Description of Aurelia pseudosolida sp. nov. (Scyphozoa, Ulmaridae) from the Adriatic Sea
Abstract
1. Introduction
2. Materials and Methods
3. Results
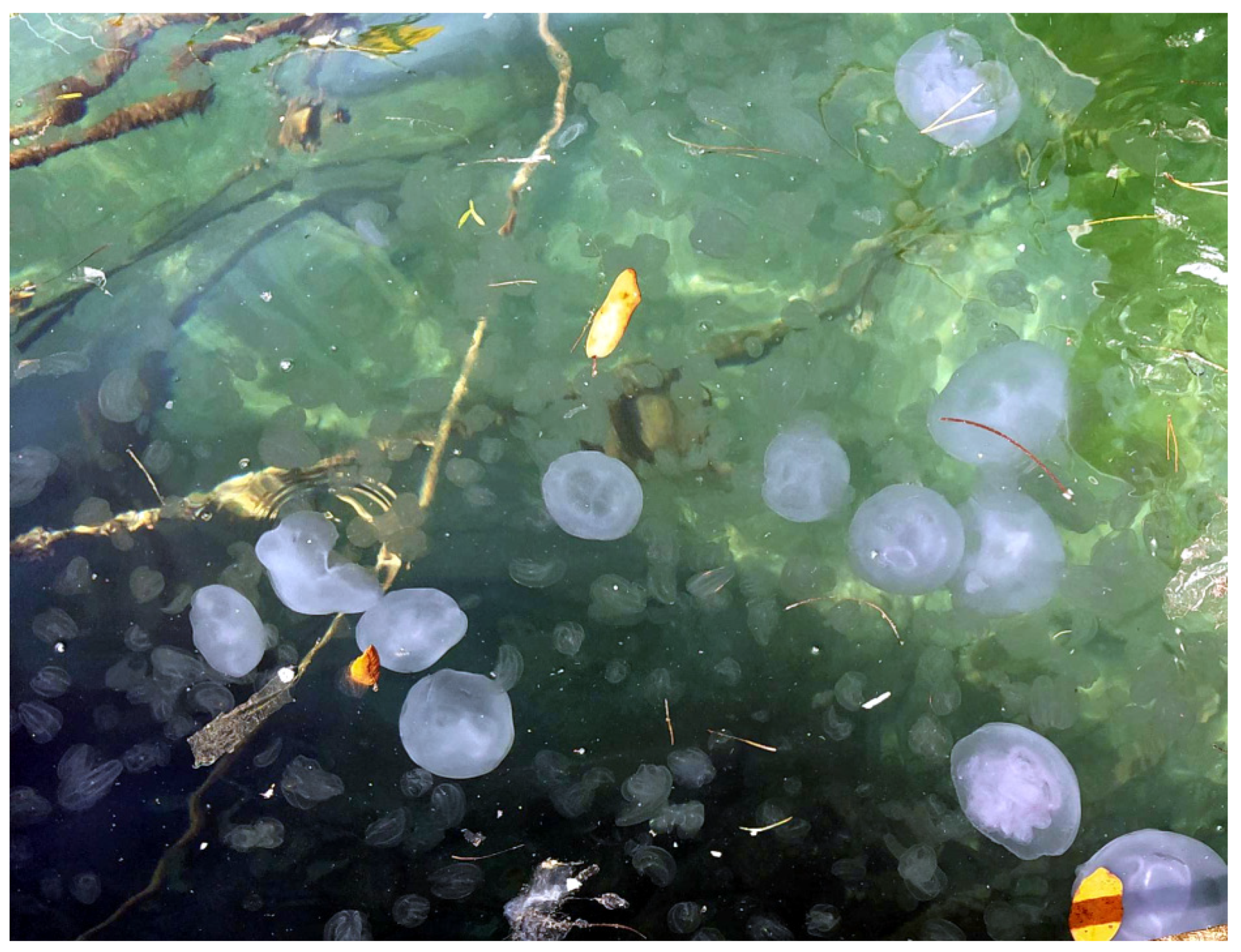
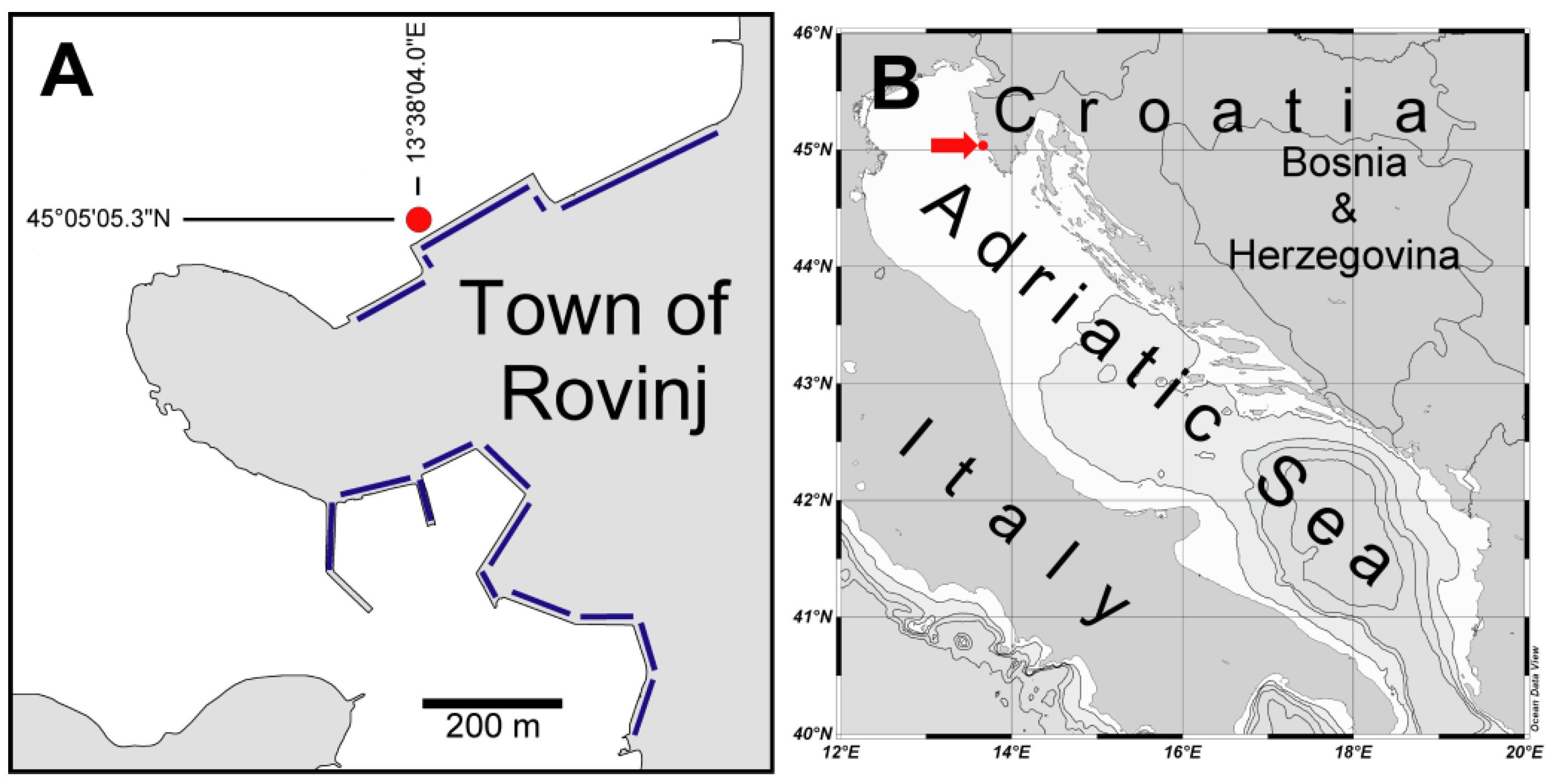
3.1. Collected Material
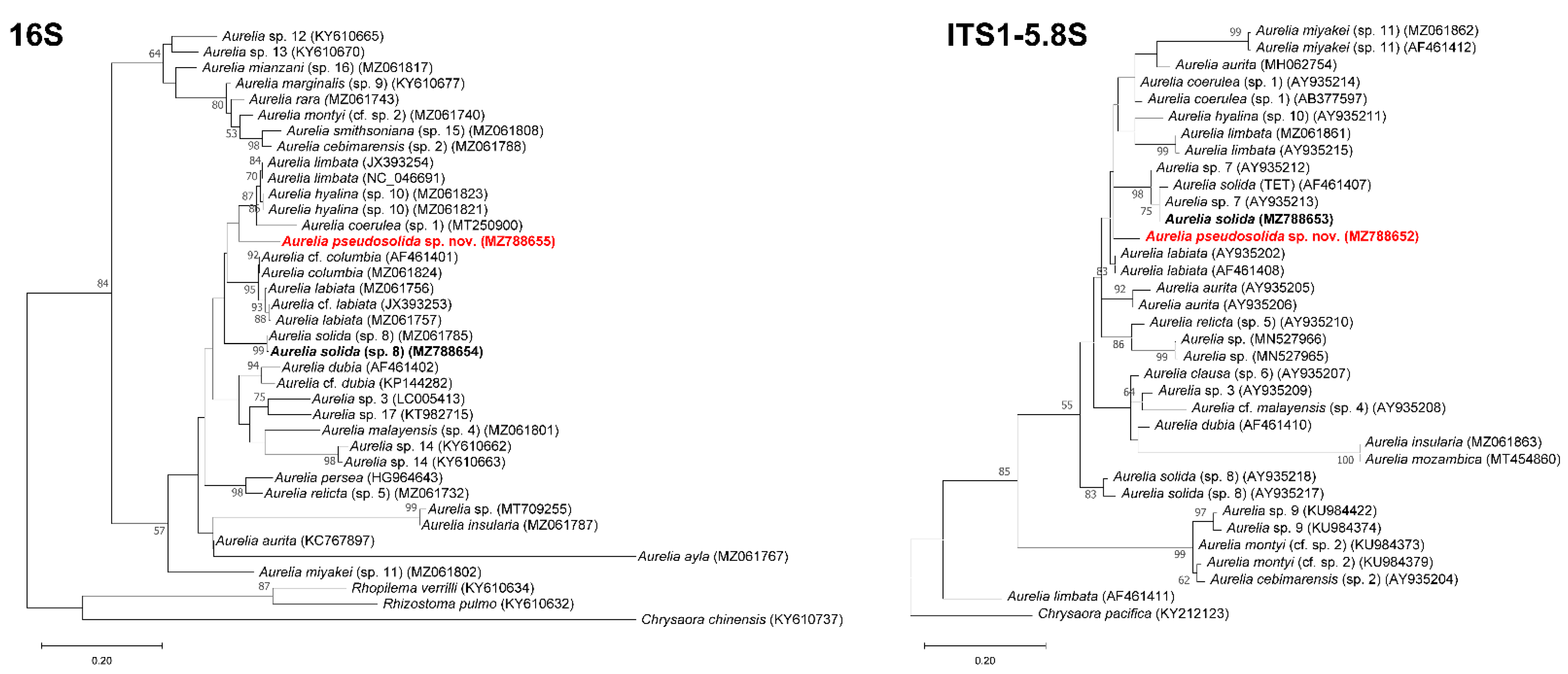
3.2. DNA Analysis
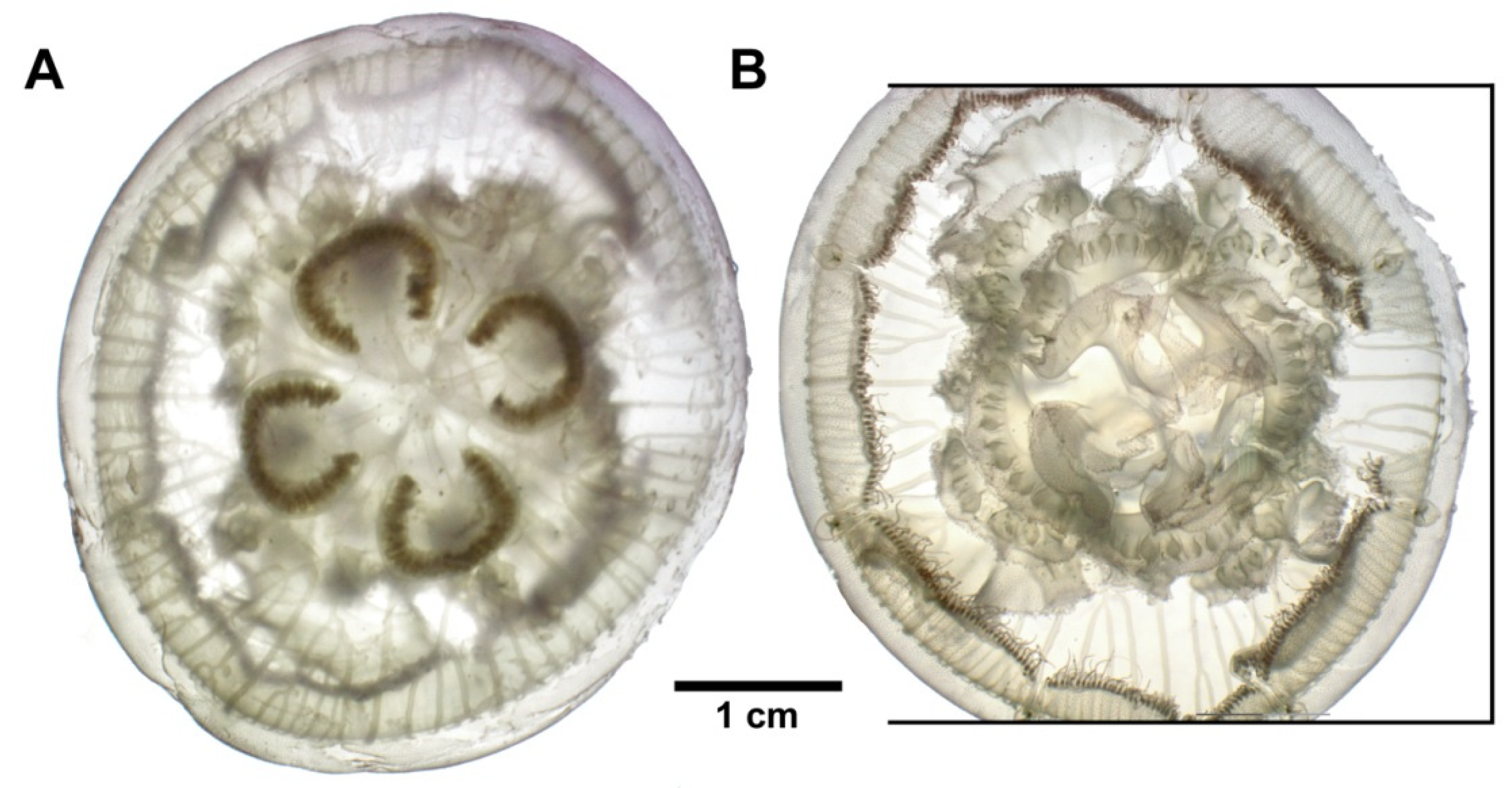
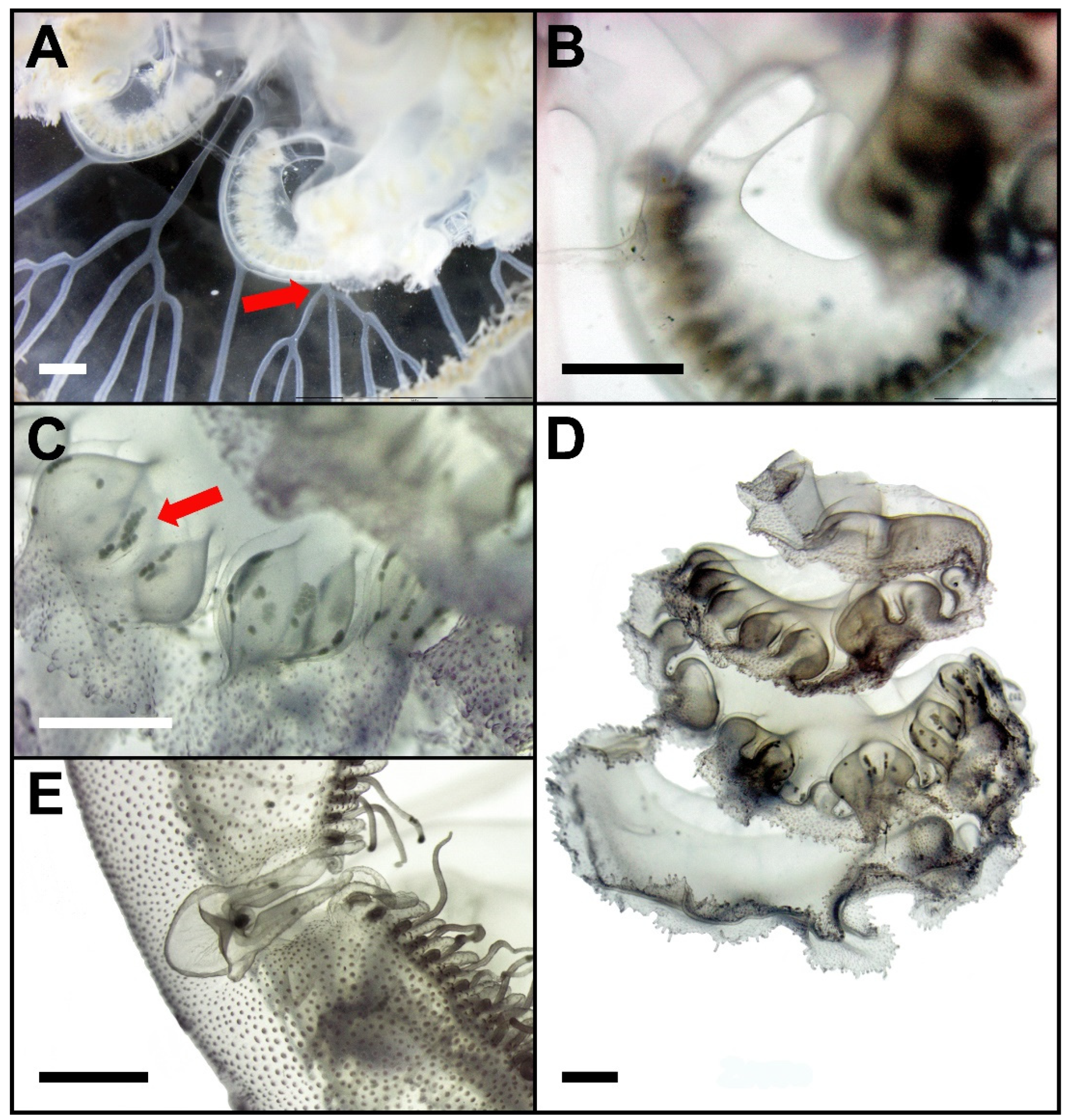
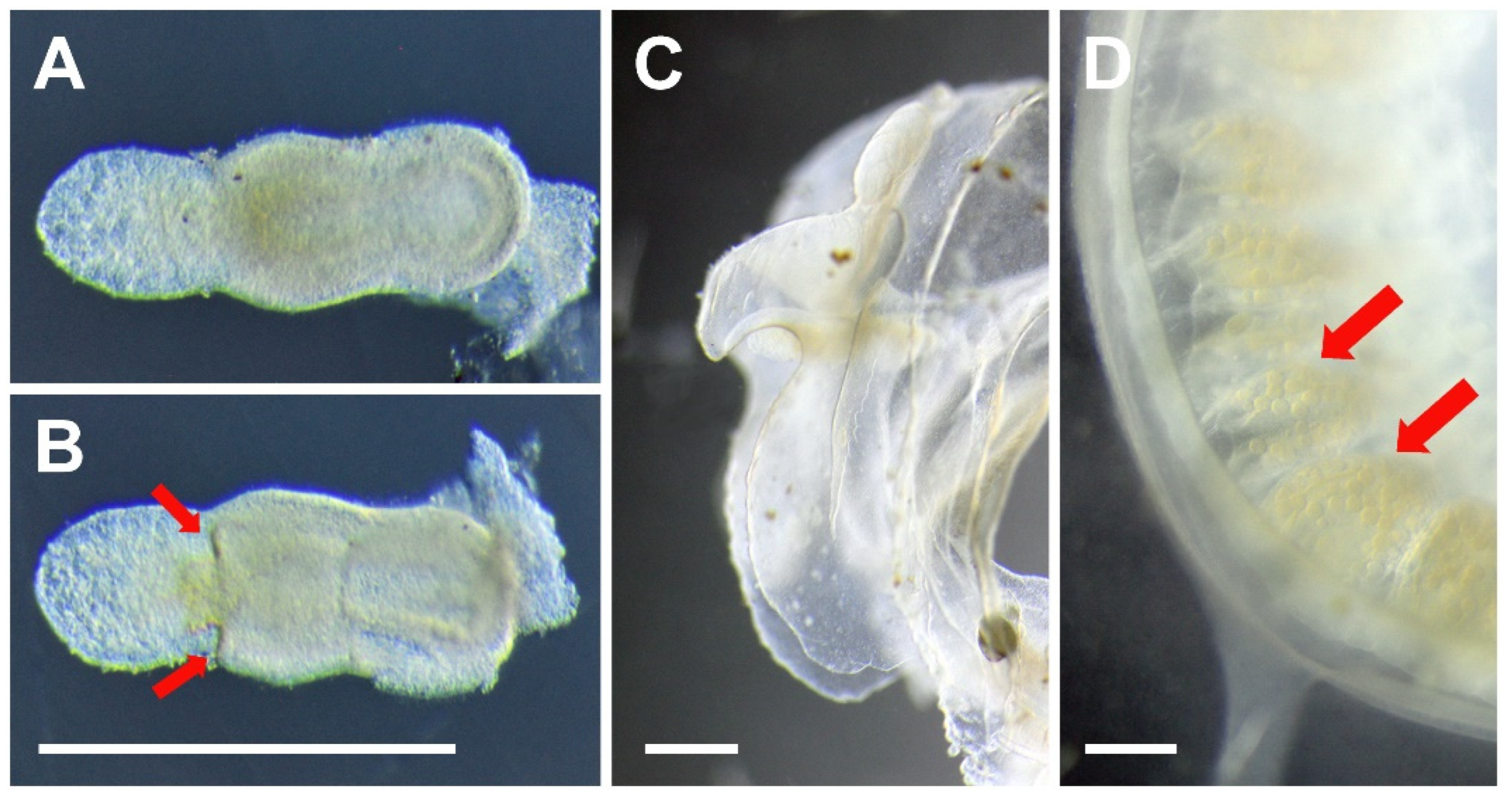
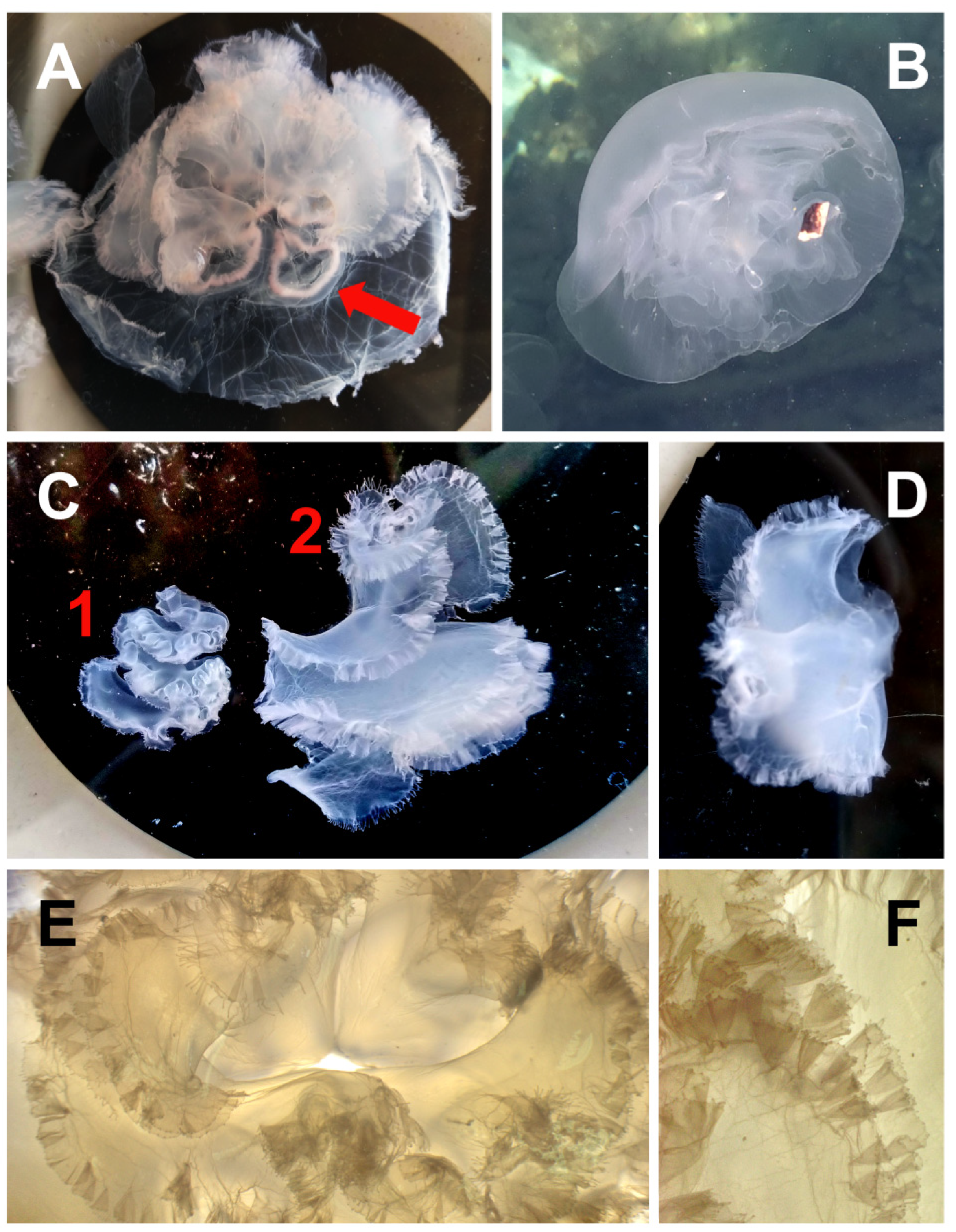
3.3. Species Description
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Purcell, J.E. Predation on Zooplankton by Large Jellyfish, Aurelia Labiata, Cyanea Capillata and Aequorea Aequorea, in Prince William Sound, Alaska. Mar. Ecol. Prog. Ser. 2003, 246, 137–152. [Google Scholar] [CrossRef]
- Purcell, J.E. Predation on Fish Eggs and Larvae by Pelagic Cnidarians and Ctenophores. Bull. Mar. Sci. 1985, 37, 739–755. [Google Scholar]
- Purcell, J.E.; Sturdevant, M.V. Prey Selection and Dietary Overlap among Zooplanktivorous Jellyfish and Juvenile Fishes in Prince William Sound, Alaska. Mar. Ecol. Prog. Ser. 2001, 210, 67–83. [Google Scholar] [CrossRef]
- Ghermandi, A.; Galil, B.; Gowdy, J.; Nunes, P.A.L.D. Jellyfish Outbreak Impacts on Recreation in the Mediterranean Sea: Welfare Estimates from a Socioeconomic Pilot Survey in Israel. Mar. Econ. Policy Relat. Ecosyst. Serv. Lessons World’s Reg. Seas. 2015, 11, 140–147. [Google Scholar] [CrossRef]
- Kim, D.-H.; Seo, J.-N.; Yoon, W.-D.; Suh, Y.-S. Estimating the Economic Damage Caused by Jellyfish to Fisheries in Korea. Fish. Sci. 2012, 78, 1147–1152. [Google Scholar] [CrossRef]
