Bioaugmentation-Enhanced Remediation of Crude Oil Polluted Water in Pilot-Scale Floating Treatment Wetlands
Abstract
:1. Introduction
2. Materials and Methods
2.1. Establishment and Operation of Pilot-Scale FTWs
- (1)
- oil-contaminated water without vegetation and bacterial inoculation (1st control, C1);
- (2)
- oil-contaminated water with bacterial inoculation only (Treatment 1, T1);
- (3)
- oil-contaminated water with vegetation only, (Treatment 2, T2);
- (4)
- oil-contaminated water with vegetation and bacterial inoculation (Treatment 3, T3);
- (5)
- Vegetation in tap water (2nd control, C2).
2.2. Chemical Analyses and Determination of Plant Growth
2.3. Bioaugmentation with a Four-Membered Bacterial Consortium
2.4. Abundance and Expression of the alkB Gene in the FTWs
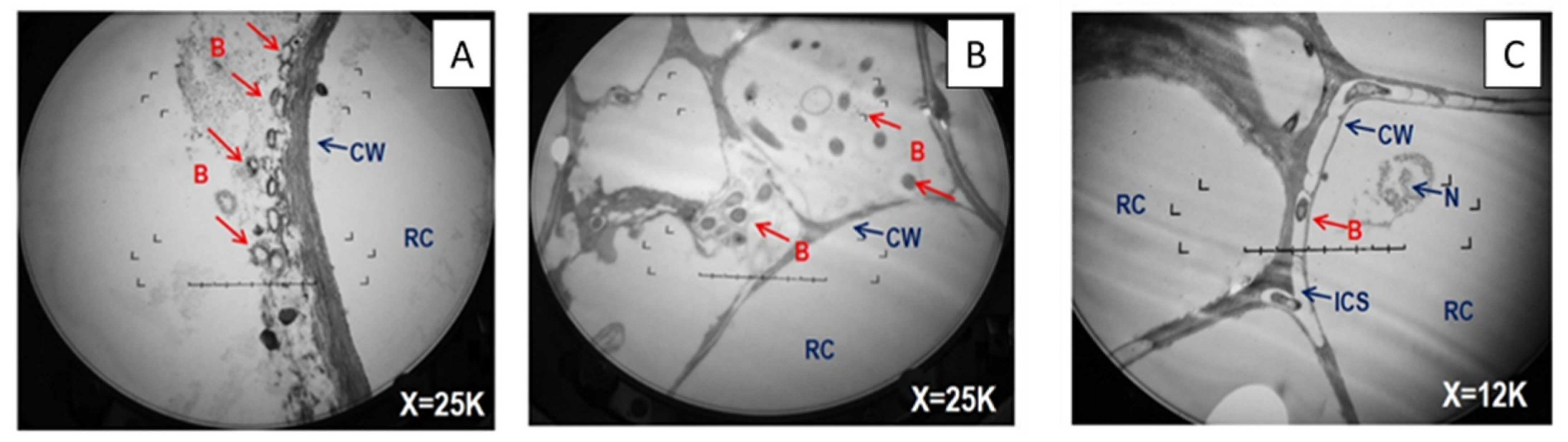
2.5. Transmission Electron Microscopy
2.6. Toxicity Bioassay
2.7. Statistical Analysis
3. Results
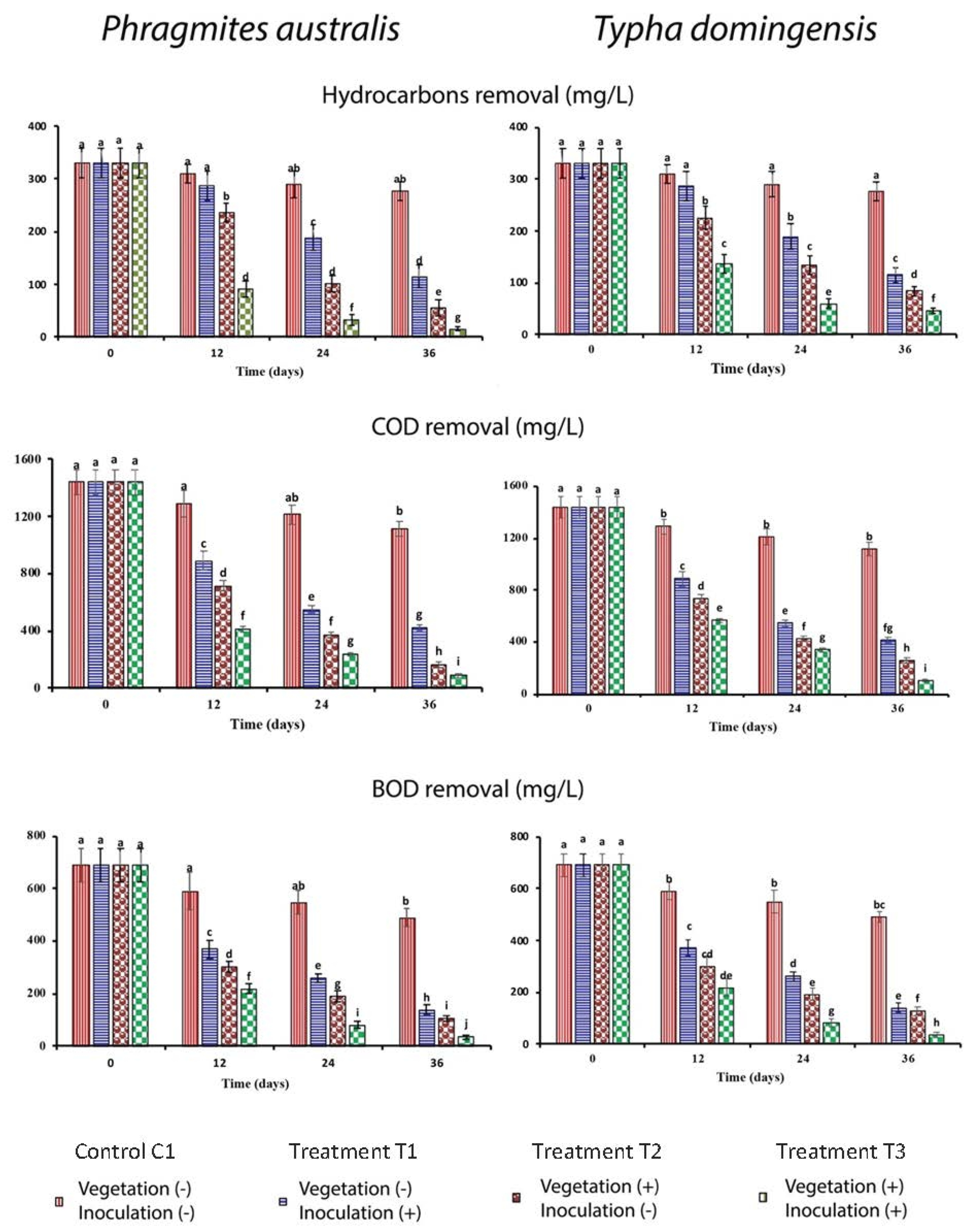
3.1. Attenuation of Hydrocarbon Concentration, COD and BOD in Pilot-Scale FTW
3.2. Plant Growth in FTWs
3.3. Abundance of Inoculated Bacteria, and of the alkB Gene and Transcripts
3.4. Fish Toxicity
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Droege, P. Urban Energy Transition: From Fossil Fuels to Renewable Power; Elsevier: Amsterdam, The Netherlands, 2011. [Google Scholar]
- Strubinger, A.; Ehrmann, U.; Leon, V.; DeSisto, A.; Gonzalez, M. Changes in Venezuelan Orinoco belt crude after different biotechnological approaches. J. Pet. Sci. Eng. 2015, 127, 421–432. [Google Scholar] [CrossRef]
- Varjani, S.; Joshi, R.; Srivastava, V.K.; Ngo, H.H.; Guo, W. Treatment of wastewater from petroleum industry: Current practices and perspectives. Environ. Sci. Pollut. Res. 2020, 27, 27172–27180. [Google Scholar] [CrossRef]
- Rehman, K.; Imran, A.; Amin, I.; Afzal, M. Enhancement of oil field-produced wastewater remediation by bacterially-augmented floating treatment wetlands. Chemosphere 2019, 217, 576–583. [Google Scholar] [CrossRef] [PubMed]
- Pichtel, J. Oil and gas production wastewater: Soil contamination and pollution prevention. Appl. Environ. Soil Sci. 2016, 2016, 1–24. [Google Scholar] [CrossRef] [Green Version]
- Afzal, M.; Rehman, K.; Shabir, G.; Tahseen, R.; Ijaz, A.; Hashmat, A.J.; Brix, H. Large-scale remediation of oil-contaminated water using floating treatment wetlands. NPJ Clean Water 2019, 2, 3. [Google Scholar] [CrossRef]
- Yu, K.L.; Show, P.L.; Ong, H.C.; Ling, T.C.; Lan, J.C.-W.; Chen, W.-H.; Chang, J.-S. Microalgae from wastewater treatment to biochar–feedstock preparation and conversion technologies. Energy Convers. Manag. 2017, 150, 1–13. [Google Scholar] [CrossRef]
