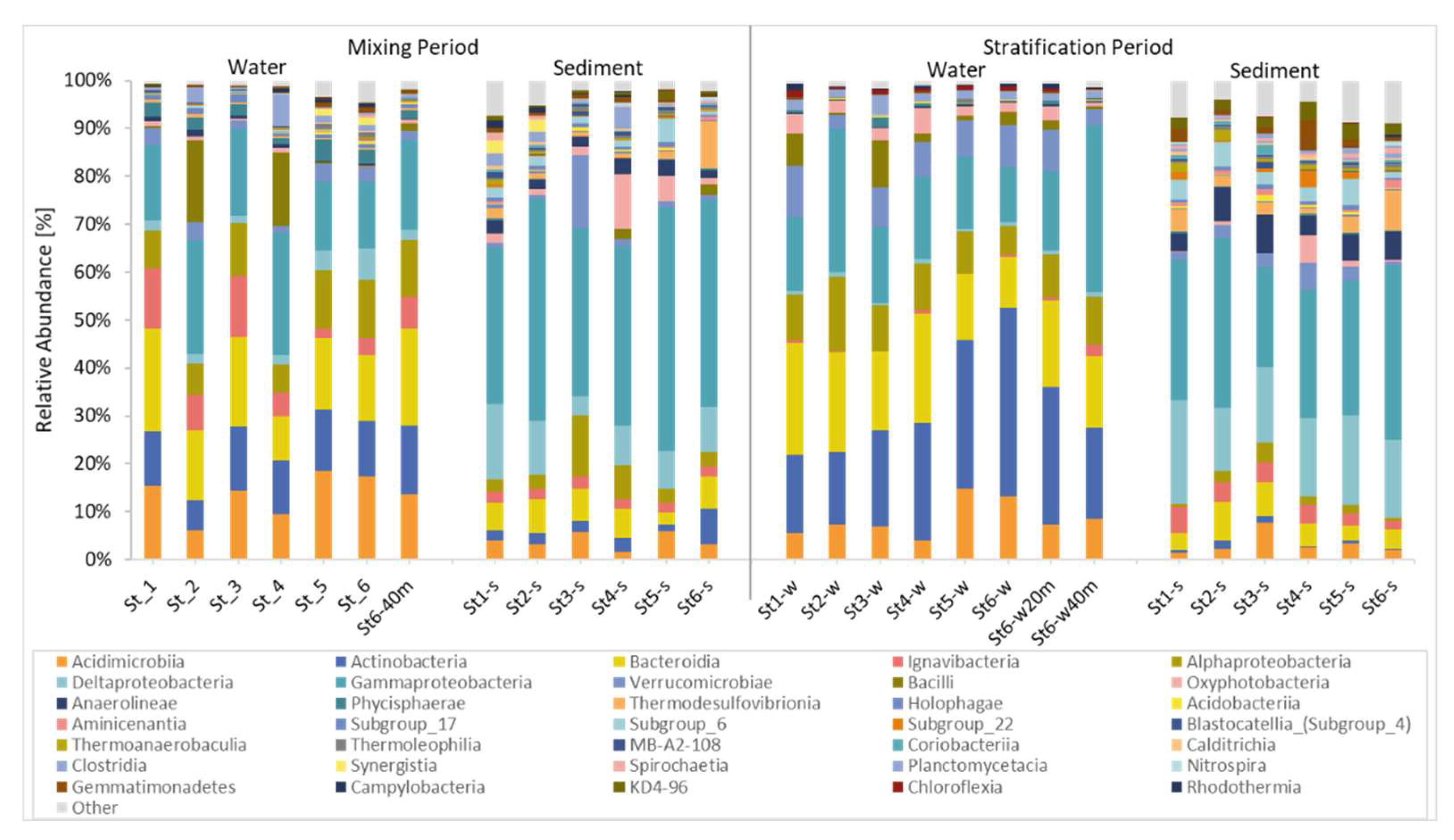
3.2. Bacterial Community Compositions
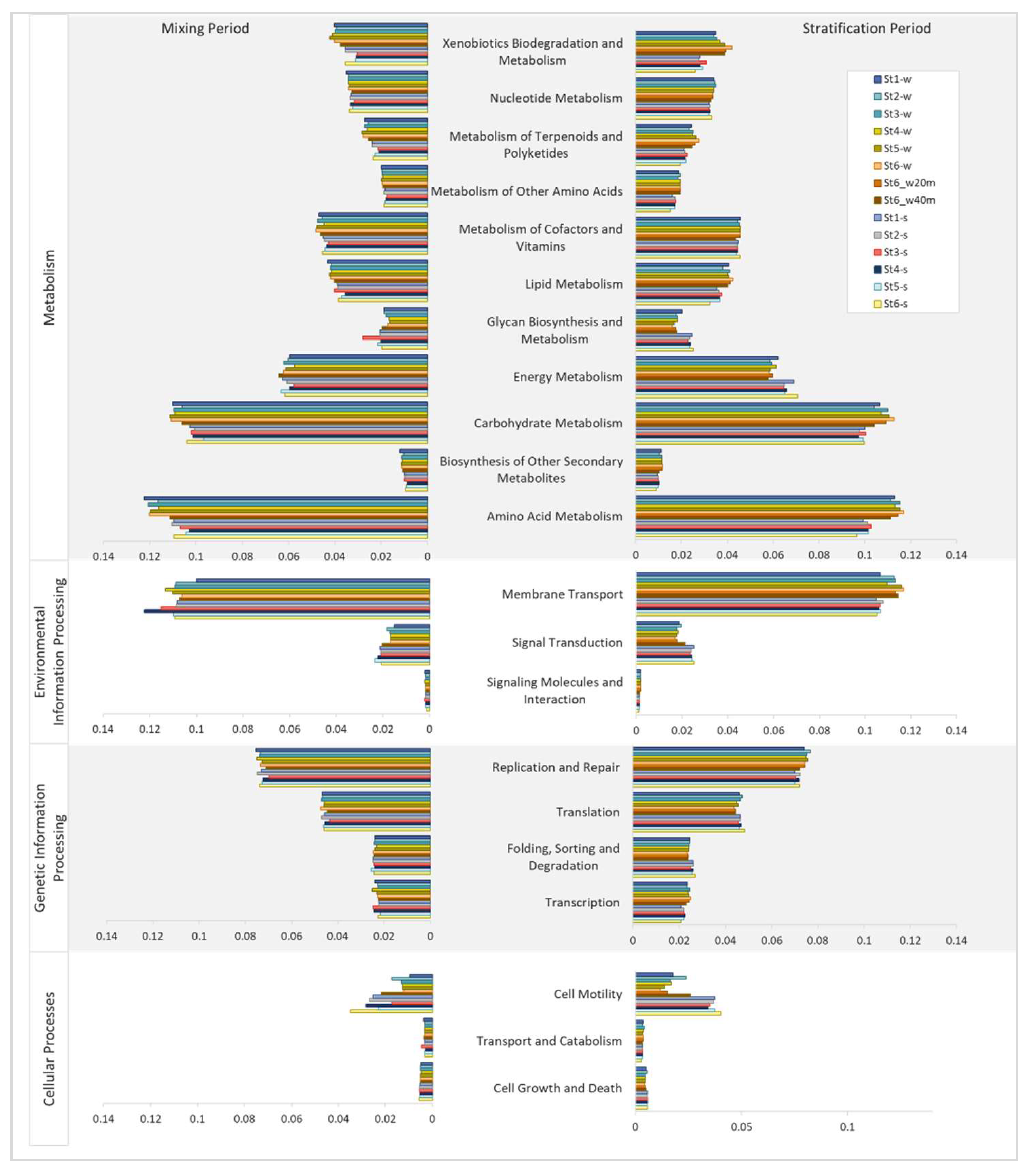
3.3. Predicted Functions of Bacterial Communities
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zhang, L.; Cheng, Y.; Gao, G.; Jiang, J. Spatial-Temporal Variation of Bacterial Communities in Sediments in Lake Chaohu, a Large, Shallow Eutrophic Lake in China. Int. J. Environ. Res. Public Health 2019, 16, 3966. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xie, G.; Tang, X.; Shao, K.; Zhu, G.; Gao, G. Bacterial Diversity, Community Composition and Metabolic Function in Lake Tianmuhu and Its Dammed River: Effects of Domestic Wastewater and Damming. Ecotoxicol. Environ. Saf. 2021, 213, 112069. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhong, Y.; Yang, S.; Zhang, W.; Xu, M.; Ma, A.; Zhuang, G.; Chen, G.; Liu, W. Diversity and Dynamics of the Microbial Community on Decomposing Wheat Straw during Mushroom Compost Production. Bioresour. Technol. 2014, 170, 183–195. [Google Scholar] [CrossRef] [PubMed]
- Gu, Z.; Liu, K.; Pedersen, M.W.; Wang, F.; Chen, Y.; Zeng, C.; Liu, Y. Community Assembly Processes Underlying the Temporal Dynamics of Glacial Stream and Lake Bacterial Communities. Sci. Total Environ. 2021, 761, 143178. [Google Scholar] [CrossRef] [PubMed]
- Ozbayram, E.G.; Koker, L.; Akçaalan, R.; Aydın, F.; Ertürk, A.; Ince, O.; Albay, M. Contrasting the Water Quality and Bacterial Community Patterns in Shallow and Deep Lakes: Manyas vs. Iznik. Environ. Manag. 2021, 67, 506–512. [Google Scholar] [CrossRef]