- Scorrano, S.; Aglieri, G.; Boero, F.; Dawson, M.N.; Piraino, S. Unmasking Aurelia Species in the Mediterranean Sea: An Integrative Morphometric and Molecular Approach. Zool. J. Linn. Soc. 2017, 180, 243–267. [Google Scholar] [CrossRef]
- Schroth, W.; Jarms, G.; Streit, B.; Schierwater, B. Speciation and Phylogeography in the Cosmopolitan Marine Moon Jelly, Aurelia Sp. BMC Evol. Biol. 2002, 2, 1. [Google Scholar] [CrossRef] [PubMed]
- Ramšak, A.; Venko, K.; Malej, A. Comparative Phylogeography of Meroplanktonic Species, Aurelia Spp. and Rhizostoma Pulmo (Cnidaria: Scyphozoa) in European Seas. Hydrobiologia 2012, 690, 69–80. [Google Scholar] [CrossRef]
- Dawson, M.N.; Jacobs, D.K. Molecular Evidence for Cryptic Species of Aurelia Aurita (Cnidaria, Scyphozoa). Biol. Bull. 2001, 200, 92–96. [Google Scholar] [CrossRef]
- Collins, A.G.; Jarms, G.; Morandini, A.C. World List of Scyphozoa. Aurelia Lamarck. 1816. Available online: https://www.marinespecies.org/aphia.php?p=taxdetails&id=135263 (accessed on 4 November 2021).
- Brown, M.; Scorrano, S.; Kuplik, Z.; Kuyper, D.; Ras, V.; Thibault, D.; Engelbrecht, A.; Gibbons, M.J. A New Macromedusa from the Coast of Mozambique: Aurelia Mozambica sp. nov. (Scyphozoa: Ulmaridae). Zootaxa 2021, 4933, 263–276. [Google Scholar] [CrossRef]
- Gershwin, L. Systematics and Biogeography of the Jellyfish Aurelia Labiata (Cnidaria: Scyphozoa). Biol. Bull. 2001, 201, 104–119. [Google Scholar] [CrossRef]