- Afzal, M.; Arslan, M.; Müller, J.A.; Shabir, G.; Islam, E.; Tahseen, R.; Anwar-ul-Haq, M.; Hashmat, A.J.; Iqbal, S.; Khan, Q.M. Floating treatment wetlands as a suitable option for large-scale wastewater treatment. Nat. Sustain. 2019, 2, 863–871. [Google Scholar] [CrossRef]
- Headley, T.; Tanner, C.C. Constructed wetlands with floating emergent macrophytes: An innovative stormwater treatment technology. Crit. Rev. Environ. Sci. Technol. 2012, 42, 2261–2310. [Google Scholar] [CrossRef]
- Chen, Z.; Cuervo, D.P.; Müller, J.A.; Wiessner, A.; Koser, H.; Vymazal, J.; Kästner, M.; Kuschk, P. Hydroponic root mats for wastewater treatment—A review. Environ. Sci. Pollut. Res. Int. 2016, 23, 15911–15928. [Google Scholar] [CrossRef] [PubMed]
- Shahid, M.J.; Arslan, M.; Ali, S.; Siddique, M.; Afzal, M. Floating wetlands: A sustainable tool for wastewater treatment. Clean Soil Air Water 2018, 46, 1800120. [Google Scholar] [CrossRef]
- Colares, G.S.; Dell’Osbel, N.; Wiesel, P.G.; Oliveira, G.A.; Lemos, P.H.Z.; da Silva, F.P.; Lutterbeck, C.A.; Kist, L.T.; Machado, Ê.L. Floating treatment wetlands: A review and bibliometric analysis. Sci. Total Environ. 2020, 714, 136776. [Google Scholar] [CrossRef]
- Ziajahromi, S.; Drapper, D.; Hornbuckle, A.; Rintoul, L.; Leusch, F.D. Microplastic pollution in a stormwater floating treatment wetland: Detection of tyre particles in sediment. Sci. Total Environ. 2020, 713, 136356. [Google Scholar] [CrossRef]
- Al-Baldawi, I.; Abdullah, S.S.; Anuar, N.; Mushrifah, I. Bioaugmentation for the enhancement of hydrocarbon phytoremediation by rhizobacteria consortium in pilot horizontal subsurface flow constructed wetlands. Int. J. Environ. Sci. Technol. 2017, 14, 75–84. [Google Scholar] [CrossRef]
- Beans, C. Core Concept: Phytoremediation advances in the lab but lags in the field. Proc. Natl. Acad. Sci. USA 2017, 114, 7475–7477. [Google Scholar] [CrossRef] [Green Version]
- Abdullah, S.R.S.; Al-Baldawi, I.A.; Almansoory, A.F.; Purwanti, I.F.; Al-Sbani, N.H.; Sharuddin, S.S.N. Plant-assisted remediation of hydrocarbons in water and soil: Application, mechanisms, challenges and opportunities. Chemosphere 2020, 247, 125932. [Google Scholar] [CrossRef]
- Keizer-Vlek, H.E.; Verdonschot, P.F.; Verdonschot, R.C.; Dekkers, D. The contribution of plant uptake to nutrient removal by floating treatment wetlands. Ecol. Eng. 2014, 73, 684–690. [Google Scholar] [CrossRef]
- Lynch, J.; Fox, L.J.; Owen, J.S., Jr.; Sample, D.J. Evaluation of commercial floating treatment wetland technologies for nutrient remediation of stormwater. Ecol. Eng. 2015, 75, 61–69. [Google Scholar] [CrossRef] [Green Version]
