Abundance and Diversity of Nitrifying Microorganisms in Marine Recirculating Aquaculture Systems
Abstract
:1. Introduction
2. Materials and Methods
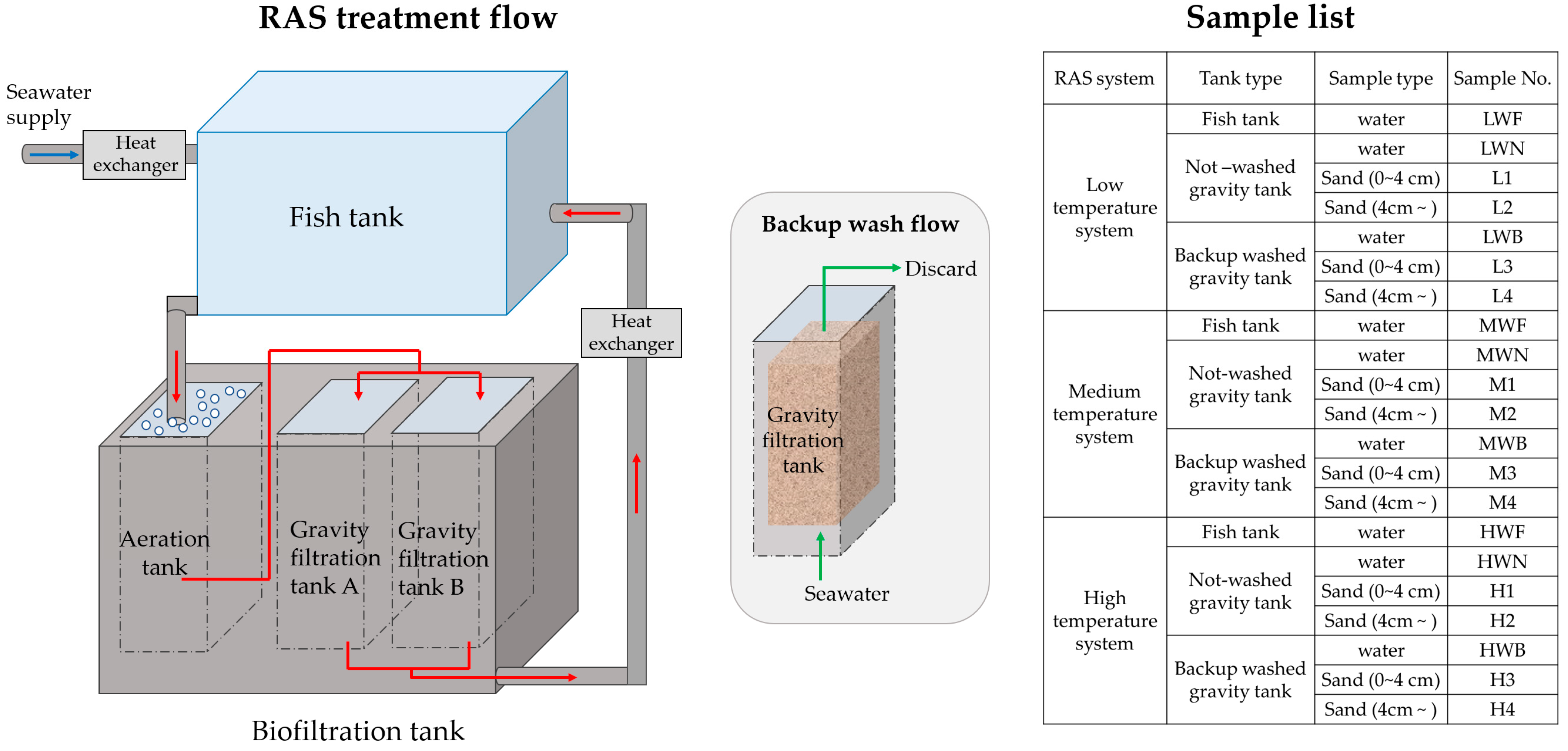
2.1. Sampling
2.2. Water Chemistry Analysis
2.3. DNA Extraction and Quantitative PCR Assays
2.4. 16 S rRNA Amplicon Sequencing
2.5. Nucleotide Sequence Accession Numbers
2.6. Statistical Analysis
3. Results
3.1. Water Quality in Each Tank
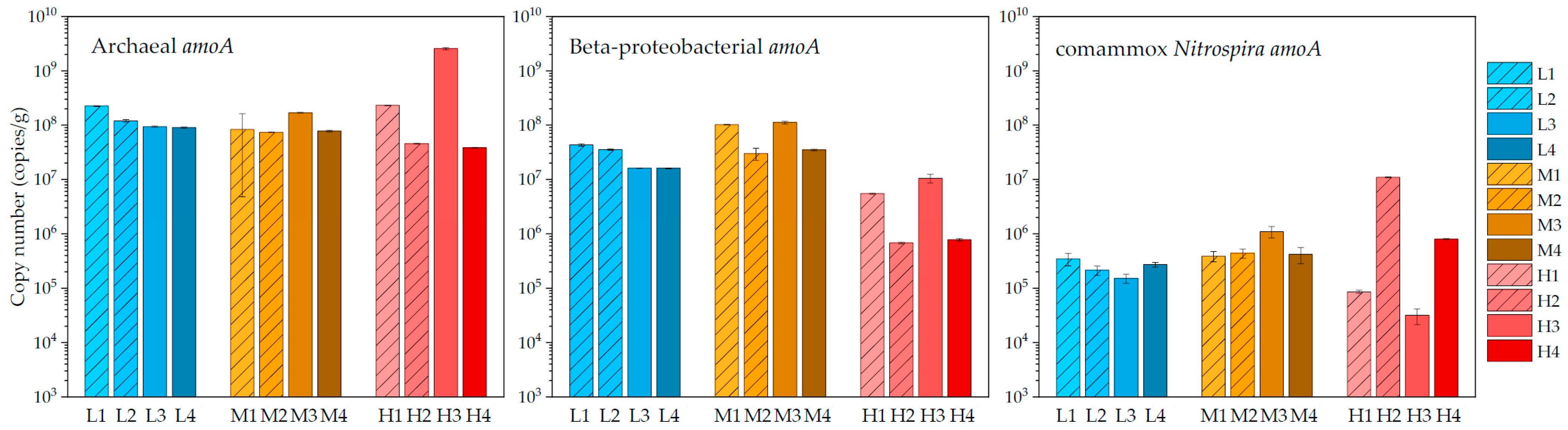
3.2. Quantification Results of Archaeal, Beta-Proteobacterial, and Comammox Nitrospira Amoa Genes
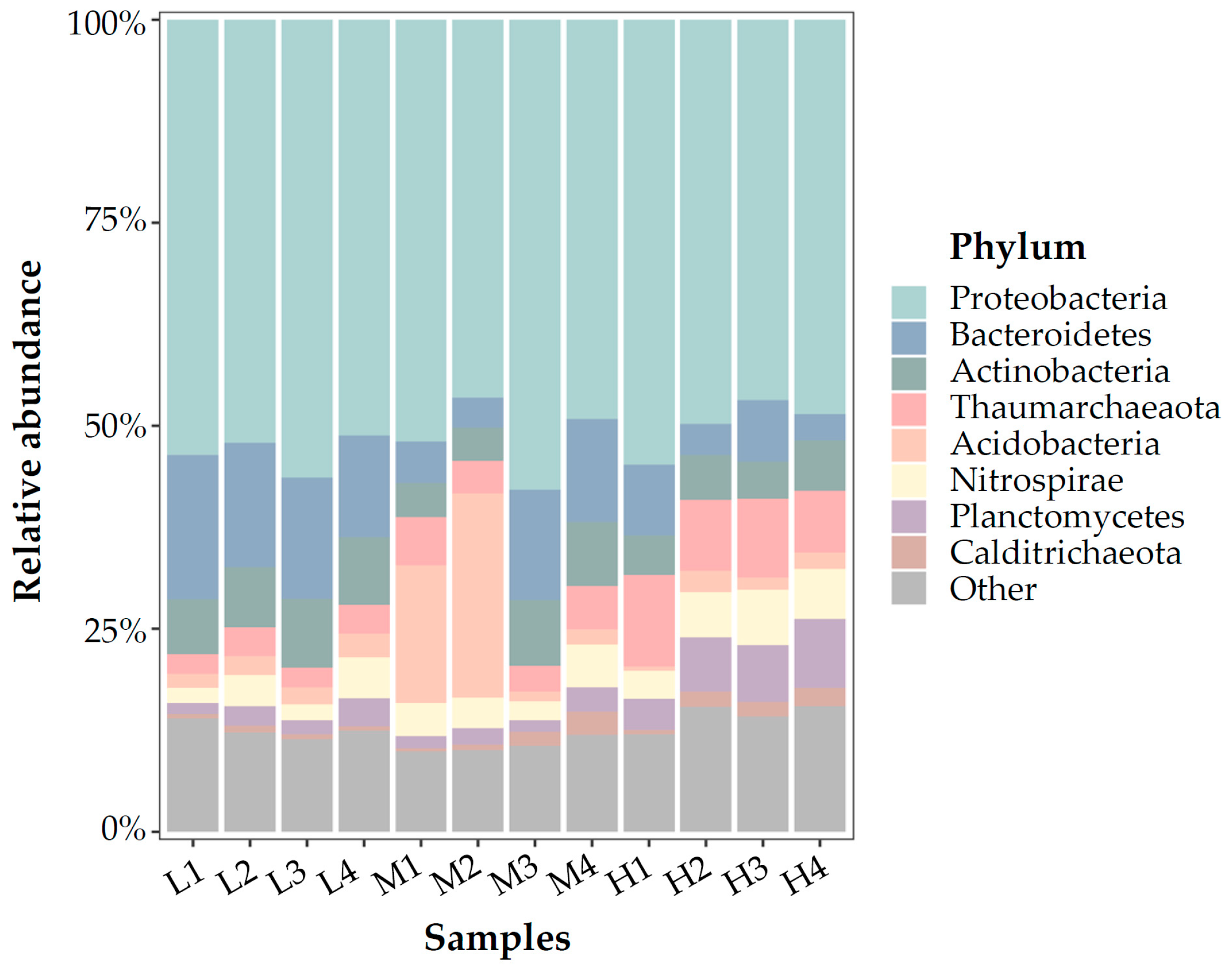
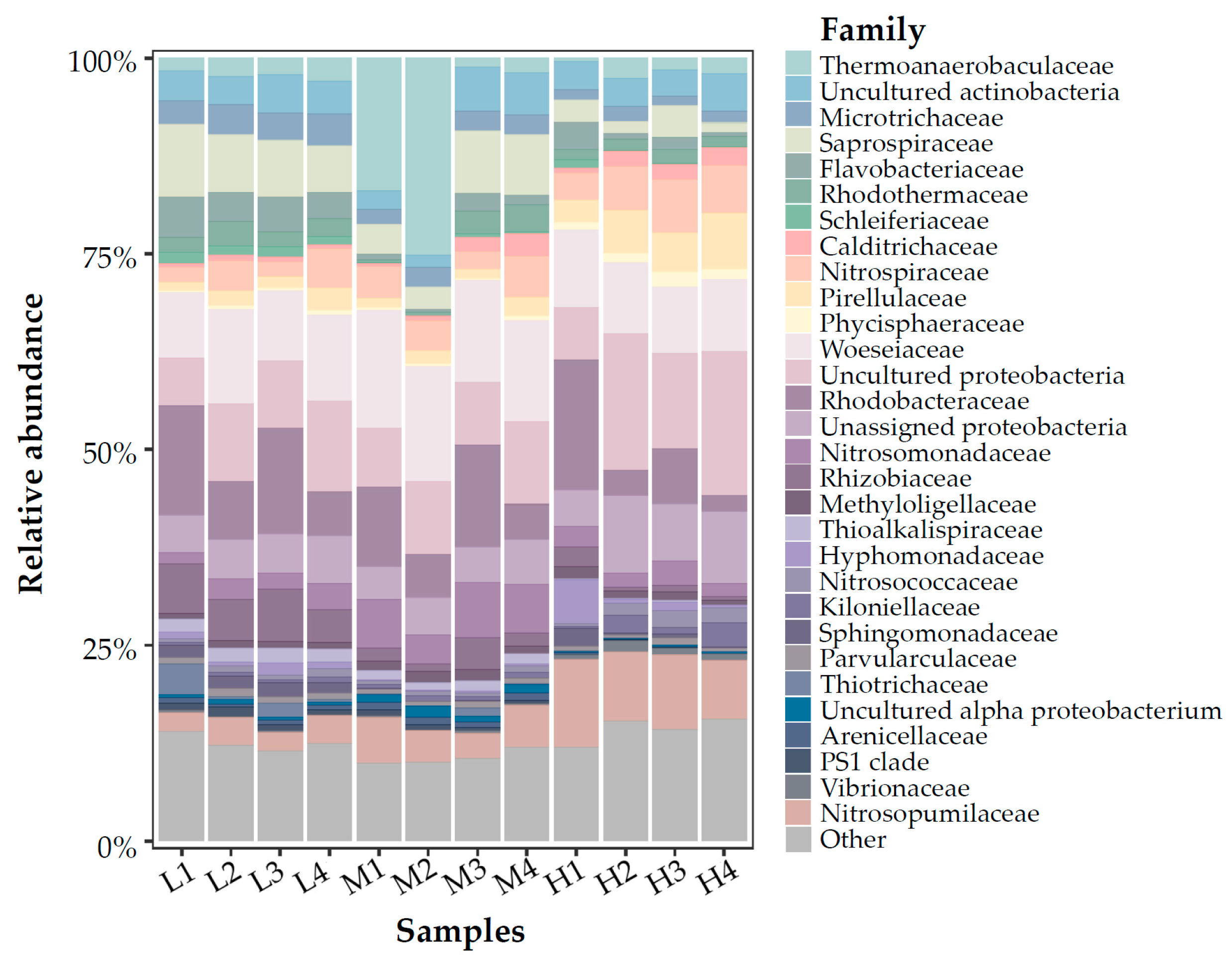
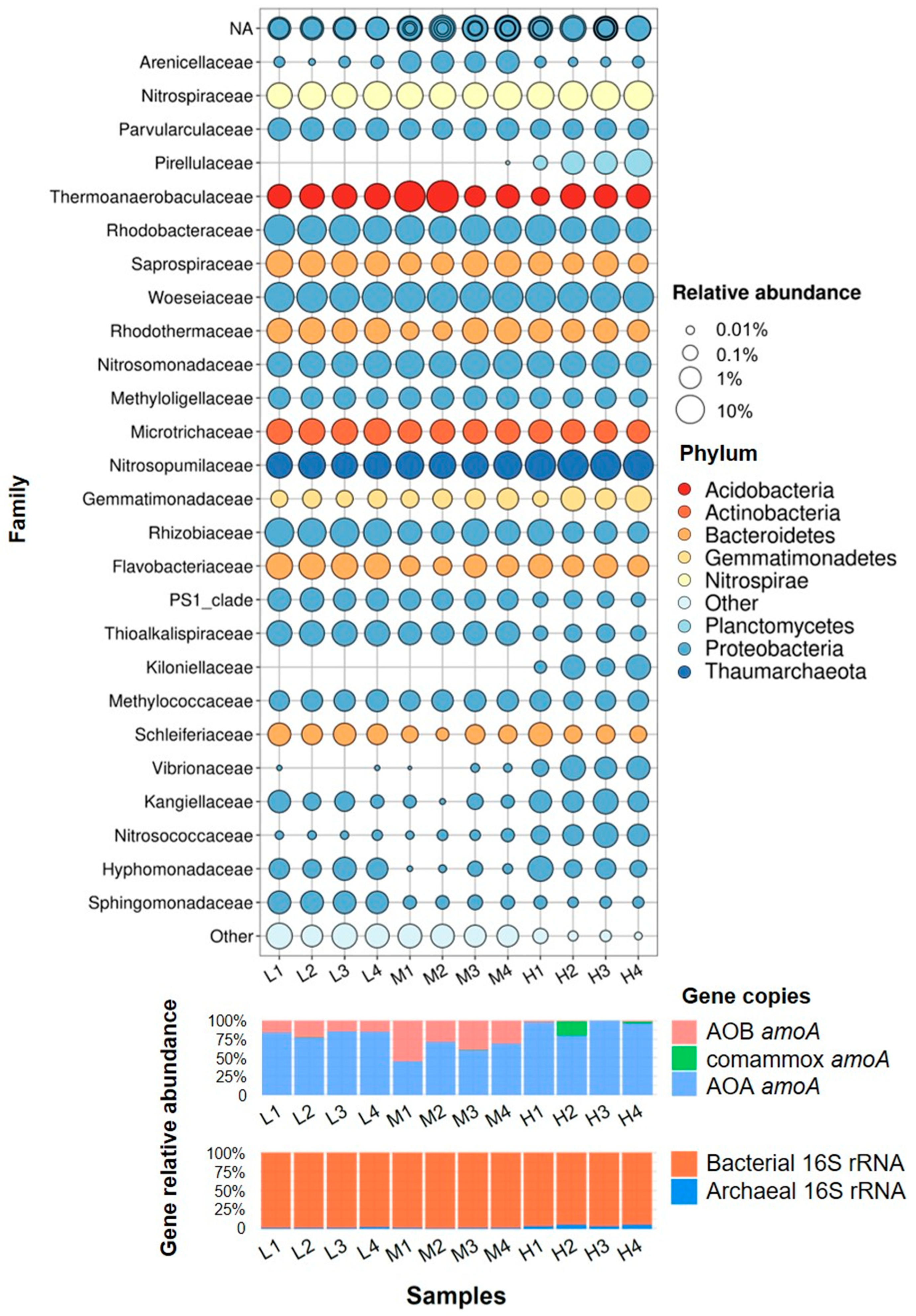
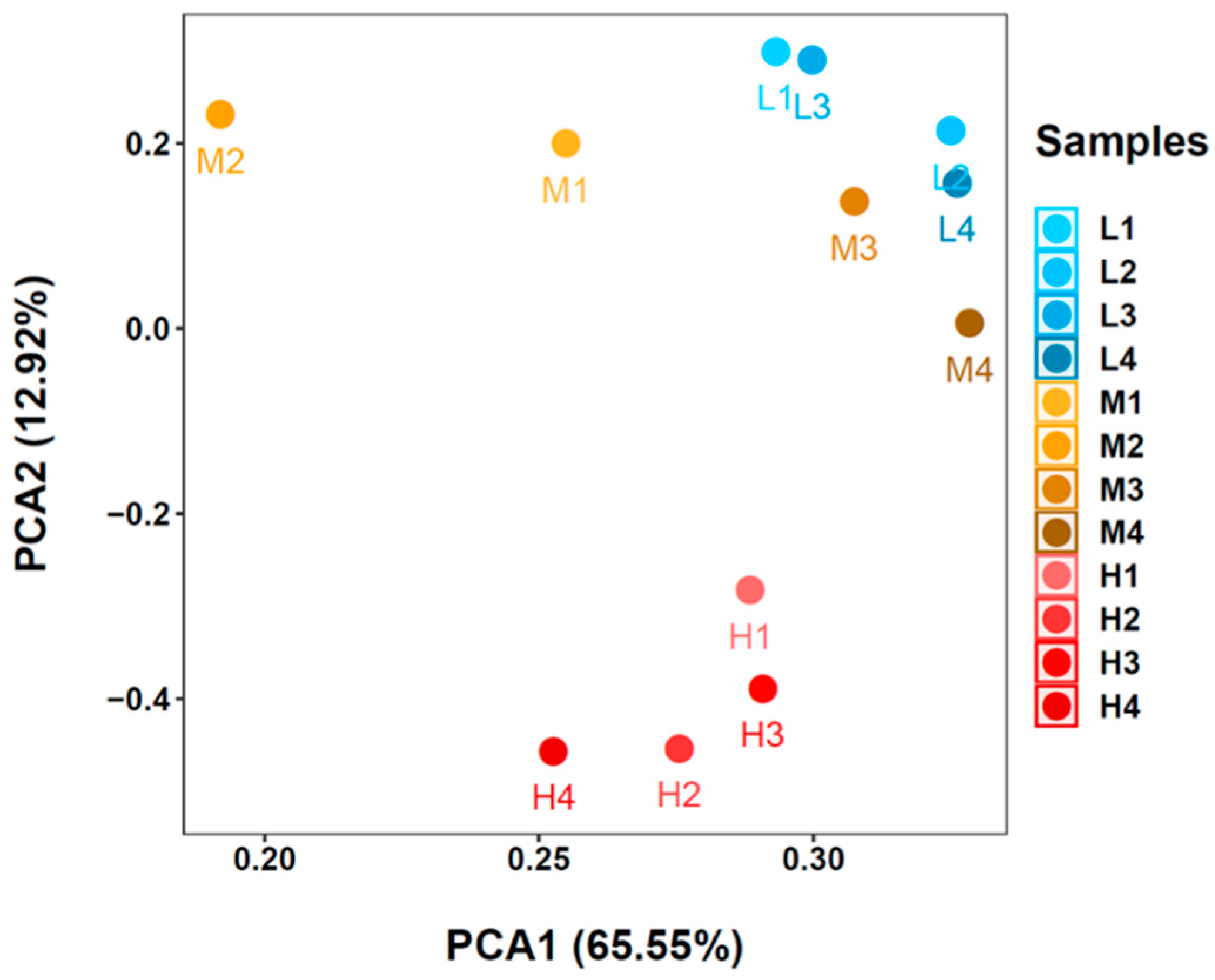
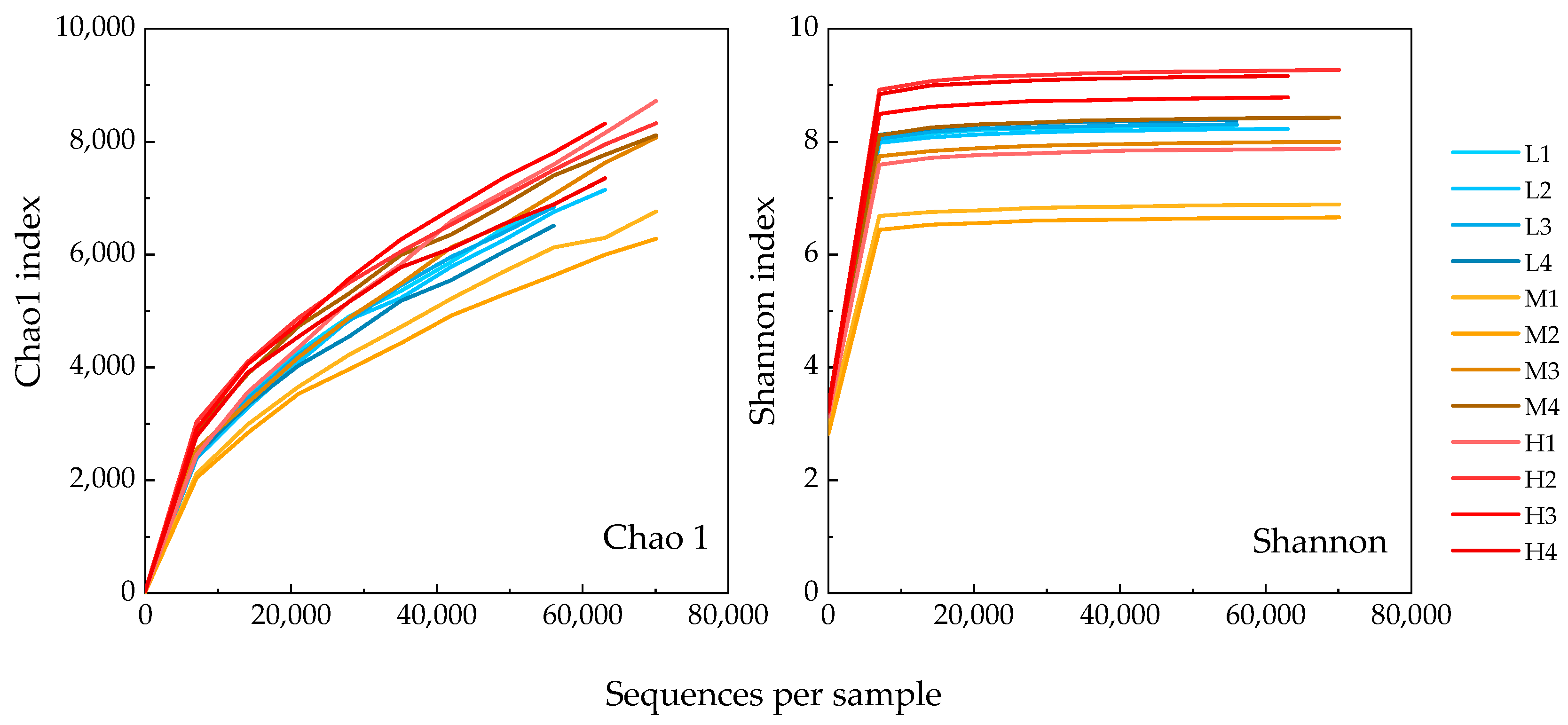
3.3. Microbial Communities Based on 16S rRNA V4 Region Amplicon Sequencing
4. Discussion
4.1. Microbial Community in Marine Aquarium Gravel Biofilm
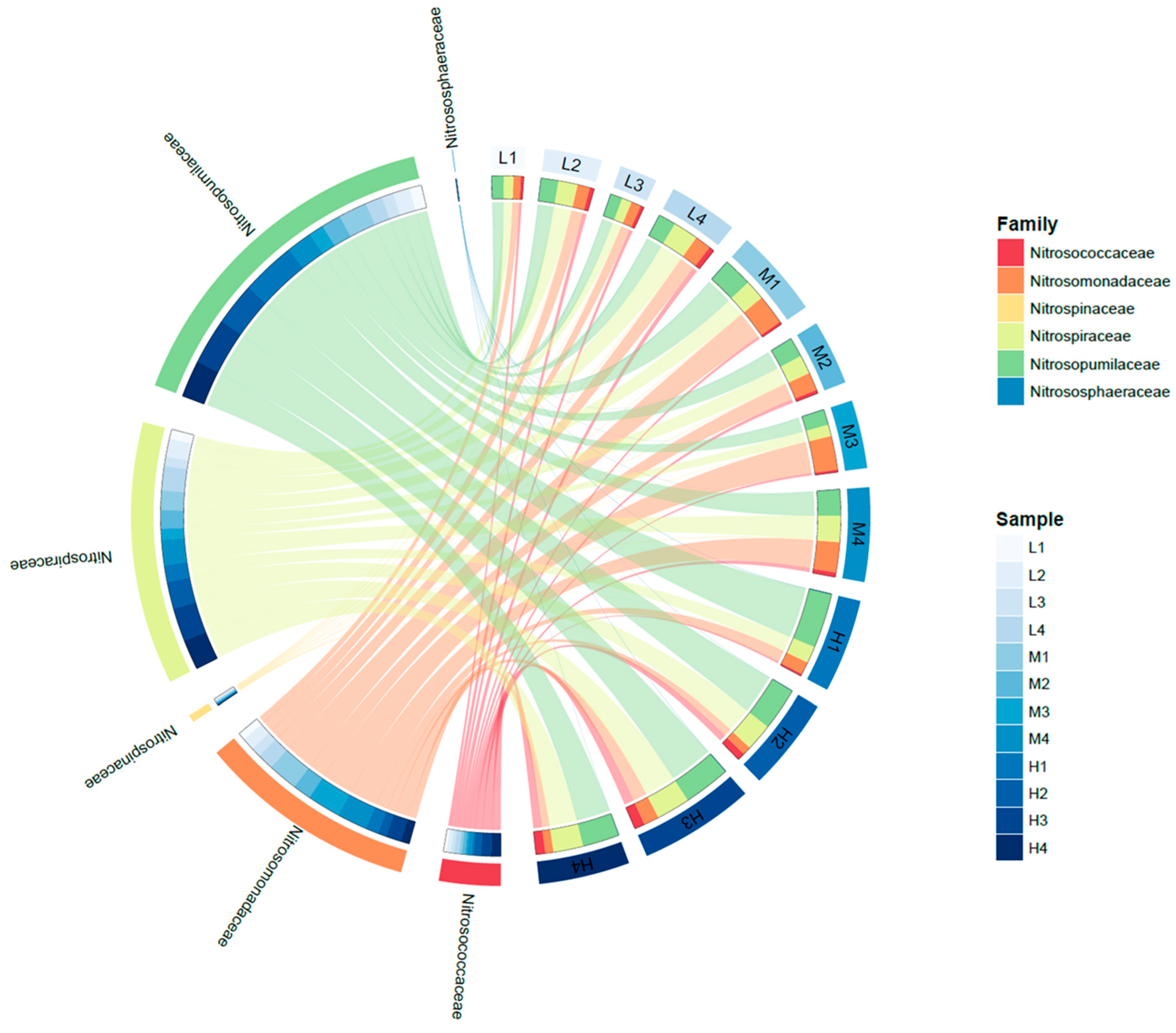
4.2. Nitrifying Microorganisms
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Dalsgaard, J.; Lund, I.; Thorarinsdottir, R.; Drengstig, A.; Arvonen, K.; Pedersen, P.B. Farming Different Species in RAS in Nordic Countries: Current Status and Future Perspectives. Aquac. Eng. 2013, 53, 2–13. [Google Scholar] [CrossRef]
- Summerfelt, S.T.; Davidson, J.; May, T.; Good, C.; Vinci, B. Emerging Trends in Salmonid RAS—Part II. System Enhancements. Glob. Aquac. Advocate 2015, 18, 64–65. [Google Scholar]
- De Guzman, M.L. New Land-Based Salmon Producer Bares Plans to Build Global RAS Facilities; Hatchery International: Simcoe, ON, Canada, 2018. [Google Scholar]
- Verdegem, M.C.J.; Bosma, R.H.; Verreth, J.A.J. Reducing Water Use for Animal Production through Aquaculture. Water Resour. Dev. 2006, 22, 101–113. [Google Scholar] [CrossRef]
- Badiola, M.; Mendiola, D.; Bostock, J. Recirculating Aquaculture Systems (RAS) Analysis: Main Issues on Management and Future Challenges. Aquac. Eng. 2012, 51, 26–35. [Google Scholar] [CrossRef]
- Piedrahita, R.H. Reducing the Potential Environmental Impact of Tank Aquaculture Effluents through Intensification and Recirculation. Aquaculture 2003, 226, 35–44. [Google Scholar] [CrossRef]
- Ruiz, P.; Vidal, J.M.; Sepúlveda, D.; Torres, C.; Villouta, G.; Carrasco, C.; Aguilera, F.; Ruiz-Tagle, N.; Urrutia, H. Overview and Future Perspectives of Nitrifying Bacteria on Biofilters for Recirculating Aquaculture Systems. Rev. Aquac. 2020, 12, 1478–1494. [Google Scholar] [CrossRef]
- Schreier, H.J.; Mirzoyan, N.; Saito, K. Microbial Diversity of Biological Filters in Recirculating Aquaculture Systems. Curr. Opin. Biotechnol. 2010, 21, 318–325. [Google Scholar] [CrossRef]