- Dawson, M.N.; Gupta, A.S.; England, M.H. Coupled Biophysical Global Ocean Model and Molecular Genetic Analyses Identify Multiple Introductions of Cryptogenic Species. Proc. Natl. Acad. Sci. USA 2005, 102, 11968. [Google Scholar] [CrossRef] [PubMed]
- Gómez Daglio, L.; Dawson, M.N. Species Richness of Jellyfishes (Scyphozoa: Discomedusae) in the Tropical Eastern Pacific: Missed Taxa, Molecules, and Morphology Match in a Biodiversity Hotspot. Invertebr. Syst. 2017, 31, 635–663. [Google Scholar] [CrossRef]
- Lawley, J.W.; Gamero-Mora, E.; Maronna, M.M.; Chiaverano, L.M.; Stampar, S.N.; Hopcroft, R.R.; Collins, A.G.; Morandini, A.C. The Importance of Molecular Characters When Morphological Variability Hinders Diagnosability: Systematics of the Moon Jellyfish Genus Aurelia (Cnidaria: Scyphozoa). PeerJ 2021, 9, e11954. [Google Scholar] [CrossRef] [PubMed]
- Slusarenko, A.J. A Rapid Miniprep for the Isolation of Total DNA from Agrobacterium Tumefaciens. Plant Mol. Biol. Report. 1990, 8, 249–252. [Google Scholar] [CrossRef]
- Osterburg, H.H.; Allen, J.K.; Finch, C.E. The Use of Ammonium Acetate in the Precipitation of Ribonucleic Acid. Biochem. J. 1975, 147, 367–368. [Google Scholar] [CrossRef]
- Medlin, L.; Elwood, H.J.; Stickel, S.; Sogin, M.L. The Characterization of Enzymatically Amplified Eukaryotic 16S-like RRNA-Coding Regions. Gene 1988, 71, 491–499. [Google Scholar] [CrossRef]
