1. Introduction
Surveys of fishery resources have become particularly important in recent years, as aquatic ecosystems are threatened by overfishing and the effects of extreme weather. A need to drive commercial fisheries towards sustainability and to improve the management of small-scale fisheries and aquaculture exists worldwide [
1]. In the past decade, the survey of aquatic organisms mainly consisted of traditional methods, such as trawling, electrofishing, angling, and acoustic measurements. However, these conventional approaches are costly and time-consuming, requiring many years of taxonomy experience, and may cause some damage to the environment. Therefore, a reliable technique must be developed to assess fishery resources for ecosystem management and production [
2,
3].
The yellowfin seabream
Acanthopagrus latus belongs to the Perciformes and Sparidae and is widely distributed along the Arabian Sea coast, Red Sea, India, Japan, Indonesia, the Philippines, Korea, Australia, and China. This pelagic fish species live in shallow inshore water [
4,
5]. Because of its delicious meat and rich nutritional value, it is frequently on the dinner table. Yellowfin seabream is an important reef-dwelling economic marine fish. In recent years, yellowfin seabream resources have substantially declined due to overfishing and habitat destruction. In addition, yellowfin seabream is also a popular target for many anglers. As a result, yellowfin seabream is vulnerable to overexploitation. Yellowfin seabream resources are in serious decline. Traditional methods of resource assessment may be ecologically damaging and biologically traumatic [
6,
7]. Monitoring with gillnets and other passive devices has produced long and valuable time series for most species of economic or socioeconomic interest, but abundance data for benthic fishes, including yellowfin seabream, are scarce. Currently, little is known about biomass assessments of yellowfin seabream in the ocean [
8,
9]. An efficient and accurate survey method must be developed to better monitor the spatial and temporal distribution and dynamic changes in yellowfin seabream, and to provide data supporting resource surveys and monitoring.
Environmental DNA (eDNA) refers to genetic material obtained directly from environmental samples, such as soil, sediment, excreta, air, and water, without any obvious biogenic material using an effective, non-invasive, and easily standardized method [
10]. As its detection is both cost-effective and nonlethal, eDNA has attracted particular interest for monitoring rare and endangered species [
11]. In addition, eDNA technology can detect very-low-abundance species or species that are undetected using other conventional methods [
12,
13,
14]. Recently, eDNA has also been shown to be useful in estimating the abundance of aquatic species [
15]. Estimating actual population abundance is critical for the sustainable management of marine species [
16]. An increasing number of studies have linked the eDNA concentration to biomass [
17,
18,
19]. Assuming that eDNA released into water by yellowfin seabream is correlated with its biomass, we could estimate the species biomass from the eDNA copy number. However, environmental factors affect the amount of eDNA detected. Several studies have reported observational or experimental results on the effects of environmental factors on eDNA decay [
20,
21]. Therefore, the environmental factors affecting eDNA concentration of yellowfin seabream must be investigated to better apply eDNA technology to the quantitative analysis of yellowfin seabream in the field. These uncertainties limit the spatiotemporal inferences that can be drawn from eDNA data, thus limiting its usefulness as a monitoring tool.
In this study, we aimed to detect the eDNA content in the water released by yellowfin seabream using quantitative fluorescence PCR (qPCR) and studied the relationship between the eDNA concentration and biomass. A factorial experiment was performed to explore the effects of temperature, salinity, and pH on the yellowfin seabream eDNA concentration. This study provides a reference for the assessment of yellowfin seabream resources and monitoring the distribution pattern of yellowfin sea bream resources in the field.
2. Materials and Methods
2.1. Experimental Materials
The yellowfin seabream used in the experiment came from Yangjiang, Guangdong, China. Fish were transported alive to the Tropical Aquatic Research and Development Center of the South China Sea Fisheries Research Institute, Chinese Academy of Fisheries Sciences, for temporary rearing. All experimental equipment was bleached with a 0.1% potassium permanganate solution and then rinsed with ddH2O several times. The experimental period was from May 2021 to September 2021.