- Diao, M.; Sinnige, R.; Kalbitz, K.; Huisman, J.; Muyzer, G. Succession of Bacterial Communities in a Seasonally Stratified Lake with an Anoxic and Sulfidic Hypolimnion. Front. Microbiol. 2017, 8, 2511. [Google Scholar] [CrossRef] [Green Version]
- Yang, X.; Huang, T.; Zhang, H. Effects of Seasonal Thermal Stratification on the Functional Diversity and Composition of the Microbial Community in a Drinking Water Reservoir. Water 2015, 7, 5525–5546. [Google Scholar] [CrossRef]
- Shilei, Z.; Yue, S.; Tinglin, H.; Ya, C.; Xiao, Y.; Zizhen, Z.; Yang, L.; Zaixing, L.; Jiansheng, C.; Xiao, L. Reservoir Water Stratification and Mixing Affects Microbial Community Structure and Functional Community Composition in a Stratified Drinking Reservoir. J. Environ. Manag. 2020, 267, 110456. [Google Scholar] [CrossRef]
- Shen, M.; Li, Q.; Ren, M.; Lin, Y.; Wang, J.; Chen, L.; Li, T.; Zhao, J. Trophic Status Is Associated With Community Structure and Metabolic Potential of Planktonic Microbiota in Plateau Lakes. Front. Microbiol. 2019, 10, 2560. [Google Scholar] [CrossRef] [Green Version]
- Xing, R.; Gao, Q.B.; Zhang, F.Q.; Wang, J.L.; Chen, S.L. Bacterial Community in Cold and Alkaline Environments of Hoh Xil Basin in Qinghai–Tibet Plateau and Isolation of Potential Sources of Microbiota. Ann. Microbiol. 2019, 69, 567–576. [Google Scholar] [CrossRef]