- White, T.; Bruns, T.; Lee, S.; Taylor, J. Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M., Gelfand, D., Sninsky, J., White, T., Eds.; Academic Press, Inc.: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- Ye, W.; Zeng, Y.; Kerns, J. Molecular Characterisation and Diagnosis of Root-Knot Nematodes (Meloidogyne Spp.) from Turfgrasses in North Carolina, USA. PLoS ONE 2015, 10, e0143556. [Google Scholar] [CrossRef]
- Folmer, O.; Black, M.; Hoeh, W.; Lutz, R.; Vrijenhoek, R. DNA Primers for Amplification of Mitochondrial Cytochrome c Oxidase Subunit I from Diverse Metazoan Invertebrates. Mol. Mar. Biol. Biotechnol. 1994, 3, 294–299. [Google Scholar]
- Cunningham, C.W.; Buss, L.W. Molecular Evidence for Multiple Episodes of Paedomorphosis in the Family Hydractiniidae. Biochem. Syst. Ecol. 1993, 21, 57–69. [Google Scholar] [CrossRef]
- Schuchert, P. Species Boundaries in the Hydrozoan Genus Coryne. Mol. Phylogenet. Evol. 2005, 36, 194–199. [Google Scholar] [CrossRef]
- Batistić, M.; Garić, R. The Case of Bougainvillia Triestina Hartlaub 1911 (Hydrozoa, Cnidaria): A 100-Year-Long Struggle for Recognition. Mar. Ecol. 2016, 37, 145–154. [Google Scholar] [CrossRef]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT Online Service: Multiple Sequence Alignment, Interactive Sequence Choice and Visualization. Brief. Bioinform. 2019, 20, 1160–1166. [Google Scholar] [CrossRef] [PubMed]
- Castresana, J. Selection of Conserved Blocks from Multiple Alignments for Their Use in Phylogenetic Analysis. Mol. Biol. Evol. 2000, 17, 540–552. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Thompson, J.D.; Higgins, D.G.; Gibson, T.J. CLUSTAL W: Improving the Sensitivity of Progressive Multiple Sequence Alignment through Sequence Weighting, Position-Specific Gap Penalties and Weight Matrix Choice. Nucleic Acids Res. 1994, 22, 4673–4680. [Google Scholar] [CrossRef] [PubMed]