- Yeh, N.; Yeh, P.; Chang, Y.-H. Artificial floating islands for environmental improvement. Renew. Sustain. Energy Rev. 2015, 47, 616–622. [Google Scholar] [CrossRef]
- Shahid, M.J.; AL-surhanee, A.A.; Kouadri, F.; Ali, S.; Nawaz, N.; Afzal, M.; Rizwan, M.; Ali, B.; Soliman, M.H. Role of microorganisms in the remediation of wastewater in Floating Treatment Wetlands: A Review. Sustainability 2020, 12, 5559. [Google Scholar] [CrossRef]
- Li, X.; Guo, R. Comparison of nitrogen removal in floating treatment wetlands constructed with Phragmites australis and Acorus calamus in a cold temperate zone. Water Air Soil Pollut. 2017, 228, 132. [Google Scholar] [CrossRef]
- Hussain, Z.; Arslan, M.; Shabir, G.; Malik, M.H.; Mohsin, M.; Iqbal, S.; Afzal, M. Remediation of textile bleaching effluent by bacterial augmented horizontal flow and vertical flow constructed wetlands: A comparison at pilot scale. Sci. Total Environ. 2019, 685, 370–379. [Google Scholar] [CrossRef] [PubMed]
- Guarino, C.; Marziano, M.; Tartaglia, M.; Prigioniero, A.; Postiglione, A.; Scarano, P.; Sciarrillo, R. Poaceae with PGPR bacteria and arbuscular mycorrhizae partnerships as a model system for plant microbiome manipulation for phytoremediation of petroleum hydrocarbons contaminated agricultural soils. Agronomy 2020, 10, 547. [Google Scholar] [CrossRef] [Green Version]
- Compant, S.; Clément, C.; Sessitsch, A. Plant growth-promoting bacteria in the rhizo- and endosphere of plants: Their role, colonization, mechanisms involved and prospects for utilization. Soil Biol. Biochem. 2010, 42, 669–678. [Google Scholar] [CrossRef] [Green Version]
- Matamoros, V.; Nguyen, L.X.; Arias, C.A.; Salvadó, V.; Brix, H. Evaluation of aquatic plants for removing polar microcontaminants: A microcosm experiment. Chemosphere 2012, 88, 1257–1264. [Google Scholar] [CrossRef]
- Sessitsch, A.; Pfaffenbichler, N.; Mitter, B. Microbiome applications from Lab to Field: Facing complexity. Trends Plant Sci. 2019, 24, 194–198. [Google Scholar] [CrossRef]
- Weyens, N.; van der Lelie, D.; Taghavi, S.; Newman, L.; Vangronsveld, J. Exploiting plant-microbe partnerships to improve biomass production and remediation. Trends Biotechnol. 2009, 27, 591–598. [Google Scholar] [CrossRef]
- Khan, S.; Afzal, M.; Iqbal, S.; Khan, Q.M. Plant-bacteria partnerships for the remediation of hydrocarbon contaminated soils. Chemosphere 2013, 90, 1317–1332. [Google Scholar] [CrossRef] [PubMed]
- Arslan, M.; Imran, A.; Khan, Q.M.; Afzal, M. Plant–bacteria partnerships for the remediation of persistent organic pollutants. Environ. Sci. Pollut. Res. 2017, 24, 4322–4336. [Google Scholar] [CrossRef]