2.2. Experimental Design
2.2.1. Biomass Experiment
The experiment was conducted at the Hainan Tropical Aquatic Research and Development Center. Three replicate groups were established in white 150-L plastic barrels; the volume of water in the barrels was 100 L. Five biomass gradients were generated, namely, 1, 3, 5, 7, and 10 yellowfin seabreams (50 ± 0.76 g). A blank control group without yellowfin seabream was also established. Aeration tubes were used for continuous aeration, and fish were not fed during the experiment. The other experimental conditions remained the same. Water samples were collected at 9:00 a.m. for 7 consecutive days, and all water quality parameters, including temperature, pH, dissolved oxygen content, and salinity, were measured. Three parallel samples of each water sample and one blank control water sample were collected for 7 consecutive days, and the samples were stored at −20 °C until DNA extraction.
Three yellowfin seabreams with an average weight of 60 g were placed in a 150-L culture tank (100 L water). Three replicate groups were established. At the end of 7 d, the fish were retrieved from the tank at 9:00 a.m. on the eighth day, and 1-L water samples were collected at 9:00 a.m. every day thereafter, with three parallel samples collected at each time point for 20 d. One litre of DNA-free distilled water prepared in the laboratory was used as a negative control, and samples were stored at −20 °C until DNA extraction.
2.2.2. Environmental Factor Experiments
This experiment used an orthogonal experimental design (
Table 1 and
Table 2). The treatment included three levels of temperature (20 °C, 25 °C and 30 °C), salinity (15‰, 25‰, and 35‰) and pH (7.5, 8.0, and 8.5). The temperature was controlled by SUNSUN heating bars, and the temperature levels represented the optimum temperature range for the native waters of yellowfin seabreams. The salinity levels were selected to represent the salinity range of the sea area in which the yellowfin seabreams were collected. Desalination was performed according to the initial salinity of seawater (35‰) in the Tropical Aquatic Center. The pH was adjusted with HCl and NaHCO