- Ortman, B.D.; Bucklin, A.; Pagès, F.; Youngbluth, M. DNA Barcoding the Medusozoa Using MtCOI. Species Divers. Mar. Zooplankton 2010, 57, 2148–2156. [Google Scholar] [CrossRef]
- Bigelow, H.B. Medusae from the Maldive Islands. Bull. Mus. Comp. Zool. Harv. Coll. 1904, 39, 245–269. [Google Scholar]
- Brandt, J.-F. Prodromus Descriptionis Animalium Ab H. Mertensio Observatorum: Fascic. I. Polypos, Acalephas Discophoras et Siphonophoras, Nec Non Echinodermata Continens. Recueil des Actes de la Séance Publique de l’Académie Impériale des Sciences de St. Pétersbourg 1835, 1834, 201–275. [Google Scholar]
- Agassiz, A.; Mayer, A.G. Acalephs from the Fiji Islands. Bull. Mus. Comp. Zoöl. Harv. Coll. 1899, 32, 157–189. [Google Scholar]
- Benović, A.; Fonda-Umani, S.; Malej, A.; Specchi, M. Net-Zooplankton Biomass of the Adriatic Sea. Mar. Biol. 1984, 79, 209–218. [Google Scholar] [CrossRef]
- Kogovšek, T.; Bogunović, B.; Malej, A. Recurrence of Bloom-Forming Scyphomedusae: Wavelet Analysis of a 200-Year Time Series. Hydrobiologia 2010, 645, 81–96. [Google Scholar] [CrossRef]
- Kršinić, F.; Njire, J. An Invasion by Muggiaea Atlantica Cunningham 1892 in the Northern Adriatic Sea in the Summer of 1997 and the Fate of Small Copepods. Acta Adriat. 2001, 42, 49–59. [Google Scholar]