- Fatima, K.; Imran, A.; Amin, I.; Khan, Q.M.; Afzal, M. Successful phytoremediation of crude-oil contaminated soil at an oil exploration and production company by plants-bacterial synergism. Int. J. Phytoremediat. 2018, 20, 675–681. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Liu, Y.; Jin, H. Nitrogen removal from polluted river by enhanced floating bed grown Canna. Ecol. Eng. 2009, 35, 135–140. [Google Scholar] [CrossRef]
- Ijaz, A.; Shabir, G.; Khan, Q.M.; Afzal, M. Enhanced remediation of sewage effluent by endophyte-assisted floating treatment wetlands. Ecol. Eng. 2015, 84, 58–66. [Google Scholar] [CrossRef]
- Rehman, K.; Imran, A.; Amin, I.; Afzal, M. Inoculation with bacteria in floating treatment wetlands positively modulates the phytoremediation of oil field wastewater. J. Hazard. Mater. 2018, 349, 242–251. [Google Scholar] [CrossRef]
- Saleem, H.; Arslan, M.; Rehman, K.; Tahseen, R.; Afzal, M. Phragmites australis—A helophytic grass—Can establish successful partnership with phenol-degrading bacteria in a floating treatment wetland. Saudi J. Biol. Sci. 2019, 26, 1179–1186. [Google Scholar] [CrossRef] [PubMed]
- Saleem, H.; Rehman, K.; Arslan, M.; Afzal, M. Enhanced degradation of phenol in floating treatment wetlands by plant-bacterial synergism. Int. J. Phytoremediat. 2018, 20, 692–698. [Google Scholar] [CrossRef]
- Ashraf, S.; Afzal, M.; Naveed, M.; Shahid, M.; Ahmad Zahir, Z. Endophytic bacteria enhance remediation of tannery effluent in constructed wetlands vegetated with Leptochloa fusca. Int. J. Phytoremediat. 2018, 20, 121–128. [Google Scholar] [CrossRef] [PubMed]
- Afzal, M.; Yousaf, S.; Reichenauer, T.G.; Sessitsch, A. The inoculation method affetcs colonization and performance of bacterial inoculant strains in the phytoremediation of soil contaminated with diesel oil. Int. J. Phytoremediat. 2012, 14, 35–47. [Google Scholar] [CrossRef] [PubMed]
- Yousaf, S.; Afzal, M.; Reichenauer, T.G.; Brady, C.L.; Sessitsch, A. Hydrocarbon degradation, plant colonization and gene expression of alkane degradation genes by endophytic Enterobacter ludwigii strains. Environ. Pollut. 2011, 159, 2675–2683. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- American Public Health Association. Standard Methods for the Examination of Water and Wastewater, 20th ed.; American Public Health Association: Washington, DC, USA, 2005. [Google Scholar]
- Fatima, K.; Afzal, M.; Imran, A.; Khan, Q. Bacterial rhizosphere and endosphere populations associated with grasses and trees to be used for phytoremediation of crude oil contaminated soil. Bull. Environ. Contam. Toxicol. 2015, 94, 314–320. [Google Scholar] [CrossRef] [PubMed]