- Koo, H.; Hakim, J.A.; Morrow, C.D.; Eipers, P.G.; Davila, A.; Andersen, D.T.; Bej, A.K. Comparison of Two Bioinformatics Tools Used to Characterize the Microbial Diversity and Predictive Functional Attributes of Microbial Mats from Lake Obersee, Antarctica. J. Microbiol. Methods 2017, 140, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Shen, T.; Cheng, Y.; Zhao, T.; Li, L.; Qi, P. Temporal and Spatial Variations in the Bacterial Community Composition in Lake Bosten, a Large, Brackish Lake in China. Sci. Rep. 2020, 10, 304. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Langille, M.G.I.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Vega Thurber, R.L.; Knight, R.; et al. Predictive Functional Profiling of Microbial Communities Using 16S RRNA Marker Gene Sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef] [PubMed]
- Oğuz, A.; Akçaalan, R.; Köker, L.; Gürevin, C.; Dorak, Z.; Albay, M. Driving Factors Affecting the Phytoplankton Functional Groups in a Deep Alkaline Lake. Turk. J. Bot. 2020, 44, 633–646. [Google Scholar] [CrossRef]
- Katip, A. The Usage of Artificial Neural Networks in Microbial Water Quality Modeling: A Case Study from the Lake İznİk. Appl. Ecol. Environ. Res. 2018, 16, 3897–3917. [Google Scholar] [CrossRef]
- Özuluǧ, M.; Altun, Ö.; Meriç, N. On the Fish Fauna of Lake İznik (Turkey). Turk. J. Zool. 2005, 29, 371–375. [Google Scholar]
