Technology for Upgrading the Tailwater of Municipal Sewage Treatment Plants: The Efficacy and Mechanism of Microbial Coupling for Nitrogen and Carbon Removal
Abstract
Highlights
- Coupling nitrogen and carbon removal microorganisms in an MBR is a promising technology for improving the tailwater of municipal sewage treatment plants.
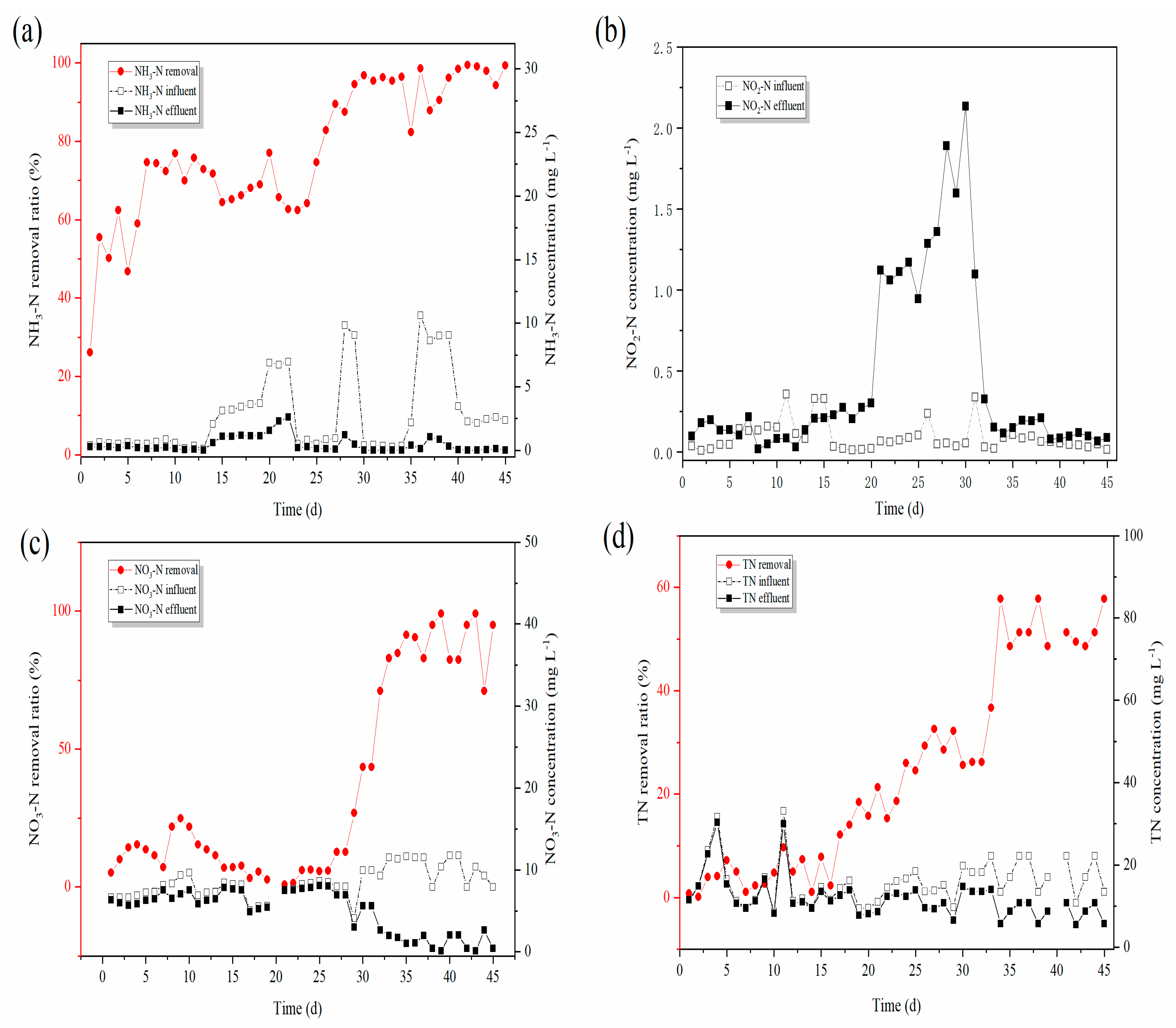
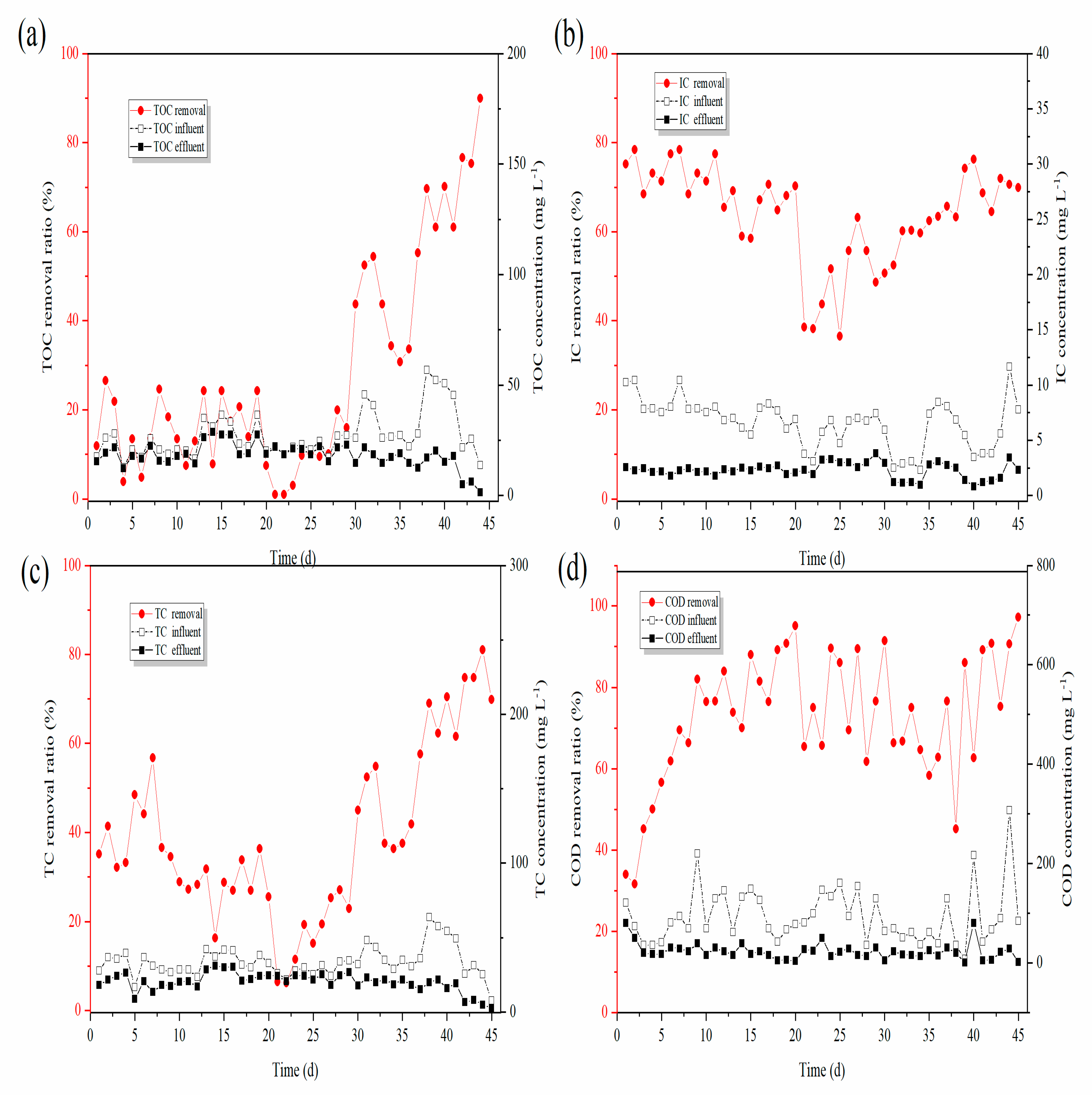
- The reactor showed stable simultaneous denitrification and carbon removal functions. The removal of NH3-N was 98.63%, and the removal of COD was 80.2%.