- Andria, V.; Reichenauer, T.G.; Sessitsch, A. Expression of alkane monooxygenase (alkB) genes by plant-associated bacteria in the rhizosphere and endosphere of Italian ryegrass (Lolium multiflorum L.) grown in diesel contaminated soil. Environ. Pollut. 2009, 157, 3347–3350. [Google Scholar] [CrossRef]
- Yousaf, S.; Andria, V.; Reichenauer, T.G.; Smalla, K.; Sessitsch, A. Phylogenetic and functional diversity of alkane degrading bacteria associated with Italian ryegrass (Lolium multiflorum) and birdsfoot trefoil (Lotus corniculatus) in a petroleum oil-contaminated environment. J. Hazard. Mater. 2010, 184, 523–532. [Google Scholar] [CrossRef]
- Arslan, M.; Afzal, M.; Amin, I.; Iqbal, S.; Khan, Q.M. Nutrients can enhance the abundance and expression of alkane hydroxylase CYP153 gene in the rhizosphere of ryegrass planted in hydrocarbon-polluted soil. PLoS ONE 2014, 9, e111208. [Google Scholar] [CrossRef] [PubMed]
- Afzal, M.; Yousaf, S.; Reichenauer, T.G.; Kuffner, M.; Sessitsch, A. Soil type affects plant colonization, activity and catabolic gene expression of inoculated bacterial strains during phytoremediation of diesel. J. Hazard. Mater. 2011, 186, 1568–1575. [Google Scholar] [CrossRef]
- Naureen, Z.; Yasmin, S.; Hameed, S.; Malik, K.; Hafeez, F.Y. Characterization and screening of bacteria from rhizosphere of maize grown in Indonesian and Pakistani soils. J. Basic Microbiol. 2005, 45, 447–459. [Google Scholar] [CrossRef]
- Fatima, K.; Imran, A.; Amin, I.; Khan, Q.; Afzal, M. Plant species affect colonization patterns and metabolic activity of associated endophytes during phytoremediation of crude oil-contaminated soil. Environ. Sci. Pollut. Res. 2016, 23, 6188–6196. [Google Scholar] [CrossRef] [PubMed]
- Afzal, M.; Shabir, G.; Hussain, I.; Khalid, Z.M. Paper and board mill effluent treatment with the combined biological-coagulation-filtration pilot scale reactor. Bioresour. Technol. 2008, 99, 7383–7387. [Google Scholar] [CrossRef] [PubMed]
- Stottmeister, U.; Wießner, A.; Kuschk, P.; Kappelmeyer, U.; Kästner, M.; Bederski, O.; Müller, R.; Moormann, H. Effects of plants and microorganisms in constructed wetlands for wastewater treatment. Biotechnol. Adv. 2003, 22, 93–117. [Google Scholar] [CrossRef]
- Faulwetter, J.L.; Gagnon, V.; Sundberg, C.; Chazarenc, F.; Burr, M.D.; Brisson, J.; Camper, A.K.; Stein, O.R. Microbial processes influencing performance of treatment wetlands: A review. Ecol. Eng. 2009, 35, 987–1004. [Google Scholar] [CrossRef]
- Afzal, M.; Khan, Q.M.; Sessitsch, A. Endophytic bacteria: Prospects and applications for the phytoremediation of organic pollutants. Chemosphere 2014, 117, 232–242. [Google Scholar] [CrossRef]
