A Virological Perspective on the Use of Bacteriophages as Hydrological Tracers
Abstract
:1. Introduction
2. Methodology
2.1. Search Strategy
2.2. Data Extraction and Analysis
3. Bacteriophages as Hydrological Tracers: An Overview
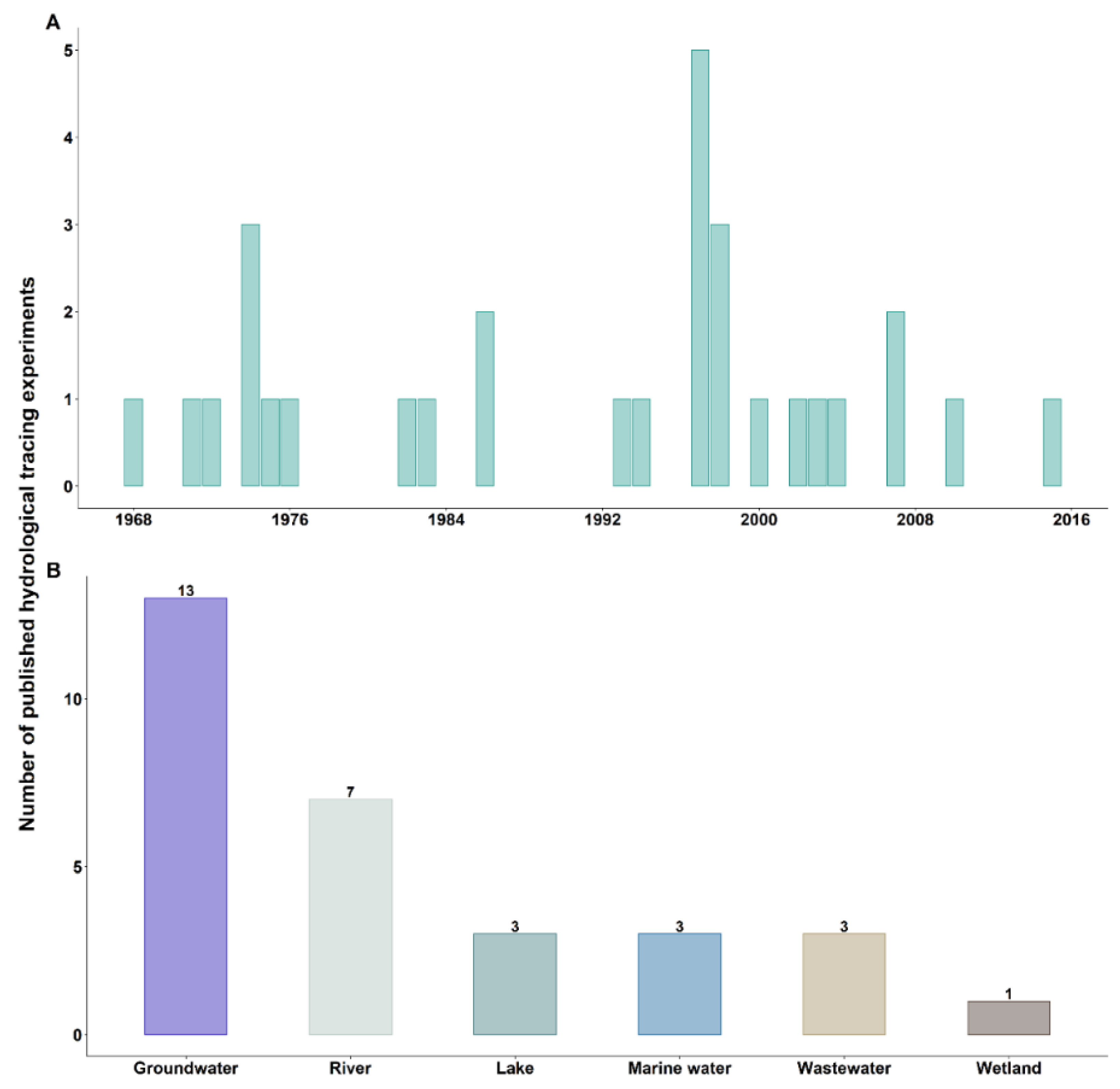
3.1. First Hydrological Tracing Experiments Using Bacteriophages
3.2. Objectives of Hydrological Tracing Experiments Using Bacteriophages
4. Bacteriophages Commonly Used as Hydrological Tracers
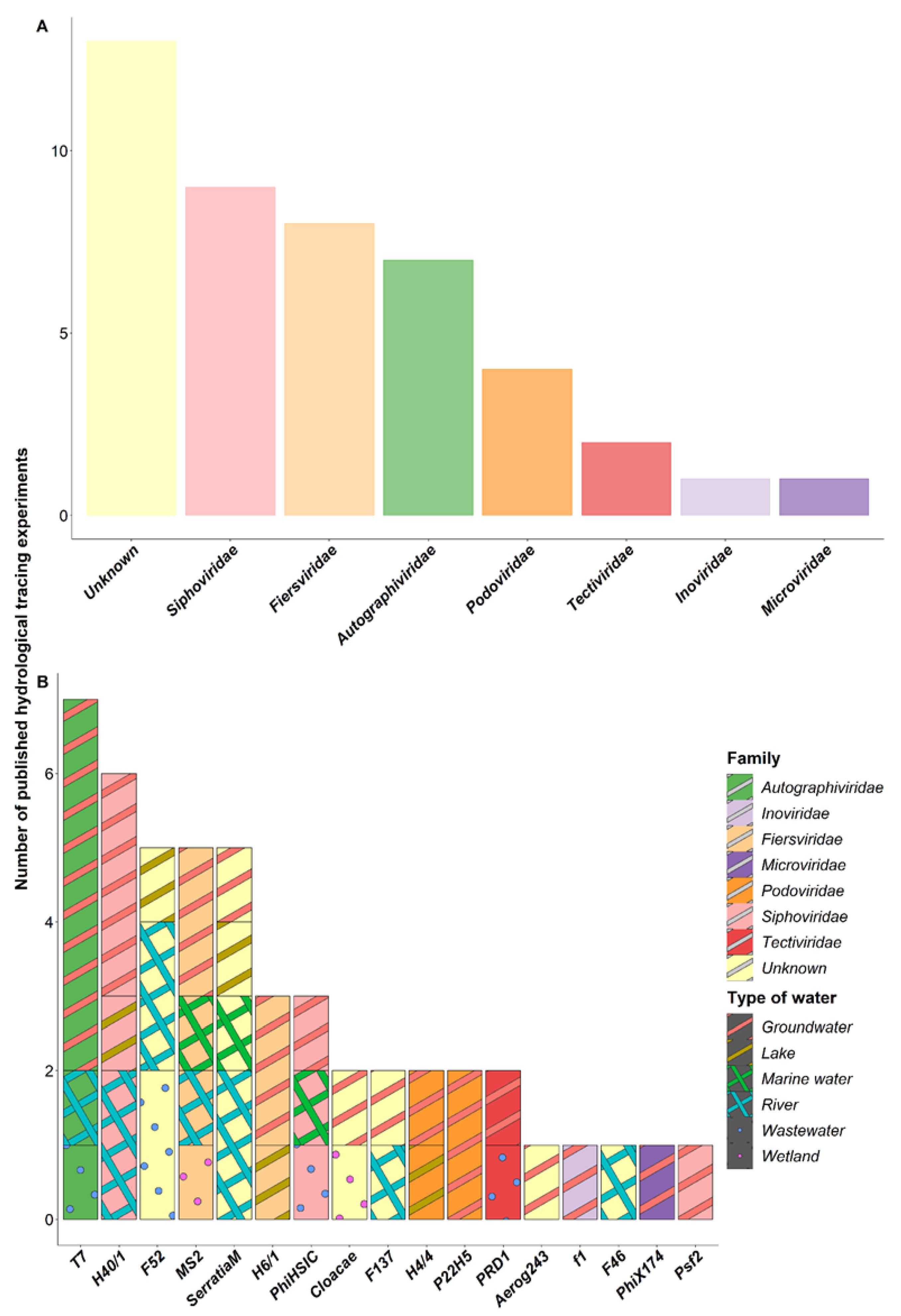
4.1. Description of the Bacteriophages Used in the Tracing Experiments
4.2. Effectiveness of Bacteriophages as Hydrological Tracers
4.3. Combination of Hydrological Tracers in Multi-Injection Experiments
5. Towards a More Adequate Use of Bacteriophages in Hydrological Tracing Experiments
5.1. Methodological Optimisation for Bacteriophage Detection
5.1.1. Limitations of the Current Detection Methods
5.1.2. The Interest in Using Several Complementary Methods for Bacteriophage Detection
5.2. “Eco-Friendly” Property of Bacteriophages
5.2.1. The Controversy of This Safe Biological Entity
5.2.2. The Interest in the Natural Populations of Bacteriophages
5.3. The Properties of the Catchment of Interest
5.3.1. Flaws in Considering the Surrounding Environments
5.3.2. Awareness of the Interactions between Bacteriophages and the Surrounding Environment
6. Conclusions and Perspectives
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Leibundgut, C.; Seibert, J. Tracer Hydrology. Treatise Water Sci. 2010, 2, 215–236. [Google Scholar] [CrossRef]
- Mcguire, K.J.; Mcdonnell, J.J. Tracer Advances in Catchment Hydrology. Hydrol. Process. 2015, 29, 5135–5138. [Google Scholar] [CrossRef]
- McGuire, K.J.; McDonnell, J.J. A Review and Evaluation of Catchment Transit Time Modeling. J. Hydrol. 2006, 330, 543–563. [Google Scholar] [CrossRef]
- Flury, M.; Wai, N.N. Dyes as Tracers for Vadose Zone Hydrology. Rev. Geophys. 2003, 41, 1–37. [Google Scholar] [CrossRef]
- Harvey, R.W. Microorganisms as Tracers in Groundwater Injection and Recovery Experiments: A Review. FEMS Microbiol. Rev. 1997, 20, 461–472. [Google Scholar] [CrossRef]
- Behrens, H.; Beims, U.; Dieter, H.; Dietze, G.; Eikmann, T.; Grummt, T.; Hanisch, H.; Henseling, H.; Käss, W.; Kerndorff, H.; et al. Toxicological and Ecotoxicological Assessment of Water Tracers. Hydrogeol. J. 2001, 9, 321–325. [Google Scholar] [CrossRef]
- Schudel, B.; Biaggi, D.; Dervey, T.; Kozel, R.; Müller, I.; Ross, J.H.; Schindler, U. Utilisation Des Traceurs Artificiels En Hydrogéologie—Guide Pratique; Série Géologique; Office fédéral des eaux et de la géologie: Washington, DC, USA, 2002; p. 77. [Google Scholar]
- Berryman, C.J. Tracer Tests for Investigating Flow and Transport in the Hyporheic Zone; Environment Agency: Bristol, UK, 2005; ISBN 9781844326983.