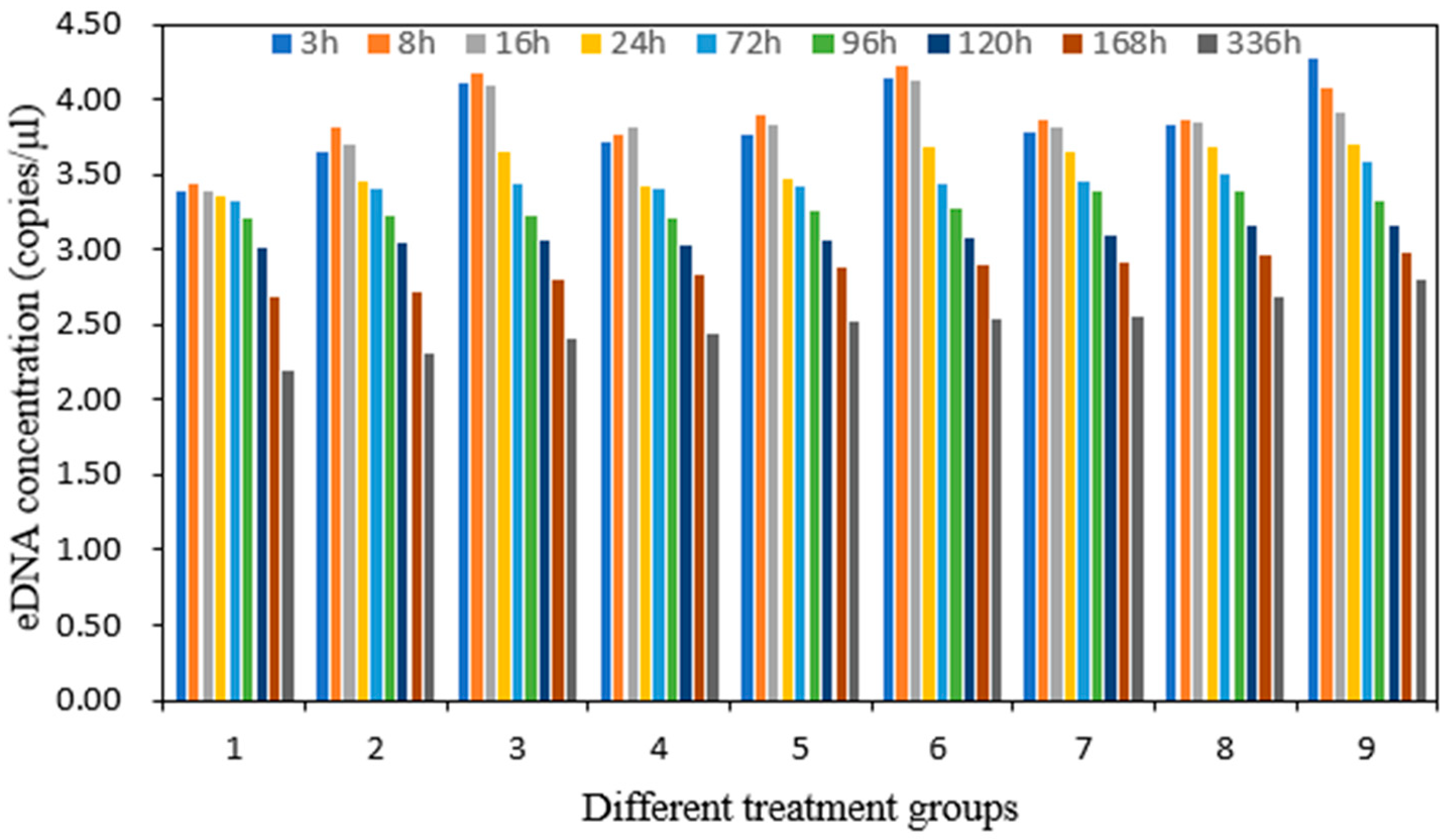
3 three times a day (morning, noon, and evening) to stabilize it. The pH levels represented a reasonable range of pH values in the experimental sea area. The experiment was conducted in white 150-L plastic buckets with a volume of water of 100 L. Each bucket contained five yellowfin seabreams. Water samples (1 L) were collected at 3, 8, 16, 24, 72, 96, 120, 168, and 336 h (0.125, 0.3, 0.7, 1, 3, 4, 5, 7, and 14 d) after transferring the yellowfin seabream into the water. Water quality parameters—namely, temperature, pH, dissolved oxygen content, and salinity—were also measured. The samples were stored at −20 °C until DNA extraction. Nine treatments were designed by orthogonal testing, and each treatment was repeated three times for a total of 27 experimental groups. Three blank groups were established per treatment for a total of 27 blank groups.
2.3. Water Sample Collection and eDNA Extraction
Before collecting water samples, the water in the bucket was gently stirred with an aeration tube to ensure that the seawater was well mixed. Water samples (1 L) were collected into 250-mL sterile polyethylene plastic bottles. The water samples were immediately extracted to obtain eDNA using glass filters with Whatman nitrocellulose membranes (47 mm diameter, 0.45 μm pore size, Cytiva, Little Chalfont, Britain). The experimental setup was wiped with 75% ethanol between each sample and then rinsed with ddH
2O. After filtration, the filter membranes were placed in sterilized 2-mL centrifuge tubes. Each membrane was wrapped separately in tinfoil after filtration, placed in a sealed bag, and marked for storage at −20 °C to avoid contamination of the samples [
22].
eDNA was extracted using the DNeasy Blood and Tissue Kit (Qiagen, Dusseldorf, Germany) with the appropriate procedure. The concentration of all extracted samples was assayed after nucleic acid purification using a Nano-300 ultramicro spectrophotometer (Allsheng, Hangzhou, China) and stored in a −20 °C freezer.