- Bock, E.; Wagner, M. Oxidation of Inorganic Nitrogen Compounds as an Energy Source. Prokaryotes 2006, 2, 457–495. [Google Scholar]
- Könneke, M.; Bernhard, A.E.; de La Torre, J.R.; Walker, C.B.; Waterbury, J.B.; Stahl, D.A. Isolation of an Autotrophic Ammonia-Oxidizing Marine Archaeon. Nature 2005, 437, 543–546. [Google Scholar] [CrossRef]
- Abeliovich, A. The Nitrite-Oxidizing Bacteria. Prokaryotes 2006, 5, 861–872. [Google Scholar]
- Daims, H.; Lebedeva, E.V.; Pjevac, P.; Han, P.; Herbold, C.; Albertsen, M.; Jehmlich, N.; Palatinszky, M.; Vierheilig, J.; Bulaev, A.; et al. Complete Nitrification by Nitrospira Bacteria. Nature 2015, 528, 504–509. [Google Scholar] [CrossRef]
- Van Kessel, M.A.; Speth, D.R.; Albertsen, M.; Nielsen, P.H.; Op den Camp, H.J.; Kartal, B.; Jetten, M.S.; Lücker, S. Complete Nitrification by a Single Microorganism. Nature 2015, 528, 555–559. [Google Scholar] [CrossRef]
- Huang, Z.; Wan, R.; Song, X.; Liu, Y.; Hallerman, E.; Dong, D.; Zhai, J.; Zhang, H.; Sun, L. Metagenomic Analysis Shows Diverse, Distinct Bacterial Communities in Biofilters among Different Marine Recirculating Aquaculture Systems. Aquacult. Int. 2016, 24, 1393–1408. [Google Scholar] [CrossRef]
- Lee, D.-E.; Lee, J.; Kim, Y.-M.; Myeong, J.-I.; Kim, K.-H. Uncultured Bacterial Diversity in a Seawater Recirculating Aquaculture System Revealed by 16S RRNA Gene Amplicon Sequencing. J. Microbiol. 2016, 54, 296–304. [Google Scholar] [CrossRef]
- Bartelme, R.P.; McLellan, S.L.; Newton, R.J. Freshwater Recirculating Aquaculture System Operations Drive Biofilter Bacterial Community Shifts around a Stable Nitrifying Consortium of Ammonia-Oxidizing Archaea and Comammox Nitrospira. Front. Microbiol. 2017, 8, 101. [Google Scholar] [CrossRef]
- Hüpeden, J.; Wemheuer, B.; Indenbirken, D.; Schulz, C.; Spieck, E. Taxonomic and Functional Profiling of Nitrifying Biofilms in Freshwater, Brackish and Marine RAS Biofilters. Aquac. Eng. 2020, 90, 102094. [Google Scholar] [CrossRef]
- Fowler, S.J.; Palomo, A.; Dechesne, A.; Mines, P.D.; Smets, B.F. Comammox Nitrospira Are Abundant Ammonia Oxidizers in Diverse Groundwater-Fed Rapid Sand Filter Communities: Comammox Nitrospira in Drinking Water Biofilters. Environ. Microbiol. 2018, 20, 1002–1015. [Google Scholar] [CrossRef]
- Wuchter, C.; Abbas, B.; Coolen, M.J.L.; Herfort, L.; van Bleijswijk, J.; Timmers, P.; Strous, M.; Teira, E.; Herndl, G.J.; Middelburg, J.J.; et al. Archaeal Nitrification in the Ocean. Proc. Natl. Acad. Sci. USA 2006, 103, 12317–12322. [Google Scholar] [CrossRef]
- Rotthauwe, J.H.; Witzel, K.P.; Liesack, W. The Ammonia Monooxygenase Structural Gene AmoA as a Functional Marker: Molecular Fine-Scale Analysis of Natural Ammonia-Oxidizing Populations. Appl. Environ. Microbiol. 1997, 63, 4704–4712. [Google Scholar] [CrossRef]