- Iamchaturapatr, J.; Yi, S.W.; Rhee, J.S. Nutrient removals by 21 aquatic plants for vertical free surface-flow (VFS) constructed wetland. Ecol. Eng. 2007, 29, 287–293. [Google Scholar] [CrossRef]
- Fahid, M.; Arslan, M.; Shabir, G.; Younus, S.; Yasmeen, T.; Rizwan, M.; Siddique, K.; Ahmad, S.R.; Tahseen, R.; Iqbal, S.; et al. Phragmites australis in combination with hydrocarbons degrading bacteria is a suitable option for remediation of diesel-contaminated water in floating wetlands. Chemosphere 2020, 240, 124890. [Google Scholar] [CrossRef]
- Shehzadi, M.; Afzal, M.; Islam, E.; Mobin, A.; Anwar, S.; Khan, Q.M. Enhanced degradation of textile effluent in constructed wetland system using Typha domingensis and textile effluent-degrading endophytic bacteria. Water Res. 2014, 58, 152–159. [Google Scholar] [CrossRef]
- Fernández-Luqueño, F.; Valenzuela-Encinas, C.; Marsch, R.; Martínez-Suárez, C.; Vázquez-Núñez, E.; Dendooven, L. Microbial communities to mitigate contamination of PAHs in soil—Possibilities and challenges: A review. Environ. Sci. Pollut. Res. 2011, 18, 12–30. [Google Scholar] [CrossRef]
- Doughari, H.J.; Ndakidemi, P.A.; Human, I.S.; Benade, S. The ecology, biology and pathogenesis of Acinetobacter spp.: An overview. Microbes Environ. 2011, 26, 101–112. [Google Scholar] [CrossRef] [Green Version]
- Stewart, P.S.; Franklin, M.J. Physiological heterogeneity in biofilms. Nat. Rev. Microbiol. 2008, 6, 199–210. [Google Scholar] [CrossRef]
- Ijaz, A.; Iqbal, Z.; Afzal, M. Remediation of sewage and industrial effluent using bacterially assisted floating treatment wetlands vegetated with Typha domingensis. Water Sci. Technol. 2016, 74, 2192–2201. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Afzal, M.; Khan, S.; Iqbal, S.; Mirza, M.S.; Khan, Q.M. Inoculation method affects colonization and activity of Burkholderia phytofirmans PsJN during phytoremediation of diesel-contaminated soil. Int. Biodeterior. Biodegrad. 2013, 85, 331–336. [Google Scholar] [CrossRef]
- Balseiro-Romero, M.; Gkorezis, P.; Kidd, P.S.; Van Hamme, J.; Weyens, N.; Monterroso, C.; Vangronsveld, J. Characterization and degradation potential of diesel-degrading bacterial strains for application in bioremediation. Int. J. Phytoremediat. 2017, 19, 955–963. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stout, L.; Nüsslein, K. Biotechnological potential of aquatic plant–microbe interactions. Curr. Opin. Biotechnol. 2010, 21, 339–345. [Google Scholar] [CrossRef]
- Srivastava, P.; Dwivedi, S.; Kumar, N.; Abbassi, R.; Garaniya, V.; Yadav, A.K. Performance assessment of aeration and radial oxygen loss assisted cathode based integrated constructed wetland-microbial fuel cell systems. Bioresour. Technol. 2017, 244, 1178–1182. [Google Scholar] [CrossRef] [PubMed]