2.4. Quantitative eDNA Analysis
The quantitative analysis was performed using a 2 × TaqMan Fast qPCR Master Mix (Low Rox) real-time fluorescent quantitative PCR kit (BBI, Shanghai, China) and a qTOWER3 fluorescence quantitative PCR instrument. The mitochondrial cytochrome c oxidative subunit I (mtDNA CO I) gene of yellowfin seabream was selected to design specific primers and probes (
Table 3), which were synthesized by Biotech Bioengineering. The PCR volume was 20 µL, including 10 µL of 2 × TaqMan Fast qPCR Master Mix, 0.4 µL of the forward primer, and 0.4 µL of the reverse primer (10 µmol/L) and 0.4 µL of probe (10 µmol/L), 6.8 µL of ddH
2O, and 2 µL of the template. The reaction started with pre-denaturation at 94 °C for 3 min, followed by 40 cycles of denaturation at 94 °C for 5 s and annealing at 60 °C for 35 s. Each sample plate contained three wells serving as a negative control. The experimental data were analysed using an absolute quantitative method. The plasmid DNA concentration was diluted from 10
8 copies/µL in a 10-fold gradient to 10 copies/µL, and the quantitative standard curve of the target gene was constructed.
All phases of molecular experiments were performed on a separate sterile ultraclean table, and the UV sterilization unit was turned on for at least 20 min before and after each use to avoid cross-contamination.
2.5. Data Analysis
The eDNA concentration in water samples was calculated to establish the relationship between the biomass and eDNA concentration and the relationship between the eDNA concentration and time. All data were processed using Origin 9.0 and R 3.6.3 software, and errors were controlled to be within the 95% confidence intervals.
The environmental factors affecting the eDNA concentration of yellowfin seabream were analysed using Excel for an extreme difference analysis and SPSS 25.0 for analysis of variance. The environmental factors affecting the concentration of yellowfin seabream eDNA were evaluated, and the best combination of factors and levels was identified by combining the advantages and disadvantages of the factors to verify whether a correlation existed between environmental factors and the eDNA concentration of yellowfin seabream. Least significant difference (LSD) analysis was performed using SPSS to compare the differences between different levels of environmental factors. An assessment of the effect of environmental factors on the eDNA concentration at different points was performed using the GAM model.
4. Discussion
Our study shows a significant positive correlation between the yellowfin seabream eDNA concentration and biomass. Apparently, the low persistence of eDNA in the aquatic environment only allows the detection of species actually present in the ecosystem. The fact that
A. latus eDNA was detected using PCR in all the experiments with different biomasses of yellowfin seabreams proves that the method is reliable. In recent years, eDNA has been used as a new tool for monitoring endangered species [
23,
24], invasive species [
25,
26], and economically valuable species [
27]. Mauvisseau et al. [
28] identified a significant positive correlation between octopus (
Octopus vulgaris) biomass and the eDNA concentration. Salter et al. [
29] quantified Atlantic cod (
Gadus morhua) eDNA and compared it with the results of standardized bottom trawl surveys. eDNA detection in Atlantic cod was highly consistent with trawl catches, and a significant positive relationship was observed between the Atlantic cod eDNA concentration and biomass. Knudsen et al. [
3] reported a correlation between the eDNA concentration and the main abundance of six Baltic Sea fish species but no significant correlation with the biomass caught in trawls. Tillotson et al. [
30] observed a strong correlation between eDNA concentrations of salmonids in streams and their biomass. Yan Huiguo et al. [
31] detected a positive linear correlation between the eDNA concentration and biomass in water bodies farmed with different numbers of rock carp (
Procypris rabaudi). These results are analogous to our findings from biomass experiments. Linear correlations between fish biomass and eDNA concentrations are usually observed in small tanks, ponds, and streams. However, in large water bodies, these relationships become more complex and less certain [
32]. In contrast, nonlinear relationships are usually described in lakes [
33,
34]. For shrimp and shellfish with hard shells, the relationship between the eDNA concentration and biomass is not significant [
35]. Several studies have shown that eDNA technology is also feasible for biomass assessments under field conditions [
36,
37]. The concentration of eDNA in the water column was negatively correlated with the time since the removal of yellowfin seabream. eDNA has been shown to exponentially degrade over a long period [
38,
39].The retention time of yellowfin seabream eDNA in the environment was approximately 20 days. Because yellowfin seabream preferentially inhabits complex offshore structures on the seafloor, effective and accurate sampling has been a challenge to studying its biomass and habitat use. Confirming and describing the positive relationship for fish in controlled environments constitutes an important first step towards using eDNA for fish monitoring [
40]. Further validation is needed for field applications.