- Hornek, R.; Pommerening-Röser, A.; Koops, H.-P.; Farnleitner, A.H.; Kreuzinger, N.; Kirschner, A.; Mach, R.L. Primers Containing Universal Bases Reduce Multiple AmoA Gene Specific DGGE Band Patterns When Analysing the Diversity of Beta-Ammonia Oxidizers in the Environment. J. Microbiol. Methods 2006, 66, 147–155. [Google Scholar] [CrossRef]
- Suzuki, M.T.; Taylor, L.T.; DeLong, E.F. Quantitative Analysis of Small-Subunit RRNA Genes in Mixed Microbial Populations via 5′-Nuclease Assays. Appl. Environ. Microbiol. 2000, 66, 4605–4614. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Lauber, C.L.; Walters, W.A.; Berg-Lyons, D.; Huntley, J.; Fierer, N.; Owens, S.M.; Betley, J.; Fraser, L.; Bauer, M. Ultra-High-Throughput Microbial Community Analysis on the Illumina HiSeq and MiSeq Platforms. ISME J. 2012, 6, 1621–1624. [Google Scholar] [CrossRef] [PubMed]
- Ruan, Y.-J.; Guo, X.-S.; Ye, Z.-Y.; Liu, Y.; Zhu, S.-M. Bacterial Community Analysis of Different Sections of a Biofilter in a Full-Scale Marine Recirculating Aquaculture System. N. Am. J. Aquac. 2015, 77, 318–326. [Google Scholar] [CrossRef]
- Gonzalez-Silva, B.M.; Jonassen, K.R.; Bakke, I.; Østgaard, K.; Vadstein, O. Nitrification at Different Salinities: Biofilm Community Composition and Physiological Plasticity. Water Res. 2016, 95, 48–58. [Google Scholar] [CrossRef]
- Smith, K.F.; Schmidt, V.; Rosen, G.E.; Amaral-Zettler, L. Microbial Diversity and Potential Pathogens in Ornamental Fish Aquarium Water. PLoS ONE 2012, 7, e39971. [Google Scholar] [CrossRef]
- Michaud, L.; Lo Giudice, A.; Troussellier, M.; Smedile, F.; Bruni, V.; Blancheton, J.-P. Phylogenetic Characterization of the Heterotrophic Bacterial Communities Inhabiting a Marine Recirculating Aquaculture System. J. Appl. Microbiol. 2009, 107, 1935–1946. [Google Scholar] [CrossRef]
- Blancheton, J.P.; Attramadal, K.J.K.; Michaud, L.; d’Orbcastel, E.R.; Vadstein, O. Insight into Bacterial Population in Aquaculture Systems and Its Implication. Aquac. Eng. 2013, 53, 30–39. [Google Scholar] [CrossRef]
- Marchant, H.K.; Tegetmeyer, H.E.; Ahmerkamp, S.; Holtappels, M.; Lavik, G.; Graf, J.; Schreiber, F.; Mussmann, M.; Strous, M.; Kuypers, M.M. Metabolic Specialization of Denitrifiers in Permeable Sediments Controls N2O Emissions. Environ. Microbiol. 2018, 20, 4486–4502. [Google Scholar] [CrossRef]
- Urakawa, H.; Tajima, Y.; Numata, Y.; Tsuneda, S. Low Temperature Decreases the Phylogenetic Diversity of Ammonia-Oxidizing Archaea and Bacteria in Aquarium Biofiltration Systems. Appl. Environ. Microbiol. 2008, 74, 894–900. [Google Scholar] [CrossRef]