- The two microorganisms coexisted in the reactor to form a relatively rich microbial community.
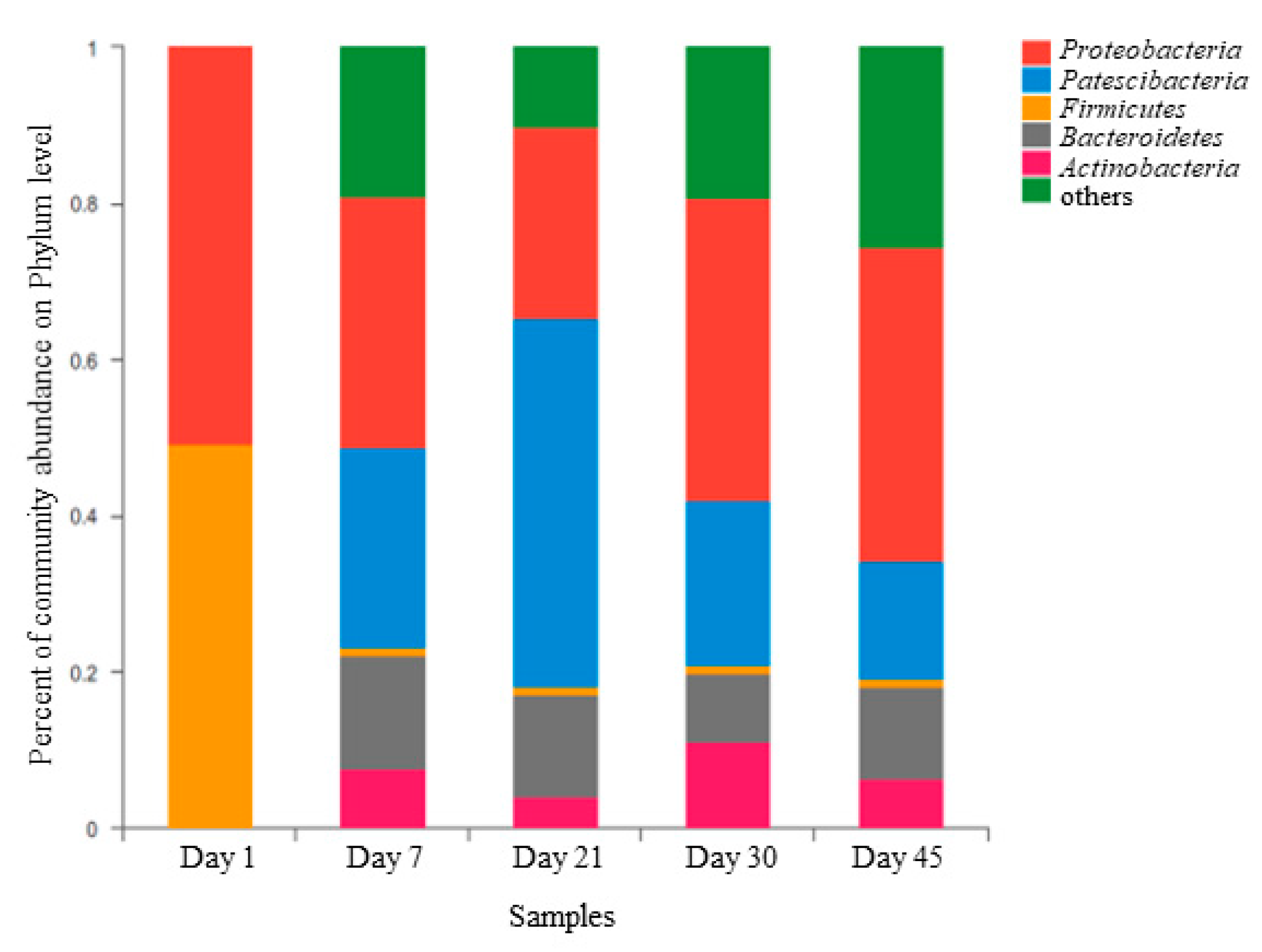
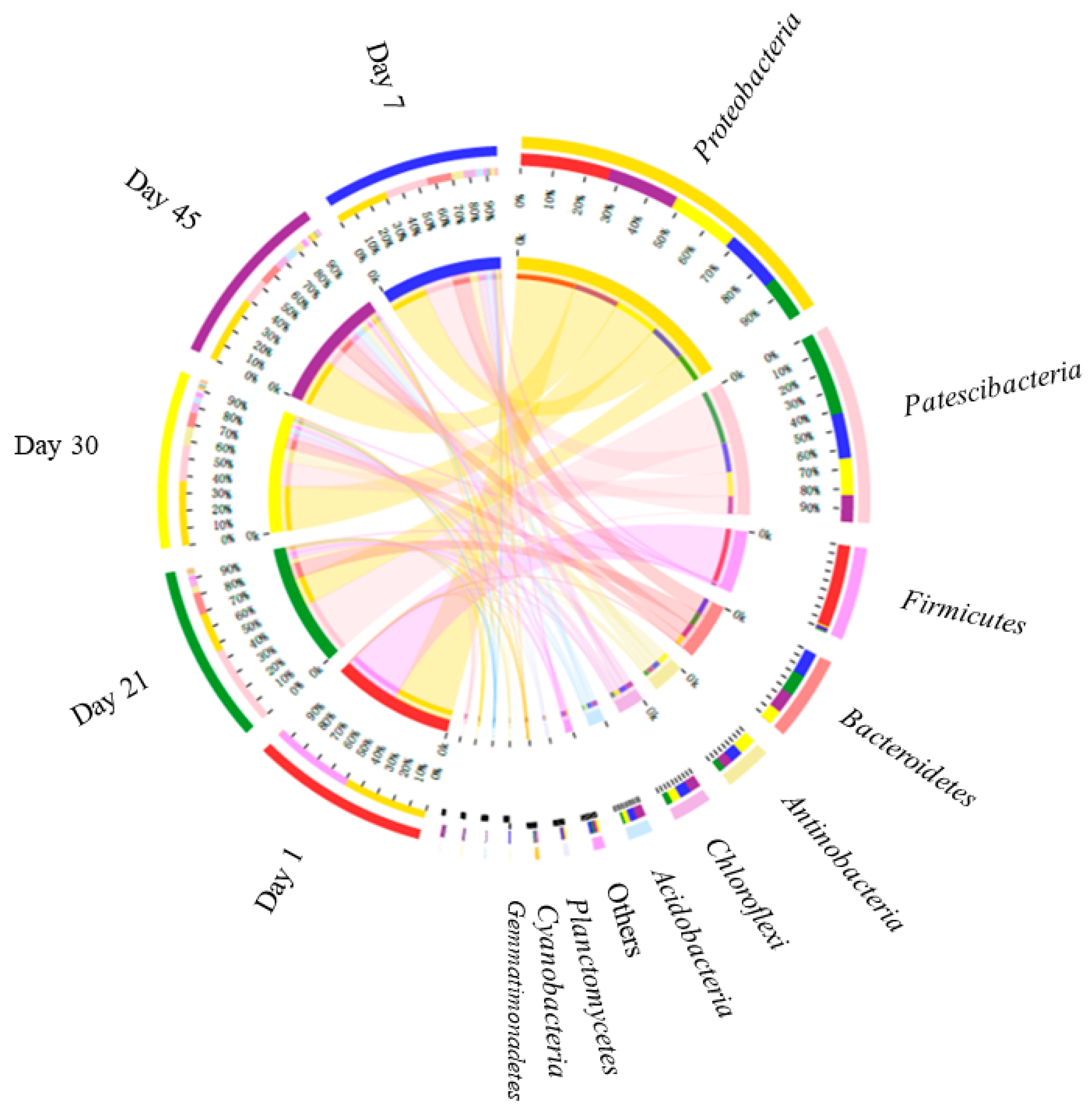
- The cooccurrence of Proteobacteria and Bacillus with Chloroflexi might have accounted for the synchronous high removal efficiency of NH3-N and COD.