Changes in the eDNA concentration are related to the rate of eDNA release from the organism itself and the rate of degradation in the environment, which results from a complex interplay [
41]. In this study, the experiment was conducted by setting three levels of three environmental factors, namely, temperature, salinity, and pH. We performed experiments examining the effects of environmental factors on eDNA in the presence of fish, distinguishing our study from other experiments assessing the influence of environmental factors on eDNA degradation without target species [
42,
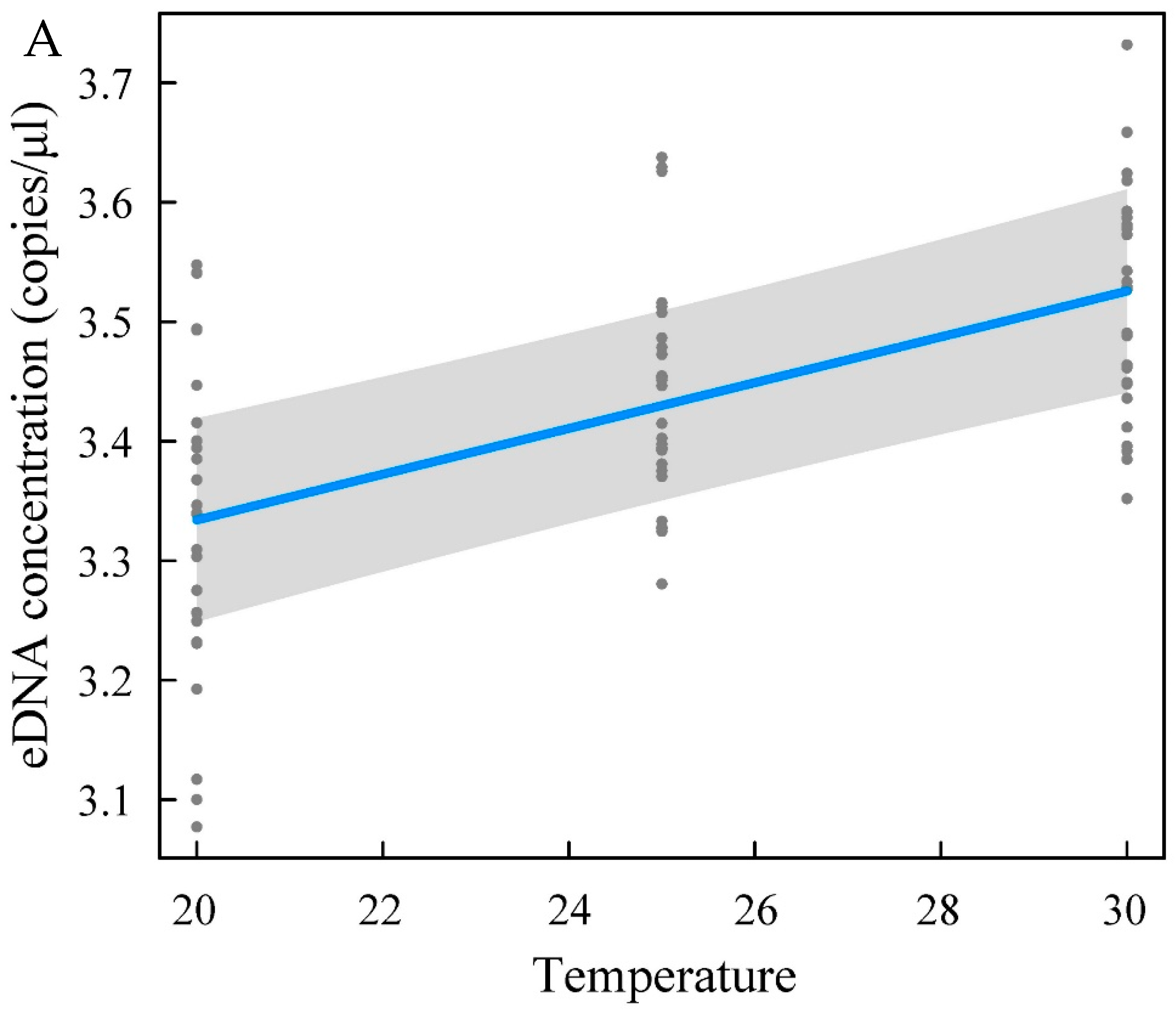
43]. The yellowfin seabream eDNA concentration increased with increasing temperature and salinity, and temperature and salinity exerted substantial effects on the eDNA concentration. The eDNA concentration in the high-temperature (30 °C) and high-salinity (35‰) groups was higher than that in the low-temperature and low-salinity groups. pH had little effect on the concentration of yellowfin seabream eDNA. An increase in water temperature within a certain range leads to an increase in the activity and metabolic rate of fish [
44]. High activity and metabolism result in epidermal cell shedding and increased excretion. This property may be one explanation for the increase in eDNA concentration. Therefore, the eDNA concentration is most likely to be affected when a large change in water temperature occurs. Water temperature must be considered when assessing biomass with the eDNA technique. Strickler et al. [
42] found that temperature differences affected eDNA decay to a greater extent than differences in UV-B exposure or pH, with the highest bullfrog eDNA concentrations detected at high temperatures (5 °C higher), low UV-B levels, and neutral pH conditions. Yellowfin seabream is a euryhaline fish. However, different salinities exert different effects on the growth performance, digestive and antioxidant enzymes, humoral immunity, and stress index of yellowfin seabream [
45]. These changes may affect the release of eDNA by yellowfin seabream living in waters with different salinities. The main effects on the concentration of biological eDNA include biotic and abiotic factors [
46,
47,
48,
49]. At different stages of organism development, the eDNA release rate in the adult group was three to four times higher than that in the juvenile group [
50]. When fish are engaged in activities such as feeding and spawning, eDNA concentrations may also change dramatically in a short period. Roussel et al. [