- Sugita, H.; Nakamura, H.; Shimada, T. Microbial Communities Associated with Filter Materials in Recirculating Aquaculture Systems of Freshwater Fish. Aquaculture 2005, 243, 403–409. [Google Scholar] [CrossRef]
- Francis, C.A.; Roberts, K.J.; Beman, J.M.; Santoro, A.E.; Oakley, B.B. Ubiquity and Diversity of Ammonia-Oxidizing Archaea in Water Columns and Sediments of the Ocean. Proc. Natl. Acad. Sci. USA 2005, 102, 14683–14688. [Google Scholar] [CrossRef]
- Agogué, H.; Brink, M.; Dinasquet, J.; Herndl, G.J. Major Gradients in Putatively Nitrifying and Non-Nitrifying Archaea in the Deep North Atlantic. Nature 2008, 456, 788–791. [Google Scholar] [CrossRef]
- Sauder, L.A.; Engel, K.; Stearns, J.C.; Masella, A.P.; Pawliszyn, R.; Neufeld, J.D. Aquarium Nitrification Revisited: Thaumarchaeota Are the Dominant Ammonia Oxidizers in Freshwater Aquarium Biofilters. PLoS ONE 2011, 6, e23281. [Google Scholar] [CrossRef] [Green Version]
- Sakami, T.; Andoh, T.; Morita, T.; Yamamoto, Y. Phylogenetic Diversity of Ammonia-Oxidizing Archaea and Bacteria in Biofilters of Recirculating Aquaculture Systems. Mar. Genom. 2012, 7, 27–31. [Google Scholar] [CrossRef]
- Brown, M.N.; Briones, A.; Diana, J.; Raskin, L. Ammonia-Oxidizing Archaea and Nitrite-Oxidizing Nitrospiras in the Biofilter of a Shrimp Recirculating Aquaculture System. FEMS Microbiol. Ecol. 2013, 83, 17–25. [Google Scholar] [CrossRef]
- Martens-Habbena, W.; Qin, W.; Horak, R.E.; Urakawa, H.; Schauer, A.J.; Moffett, J.W.; Armbrust, E.V.; Ingalls, A.E.; Devol, A.H.; Stahl, D.A. The Production of Nitric Oxide by Marine Ammonia-oxidizing Archaea and Inhibition of Archaeal Ammonia Oxidation by a Nitric Oxide Scavenger. Environ. Microbiol. 2015, 17, 2261–2274. [Google Scholar] [CrossRef]
- Hatzenpichler, R. Diversity, Physiology, and Niche Differentiation of Ammonia-Oxidizing Archaea. Appl. Environ. Microbiol. 2012, 78, 7501–7510. [Google Scholar] [CrossRef]
- Roalkvam, I.; Drønen, K.; Dahle, H.; Wergeland, H.I. Comparison of Active Biofilm Carriers and Commercially Available Inoculum for Activation of Biofilters in Marine Recirculating Aquaculture Systems (RAS). Aquaculture 2020, 514, 734480. [Google Scholar] [CrossRef]
- Liu, S.; Hu, J.-J.; Shen, J.-X.; Chen, S.; Tian, G.-M.; Zheng, P.; Lou, L.-P.; Ma, F.; Hu, B.-L. Potencial Correlated Environmental Factors Leading to the Niche Segregation of Ammonia-Oxidizing Archaea and Ammonia-Oxidizing Bacteria: A Review. Appl. Environ. Biotechnol. 2017, 2, 11–19. [Google Scholar] [CrossRef]
- Qin, W.; Amin, S.A.; Martens-Habbena, W.; Walker, C.B.; Urakawa, H.; Devol, A.H.; Ingalls, A.E.; Moffett, J.W.; Armbrust, E.V.; Stahl, D.A. Marine Ammonia-Oxidizing Archaeal Isolates Display Obligate Mixotrophy and Wide Ecotypic Variation. Proc. Natl. Acad. Sci. USA 2014, 111, 12504–12509. [Google Scholar] [CrossRef]