- Calheiros, C.S.C.; Pereira, S.I.A.; Brix, H.; Rangel, A.O.S.S.; Castro, P.M.L. Assessment of culturable bacterial endophytic communities colonizing Canna flaccida inhabiting a wastewater treatment constructed wetland. Ecol. Eng. 2017, 98, 418–426. [Google Scholar] [CrossRef]

| Length (cm) | Fresh Biomass (g m−3) | Dry Biomass (g m−3) | ||||
|---|---|---|---|---|---|---|
| Roots | Shoots | Roots | Shoots | Roots | Shoots | |
| P. australis | ||||||
| Tap water | 58.3 a ± 3.4 | 218 a ± 20 | 654 a ± 17 | 187 a ± 9 | 183 a ±13 | 112 a ± 7 |
| Inoculation (−) | 34.7 c ± 3.8 | 135 c ± 15 | 448 c ± 31 | 130 c ± 7 | 119 c ± 8 | 78 c ± 6 |
| Inoculation (+) | 45.1 b ± 3.1 | 172 b ± 16 | 511 b ± 24 | 151 b ± 6 | 145 b ± 7 | 94 b ± 6 |
| T. domingensis | ||||||
| Tap water | 52.1 a ± 4.2 | 178 a ± 12 | 576 a ± 16 | 164 a ± 7 | 116 a± 7 | 107 a ± 9 |
| Inoculation (−) | 33.4 c ± 3.7 | 124 c ± 8.2 | 384 c ± 21 | 119 c ± 6 | 81 c ± 5 | 68 c ± 4 |
| Inoculation (+) | 42.2 b ± 3.8 | 150 b ± 9.5 | 482 b ± 26 | 142 b ± 7 | 96 b ± 6 | 87 b ± 5 |
| Time Days | CFU mL−1 or g−1 × 106 | |||
|---|---|---|---|---|
| Water | RP | RI | SI | |
| P. australis | ||||
| 0 | 59 a ± 4 | 55 c ± 9 | 5.6 c ± 0.2 | 1.3 d ± 0.1 |
| 12 | 55 ab ± 7 | 153 b ± 7 | 7.8 b ± 0.5 | 1.4 c ± 0.1 |
| 24 | 53 ab ± 4 | 156 ab ± 9 | 12.3 a ± 0.9 | 2.6 b ± 0.1 |
| 36 | 46 b ± 8 | 168 a ± 2 | 12.5 a ± 0.7 | 2.8 a± 0.1 |
| T. domingensis | ||||
| 0 | 64 a ± 8 | 27 b ± 1 | 2.8 e ± 0.2 | 0.7 b ± 0.1 |
| 12 | 58 ab ± 4 | 46 a ± 4 | 3.7 b ± 0.4 | 0.8 a ± 0.1 |
| 24 | 51 b ± 5 | 52 a ± 2 | 4.3 d ± 0.5 | 0.8 a ± 0.1 |
| 36 | 43 c ± 5 | 49 a ± 1 | 4.6 h ± 0.5 | 0.8 a ± 0.1 |
| Strain | CFU mL−1 or g−1 × 106 | |||
|---|---|---|---|---|
| Water | RP | RI | SI | |
| Phragmites australis | ||||
| Acinetobacter junii TYRH47 | 6.9 ± 1.2 | 58.8 ± 4.2 | 1.3 ± 0.1 | 0.2 ± 0.004 |
| Acinetobacter sp. LCRH81 | 9.2 ± 1.6 | 81.2 ± 5.8 | 1.5 ± 0.1 | 0.2 ± 0.004 |
| Bacillus subtilis LORI66 | 6.9 ± 4.2 | 61.6 ± 4.4 | 3.8 ± 0.21 | 0.4 ± 0.007 |
| Klebsiella sp. LCRI87 | 4.6 ± 0.8 | 39.2 ± 2.8 | 1.7 ± 0.1 | 0.3± 0.005 |
| Typha domingensis | ||||
| Acinetobacter junii TYRH47 | 10 ± 1.1 | 13.9 ± 0.4 | 0.8 ± 0.08 | 0.2 ± 0.002 |
| Acinetobacter sp. LCRH81 | 9.3 ± 1.1 | 12.3 ± 0.4 | 0.4 ± 0.04 | 0.1 ± 0.002 |
| Bacillus subtilis LORI66 | 5.7 ± 0.7 | 21.2 ± 0.6 | 0.8 ± 0.08 | 0.2 ± 0.002 |
| Klebsiella sp. LCRI87 | 4.3 ± 0.5 | 21.2 ± 0.6 | 1.0 ± 0.1 | 0.1 ± 0.002 |
| Time Days | alkB Abundance | alkB Expression | ||||||
|---|---|---|---|---|---|---|---|---|
| Copy Number mL−1 or g−1 × 104 | Copy Number mL−1 or g−1 × 104 | |||||||
| Water | RP | RI | SI | Water | RP | RI | SI | |