- Coles, A.E.; Wetzel, C.E.; Martínez-Carreras, N.; Ector, L.; Mcdonnell, J.J.; Frentress, J.; Klaus, J.; Hoffmann, L.; Pfister, L. Diatoms as a Tracer of Hydrological Connectivity: Are They Supply Limited? Ecohydrology 2016, 9, 631–645. [Google Scholar] [CrossRef]
- Kirchner, J.W.; Tetzlaff, D.; Soulsby, C. Comparing Chloride and Water Isotopes as Hydrological Tracers in Two Scottish Catchments. Hydrol. Process. 2010, 24, 1631–1645. [Google Scholar] [CrossRef]
- Klaus, J.; McDonnell, J.J. Hydrograph Separation Using Stable Isotopes: Review and Evaluation. J. Hydrol. 2013, 505, 47–64. [Google Scholar] [CrossRef]
- Burns, D.A. Stormflow-Hydrograph Separation Based on Isotopes: The Thrill Is Gone-What’s Next? Hydrol. Process. 2002, 16, 1515–1517. [Google Scholar] [CrossRef]
- Cartwright, I.; Cendón, D.; Currell, M.; Meredith, K. A Review of Radioactive Isotopes and Other Residence Time Tracers in Understanding Groundwater Recharge: Possibilities, Challenges, and Limitations. J. Hydrol. 2017, 555, 797–811. [Google Scholar] [CrossRef]
- Skilton, H.; Wheeler, D. Bacteriophage Tracer Experiments in Groundwater. J. Appl. Bacteriol. 1988, 65, 387–395. [Google Scholar] [CrossRef] [PubMed]
- Martin, C. The Application of Bacteriophage Tracer Techniques in South West Water. Water Environ. J. 1988, 2, 638–642. [Google Scholar] [CrossRef]
- Sklash, M.G.; Farvolden, R.N.; Fritz, P. A Conceptual Model of Watershed Response to Rainfall, Developed through the Use of Oxygen-18 as a Natural Tracer. Can. J. Earth Sci. 1976, 13, 271–283. [Google Scholar] [CrossRef]
- McGlynn, B.; McDonnell, J.; Stewart, M.; Seibert, J. On the Relationships between Catchment Scale and Streamwater Mean Residence Time. Hydrol. Process. 2003, 17, 175–181. [Google Scholar] [CrossRef]
- Burt, T.P.; McDonnell, J.J. Whither Field Hydrology? The Need for Discovery Science and Outrageous Hydrological Hypotheses. Water Resour. Res. 2015, 51, 5919–5928. [Google Scholar] [CrossRef] [Green Version]
- Vitvar, T.; Aggarwal, P.K.; McDonnell, J.J. A Review of Isotope Applications in Catchment Hydrology. In Isotopes in the Water Cycle: Past, Present and Future of a Developing Science; Springer: Dordrecht, The Netherlands, 2005; pp. 151–169. ISBN 140203010X. [Google Scholar]
- Gibson, J.J.; Aggarwal, P.; Hogan, J.; Kendall, C.; Martinelli, L.A.; Stichler, W.; Rank, D.; Goni, I.; Choudhry, M.; Gat, J.; et al. Isotope Studies in Large River Basins: A New Global Research Focus. Eos Trans. Am. Geophys. Union 2002, 83, 613–620. [Google Scholar] [CrossRef]
- Selker, J.S.; Thévenaz, L.; Huwald, H.; Mallet, A.; Luxemburg, W.; Van De Giesen, N.; Stejskal, M.; Zeman, J.; Westhoff, M.; Parlange, M.B. Distributed Fiber-Optic Temperature Sensing for Hydrologic Systems. Water Resour. Res. 2006, 42, 1–8. [Google Scholar] [CrossRef] [Green Version]
- Westhoff, M.C.; Savenjie, H.H.G.; Luxemburg, W.M.J.; Stelling, G.S.; Van De Giesen, N.C.; Selker, J.S.; Pfister, L.; Uhlenbrook, S. A Distributed Stream Temperature Model Using High Resolution Temperature Observations. Hydrol. Earth Syst. Sci. 2007, 11, 1469–1480. [Google Scholar] [CrossRef] [Green Version]
- Pfister, L.; McDonnell, J.J.; Hissler, C.; Hoffmann, L. Ground-Based Thermal Imagery as a Simple, Practical Tool for Mapping Saturated Area Connectivity and Dynamics. Hydrol. Process. 2010, 24, 3123–3132. [Google Scholar] [CrossRef]
- Glaser, B.; Antonelli, M.; Chini, M.; Pfister, L.; Klaus, J. Technical Note: Mapping Surface-Saturation Dynamics with Thermal Infrared Imagery. Hydrol. Earth Syst. Sci. 2018, 22, 5987–6003. [Google Scholar] [CrossRef] [Green Version]
- Pfister, L.; McDonnell, J.J.; Wrede, S.; Hlúbiková, D.; Matgen, P.; Fenicia, F.; Ector, L.; Hoffmann, L. The Rivers Are Alive: On the Potential for Diatoms as a Tracer of Water Source and Hydrological Connectivity. Hydrol. Process. 2009, 23, 2841–2845. [Google Scholar] [CrossRef]
- Klaus, J.; Wetzel, C.E.; Martínez-Carreras, N.; Ector, L.; Pfister, L. A Tracer to Bridge the Scales: On the Value of Diatoms for Tracing Fast Flow Path Connectivity from Headwaters to Meso-Scale Catchments. Hydrol. Process. 2015, 29, 5275–5289. [Google Scholar] [CrossRef]
- Foppen, J.W.; Orup, C.; Adell, R.; Poulalion, V.; Uhlenbrook, S. Using Multiple Artificial DNA Tracers in Hydrology. Hydrol. Process. 2011, 25, 3101–3106. [Google Scholar] [CrossRef]
- Tang, Y.; Foppen, J.W.; Bogaard, T.A. Transport of Silica Encapsulated DNA Microparticles in Controlled Instantaneous Injection Open Channel Experiments. J. Contam. Hydrol. 2021, 242, 103880. [Google Scholar] [CrossRef]
- Harvey, R.W.; Ryan, J.N. Use of PRD1 Bacteriophage in Groundwater Viral Transport, Inactivation, and Attachment Studies. FEMS Microbiol. Ecol. 2004, 49, 3–16. [Google Scholar] [CrossRef]
- Jończyk, E.; Kłak, M.; Miedzybrodzki, R.; Górski, A. The Influence of External Factors on Bacteriophages-Review. Folia Microbiol. 2011, 56, 191–200. [Google Scholar] [CrossRef] [Green Version]
- Wommack, K.E.; Hill, R.T.; Kessel, M.; Russek-Cohen, E.; Colwell, R.R. Distribution of Viruses in the Chesapeake Bay. Appl. Environ. Microbiol. 1992, 58, 2965–2970. [Google Scholar] [CrossRef] [Green Version]
- Sinton, L.W.; Ching, S.B. An Evaluation of Two Bacteriophages as Sewage Tracers. Water Air Soil Pollut. 1987, 35, 347–356. [Google Scholar] [CrossRef]
- Harden, H.S.; Chanton, J.P.; Rose, J.B.; John, D.E.; Hooks, M.E. Comparison of Sulfur Hexafluoride, Fluorescein and Rhodamine Dyes and the Bacteriophage PRD-1 in Tracing Subsurface Flow. J. Hydrol. 2003, 277, 100–115. [Google Scholar] [CrossRef]
- Hodgson, C.J.; Perkins, J.; Labadz, J.C. Evaluation of Biotracers to Monitor Effluent Retention Time in Constructed Wetlands. Lett. Appl. Microbiol. 2003, 36, 362–371. [Google Scholar] [CrossRef]
- Drury, D.F.; Wheeler, D.C. Applications of a Serratia Marcescens Bacteriophage as a New Microbial Tracer of Aqueous Environments. J. Appl. Bacteriol. 1982, 53, 137–142. [Google Scholar] [CrossRef]
- Rossi, P.; de Carvalho-Dill, A.; Müller, I.; Aragno, M. Comparative Tracing Experiments in a Porous Aquifer Using Bacteriophages and Fluorescent Dye on a Test Field Located at Wilerwald (Switzerland) and Simultaneously Surveyed in Detail on a Local Scale by Radio-Magneto-Tellury (12–240 KHz). Environ. Geol. 1994, 23, 192–200. [Google Scholar] [CrossRef]