- Kim, J.-G.; Jung, M.-Y.; Park, S.-J.; Rijpstra, W.I.C.; Sinninghe Damsté, J.S.; Madsen, E.L.; Min, D.; Kim, J.-S.; Kim, G.-J.; Rhee, S.-K. Cultivation of a Highly Enriched Ammonia-oxidizing Archaeon of Thaumarchaeotal Group I. 1b from an Agricultural Soil. Environ. Microbiol. 2012, 14, 1528–1543. [Google Scholar] [CrossRef] [PubMed]
- De la Torre, J.R.; Walker, C.B.; Ingalls, A.E.; Könneke, M.; Stahl, D.A. Cultivation of a Thermophilic Ammonia Oxidizing Archaeon Synthesizing Crenarchaeol. Environ. Microbiol. 2008, 10, 810–818. [Google Scholar] [CrossRef]
- Lin, Z.; Huang, W.; Zhou, J.; He, X.; Wang, J.; Wang, X.; Zhou, J. The Variation on Nitrogen Removal Mechanisms and the Succession of Ammonia Oxidizing Archaea and Ammonia Oxidizing Bacteria with Temperature in Biofilm Reactors Treating Saline Wastewater. Bioresour. Technol. 2020, 314, 123760. [Google Scholar] [CrossRef]
- Taylor, A.E.; Giguere, A.T.; Zoebelein, C.M.; Myrold, D.D.; Bottomley, P.J. Modeling of Soil Nitrification Responses to Temperature Reveals Thermodynamic Differences between Ammonia-Oxidizing Activity of Archaea and Bacteria. ISME J. 2017, 11, 896–908. [Google Scholar] [CrossRef] [PubMed]
- Daims, H.; Lücker, S.; Wagner, M. A New Perspective on Microbes Formerly Known as Nitrite-Oxidizing Bacteria. Trends Microbiol. 2016, 24, 699–712. [Google Scholar] [CrossRef] [PubMed]
- Füssel, J.; Lam, P.; Lavik, G.; Jensen, M.M.; Holtappels, M.; Günter, M.; Kuypers, M.M. Nitrite Oxidation in the Namibian Oxygen Minimum Zone. ISME J. 2012, 6, 1200–1209. [Google Scholar] [CrossRef]
- Jorgensen, S.L.; Hannisdal, B.; Lanzén, A.; Baumberger, T.; Flesland, K.; Fonseca, R.; Øvreås, L.; Steen, I.H.; Thorseth, I.H.; Pedersen, R.B. Correlating Microbial Community Profiles with Geochemical Data in Highly Stratified Sediments from the Arctic Mid-Ocean Ridge. Proc. Natl. Acad. Sci. USA 2012, 109, E2846–E2855. [Google Scholar] [CrossRef] [PubMed]
- Beman, J.M.; Leilei Shih, J.; Popp, B.N. Nitrite Oxidation in the Upper Water Column and Oxygen Minimum Zone of the Eastern Tropical North Pacific Ocean. ISME J. 2013, 7, 2192–2205. [Google Scholar] [CrossRef]
- Nunoura, T.; Takaki, Y.; Hirai, M.; Shimamura, S.; Makabe, A.; Koide, O.; Kikuchi, T.; Miyazaki, J.; Koba, K.; Yoshida, N. Hadal Biosphere: Insight into the Microbial Ecosystem in the Deepest Ocean on Earth. Proc. Natl. Acad. Sci. USA 2015, 112, E1230–E1236. [Google Scholar] [CrossRef]
- Gruber-Dorninger, C.; Pester, M.; Kitzinger, K.; Savio, D.F.; Loy, A.; Rattei, T.; Wagner, M.; Daims, H. Functionally Relevant Diversity of Closely Related Nitrospira in Activated Sludge. ISME J. 2015, 9, 643–655. [Google Scholar] [CrossRef]