- Katip, A. Meteorological Drought Analysis Using Artificial Neural Networks for Bursa City, Turkey. Appl. Ecol. Environ. Res. 2018, 16, 3315–3332. [Google Scholar] [CrossRef]
- Akcaalan, R.; Köker, L.; Oğuz, A.; Spoof, L.; Meriluoto, J.; Albay, M. First Report of Cylindrospermopsin Production by Two Cyanobacteria (Dolichospermum mendotae and Chrysosporum ovalisporum) in Lake Iznik, Turkey. Toxins 2014, 6, 3173–3186. [Google Scholar] [CrossRef] [Green Version]
- Franz, S.O.; Schwark, L.; Brüchmann, C.; Scharf, B.; Klingel, R.; Van Alstine, J.D.; Çagatay, N.; Ülgen, U.B. Results from a Multi-Disciplinary Sedimentary Pilot Study of Tectonic Lake Iznik (NW Turkey)—Geochemistry and Paleolimnology of the Recent Past. J. Paleolimnol. 2006, 35, 715–736. [Google Scholar] [CrossRef]
- Akcaalan, R.; Mazur-Marzec, H.; Zalewska, A.; Albay, M. Phenotypic and Toxicological Characterization of Toxic Nodularia Spumigena from a Freshwater Lake in Turkey. Harmful Algae 2009, 8, 273–278. [Google Scholar] [CrossRef]
- Albay, M.; Aykulu, G. Invertebrate Grazer—Epiphytic Algae Interactions on Submerged Macrophytes in a Mesotrophic Turkish Lake. J. Fish. Aquat. Sci. 2002, 19, 247–258. [Google Scholar]
- APHA/AWWA/WEF. Standard Methods for the Examination of Water and Waste Water, 17th ed.; APHA, AWAA, WPFC: Washington DC, USA, 1989; p. 113. [Google Scholar]
- ISO 10260; Water Quality—Measurement of Biochemical Parameters—Spectrometric Determination of the Chlorophyll-a Concentration. International Standards Organization: Geneva, Switzerland, 1992.