- Rstudio Team. RStudio: Integrated Development for R; RStudio, PBC.: Boston, MA, USA, 2020. [Google Scholar]
- Kinnunen, K. Tracing Water Movement by Means of Eschirichia Coli Bacteriophages; The water research institute: Helsinki, Finland, 1978; ISBN 951463344X. [Google Scholar]
- Wimpenny, J.W.T.; Cotton, N.; Statham, M. Microbes as Tracers of Water Movement. Water Res. 1972, 6, 731–739. [Google Scholar] [CrossRef]
- Schaub, S.A.; Sorber, C.A. Virus and Bacteria Removal from Wastewater by Rapid Infiltration through Soil. Appl. Environ. Microbiol. 1977, 33, 609–619. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martin, R.; Thomas, A. An Example of the Use of Bacteriophage as a Groundwater Tracer. J. Hydrol. 1974, 23, 73–78. [Google Scholar] [CrossRef]
- Drew, D.; Doerfliger, N.; Formentin, K. The Use of Bacteriophages for Multi-Tracing in a Lowland Karst Aquifer in Western Ireland. Tracer Hydrol. 1997, 97, 33–37. [Google Scholar]
- Bales, R.C.; Li, S.; Yeh, T.J.; Lenczewski, M.E.; Gerba, P. Bacteriophage and Microsphere Transport in Saturated Porous Media: Forced-Gradient Experiment at Borden, Ontario. Water Resour. Res. 1997, 33, 639–648. [Google Scholar] [CrossRef] [Green Version]
- Paul, J.H.; Rose, J.B.; Jiang, S.C.; Zhou, X.; Cochran, P.; Kellogg, C.; Kang, J.B.; Griffin, D.; Farrah, S.; Lukasik, J. Evidence for Groundwater and Surface Marine Water Contamination by Waste Disposal Wells in the Florida Keys. Water Res. 1997, 31, 1448–1454. [Google Scholar] [CrossRef]
- Rossi, P.; Doerfliger, N.; Kennedy, K.; Müller, I.; Aragno, M. Bacteriophages as Surface and Ground Water Tracers. Hydrol. Earth Syst. Sci. 1998, 2, 101–110. [Google Scholar] [CrossRef]
- Antonelli, M.; Balasubramanian, M.N.; Ogorzaly, L.; Pfister, L. QPCR (Quantitative Polymerase Chain Reaction) for the Quantification of Bacteriophages in Stream Water Samples to Investigate Hydrological Processes: A Proof-of-Concept Study in the Huewelerbach Experimental Catchment (Luxembourg). In Proceedings of the EGU General assembly, Vienna, Austria, 17–22 April 2016; Volume 18, p. 1. [Google Scholar]
- Maurice, L.; Atkinson, T.C.; Williams, A.T.; Barker, J.A.; Farrant, A.R. Catchment Scale Tracer Testing from Karstic Features in a Porous Limestone. J. Hydrol. 2010, 389, 31–41. [Google Scholar] [CrossRef]
- McKay, L.D.; Cherry, J.A.; Bales, R.C.; Yahya, M.T.; Gerba, C.P. A Field Example of Bacteriophage as Tracers of Fracture Flow. Environ. Sci. Technol. 1993, 27, 1075–1079. [Google Scholar] [CrossRef]
- Bricelj, M. Microbial Tracers in Groundwater Research. RMZ—Mater. Geoenviron. 2003, 1, 67–70. [Google Scholar]
- Curk, B.C.; Stichler, W. Deuterium Tracer Experiment in the Unsaturated Zone of Fractured Karst Aquifers. In Proceedings of the International Symposium On Advances In Isotope Hydrology, Vienna, Austria, 21–25 May 2007; Volume 1, pp. 597–604. [Google Scholar]
- Paul, J.H.; McLaughlin, M.R.; Griffin, D.W.; Lipp, E.K.; Stokes, R.; Rose, J.B. Rapid Movement of Wastewater from On-Site Disposal Systems into Surface Waters in the Lower Florida Keys. Estuaries Coasts 2000, 23, 662–668. [Google Scholar] [CrossRef]
- Horan, N.J.; Parr, J.; Naylor, P.J. Evaluation of Tracers for the Determination of the Mixing Characteristics of Activated Sludge Reactors. Environ. Technol. 1991, 12, 603–608. [Google Scholar] [CrossRef]
- Langlet, J.; Gaboriaud, F.; Gantzer, C. Effects of PH on Plaque Forming Unit Counts and Aggregation of MS2 Bacteriophage. J. Appl. Microbiol. 2007, 103, 1632–1638. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.Y.; Ong, S.L.; Hu, J.Y.; Tan, X.L.; Ng, W.J. Effects of PH and Temperature on the Survival of Coliphages MS2 and Qβ. J. Ind. Microbiol. Biotechnol. 2003, 30, 549–552. [Google Scholar] [CrossRef]
- Elhadidy, A.M.; Peldszus, S.; van Dyke, M.I. An Evaluation of Virus Removal Mechanisms by Ultrafiltration Membranes Using MS2 and ΦX174 Bacteriophage. Sep. Purif. Technol. 2013, 120, 215–223. [Google Scholar] [CrossRef]
- Ogorzaly, L.; Tissier, A.; Bertrand, I.; Maul, A.; Gantzer, C. Relationship between F-Specific RNA Phage Genogroups, Faecal Pollution Indicators and Human Adenoviruses in River Water. Water Res. 2009, 43, 1257–1264. [Google Scholar] [CrossRef]
- Ogorzaly, L.; Bertrand, I.; Paris, M.; Maul, A.; Gantzer, C. Occurrence, Survival, and Persistence of Human Adenoviruses and F-Specific RNA Phages in Raw Groundwater. Appl. Environ. Microbiol. 2010, 76, 8019–8025. [Google Scholar] [CrossRef] [Green Version]
- Ogorzaly, L.; Gantzer, C. Development of Real-Time RT-PCR Methods for Specific Detection of F-Specific RNA Bacteriophage Genogroups: Application to Urban Raw Wastewater. J. Virol. Methods 2006, 138, 131–139. [Google Scholar] [CrossRef]
- Jain, R.; Srivastava, R. Metabolic Investigation of Host/Pathogen Interaction Using MS2-Infected Escherichia Coli. BMC Syst. Biol. 2009, 3, 121. [Google Scholar] [CrossRef] [PubMed]
- Mayotte, J.M.; Hölting, L.; Bishop, K. Reduced Removal of Bacteriophage MS2 in during Basin Infiltration Managed Aquifer Recharge as Basin Sand Is Exposed to Infiltration Water. Hydrol. Process. 2017, 31, 1690–1701. [Google Scholar] [CrossRef]
- Bockoven, R.; Gutierrez, J.; Newkirk, H.; Liu, M.; Cahill, J.; Ramsey, J. Complete Genome Sequence of Serratia Marcescens Podophage Parlo. Microbiol. Resour. Announc. 2019, 8, e00569-19. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Denyes, J.M.; Krell, P.J.; Manderville, R.A.; Ackermann, H.W.; She, Y.M.; Kropinski, A.M. The Genome and Proteome of Serratia Bacteriophage η Which Forms Unstable Lysogens. Virol. J. 2014, 11, 6. [Google Scholar] [CrossRef] [Green Version]
- Bujak, K.; Decewicz, P.; Kaminski, J.; Radlinska, M. Identification, Characterization, and Genomic Analysis of Novel Serratia Temperate Phages from a Gold Mine. Int. J. Mol. Sci. 2020, 21, 6709. [Google Scholar] [CrossRef]
- Tian, C.; Zhao, J.; Zhang, Z.; Chen, X.; Wei, X.; Li, H.; Lin, W.; Ke, Y.; Hu, L.; Jiang, A.; et al. Identification and Molecular Characterization of Serratia Marcescens Phages VB_SmaA_2050H1 and VB_SmaM_2050HW. Arch. Virol. 2019, 164, 1085–1094. [Google Scholar] [CrossRef]