- Cao, Y.; van Loosdrecht, M.C.M.; Daigger, G.T. Mainstream Partial Nitritation–Anammox in Municipal Wastewater Treatment: Status, Bottlenecks, and Further Studies. Appl. Microbiol. Biotechnol. 2017, 101, 1365–1383. [Google Scholar] [CrossRef]
- Palomo, A.; Jane Fowler, S.; Gülay, A.; Rasmussen, S.; Sicheritz-Ponten, T.; Smets, B.F. Metagenomic Analysis of Rapid Gravity Sand Filter Microbial Communities Suggests Novel Physiology of Nitrospira Spp. ISME J. 2016, 10, 2569–2581. [Google Scholar] [CrossRef]
- Liu, S.; Wang, H.; Chen, L.; Wang, J.; Zheng, M.; Liu, S.; Chen, Q.; Ni, J. Comammox Nitrospira within the Yangtze River Continuum: Community, Biogeography, and Ecological Drivers. ISME J. 2020, 14, 2488–2504. [Google Scholar] [CrossRef]
- Sun, D.; Zhao, M.; Tang, X.; Liu, M.; Hou, L.; Zhao, Q.; Li, J.; Gu, J.-D.; Han, P. Niche Adaptation Strategies of Different Clades of Comammox Nitrospira in the Yangtze Estuary. Int. Biodeterior. Biodegrad. 2021, 164, 105286. [Google Scholar] [CrossRef]
- Sun, D.; Tang, X.; Zhao, M.; Zhang, Z.; Hou, L.; Liu, M.; Wang, B.; Klümper, U.; Han, P. Distribution and Diversity of Comammox Nitrospira in Coastal Wetlands of China. Front. Microbiol. 2020, 11, 589268. [Google Scholar] [CrossRef]
- Pjevac, P.; Schauberger, C.; Poghosyan, L.; Herbold, C.W.; van Kessel, M.A.H.J.; Daebeler, A.; Steinberger, M.; Jetten, M.S.M.; Lücker, S.; Wagner, M.; et al. AmoA-Targeted Polymerase Chain Reaction Primers for the Specific Detection and Quantification of Comammox Nitrospira in the Environment. Front. Microbiol. 2017, 8, 1508. [Google Scholar] [CrossRef]
- Osburn, E.D.; Barrett, J.E. Abundance and Functional Importance of Complete Ammonia-Oxidizing Bacteria (Comammox) versus Canonical Nitrifiers in Temperate Forest Soils. Soil Biol. Biochem. 2020, 145, 107801. [Google Scholar] [CrossRef]
- Roots, P.; Wang, Y.; Rosenthal, A.F.; Griffin, J.S.; Sabba, F.; Petrovich, M.; Yang, F.; Kozak, J.A.; Zhang, H.; Wells, G.F. Comammox Nitrospira Are the Dominant Ammonia Oxidizers in a Mainstream Low Dissolved Oxygen Nitrification Reactor. Water Res. 2019, 157, 396–405. [Google Scholar] [CrossRef]
- Cotto, I.; Dai, Z.; Huo, L.; Anderson, C.L.; Vilardi, K.J.; Ijaz, U.; Khunjar, W.; Wilson, C.; De Clippeleir, H.; Gilmore, K.; et al. Long Solids Retention Times and Attached Growth Phase Favor Prevalence of Comammox Bacteria in Nitrogen Removal Systems. Water Res. 2020, 169, 115268. [Google Scholar] [CrossRef]
- Kits, K.D.; Sedlacek, C.J.; Lebedeva, E.V.; Han, P.; Bulaev, A.; Pjevac, P.; Daebeler, A.; Romano, S.; Albertsen, M.; Stein, L.Y.; et al. Kinetic Analysis of a Complete Nitrifier Reveals an Oligotrophic Lifestyle. Nature 2017, 549, 269–272. [Google Scholar] [CrossRef]