- Piraino, S.; Aglieri, G.; Martell, L.; Mazzoldi, C.; Melli, V.; Milisenda, G.; Scorrano, S.; Boero, F. Pelagia Benovici sp. nov. (Cnidaria, Scyphozoa): A New Jellyfish in the Mediterranean Sea. Zootaxa 2014, 3794, 455–468. [Google Scholar] [CrossRef] [PubMed]
- Malej, A.; Tirelli, V.; Lučić, D.; Paliaga, P.; Vodopivec, M.; Goruppi, A.; Ancona, S.; Benzi, M.; Bettoso, N.; Camatti, E.; et al. Mnemiopsis Leidyi in the Northern Adriatic: Here to Stay? J. Sea Res. 2017, 124. [Google Scholar] [CrossRef]
- Fiori, E.; Benzi, M.; Ferrari, C.R.; Mazziotti, C. Zooplankton Community Structure before and after Mnemiopsis Leidyi Arrival. J. Plankton Res. 2019, 41, 803–820. [Google Scholar] [CrossRef]
- Malej, A.; Kogovšek, T.; Ramšak, A.; Catenacci, L. Blooms and Population Dynamics of Moon Jellyfish in the Northern Adriatic. Cah. Biol. Mar. 2012, 53, 337–342. [Google Scholar]
- Batistić, M.; Garić, R.; Molinero, J.C. Interannual Variations in Adriatic Sea Zooplankton Mirror Shifts in Circulation Regimes in the Ionian Sea. Clim. Res. 2014, 61, 231–240. [Google Scholar] [CrossRef]
- Gačić, M.; Borzelli, G.L.E.; Civitarese, G.; Cardin, V.; Yari, S. Can Internal Processes Sustain Reversals of the Ocean Upper Circulation? The Ionian Sea Example. Geophys. Res. Lett. 2010, 37. [Google Scholar] [CrossRef]
- Civitarese, G.; Gačić, M.; Lipizer, M.; Eusebi Borzelli, G.L. On the Impact of the Bimodal Oscillating System (BiOS) on the Biogeochemistry and Biology of the Adriatic and Ionian Seas (Eastern Mediterranean). Biogeosciences 2010, 7, 3987–3997. [Google Scholar] [CrossRef]