- Carlson, R.E. A Trophic State Index for Lakes. Limnol. Oceanogr. 1977, 22, 361–369. [Google Scholar] [CrossRef]
- Comeau, A.M.; Li, W.K.W.; Tremblay, J.É.; Carmack, E.C.; Lovejoy, C. Arctic Ocean Microbial Community Structure before and after the 2007 Record Sea Ice Minimum. PLoS ONE 2011, 6, e27492. [Google Scholar] [CrossRef] [PubMed]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, Interactive, Scalable and Extensible Microbiome Data Science Using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef] [PubMed]
- Martin, M. Cutadapt Removes Adapter Sequences from High-Throughput Sequencing Reads. EMBnet. J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Amir, A.; Daniel, M.; Navas-Molina, J.; Kopylova, E.; Morton, J.; Xu, Z.Z.; Eric, K.; Thompson, L.; Hyde, E.; Gonzalez, A.; et al. Deblur Rapidly Resolves Single-Nucleotide Community Sequence Patterns. Am. Soc. Microbiol. 2017, 2, e00191-16. [Google Scholar] [CrossRef] [Green Version]
- Comeau, A.M.; Douglas, G.M.; Langille, M.G.I. Microbiome Helper: A Custom and Streamlined Workflow for Microbiome Research. mSystems 2017, 2, e00127-16. [Google Scholar] [CrossRef] [Green Version]
- Wu, D.M.; Dai, Q.P.; Liu, X.Z.; Fan, Y.P.; Wang, J.X. Comparison of Bacterial Community Structure and Potential Functions in Hypoxic and Non-Hypoxic Zones of the Changjiang Estuary. PLoS ONE 2019, 14, e0217431. [Google Scholar] [CrossRef] [Green Version]
- Parks, D.H.; Beiko, R.G. Identifying Biologically Relevant Differences between Metagenomic Communities. Bioinformatics 2010, 26, 715–721. [Google Scholar] [CrossRef] [Green Version]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021. [Google Scholar]
- Li, W.; Morgan-Kiss, R.M. Influence of Environmental Drivers and Potential Interactions on the Distribution of Microbial Communities from Three Permanently Stratified Antarctic Lakes. Front. Microbiol. 2019, 10, 1067. [Google Scholar] [CrossRef] [Green Version]
- Shade, A.; Jones, S.E.; McMahon, K.D. The Influence of Habitat Heterogeneity on Freshwater Bacterial Community Composition and Dynamics. Environ. Microbiol. 2008, 10, 1057–1067. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.; Yang, J.; Amalfitano, S.; Yu, X.; Liu, L. Effects of Water Stratification and Mixing on Microbial Community Structure in a Subtropical Deep Reservoir. Sci. Rep. 2014, 4, srep05821. [Google Scholar] [CrossRef] [PubMed]
- Chimney, M.J.; Wenkert, L.; Pietro, K.C. Patterns of Vertical Stratification in a Subtropical Constructed Wetland in South Florida (USA). Ecol. Eng. 2006, 27, 322–330. [Google Scholar] [CrossRef]
- Jin, J.; Wells, S.A.; Liu, D.; Yang, G. Thermal Stratification and Its Relationship with Water Quality in the Typical Tributary Bay of the Three Gorges Reservoir. Water Sci. Technol. Water Supply 2019, 19, 918–925. [Google Scholar] [CrossRef] [Green Version]