- Chan, L.Y.; Kosuri, S.; Endy, D. Refactoring Bacteriophage T7. Mol. Syst. Biol. 2005, 1, 1–10. [Google Scholar] [CrossRef] [Green Version]
- Huss, P.; Meger, A.; Leander, M.; Nishikawa, K.; Raman, S. Mapping the Functional Landscape of the Receptor Binding Domain of T7 Bacteriophage by Deep Mutational Scanning. Elife 2021, 10, e63775. [Google Scholar] [CrossRef]
- Rossi, P.; Aragno, M. Analysis of Bacteriophage Inactivation and Its Attenuation by Adsorption onto Colloidal Particles by Batch Agitation Techniques. Can. J. Microbiol. 1999, 45, 9–17. [Google Scholar] [CrossRef]
- Kallies, R.; Kiesel, B.; Zopfi, J.; Wick, L.Y.; Chatzinotas, A. Complete Genome Sequence of Alteromonas Virus VB_AspP-H4/4. Genome Announc. 2017, 5, e00914-17. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dedeo, M.T.; Finley, D.T.; Francis, M.B. Viral Capsids as Self-Assembling Templates for New Materials, 1st ed.; Elsevier Inc.: Amsterdam, The Netherlands, 2011; Volume 103, ISBN 9780124159068. [Google Scholar]
- Carr, E.L.; Wilson, M.E.; Adams, S.T.; Arens, D.K.; Ayala, M.; Ayers, H.; Barker, A.; Beecroft, V.; Bishop, E.; Brundage, B.; et al. Genome Sequences of 14 Siphophages That Infect Serratia Marcescens. Microbiol. Resour. Announc. 2022, 11, e0121221. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.C.; Kellogg, C.A.; Paul, J.H. Characterization of Marine Temperate Phage-Host Systems Isolated from Mamala Bay, Oahu, Hawaii. Appl. Environ. Microbiol. 1998, 64, 535–542. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kallies, R.; Kiesel, B.; Schmidt, M.; Ghanem, N.; Zopfi, J.; Hackermüller, J.; Harms, H.; Wick, L.Y.; Chatzinotas, A. Complete Genome Sequence of Pseudoalteromonas Virus VB_PspP-H6/1 That Infects Pseudoalteromonas Sp. Strain H6. Mar. Genom. 2019, 47, 100667. [Google Scholar] [CrossRef]
- Olsen, R.H.; Siak, J.-S.; Gray, R.H. Characteristics of PRD1, a Plasmid-Dependent Broad Host Range DNA Bacteriophage. J. Virol 1974, 14, 689–699. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ziedaite, G.; Daugelavic, R.; Bamford, J.K.H.; Bamford, D.H. The Holin Protein of Bacteriophage PRD1 Forms a Pore for Small-Molecule and Endolysin Translocation. J. Bacteriol. 2005, 187, 5397–5405. [Google Scholar] [CrossRef] [Green Version]
- Hodgson, C.J.; Perkins, J.; Labadz, J.C. The Use of Microbial Tracers to Monitor Seasonal Variations in Effluent Retention in a Constructed Wetland. Water Res. 2004, 38, 3833–3844. [Google Scholar] [CrossRef]
- Morozova, V.; Jdeed, G.; Kozlova, Y.; Babkin, I.; Tikunov, A.; Tikunova, N. A New Enterobacter Cloacae Bacteriophage Ec151 Encodes the Deazaguanine Dna Modification Pathway and Represents a New Genus within the Siphoviridae Family. Viruses 2021, 13, 1372. [Google Scholar] [CrossRef]
- McKenna, R.; Xia, D.; Willingmann, P.; Ilag, L.L.; Rossmann, M.G. Structure Determination of the Bacteriophage PhiX174. Acta Cryst. B 1992, 48, 499–511. [Google Scholar] [CrossRef]
- Schnegg, P.-A.; Flynn, R. Online Field Fluorometers for Hydrogeological Tracer Tests. Isot. Und Tracer Der Wasserforsch. 2002, 1879, 29–36. [Google Scholar]
- Goldscheider, N.; Haller, L.; Poté, J.; Wildi, W.; Zopfi, J. Characterizing Water Circulation and Contaminant Transport in Lake Geneva Using Bacteriophage Tracer Experiments and Limnological Methods. Environ. Sci. Technol. 2007, 41, 5252–5258. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McKay, L.D.; Sanford, W.E.; Strong, J.M. Field-Scale Migration of Colloidal Tracers in a Fractured Shale Saprolite. Ground Water 2000, 38, 139–147. [Google Scholar] [CrossRef]
- Flynn, R.; Hacini, Y.; Schnegg, P.-A.; Costa, R.; Diomande, K.A. Use of Tracer Tests and Geophysical Logging to Understand Solute and Micro-Organism Tracer Responses in Monitoring Wells with Long Screen Intervals in a Gravel Aquifer. Beiträge Zur. Hydrogeol. 2006, 55, 5–20. [Google Scholar]
- Ghanem, N.; Kiesel, B.; Kallies, R.; Harms, H.; Chatzinotas, A.; Wick, L.Y. Marine Phages as Tracers: Effects of Size, Morphology, and Physico− Chemical Surface Properties on Transport in a Porous Medium. Environ. Sci. Technol. 2016, 50, 12816–12824. [Google Scholar] [CrossRef] [PubMed]
- Flynn, R.M.; Sinreich, M. Characterisation of Virus Transport and Attenuation in Epikarst Using Short Pulse and Prolonged Injection Multi-Tracer Testing. Water Res. 2010, 44, 1138–1149. [Google Scholar] [CrossRef] [PubMed]
- Cooper, P.D. The Plaque Assay of Animal Viruses. Adv. Virus Res. 1961, 8, 319–378. [Google Scholar] [CrossRef]
- Dulbecco, R.; Vogt, M. Some Problems of Animal Virology as Studied by the Plaque Technique. Cold Spring Harb. Symp. Quant. Biol. 1953, 18, 273–279. [Google Scholar] [CrossRef]
- Roux, M. On an Invisible Microbe Antagonistic to Dysentery Bacilli. Note by M. F. d’Herelle, Presented by M. Roux. Comptes Rendus Acad. Des Sci. 1917, 165, 373–375, Erratum in Bacteriophage 2011, 1, 3–5.. [Google Scholar] [CrossRef] [Green Version]
- Shen, C.; Phanikumar, M.S.; Fong, T.T.; Aslam, I.; Mcelmurry, S.P.; Molloy, S.L.; Rose, J.B. Evaluating Bacteriophage P22 as a Tracer in a Complex Surface Water System: The Grand River, Michigan. Environ. Sci. Technol. 2008, 42, 2426–2431. [Google Scholar] [CrossRef]
- Wyer, M.D.; Kay, D.; Watkins, J.; Davies, C.; Kay, C.; Thomas, R.; Porter, J.; Stapleton, C.M.; Moore, H. Evaluating Short-Term Changes in Recreational Water Quality during a Hydrograph Event Using a Combination of Microbial Tracers, Environmental Microbiology, Microbial Source Tracking and Hydrological Techniques: A Case Study in Southwest Wales, UK. Water Res. 2010, 44, 4783–4795. [Google Scholar] [CrossRef]
- Hunt, R.J.; Borchardt, M.A.; Bradbury, K.R. Viruses as Groundwater Tracers: Using Ecohydrology to Characterize Short Travel Times in Aquifers. Groundwater 2014, 52, 187–193. [Google Scholar] [CrossRef]
- Ács, N.; Gambino, M.; Brøndsted, L. Bacteriophage Enumeration and Detection Methods. Front. Microbiol. 2020, 11, 594868. [Google Scholar] [CrossRef]