- Garić, R.; Batistić, M. Description of Brooksia Lacromae sp. nov. (Tunicata, Thaliacea) from the Adriatic Sea. Eur. J. Taxon. 2016, 196. [Google Scholar] [CrossRef]
- Garić, R.; Batistić, M. Fritillaria Ragusina sp. nov., a New Species of Appendicularia (Tunicata) from the Adriatic Sea. J. Mar. Biol. Assoc. UK 2011, 91, 555–559. [Google Scholar] [CrossRef]

| Primer | 5′-3′ Sequence | PCR | Seq | Reference |
|---|---|---|---|---|
| 18S-ITS1-5.8S-ITS2-28S | ||||
| app2f | ATCTGGTTGATCCTGCCAGT | X | Modified from Medlin et al., 1988 [18] | |
| n1800r | GATCCTTCCGCAGGTTCACCT | X | ||
| its5 | GGAAGTAAAAGTCGTAACAAGG | X | White et al., 1990 [19] | |
| 28sr | TGGTTCGATTAGTCTTTCGCC | X | Modified from Ye et al., 2015 [20] | |
| 500f | ATTGGAGGGCAAGTCTGGTG | X | Universal | |
| 1300f | GGTGGTGCATGGCCGTTCTTAG | X | ||
| omm1100r | ATCCAAGAATTTCACCTCTAACG | X | This study | |
| oiko800r | CTGCTTTGAACACTCTAATTTTTTC | X | ||
| aur550f | GGTAATTCCAGCTCCAATAGC | X | ||
| aur750f | TGTGCTCTTAACTGAGTGTGC | X | ||
| omm1100r | ATCCAAGAATTTCACCTCTAACG | X | ||
| aur58sr | TTGACATGACGCTCAGACAG | X | ||
| 28sinf | ACCCGCTGAATTTAAGCATATTA | X | X | |
| ilitis2r | GCATTCCCAAACAACCCGACTC | X | ||
| aur28s200r | TCTCTGATGTGCTGTTCCAAG | X | ||
| aur28s600r | TCTAGGCACGAGTGGATATAAC | X | ||
| COI | ||||
| LCO1490 | GGTCAACAAATCATAAAGATATTGG | X | X | Folmer et al., 1994 [21] |
| HCO2198 | TAAACTTCAGGGTGACCAAAAAATCA | X | X | |
| 16S | ||||
| SHB | TCGACTGTTTACCAAAAACATA | X | Cunningham and Buss (1993) [22]; Schuchert (2005) [23] | |