- Lindström, E.S.; Kamst-Van Agterveld, M.P.; Zwart, G. Distribution of Typical Freshwater Bacterial Groups Is Associated with PH, Temperature, and Lake Water Retention Time. Appl. Environ. Microbiol. 2005, 71, 8201–8206. [Google Scholar] [CrossRef] [Green Version]
- Garcia, S.L.; Salka, I.; Grossart, H.P.; Warnecke, F. Depth-Discrete Profiles of Bacterial Communities Reveal Pronounced Spatio-Temporal Dynamics Related to Lake Stratification. Environ. Microbiol. Rep. 2013, 5, 549–555. [Google Scholar] [CrossRef]
- Llorens-Marès, T.; Catalan, J.; Casamayor, E.O. Taxonomy and Functional Interactions in Upper and Bottom Waters of an Oligotrophic High-Mountain Deep Lake (Redon, Pyrenees) Unveiled by Microbial Metagenomics. Sci. Total Environ. 2020, 707, 135929. [Google Scholar] [CrossRef]
- Paruch, L.; Paruch, A.M.; Eiken, H.G.; Skogen, M.; Sørheim, R. Seasonal Dynamics of Lotic Bacterial Communities Assessed by 16S RRNA Gene Amplicon Deep Sequencing. Sci. Rep. 2020, 10, 16399. [Google Scholar] [CrossRef]
- Ji, B.; Liang, J.; Ma, Y.; Zhu, L.; Liu, Y. Bacterial Community and Eutrophic Index Analysis of the East Lake. Environ. Pollut. 2019, 252, 682–688. [Google Scholar] [CrossRef]
- Alotaibi, M.O.; Mohammed, A.E.; Eltom, K.H. Metagenomic Analysis of Bacterial Communities of Wadi Namar Lake, Riyadh, Saudi Arabia. Saudi J. Biol. Sci. 2022, 29, 3749–3758. [Google Scholar] [CrossRef]
- Shang, Y.; Wu, X.; Wang, X.; Wei, Q.; Ma, S.; Sun, G.; Zhang, H.; Wang, L.; Dou, H.; Zhang, H. Factors Affecting Seasonal Variation of Microbial Community Structure in Hulun Lake, China. Sci. Total Environ. 2022, 805, 150294. [Google Scholar] [CrossRef] [PubMed]
- Hoke, A.; Woodhouse, J.; Zoccarato, L.; McCarthy, V.; de Eyto, E.; Calderó-Pascual, M.; Geffroy, E.; Dillane, M.; Grossart, H.P.; Jennings, E. Impacts of Extreme Weather Events on Bacterial Community Composition of a Temperate Humic Lake. Water 2020, 12, 2757. [Google Scholar] [CrossRef]
- Rosenberg, E. The Prokaryotes: Alphaproteobacteria and Betaproteobacteria; Springer: Berlin/Heidelberg, Germany, 2013; ISBN 9783642301971. [Google Scholar]
- Pineda-Mendoza, R.M.; Briones-Roblero, C.I.; Gonzalez-Escobedo, R.; Rivera-Orduña, F.N.; Martínez-Jerónimo, F.; Zúñiga, G. Seasonal Changes in the Bacterial Community Structure of Three Eutrophicated Urban Lakes in Mexico City, with Emphasis on Microcystis spp. Toxicon 2020, 179, 8–20. [Google Scholar] [CrossRef] [PubMed]
- Yue, Y.; Cai, L.; Tang, Y.; Zhang, Y.; Yang, M.; Wang, F. Vertical Distribution of Bacterial Community in Water Columns of Reservoirs With Different Trophic Conditions During Thermal Stratification. Front. Environ. Sci. 2021, 9, 632089. [Google Scholar] [CrossRef]
- Garlapati, D.; Kumar, B.C.; Muthukumar, C.; Madeswaran, P.; Ramu, K.; Murthy, M.V.R. Assessing the in Situ Bacterial Diversity and Composition at Anthropogenically Active Sites Using the Environmental DNA (EDNA). Mar. Pollut. Bull. 2021, 170, 112593. [Google Scholar] [CrossRef]