- Watzinger, F.; Suda, M.; Preuner, S.; Baumgartinger, R.; Ebner, K.; Baskova, L.; Niesters, H.G.M.; Lawitschka, A.; Lion, T. Real-Time Quantitative PCR Assays for Detection and Monitoring of Pathogenic Human Viruses in Immunosuppressed Pediatric Patients. J. Clin. Microbiol. 2004, 42, 5189. [Google Scholar] [CrossRef] [Green Version]
- Toussaint, J.F.; Sailleau, C.; Breard, E.; Zientara, S.; De Clercq, K. Bluetongue Virus Detection by Two Real-Time RT-QPCRs Targeting Two Different Genomic Segments. J. Virol. Methods 2007, 140, 115–123. [Google Scholar] [CrossRef]
- Vester, D.; Lagoda, A.; Hoffmann, D.; Seitz, C.; Heldt, S.; Bettenbrock, K.; Genzel, Y.; Reichl, U. Real-Time RT-QPCR Assay for the Analysis of Human Influenza A Virus Transcription and Replication Dynamics. J. Virol. Methods 2010, 168, 63–71. [Google Scholar] [CrossRef]
- Arya, M.; Shergill, I.S.; Williamson, M.; Gommersall, L.; Arya, N.; Patel, H.R.H. Basic Principles of Real-Time Quantitative PCR. Expert Rev. Mol. Diagn. 2005, 5, 209–219. [Google Scholar] [CrossRef]
- Klymus, K.E.; Ruiz Ramos, D.V.; Thompson, N.L.; Richter, C.A.; Ramos, R.; Development, C. Development and Testing of Species-Specific Quantitative PCR Assays for Environmental DNA Applications. J. Vis. Exp. 2020, 165, e61825. [Google Scholar] [CrossRef]
- Knudsen, S.W.; Ebert, R.B.; Hesselsøe, M.; Kuntke, F.; Hassingboe, J.; Mortensen, P.B.; Thomsen, P.F.; Sigsgaard, E.E.; Hansen, B.K.; Nielsen, E.E.; et al. Species-Specific Detection and Quantification of Environmental DNA from Marine Fishes in the Baltic Sea. J. Exp. Mar. Biol. Ecol. 2019, 510, 31–45. [Google Scholar] [CrossRef]
- Bofill-Mas, S.; Rusiñol, M. Recent Trends on Methods for the Concentration of Viruses from Water Samples. Curr. Opin. Environ. Sci. Health 2020, 16, 7–13. [Google Scholar] [CrossRef]
- Forés, E.; Rusiñol, M.; Itarte, M.; Martínez-Puchol, S.; Calvo, M.; Bofill-Mas, S. Evaluation of a Virus Concentration Method Based on Ultrafiltration and Wet Foam Elution for Studying Viruses from Large-Volume Water Samples. Sci. Total Environ. 2022, 829, 154431. [Google Scholar] [CrossRef]
- Cashdollar, J.L.; Wymer, L. Methods for Primary Concentration of Viruses from Water Samples: A Review and Meta-Analysis of Recent Studies. J. Appl. Microbiol. 2013, 115, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Chevallereau, A.; Pons, B.J.; van Houte, S.; Westra, E.R. Interactions between Bacterial and Phage Communities in Natural Environments. Nat. Rev. Microbiol. 2022, 20, 49–62. [Google Scholar] [CrossRef] [PubMed]
- Maslov, S.; Sneppen, K. Population Cycles and Species Diversity in Dynamic Kill-the-Winner Model of Microbial Ecosystems. Sci. Rep. 2017, 7, 39642. [Google Scholar] [CrossRef]
- Oechslin, F. Resistance Development to Bacteriophages Occurring during Bacteriophage Therapy. Viruses 2018, 10, 351. [Google Scholar] [CrossRef] [PubMed]
- Koskella, B.; Vos, M. Adaptation in Natural Microbial Populations. Annu Rev. Ecol. Evol. Syst. 2015, 46, 503–522. [Google Scholar] [CrossRef] [Green Version]
- Weitz, J.S.; Wilhelm, S.W. Ocean Viruses and Their Effects on Microbial Communities and Biogeochemical Cycles. F1000 Biol. Rep. 2012, 4, 2–9. [Google Scholar] [CrossRef] [Green Version]
- Clokie, M.R.J.; Millard, A.D.; Letarov, A.V.; Heaphy, S. Phages in Nature. Bacteriophage 2011, 1, 31–45. [Google Scholar] [CrossRef] [Green Version]
- Braga, L.P.P.; Spor, A.; Kot, W.; Breuil, M.C.; Hansen, L.H.; Setubal, J.C.; Philippot, L. Impact of Phages on Soil Bacterial Communities and Nitrogen Availability under Different Assembly Scenarios. Microbiome 2020, 8, 52. [Google Scholar] [CrossRef] [Green Version]
- Roux, S.; Brum, J.R.; Dutilh, B.E.; Sunagawa, S.; Duhaime, M.B.; Loy, A.; Poulos, B.T.; Solonenko, N.; Lara, E.; Poulain, J.; et al. Ecogenomics and Potential Biogeochemical Impacts of Globally Abundant Ocean Viruses. Nature 2016, 537, 689–693. [Google Scholar] [CrossRef] [Green Version]
- Chen, I.; Dubnau, D. DNA Uptake during Bacterial Transformation. Nat. Rev. Microbiol. 2004, 2, 241–249. [Google Scholar] [CrossRef]
- Johnston, C.; Martin, B.; Fichant, G.; Polard, P.; Claverys, J.P. Bacterial Transformation: Distribution, Shared Mechanisms and Divergent Control. Nat. Rev. Microbiol. 2014, 12, 181–196. [Google Scholar] [CrossRef] [PubMed]
- Ye, M.; Zhang, Z.; Sun, M.; Shi, Y. Dynamics, Gene Transfer, and Ecological Function of Intracellular and Extracellular DNA in Environmental Microbiome. iMeta 2022, 1, e34. [Google Scholar] [CrossRef]
- Woegerbauer, M.; Bellanger, X.; Merlin, C. Cell-Free DNA: An Underestimated Source of Antibiotic Resistance Gene Dissemination at the Interface Between Human Activities and Downstream Environments in the Context of Wastewater Reuse. Front. Microbiol. 2020, 11, 671. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Muniesa, M.; Colomer-Lluch, M.; Jofre, J. Could Bacteriophages Transfer Antibiotic Resistance Genes from Environmental Bacteria to Human-Body Associated Bacterial Populations? Mob. Genet. Elem. 2013, 3, 739–751. [Google Scholar] [CrossRef]
- Colavecchio, A.; Cadieux, B.; Lo, A.; Goodridge, L.D. Bacteriophages Contribute to the Spread of Antibiotic Resistance Genes among Foodborne Pathogens of the Enterobacteriaceae Family—A Review. Front. Microbiol. 2017, 8, 1108. [Google Scholar] [CrossRef] [Green Version]
- Sala-Comorera, L.; Nolan, T.M.; Reynolds, L.J.; Venkatesh, A.; Cheung, L.; Martin, N.A.; Stephens, J.H.; Gitto, A.; O’Hare, G.M.P.; O’Sullivan, J.J.; et al. Bacterial and Bacteriophage Antibiotic Resistance in Marine Bathing Waters in Relation to Rivers and Urban Streams. Front. Microbiol. 2021, 12, 2017. [Google Scholar] [CrossRef]
- Zare, S.; Derakhshandeh, A.; Mohammadi, A.; Noshadi, M. Abundance of Antibiotic Resistance Genes in Bacteria and Bacteriophages Isolated from Wastewater in Shiraz. Mol. Biol. Res. Commun. 2021, 10, 73–83. [Google Scholar] [CrossRef]
- Moon, K.; Jeon, J.H.; Kang, I.; Park, K.S.; Lee, K.; Cha, C.J.; Lee, S.H.; Cho, J.C. Freshwater Viral Metagenome Reveals Novel and Functional Phage-Borne Antibiotic Resistance Genes. Microbiome 2020, 8, 75. [Google Scholar] [CrossRef]
- Sinke, L. Viruses in Soils: What Do We Know and Where Do We Go from Here ? Available online: https://www.labroots.com/webinar/viruses-soils-know-here (accessed on 24 June 2019).