| SHA2 | ATTGTAGATAGAAACCTTCCTG | X | Batistić and Garić, 2016 [24] |
| Species | Isolate | Accession Number |
|---|---|---|
| 18S-ITS1- 5.8S-ITS2-28S | ||
| Aurelia pseudosolida sp. nov. | ASP2 | MZ788652 |
| Aurelia solida | ASP6 | MZ788653 |
| COI | ||
| Aurelia pseudosolida sp. nov. | ASP2 | MZ782092 |
| Aurelia solida | ASP1 | MZ782091 |
| Aurelia solida | ASP3 | MZ782093 |
| Aurelia solida | ASP4 | MZ782094 |
| Aurelia solida | ASP5 | MZ782095 |
| Aurelia solida | ASP6 | MZ782096 |
| 16S | ||
| Aurelia pseudosolida sp. nov. | ASP2 | MZ788655 |
| Aurelia solida | ASP1 | MZ788654 |
| Aurelia solida | ASP3 | MZ788656 |
| Aurelia solida | ASP4 | MZ788657 |
| Aurelia solida | ASP5 | MZ788658 |
| Aurelia solida | ASP6 | MZ788659 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Garić, R.; Batistić, M. Description of Aurelia pseudosolida sp. nov. (Scyphozoa, Ulmaridae) from the Adriatic Sea. Water 2022, 14, 135. https://doi.org/10.3390/w14020135
Garić R, Batistić M. Description of Aurelia pseudosolida sp. nov. (Scyphozoa, Ulmaridae) from the Adriatic Sea. Water. 2022; 14(2):135. https://doi.org/10.3390/w14020135
Chicago/Turabian StyleGarić, Rade, and Mirna Batistić. 2022. "Description of Aurelia pseudosolida sp. nov. (Scyphozoa, Ulmaridae) from the Adriatic Sea" Water 14, no. 2: 135. https://doi.org/10.3390/w14020135
APA StyleGarić, R., & Batistić, M. (2022). Description of Aurelia pseudosolida sp. nov. (Scyphozoa, Ulmaridae) from the Adriatic Sea. Water, 14(2), 135. https://doi.org/10.3390/w14020135