- Koch, H.; van Kessel, M.A.H.J.; Lücker, S. Complete Nitrification: Insights into the Ecophysiology of Comammox Nitrospira. App.l Microbiol. Biotechnol. 2019, 103, 177–189. [Google Scholar] [CrossRef] [PubMed]
- Daims, H.; Nielsen, J.L.; Nielsen, P.H.; Schleifer, K.-H.; Wagner, M. In Situ Characterization of Nitrospira -Like Nitrite-Oxidizing Bacteria Active in Wastewater Treatment Plants. Appl. Environ. Microbiol. 2001, 67, 5273–5284. [Google Scholar] [CrossRef] [PubMed]
- Juretschko, S.; Timmermann, G.; Schmid, M.; Schleifer, K.-H.; Pommerening-Röser, A.; Koops, H.-P.; Wagner, M. Combined Molecular and Conventional Analyses of Nitrifying Bacterium Diversity in Activated Sludge: Nitrosococcus Mobilis and Nitrospira-Like Bacteria as Dominant Populations. Appl. Environ. Microbiol. 1998, 64, 3042–3051. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bartosch, S.; Wolgast, I.; Spieck, E.; Bock, E. Identification of Nitrite-Oxidizing Bacteria with Monoclonal Antibodies Recognizing the Nitrite Oxidoreductase. Appl. Environ. Microbiol. 1999, 65, 4126–4133. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Zhang, C.; Wei, Q.; Zhang, S.; Quan, Z.; Li, M. Temperature and Salinity Drive Comammox Community Composition in Mangrove Ecosystems across Southeastern China. Sci. Total Environ. 2020, 742, 140456. [Google Scholar] [CrossRef] [PubMed]
| Target Gene | Primer/Probe | Sequence | Ref. |
|---|---|---|---|
| Thaumarchaeal amoA | amoA-for | STAATGGTCTGGCTTAGACG | [19] |
| amoA-rev | GCGGCCATCCATCTGTATGT | ||
| Beta-proteobacterial amoA | amoA-1F | GGGGTTTCTACTGGTGGT | [20] |
| amoA-rNew | CCCCTCBGSAAAVCCTTCTTC | [21] | |
| Comammox Nitrospira amoA | Ntsp-amoA162F | GGATTTCTGGNTSGATTGGA | [18] |
| Ntsp-amoA359R | WAGTTNGACCACCASTACCA | ||
| Archaeal 16S rRNA | ARCH1-1369F | CGGTGAATACGTCCCTGC | [22] |
| ARCH2-1369F | CGGTGAATATGCCCCTGC | ||
| PROK 1541R | AAGGAGGTGATCCRGCCGCA | ||
| Tm1389F | CTTGTACACACCGCCCGTC | ||
| Bacterial 16S rRNA | BACT1369F | CGGTGAATACGTTCYCGG | [22] |
| PROK1492R | GGWTACCTTGTTACGACTT | ||
| Tm1389F | CTTGTACACACCGCCCGTC |
| Sample No. | Temp (°C) | pH | DO (mg/L) | NH4+-N (μg/L) | NO3− -N (μg/L) | PO43-P (μg/L) |
|---|---|---|---|---|---|---|
| LWF | 10.0 | 7.6 | 8.90 | 2.54 (±0.68) | 41.71 (±3.37) | 1.07 (±0.03) |
| LWN | 10.1 | 7.6 | 9.03 | 2.35 (±1.11) | 32.43 (±3.79) | 1.28 (±0.13) |
| LWB | 10.1 | 7.6 | 8.87 | 5.07 (±1.33) | 30.29 (±2.67) | 1.21 (±0.19) |
| MWF | 19.0 | 7.5 | 7.90 | 1.86 (±4.07) | 138.50 (±15.35) | 3.55 (±0.06) |
| MWN | 18.4 | 7.7 | 8.04 | 6.43 (±0.09) | 113.86 (±6.93) | 3.84 (±0.03) |
| MWB | 18.4 | 7.7 | 7.70 | 1.96 (±0.32) | 125.29 (±2.77) | 3.77 (±0.12) |