| P. australis | ||||||||
| 0 | 18.6 a ± 0.5 | 1.3 c ± 1.2 | 0.3 d ± 0.02 | 0.2 d ± 0.1 | 7.3 a ± 0.04 | 0.7 c ±0.2 | 0.04 c ± 0.1 | 0.01 d ± 0.02 |
| 12 | 17.3 b ± 0.7 | 14.7 b ± 0.9 | 4.8 c ± 0.1 | 2.3 c ± 0.1 | 6.6 b ± 0.03 | 10.5 b ± 0.2 | 3.7 a ±0.1 | 0.4 c ±0.03 |
| 24 | 13.8 c ± 0.04 | 15.2 b ± 0.7 | 6.6 b ± 0.1 | 2.8 a ± 0.03 | 3.2 c ± 0.02 | 11.2 b ± 0.1 | 3.0 b ±0.02 | 0.6 b ±0.01 |
| 36 | 11.5 d ± 0.02 | 15.8 a ± 0.1 | 7.2 a ± 0.04 | 2.7 b ± 0.01 | 2.2 d ± 0.01 | 12.5 a ± 0.1 | 3.5 a ±0.01 | 0.8 a ± 0.02 |
| T. domingensis | ||||||||
| 0 | 13.3 a ± 0.2 | 1.1 d ± 0.9 | 0.05 d ± 0.1 | 0.02 c ± 0.04 | 6.1 a ± 0.2 | 0.3 d ± 0.02 | 0.8 d± 0.1 | 0.03 b ± 0.03 |
| 12 | 12.3 b ± 0.1 | 3.3 c ± 0.3 | 1.1 c ± 0.04 | 0.4 b ± 0.03 | 5.3 b ± 0.3 | 2.8 c± 0.2 | 1.1 c± 0.1 | 0.1 a ±0.02 |
| 24 | 13.8 c ± 0.1 | 4.3 b ± 0.2 | 2.1 b ± 0.02 | 0.7 a ± 0.1 | 2.1 c ± 0.7 | 3.8 b± 0.3 | 1.3 a ± 0.01 | 0.1 a ±0.01 |
| 36 | 12.5 d ± 0.01 | 4.7 a ± 0.1 | 2.5 a ± 0.01 | 0.64 a ± 0.1 | 1.4 d ± 0.02 | 4.2 a± 0.2 | 1.1 b ± 0.01 | 0.2 b ±0.02 |
| Treatment | Fish Mortality up to Time Point | |||
|---|---|---|---|---|
| 24 h | 48 h | 72 h | 96 h | |
| Wastewater only | 10 | - | - | - |
| with bioaugmentation | 0 | 3 | 4 | 5 |
| Phragmites australis | ||||
| without bioaugmentation | 0 | 0 | 2 | 3 |
| with bioaugmentation | 0 | 0 | 0 | 0 |
| Typha domingensis | ||||
| without bioaugmentation | 0 | 0 | 2 | 4 |
| with bioaugmentation | 0 | 0 | 0 | 0 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rehman, K.; Arslan, M.; Müller, J.A.; Saeed, M.; Imran, A.; Amin, I.; Mustafa, T.; Iqbal, S.; Afzal, M. Bioaugmentation-Enhanced Remediation of Crude Oil Polluted Water in Pilot-Scale Floating Treatment Wetlands. Water 2021, 13, 2882. https://doi.org/10.3390/w13202882
Rehman K, Arslan M, Müller JA, Saeed M, Imran A, Amin I, Mustafa T, Iqbal S, Afzal M. Bioaugmentation-Enhanced Remediation of Crude Oil Polluted Water in Pilot-Scale Floating Treatment Wetlands. Water. 2021; 13(20):2882. https://doi.org/10.3390/w13202882
Chicago/Turabian StyleRehman, Khadeeja, Muhammad Arslan, Jochen A. Müller, Muhammad Saeed, Asma Imran, Imran Amin, Tanveer Mustafa, Samina Iqbal, and Muhammad Afzal. 2021. "Bioaugmentation-Enhanced Remediation of Crude Oil Polluted Water in Pilot-Scale Floating Treatment Wetlands" Water 13, no. 20: 2882. https://doi.org/10.3390/w13202882
APA StyleRehman, K., Arslan, M., Müller, J. A., Saeed, M., Imran, A., Amin, I., Mustafa, T., Iqbal, S., & Afzal, M. (2021). Bioaugmentation-Enhanced Remediation of Crude Oil Polluted Water in Pilot-Scale Floating Treatment Wetlands. Water, 13(20), 2882. https://doi.org/10.3390/w13202882