- Williamson, K.E.; Fuhrmann, J.J.; Wommack, K.E.; Radosevich, M. Viruses in Soil Ecosystems: An Unknown Quantity Within an Unexplored Territory. Annu. Rev. Virol. 2017, 4, 201–219. [Google Scholar] [CrossRef]
- Florent, P.; Cauchie, H.-M.; Herold, M.; Jacquet, S.; Ogorzaly, L. Soil PH, Calcium Content and Bacteria as Major Factors Responsible for the Distribution of the Known Fraction of the DNA Bacteriophage Populations in Soils of Luxembourg. Microorganisms 2022, 10, 1458. [Google Scholar] [CrossRef]
- Børsheim, K.Y. Native Marine Bacteriophages. FEMS Microbiol. Ecol. 1993, 102, 141–159. [Google Scholar] [CrossRef]
- Bergh, O.; Børsheim, K.Y.; Bratbak, G.; Heldal, M. High Abundance of Viruses Found in Aquatic Environments. Nature 1989, 340, 467–468. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Carreras, N.; Wetzel, C.E.; Frentress, J.; Ector, L.; McDonnell, J.J.; Hoffmann, L.; Pfister, L. Hydrological Connectivity Inferred from Diatom Transport through the Riparian-Stream System. Hydrol. Earth Syst. Sci. 2015, 19, 3133–3151. [Google Scholar] [CrossRef] [Green Version]
- Kieft, K.; Zhou, Z.; Anantharaman, K. VIBRANT: Automated Recovery, Annotation and Curation of Microbial Viruses, and Evaluation of Viral Community Function from Genomic Sequences. Microbiome 2020, 8, 90. [Google Scholar] [CrossRef] [PubMed]
- Roux, S.; Enault, F.; Hurwitz, B.L.; Sullivan, M.B. VirSorter: Mining Viral Signal from Microbial Genomic Data. PeerJ 2015, 2015, e985. [Google Scholar] [CrossRef]
- Trubl, G.; Hyman, P.; Roux, S.; Abedon, S.T. Coming-of-Age Characterization of Soil Viruses: A User’s Guide to Virus Isolation, Detection within Metagenomes, and Viromics. Soil Syst. 2020, 4, 23. [Google Scholar] [CrossRef] [Green Version]
- Mächler, E.; Salyani, A.; Walser, J.-C.; Larsen, A.; Schaefli, B.; Altermatt, F.; Ceperley, N. Environmental DNA Simultaneously Informs Hydrological and Biodiversity Characterization of an Alpine Catchment. Hydrol. Earth Syst. Sci. 2021, 25, 735–753. [Google Scholar] [CrossRef]
- Peters, J. Fate and Transport of Synthetic DNA in Surface Water; University of Waterloo: Waterloo, ON, Canada, 2021. [Google Scholar]
- Middelboe, M.; Jacquet, S.; Weinbauer, M. Viruses in Freshwater Ecosystems: An Introduction to the Exploration of Viruses in New Aquatic Habitats. Freshw. Biol. 2008, 53, 1069–1075. [Google Scholar] [CrossRef]
- Hurst, C.J.; Gerba, C.P.; Cech, I. Effects of Environmental Variables and Soil Characteristics on Virus Survival in Soil. Appl. Environ. Microbiol. 1980, 40, 1067–1079. [Google Scholar] [CrossRef] [Green Version]
- Drulis-Kawa, Z.; Majkowska-Skrobek, G.; Maciejewska, B. Bacteriophages and Phage-Derived Proteins—Application Approaches. Curr. Med. Chem. 2015, 22, 1757. [Google Scholar] [CrossRef]
- Wigginton, K.R.; Pecson, B.M.; Sigstam, T.; Bosshard, F.; Kohn, T. Virus Inactivation Mechanisms: Impact of Disinfectants on Virus Function and Structural Integrity. Environ. Sci. Technol. 2012, 46, 12069–12078. [Google Scholar] [CrossRef] [PubMed]
- Bertrand, I.; Schijven, J.F.; Sánchez, G.; Wyn-Jones, P.; Ottoson, J.; Morin, T.; Muscillo, M.; Verani, M.; Nasser, A.; de Roda Husman, A.M.; et al. The Impact of Temperature on the Inactivation of Enteric Viruses in Food and Water: A Review. J. Appl. Microbiol. 2012, 112, 1059–1074. [Google Scholar] [CrossRef] [PubMed]
- Fauvel, B.; Ogorzaly, L.; Cauchie, H.M.; Gantzer, C. Interactions of Infectious F-Specific RNA Bacteriophages with Suspended Matter and Sediment: Towards an Understanding of FRNAPH Distribution in a River Water System. Sci. Total Environ. 2017, 574, 960–968. [Google Scholar] [CrossRef] [Green Version]
- Gitis, V.; Dlugy, C.; Gun, J.; Lev, O. Studies of Inactivation, Retardation and Accumulation of Viruses in Porous Media by a Combination of Dye Labeled and Native Bacteriophage Probes. J. Contam. Hydrol. 2011, 124, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.; Chetochine, A.S.; Pepper, I.L.; Brusseau, M.L. Influence of DOC on MS-2 Bacteriophage Transport in a Sandy Soil. Water Air Soil Pollut. 2007, 178, 315–322. [Google Scholar] [CrossRef]
- Weinbauer, M.G.; Wilhelm, S.W.; Suttle, C.A.; Pledger, R.J.; Mitchell, D.L. Sunlight-Induced DNA Damage and Resistance in Natural Viral Communities. Aquat. Microb. Ecol. 1999, 17, 111–120. [Google Scholar] [CrossRef]
- Beck, S.E.; Rodriguez, R.A.; Hawkins, M.A.; Hargy, T.M.; Larason, T.C.; Linden, K.G. Comparison of UV-Induced Inactivation and RNA Damage in MS2 Phage across the Germicidal UV Spectrum. Appl. Environ. Microbiol. 2016, 82, 1468. [Google Scholar] [CrossRef] [Green Version]
- Kim, S.I.; Jin, S.G.; Pfeifer, G.P. Formation of Cyclobutane Pyrimidine Dimers at Dipyrimidines Containing 5-Hydroxymethylcytosine. Photochem. Photobiol. Sci. 2013, 12, 1409. [Google Scholar] [CrossRef] [Green Version]
- Rochette, P.J.; Therrien, J.P.; Drouin, R.; Perdiz, D.; Bastien, N.; Drobetsky, E.A.; Sage, E. UVA-Induced Cyclobutane Pyrimidine Dimers Form Predominantly at Thymine–Thymine Dipyrimidines and Correlate with the Mutation Spectrum in Rodent Cells. Nucleic Acids. Res. 2003, 31, 2786. [Google Scholar] [CrossRef] [Green Version]
- Silverman, A.I.; Peterson, B.M.; Boehm, A.B.; McNeill, K.; Nelson, K.L. Sunlight Inactivation of Human Viruses and Bacteriophages in Coastal Waters Containing Natural Photosensitizers. Environ. Sci. Technol. 2013, 47, 1870–1878. [Google Scholar] [CrossRef]
- Taj, M.K.; Ling, J.X.; Bing, L.L.; Qi, Z.; Taj, I.; Hassani, T.M.; Samreen, Z.; Yunlin, W. Effect of Dilution, Temperature and PH on the Lysis Activity of T4 Phage against E. Coli BL21. J. Anim. Plant. Sci. 2014, 24, 1252–1255. [Google Scholar]
- Jurczak-Kurek, A.; Gasior, T.; Nejman-Faleńczyk, B.; Bloch, S.; Dydecka, A.; Topka, G.; Necel, A.; Jakubowska-Deredas, M.; Narajczyk, M.; Richert, M.; et al. Biodiversity of Bacteriophages: Morphological and Biological Properties of a Large Group of Phages Isolated from Urban Sewage. Sci. Rep. 2016, 6, 34338. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zemb, O.; Manefield, M.; Thomas, F.; Jacquet, S. Phage Adsorption to Bacteria in the Light of the Electrostatics: A Case Study Using E. Coli, T2 and Flow Cytometry. J. Virol. Methods 2013, 189, 283–289. [Google Scholar] [CrossRef]
- Schijven, J.F.; Hassanizadeh, S.M. Removal of Viruses by Soil Passage: Overview of Modeling, Processes, and Parameters. Crit Rev. Environ. Sci. Technol. 2000, 30, 49–127. [Google Scholar] [CrossRef]
- Germann, P.F.; Alaoui, A.; Riesen, D. Drag Force Approach to the Transport of Colloids in Unsaturated Soils. Water Resour. Res. 2002, 38, 18-1–18-15. [Google Scholar] [CrossRef]
- Flynn, R.; Hunkeler, D.; Guerin, C.; Burn, C.; Rossi, P.; Aragno, M. Geochemical Influences on H40/1 Bacteriophage Inactivation in Glaciofluvial Sands. Environ. Geol. 2004, 45, 504–517. [Google Scholar] [CrossRef] [Green Version]