| HWB | 26.8 | 7.4 | 5.28 | 3.32 (±0.05) | 98.86 (±5.15) | 1.35 (±0.17) |
| HWN | 24.7 | 7.5 | 6.58 | 2.64 (±0.17) | 81.71 (±8.37) | 1.14 (±0.09) |
| HWB | 24.7 | 7.5 | 6.15 | 3.51 (±0.80) | 88.86 (±7.16) | 1.28 (±0.03) |
| Sample No. | AOA | AOB | Comammox Nitrospira |
|---|---|---|---|
| LWF | 1.76 × 104 (±659.14) | 1.50 × 102 (±23.54) | 1.6 × 101 (±2.40) |
| LWN | 5.04 × 103 (±49.83) | 1.17 × 102 (±19.01) | 3.81 × 101 (±7.91) |
| LWB | 2.40 × 104 (±678.11) | 1.37 × 102 (±3.33) | 2.23 × 101 (±0.26) |
| MWF | 2.13 × 104 (±747.40) | 5.94 × 102 (±86.85) | 3.87 × 100 (±1.01) |
| MWN | 1.84 × 104 (±260.69) | 5.52 × 102 (±23.31) | 8.7 × 100 (±0.44) |
| MWB | 2.03 × 104 (±253.31) | 6.57 × 102 (±12.38) | 7.54 × 100 (±0.32) |
| HWF | 5.2 × 104 (±1177.35) | 1.82 × 102 (±19.52) | 1.18 × 101 (±3.7) |
| HWN | 4.17 × 104 (±1122.59) | 1.55 × 102 (±10.35) | 9.91 × 100 (±1.08) |
| HWB | 3.36 × 104 (±686.65) | 8.13 × 101 (±11.69) | 5.33 × 100 (±1.56) |
| Raw Reads | Clean Reads | AOA Reads | AOB Reads | NOB Reads | OTU No. | AOA OTU No. | AOB OTU No. | NOB OTU No. | |
|---|---|---|---|---|---|---|---|---|---|
| L1 | 69,800 | 59,133 | 1443 | 1220 | 1177 | 3543 | 36 | 32 | 17 |
| L2 | 77,781 | 65,949 | 2328 | 2275 | 2645 | 3732 | 41 | 49 | 28 |
| L3 | 68,202 | 57,731 | 1401 | 1569 | 1193 | 3513 | 34 | 36 | 24 |
| L4 | 69,915 | 59,308 | 2089 | 2631 | 3098 | 3565 | 50 | 40 | 28 |
| M1 | 87,868 | 74,951 | 4433 | 5026 | 3135 | 3450 | 60 | 64 | 25 |
| M2 | 82,719 | 70,304 | 2813 | 3114 | 2756 | 3281 | 38 | 47 | 24 |
| M3 | 85,189 | 72,402 | 2283 | 5402 | 1733 | 4011 | 34 | 60 | 23 |
| M4 | 101,204 | 85,612 | 4569 | 6095 | 4666 | 4961 | 69 | 72 | 33 |
| H1 | 96,814 | 77,850 | 8808 | 2368 | 2689 | 4265 | 87 | 37 | 49 |
| H2 | 90,945 | 76,243 | 6676 | 2503 | 4300 | 4937 | 74 | 63 | 47 |
| H3 | 77,410 | 64,298 | 6222 | 3390 | 4492 | 4412 | 79 | 70 | 59 |
| H4 | 83,213 | 69,846 | 5273 | 2521 | 4313 | 4470 | 63 | 60 | 43 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Q.; Hasezawa, R.; Saito, R.; Okano, K.; Shimizu, K.; Utsumi, M. Abundance and Diversity of Nitrifying Microorganisms in Marine Recirculating Aquaculture Systems. Water 2022, 14, 2744. https://doi.org/10.3390/w14172744
Li Q, Hasezawa R, Saito R, Okano K, Shimizu K, Utsumi M. Abundance and Diversity of Nitrifying Microorganisms in Marine Recirculating Aquaculture Systems. Water. 2022; 14(17):2744. https://doi.org/10.3390/w14172744
Chicago/Turabian StyleLi, Qintong, Ryo Hasezawa, Riho Saito, Kunihiro Okano, Kazuya Shimizu, and Motoo Utsumi. 2022. "Abundance and Diversity of Nitrifying Microorganisms in Marine Recirculating Aquaculture Systems" Water 14, no. 17: 2744. https://doi.org/10.3390/w14172744