- Wang, Y.; Guo, M.; Li, X.; Liu, G.; Hua, Y.; Zhao, J.; Huguet, A.; Li, S. Shifts in Microbial Communities in Shallow Lakes Depending on Trophic States: Feasibility as an Evaluation Index for Eutrophication. Ecol. Indic. 2022, 136, 108691. [Google Scholar] [CrossRef]
- Williams, K.P.; Gillespie, J.J.; Sobral, B.W.S.; Nordberg, E.K.; Snyder, E.E.; Shallom, J.M.; Dickerman, A.W. Phylogeny of Gammaproteobacteria. J. Bacteriol. 2010, 192, 2305–2314. [Google Scholar] [CrossRef] [Green Version]
- Chen, Y.; Li, S.; Xu, X.; Ma, M.; Mi, T.; Zhen, Y.; Yu, Z. Characterization of Microbial Communities in Sediments of the South Yellow Sea. J. Oceanol. Limnol. 2021, 39, 846–864. [Google Scholar] [CrossRef]
- Huang, W.; Chen, X.; Jiang, X.; Zheng, B. Characterization of Sediment Bacterial Communities in Plain Lakes with Different Trophic Statuses. Microbiologyopen 2017, 6, e00503. [Google Scholar] [CrossRef]
- Chen, J.; Wu, J.; Liu, M.; Li, L.; Zhang, W.; Wang, D.; Ma, T. Bacterial Community Structure in the Surface Sediments of Different Habitats of Baiyangdian Lake, Northern China: Effects of Nutrient Conditions. J. Soils Sediments 2021, 21, 1866–1874. [Google Scholar] [CrossRef]
- Custodio, M.; Espinoza, C.; Peñaloza, R.; Peralta-Ortiz, T.; Sánchez-Suárez, H.; Ordinola-Zapata, A.; Vieyra-Peña, E. Microbial Diversity in Intensively Farmed Lake Sediment Contaminated by Heavy Metals and Identification of Microbial Taxa Bioindicators of Environmental Quality. Sci. Rep. 2022, 12, 80. [Google Scholar] [CrossRef] [PubMed]
- Aydın, F.; Albay, M. Accumulation of Organochlorine Pesticide (OCP) Residues in Surface Water and Sediment from the İznik Lake in Turkey. Environ. Monit. Assess. 2022, 194, 872. [Google Scholar] [CrossRef] [PubMed]
- Zeng, J.; Jiao, C.; Zhao, D.; Xu, H.; Huang, R.; Cao, X.; Yu, Z.; Wu, Q.L. Patterns and Assembly Processes of Planktonic and Sedimentary Bacterial Community Differ along a Trophic Gradient in Freshwater Lakes. Ecol. Indic. 2019, 106, 105491. [Google Scholar] [CrossRef]
- Hu, A.; Yang, X.; Chen, N.; Hou, L.; Ma, Y.; Yu, C.P. Response of Bacterial Communities to Environmental Changes in a Mesoscale Subtropical Watershed, Southeast China. Sci. Total Environ. 2014, 472, 746–756. [Google Scholar] [CrossRef] [PubMed]
- Andrei, A.Ş.; Robeson, M.S.; Baricz, A.; Coman, C.; Muntean, V.; Ionescu, A.; Etiope, G.; Alexe, M.; Sicora, C.I.; Podar, M.; et al. Contrasting Taxonomic Stratification of Microbial Communities in Two Hypersaline Meromictic Lakes. ISME J. 2015, 9, 2642–2656. [Google Scholar] [CrossRef] [PubMed]

| Parameter | Mixing Period | Stratification Period | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| St_1 | St_2 | St_3 | St_4 | St_5 | St_6 | St_6_40 m | St_1 | St_2 | St_3 | St_4 | St_5 | St_6 | St_6_40 m | ||
| Temperature | °C | 9.38 | 10.17 | 9.54 | 10.23 | 10.12 | 9.99 | 9.19 | 25.28 | 29.11 | 27.79 | 29.22 | 25.06 | 25.76 | 9.37 |
| EC | µS/cm | 748 | 996 | 928 | 937 | 760 | 748 | 734 | 875 | 756 | 746 | 761 | 708 | 1035 | 398 |
| pH | 8.29 | 9.26 | 9.05 | 9.64 | 8.56 | 8.58 | 8.39 | 8.74 | 9.39 | 9.57 | 9.67 | 9.19 | 9.1 | 7.7 | |