- Kvitsand, H.M.L.; Ilyas, A.; Østerhus, S.W. Rapid Bacteriophage MS2 Transport in an Oxic Sandy Aquifer in Cold Climate: Field Experiments and Modeling. Water Resour. Res. 2015, 51, 9725–9745. [Google Scholar] [CrossRef]
- Flynn, R.M.; Mallèn, G.; Engel, M.; Ahmed, A.; Rossi, P. Characterizing Aquifer Heterogeneity Using Bacterial and Bacteriophage Tracers. J. Environ. Qual. 2015, 44, 1448–1458. [Google Scholar] [CrossRef] [Green Version]
- Chattopadhyay, S.; Puls, R.W. Forces Dictating Colloidal Interactions between Viruses and Soil. Chemosphere 2000, 41, 1279–1286. [Google Scholar] [CrossRef]
- Sautrey, G.; Brié, A.; Gantzer, C.; Walcarius, A. MS2 and Qβ Bacteriophages Reveal the Contribution of Surface Hydrophobicity on the Mobility of Non-Enveloped Icosahedral Viruses in SDS-Based Capillary Zone Electrophoresis. Electrophoresis 2018, 39, 377–385. [Google Scholar] [CrossRef] [PubMed]
- Schijven, J.F.; Hoogenboezem, W.; Hassanizadeh, S.M.; Peters, J.H. Modeling Removal of Bacteriophages MS2 and PRD1 by Dune Recharge at Castricum, Netherlands. Water Resour. Res. 1999, 35, 1101–1111. [Google Scholar] [CrossRef]
- Bales, R.C.; Li, S.; Maguire, K.M.; Yahya, M.T.; Gerba, C.P.; Harvey, R.W. Virus and Bacteria Transport in a Sandy Aquifer, Cape Cod, MA. Groundwater 1995, 33, 653–661. [Google Scholar] [CrossRef]
- Flynn, R.; Cornaton, F.; Hunkeler, D.; Rossi, P. Bacteriophage Transport through a Fining-Upwards Sedimentary Sequence: Laboratory Experiments and Simulation. J. Contam. Hydrol. 2004, 74, 231–252. [Google Scholar] [CrossRef] [PubMed]
- Sinton, L.W.; Noonan, M.J.; Finlay, R.K.; Pang, L.; Close, M.E. Transport and Attenuation of Bacteria and Bacteriophages in an Alluvial Gravel Aquifer. N. Z. J. Mar. Freshw. Res. 2000, 34, 175–186. [Google Scholar] [CrossRef] [Green Version]
- Ghanem, N.; Trost, M.; Sánchez Fontanet, L.; Harms, H.; Chatzinotas, A.; Wick, L.Y. Changes of the Specific Infectivity of Tracer Phages during Transport in Porous Media. Environ. Sci. Technol. 2018, 52, 3486–3492. [Google Scholar] [CrossRef] [PubMed]
- Lisovski, S.; Ramenofsky, M.; Wingfield, J.C. Defining the Degree of Seasonality and Its Significance for Future Research. Integr. Comp. Biol. 2017, 57, 934–942. [Google Scholar] [CrossRef] [Green Version]
- Kwiecien, O.; Braun, T.; Brunello, C.F.; Faulkner, P.; Hausmann, N.; Helle, G.; Hoggarth, J.A.; Ionita, M.; Jazwa, C.S.; Kelmelis, S.; et al. What We Talk about When We Talk about Seasonality—A Transdisciplinary Review. Earth Sci. Rev. 2022, 225, 103843. [Google Scholar] [CrossRef]
- Lembrechts, J.J.; van den Hoogen, J.; Aalto, J.; Ashcroft, M.B.; De Frenne, P.; Kemppinen, J.; Kopecký, M.; Luoto, M.; Maclean, I.M.D.; Crowther, T.W.; et al. Global Maps of Soil Temperature. Glob. Chang. Biol. 2022, 28, 3110–3144. [Google Scholar] [CrossRef]
- Zhan, M.J.; Xia, L.; Zhan, L.; Wang, Y. Recognition of Changes in Air and Soil Temperatures at a Station Typical of China’s Subtropical Monsoon Region (1961–2018). Adv. Meteorol. 2019, 9, 6927045. [Google Scholar] [CrossRef] [Green Version]
- Tiwari, G.N. Madhuri Effect of Water Depth on Daily Yield of the Still. Desalination 1987, 61, 67–75. [Google Scholar] [CrossRef]
- Caldarescu, D.E.; Brey, T.; Abele, D.; Beierlein, L.; Lohmann, G.; Ionita, M. The Influence of Depth-Dependent Seasonal Temperature Variability on Growth Signal in Arctica Islandica. Front. Mar. Sci. 2021, 8, 823. [Google Scholar] [CrossRef]
- Caldentey, J.; Bamford, J.K.H.; Bamford, D.H. Structure and Assembly of Bacteriophage PRD1, an Escherichia Coli Virus with a Membrane. J. Struct. Biol. 1990, 104, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Kinoshita, T.; Bales, R.C.; Maguire, K.M.; Gerba, C.P. Effect of PH on Bacteriophage Transport through Sandy Soils. J. Contam. Hydrol. 1993, 14, 55–70. [Google Scholar] [CrossRef]
- Schijven, J.F.; Hassanizadeh, S.M.; De Bruin, R.H.A.M. Two-Site Kinetic Modeling of Bacteriophages Transport through Columns of Saturated Dune Sand. J. Contam. Hydrol. 2002, 57, 259–279. [Google Scholar] [CrossRef] [PubMed]
- Michen, B.; Graule, T. Isoelectric Points of Viruses. J. Appl. Microbiol. 2010, 109, 388–397. [Google Scholar] [CrossRef]
- Rossi, P.; Käss, W. Phagen. In Geohydrologische Markierungstechnik; Gebruüder Borntraeger: Berlin, Stuttgart, 1998; pp. 209–232. [Google Scholar]
- Hijnen, W.A.M.; Brouwer-Hanzens, A.J.; Charles, K.J.; Medema, G.J. Transport of MS2 Phage, Escherichia Coli, Clostridium Perfringens, Cryptosporidium Parvum and Giardia Intestinalis in a Gravel and a Sandy Soil. Environ. Sci. Technol. 2005, 39, 7860–7868. [Google Scholar] [CrossRef]
- Cao, H.; Tsai, F.T.C.; Rusch, K.A. Salinity and Soluble Organic Matter on Virus Sorption in Sand and Soil Columns. Ground Water 2010, 48, 42–52. [Google Scholar] [CrossRef]
- Chu, Y.; Jin, Y.; Flury, M.; Yates, M.V. Mechanisms of Virus Removal during Transport in Unsaturated Porous Media. Water Resour. Res. 2001, 37, 253–263. [Google Scholar] [CrossRef]
- Chu, Y.; Jin, Y.; Baumann, T.; Yates, M.V. Effect of Soil Properties on Saturated and Unsaturated Virus Transport through Columns. Adsorption 2003, 32, 2017–2025. [Google Scholar] [CrossRef]
- Bitton, G. Adsorption of Viruses Onto Surfaces Soil and Water. Water Res. 1975, 9, 473–484. [Google Scholar] [CrossRef]
- Bitton, G.; Mitchell, R. Effect of Colloids on the Survival of Bacteriophages in Seawater. Water Res. 1974, 8, 227–229. [Google Scholar] [CrossRef]
- Babich, H.; Stotzky, G. Reductions in Inactivation Rates of Bacteriophages by Clay Minerals in Lake Water. Water Res. 1980, 14, 185–187. [Google Scholar] [CrossRef]
- Auckenthaler, A.; Raso, G.; Huggenberger, P. Particle Transport in a Karst Aquifer: Natural and Artificial Tracer Experiments with Bacteria, Bacteriophages and Microspheres. Water Sci. Technol. 2002, 46, 131–138. [Google Scholar] [CrossRef] [PubMed]

| Bacteriophages | Family | Structure | Genome | Isoelectric Point | Bacterial Host | References |
|---|---|---|---|---|---|---|
| THE FIVE MOST USED BACTERIOPHAGES AS HYDROLOGICAL TRACERS | ||||||
| Enterobacteria phage T7 (Teseptimavirus T7) | Autographiviridae | 55-nm capsid with 19-nm tail | dsDNA of 40 kb | 4.85 | Most strains of Escherichia coli | [65,66,67] |
| Pseudoalteromonas phage vB_PspS-H40/1 | Siphoviridae | Non-contractile 68-nm tail and an icosahedral capsid of 45-nm diameter | dsDNA of 45 kb | Unknown | Pseudoalteromonas sp. | [68] |
| Coliphage F52 | Unknown | Unknown | Unknown | Unknown | Escherichia coli | [38] |
| Lessievirus MS2 (Emesvirus zinderi) | Fiersviridae | Small icosahedral virus (28 nm) | (+) ssRNA of 3 kb | 3.9 | Escherichia coli | [59,69] |