41] showed that fish release more eDNA in warm water than in cold water. In contrast, Klymus et al. [
27] found that although feeding increased eDNA shedding by 10-fold, water temperature had no effect. Collins et al. [
51] reported that marine systems with higher salinity, ion concentrations, and pH and more stable temperature exert favourable effects on the release and preservation of eDNA. Many factors affect the concentration of eDNA, but the roles of these factors remain somewhat controversial, and the correlation between these factors must be further explored to inform biomass assessment efforts.
Most studies assessing the effects of environmental factors on eDNA concentrations used one-way ANOVA [
16,
27,
42]. Orthogonal tests are indicators used to measure the effectiveness of an experiment [
52], and most studies have used orthogonal experiments to optimize a particular process [
53,
54]. No studies have applied orthogonal experiments to detect eDNA of different species and more efficiently obtain a representative group of environmental factor impact levels. The analysis of selected experimental results may provide a comprehensive understanding of the experimental situation to determine the optimal combination of levels and avoid large, repetitive sampling efforts. The conservation of endangered species and the management of economic species are important components of sustainable fishery development. Compared with traditional direct trapping and other invasive monitoring methods, eDNA technology is a novel method for monitoring and surveying aquatic organisms and is widely used for target-species monitoring and biomass assessment. With low invasiveness, high sensitivity, and wide applicability, eDNA monitoring methods have been widely used in recent years for qualitative, quantitative, and species diversity studies and surveys of different species [
55,
56]. This study only detected eDNA from a single species in a limited environment, but it shows the potential of a quantitative eDNA assessment in monitoring populations. Future research on eDNA detection from multiple species and understanding the key factors controlling eDNA production and persistence will potentially help translate the relationship between the eDNA concentration and target-species bioabundance into a general tool for ecological research, natural resource management, and conservation [
31].