| DO | mg/L | 10.3 | 10.12 | 10.77 | 11.65 | 10.53 | 11.08 | 9.72 | 8.92 | 12.76 | 11.48 | 9.49 | 9.01 | 9.26 | 1.04 |
| SRP | µg/L | 22.97 | 13.35 | 16.90 | 10.48 | 17.91 | 18.25 | 19.77 | 2.12 | 2.27 | 1.53 | 1.68 | 3.15 | 1.09 | 71.2 |
| TP | µg/L | 34.21 | 38.63 | 31.75 | 30.89 | 32.49 | 29.91 | 30.77 | 28.43 | 50.78 | 32.31 | 34.72 | 27.49 | 25.75 | 119.7 |
| DIN | mg/L | 0.35 | 0.35 | 0.25 | 0.39 | 0.34 | 0.31 | 0.35 | 0.17 | 0.15 | 0.14 | 0.17 | 0.19 | 0.14 | 0.25 |
| SiO2 | mg/L | 1.59 | 1.48 | 0.93 | 2.92 | 1.51 | 1.58 | 1.79 | 1.33 | 0.89 | 1.22 | 1.16 | 1.28 | 1.32 | 3.05 |
| Chl-a | µg/L | 9.32 | 8.44 | 8.88 | 8.44 | 2.66 | 5.33 | 5.33 | 14.21 | 14.80 | 14.21 | 13.32 | 14.21 | 11.84 | 2.22 |
| SO4 | mg/L | 9.92 | 10.26 | 9.34 | 13.95 | 9.13 | 9.52 | 9.39 | 11.30 | 10.97 | 10.86 | 10.84 | 10.46 | 10.62 | 9.42 |
| Alkalinity | mg/L CaCO3 | 455 | 453 | 451 | 454 | 478 | 475 | 443 | 366 | 357 | 358 | 354 | 355 | 352 | 325 |
| Sampling Period | Sample Type | Sample | Number of OTUs | Shannon | Pielou’s Evenness | Faith’s PD |
|---|---|---|---|---|---|---|
| Mixing Period | Water | St_1 | 466 | 6.74 | 0.76 | 50.11 |
| St_2 | 548 | 7.52 | 0.83 | 53.81 | ||
| St_3 | 429 | 6.47 | 0.74 | 41.92 | ||
| St_4 | 765 | 7.82 | 0.82 | 67.34 | ||
| St_5 | 653 | 7.56 | 0.81 | 61.49 | ||
| St_6 | 663 | 7.43 | 0.79 | 63.79 | ||
| St_6_40 m | 766 | 7.80 | 0.81 | 66.72 | ||
| Sediment | St_1 | 801 | 8.64 | 0.90 | 70.69 | |
| St_2 | 815 | 8.48 | 0.88 | 71.84 | ||
| St_3 | 885 | 8.44 | 0.86 | 80.68 | ||
| St_4 | 723 | 8.23 | 0.87 | 70.79 | ||
| St_5 | 791 | 7.90 | 0.82 | 71.25 | ||
| St_6 | 653 | 7.62 | 0.81 | 57.17 | ||
| Stratification Period | Water | St_1 | 212 | 6.13 | 0.79 | 29.14 |
| St_2 | 390 | 7.08 | 0.82 | 42.55 | ||
| St_3 | 1068 | 8.47 | 0.84 | 91.46 | ||
| St_4 | 334 | 6.79 | 0.81 | 38.52 | ||
| St_5 | 324 | 6.47 | 0.78 | 37.93 | ||
| St_6 | 160 | 5.54 | 0.76 | 25.07 | ||
| St_6_20 m | 345 | 6.66 | 0.79 | 41.21 | ||
| St_6_40 m | 434 | 7.43 | 0.85 | 45.56 | ||
| Sediment | St_1 | 699 | 8.13 | 0.86 | 66.65 | |
| St_2 | 544 | 8.05 | 0.89 | 49.95 | ||
| St_3 | 859 | 8.83 | 0.91 | 81.32 | ||
| St_4 | 607 | 7.74 | 0.84 | 61.15 | ||
| St_5 | 892 | 8.63 | 0.88 | 86.35 | ||
| St_6 | 666 | 7.88 | 0.84 | 63.36 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ozbayram, E.G.; Köker, L.; Çam, A.O.; Akçaalan, R.; Albay, M. Temporal and Spatial Variations of the Bacterial Diversity in a Deep Alkaline Lake. Water 2022, 14, 4097. https://doi.org/10.3390/w14244097
Ozbayram EG, Köker L, Çam AO, Akçaalan R, Albay M. Temporal and Spatial Variations of the Bacterial Diversity in a Deep Alkaline Lake. Water. 2022; 14(24):4097. https://doi.org/10.3390/w14244097
Chicago/Turabian StyleOzbayram, E. Gozde, Latife Köker, Ayça Oğuz Çam, Reyhan Akçaalan, and Meriç Albay. 2022. "Temporal and Spatial Variations of the Bacterial Diversity in a Deep Alkaline Lake" Water 14, no. 24: 4097. https://doi.org/10.3390/w14244097