| Serratia phage strains * | Myoviridae, Siphoviridae, Podoviridae, Ackermannviridae | Head-tailed structure | dsDNA of 44–350 kb | Unknown | Strains of Serratia marcescens | [63,70] |
| ALL THE OTHER BACTERIOPHAGES USED AS HYDROLOGICAL TRACERS | ||||||
| Listonella phage phiHSIC | Siphoviridae | 47-nm capsid diameter with a non-contractile tail of 146 nm | dsDNA genome of 37,966 bp | Unknown | Vibrio pelagius | [71] |
| Pseudoalteromonasvirus vB_PspP-H6/1 | Podoviridae | 56-nm diameter icosahedral capsid and a short (15 nm) tail | linear dsDNA genome of a size of 36,753 bp | <4 | Pseudoalteromonas sp. | [72,73] |
| Alteromonasvirus vB_AspP-H4/4 | Podoviridae | Icosahedral capsid of 41 nm with a short tail of 6.6 nm | dsDNA genome of 47,631 bp | Unknown | Alteromonas sp. | [68] |
| Enterobacteria phage PRD1 (Alphatectivirus PRD1) | Tectiviridae | 62-nm capsid | dsDNA genome of 15 kb | 3.8–4.2 | Gram-negative bacterial species (i.e., Escherichia coli, Salmonella enterica, Pseudomonas aeruginosa) | [74,75] |
| Enterobacter cloacae phage strains * | Unknown | Isometric head of about 55–93 nm with a 103 nm long | DNA genomes | Unknown | Enterobacter cloacae | [76,77] |
| Coliphages F137 and F46 | Unknown | Unknown | Unknown | Unknown | Escherichia coli | [38] |
| Salmonella phage P22H5 | Podoviridae | T7-like structure | dsDNA | Unknown | Salmonella typhimurium | [49] |
| Enterobacteria phage f1 | Inoviridae | Filamentous 850 nm long | ssDNA | Unknown | Escherichia coli | [36] |
| Pseudomonas phage Psf2 (Tunavirus Psf2) | Drexlerviridae (former Siphoviridae) | T1-like structure (60-nm head and 151-nm tail) | Circular genome of about 50 kb | Unknown | Pseudomonas fluorescens | [42] |
| Enterobacteria phage phiX174 (Sinsheimervirus phiX174) | Microviridae | Icosahedral capsid | Circular ssDNA | 6.6–7 | Escherichia sp. | [78] |
| Aerobacter aerogenose 243 phages | Unknown | Unknown | Unknown | Unknown | Aerobacter aerogenes NCTC 243 | [41] |
| Bacteriophages | Reservoirs | Distance Travelled | Time Travelled | Velocity | Loss at the End of the Experiment | Number of Experiments | Other Tracers Used in Complement | References |
|---|---|---|---|---|---|---|---|---|
| THE FIVE MOST USED BACTERIOPHAGES AS HYDROLOGICAL TRACERS | ||||||||
| Enterobacteria phage T7 (Teseptimavirus T7) | Groundwater and lake | 1–160 m (24 km in one experiment) | 20–70 h | 1–350 m/h | 99.9% | 7 | H40/1, f1, F52, F137, F46, naphthionate | [36,38,45] |
| Pseudoalteromonas phage vB_PspS-H40/1 | Freshwater (especially lakes) | 64 m–6 km | 8–48 h | 8–2000 m/h | 40–99% | 6 | T7, H6/1, uranine | [45] |
| Coliphage F52 | River and lake | 8–62 km | 4–119 h | 900–2000 m/h | - | 5 | T7, F137, F46 | [38] |
| Lessievirus MS2 (Emesvirus zinderi) | Water surface treatment wetlands, aquifers | 4–15 m | 4 h–5 d | 0.2–19 m/h | 10–77% | 5 | PhiHSIC, PRD1, bromide | [44,46,48] |
| Serratia phage strains * | Surface water | 3–10 km | 1–6 h | - | 30–62% | 5 | MS2, fluorescein, Enterobacteria phage, Bacillus spores, Lithium | [15,35] |
| ALL THE OTHER BACTERIOPHAGES USED AS HYDROLOGICAL TRACERS | ||||||||
| Listonella phage phiHSIC | From wastewater to surface water and groundwater | 10 m–4 km | 3–10 h | 0.1–2 m/d 1–140 m/h | - | 3 | PRD1, MS2 | [44,51] |
| Pseudoalteromonasvirus vB_PspP-H6/1 | Freshwater (especially lakes) | 6 km | 5–26 h to 4 d | 23–210 m/h | 75–99.9% | 3 | H40/1, T7, Psf2, H4/4 | [42,45,79] |
| Alteromonasvirus vB_AspP-H4/4 | Freshwater (especially lakes) | - | 1–48 h | - | 99.9% | 2 | H40/1, T7, Psf2, H6/1 | [42,79] |
| Enterobacteria phage PRD1 (Alphatectivirus PRD1) | Low clay-content media | 4 m–4 km | 6 h–5 d | 0.2–5 m/d 1–57 m/h | 50% | 2 | MS2, bromide, PhiHSIC | [48,51] |
| Enterobacter cloacae phage strains * | Constructed wetlands and groundwater | - | 3–14 d | 0.8–4 L/s | 9–64% in wetlands 100% in groundwater | 2 | Photine, fluorescein, Serratia phage | [47] |
| Coliphages F137 and F46 | River and groundwater | 31–46 km | 78–111 h | 300–400 m/h | - | 2 and 1 | T7, F52 | [38] |
| Salmonella phage P22H5 | Karstic aquifer, groundwater | - | 4–28 d | 4–36 m/d | 99.2–99.9% | 2 | Deuterium, bromide, chloride, sulfate, pyranine, naphthionate, uranine, sulforhoramine, microspheres | [49,50] |
| Enterobacteria phage f1 | Permeable aquifers | 11–110 m | 20–70 h | - | 100% | 1 | T7, naphthionate | [36] |
| Pseudomonas phage Psf2 (Tunavirus Psf2) | Groundwater | - | 11 d | - | - | 1 | T7, H6/1, H40/1, H4/4 | [42] |
| Enterobacteria phage phiX174 (Sinsheimervirus phiX174) | Karstic aquifer | - | 10–14 d | - | 99% | 1 | Photine, fluorescein, Serratia phage, Enterobacteria phage | [47] |
| Aerobacter aerogenose 243 phages | Groundwater | 200–700 m | 2–8 d | 1–8 m/h | 99.9% | 1 | - | [41] |
| Status Quo | Encountered Problems | Proposed Solutions | |
|---|---|---|---|
| Detection methods | Use of culture-based methods only | This only detects the infectious bacteriophages, which mean they need to be intact to be detected. | The complementary use of cultured-based methods with molecular techniques, which would allow for the detection of the nucleic acid. Thus, all bacteriophages, intact and damaged, can be detected. |
| Eco-friendly property | Bacteriophages are biological entities and thus are safe for the environment. In addition, they can be rapidly eliminated from the environment. | The injection of large volumes of highly concentrated bacteriophage solutions into the environment. Their genetic material can be released and persist for a long period in the environment. | The natural populations of bacteriophages could be considered since they are numerously abundant. |
| Surrounding environment | Few of the reported tracing experiments using bacteriophages considered the characteristics of the ecosystems of interest and the environmental conditions before tracer injection. | Bacteriophage inactivation is highly dependent on environmental factors, which can lead to their detection using cultured-based methods being missed. | Characterise the catchment of interest before launching the tracing experiment and select the bacteriophage species according to the relationship between the virus and the catchment properties. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Florent, P.; Cauchie, H.-M.; Ogorzaly, L. A Virological Perspective on the Use of Bacteriophages as Hydrological Tracers. Water 2022, 14, 3991. https://doi.org/10.3390/w14243991
Florent P, Cauchie H-M, Ogorzaly L. A Virological Perspective on the Use of Bacteriophages as Hydrological Tracers. Water. 2022; 14(24):3991. https://doi.org/10.3390/w14243991
Chicago/Turabian StyleFlorent, Perrine, Henry-Michel Cauchie, and Leslie Ogorzaly. 2022. "A Virological Perspective on the Use of Bacteriophages as Hydrological Tracers" Water 14, no. 24: 3991. https://doi.org/10.3390/w14243991