1. Introduction
2. Materials and Methods
2.1. MBR Setup and Operation
2.2. Seed Microbial Community
2.3. Water Quality Analytical Parameters
2.4. Microbial Community Analysis
3. Results
3.1. The Performance of the MBR in Nitrogen and Carbon Removal
3.1.1. Changes in Denitrification Performance
3.1.2. Changes in Decarburization Performance
3.2. Analysis of the Microbial Community
3.2.1. Diversity of the Microbial Community
3.2.2. Composition of the Microbial Community
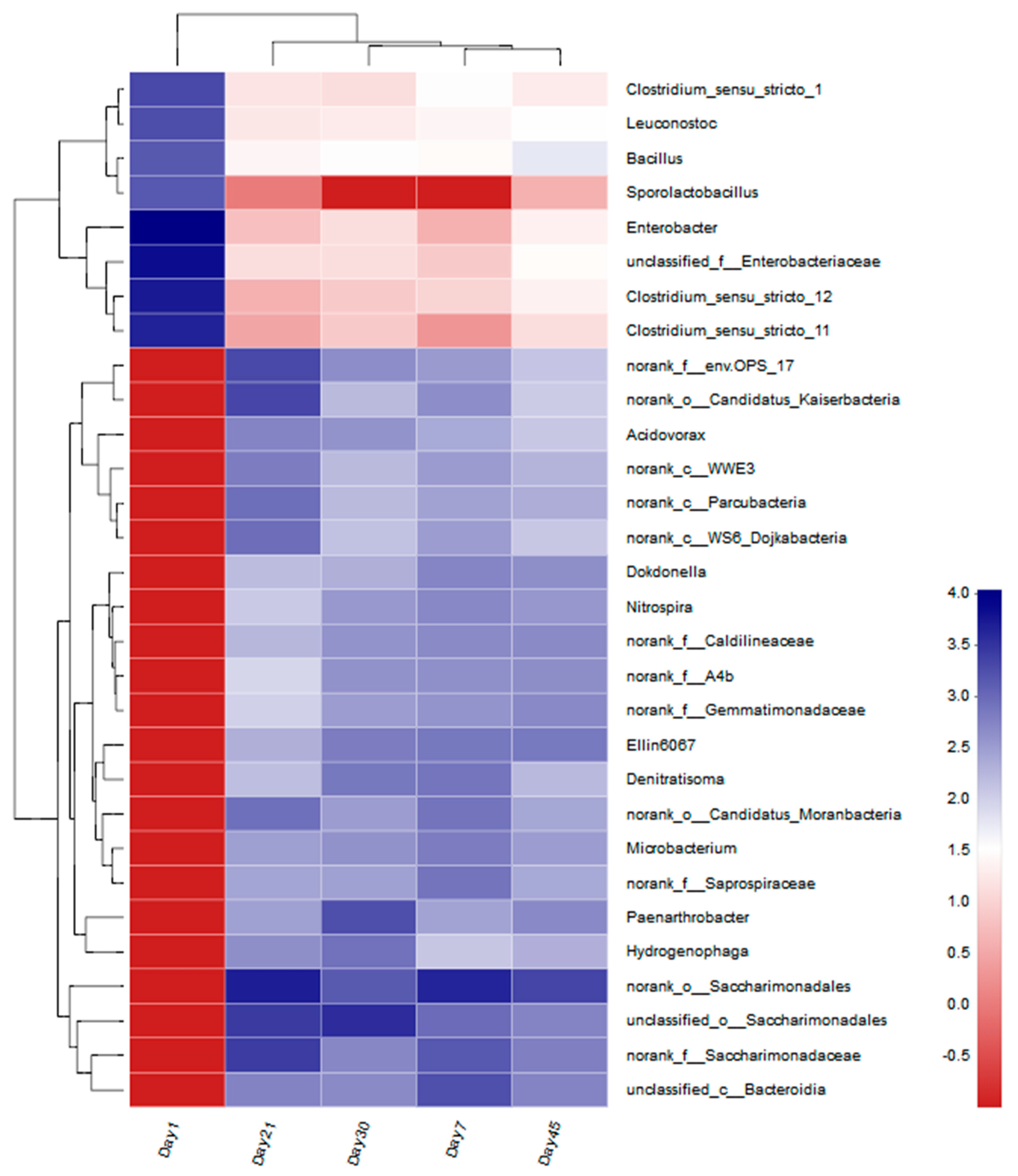
3.2.3. Microbial Genetic Relationship Analysis at the Genus Level
3.3. Microbial Environment Relevance
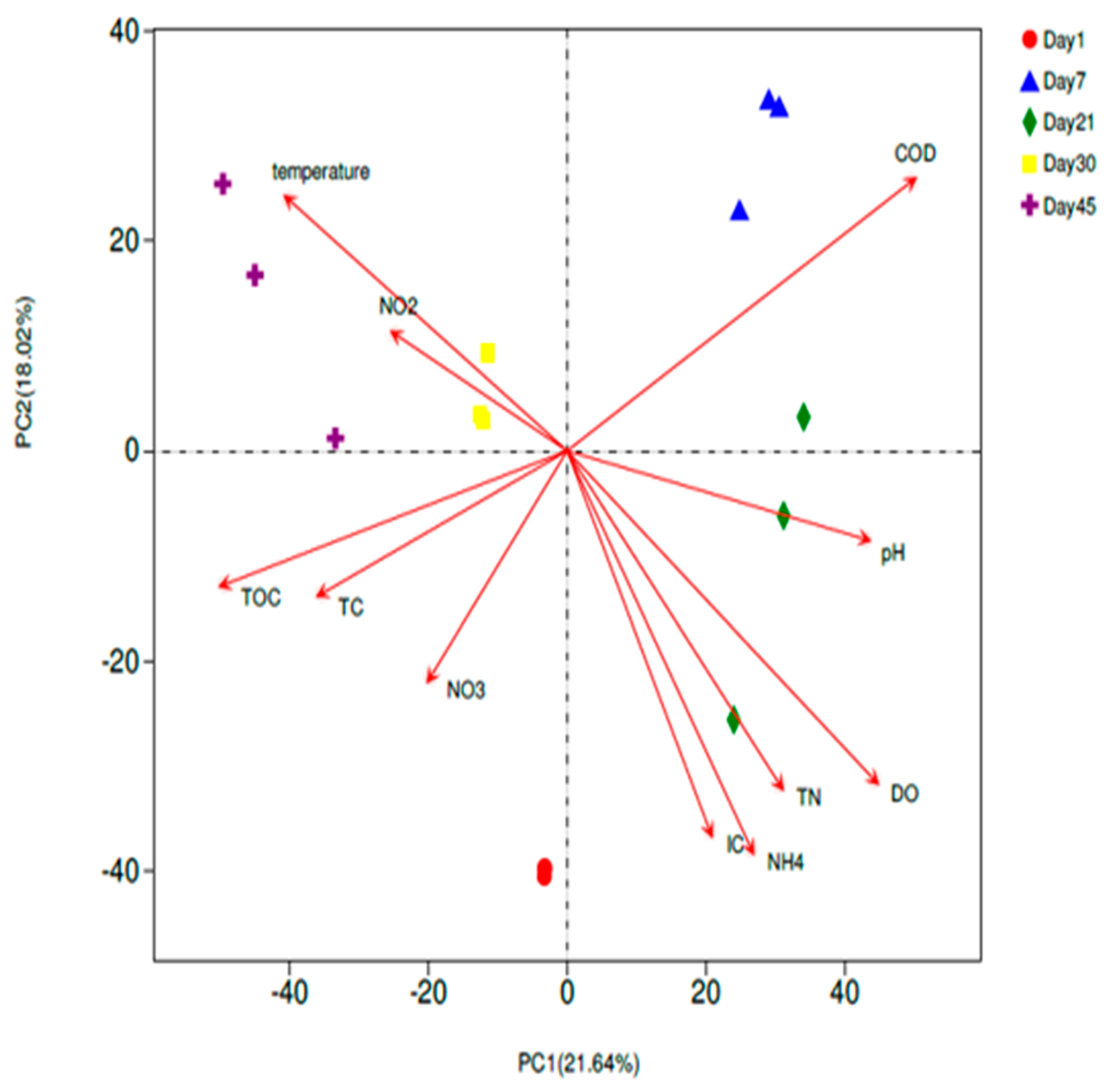
3.3.1. Principal Component Analysis (PCA)
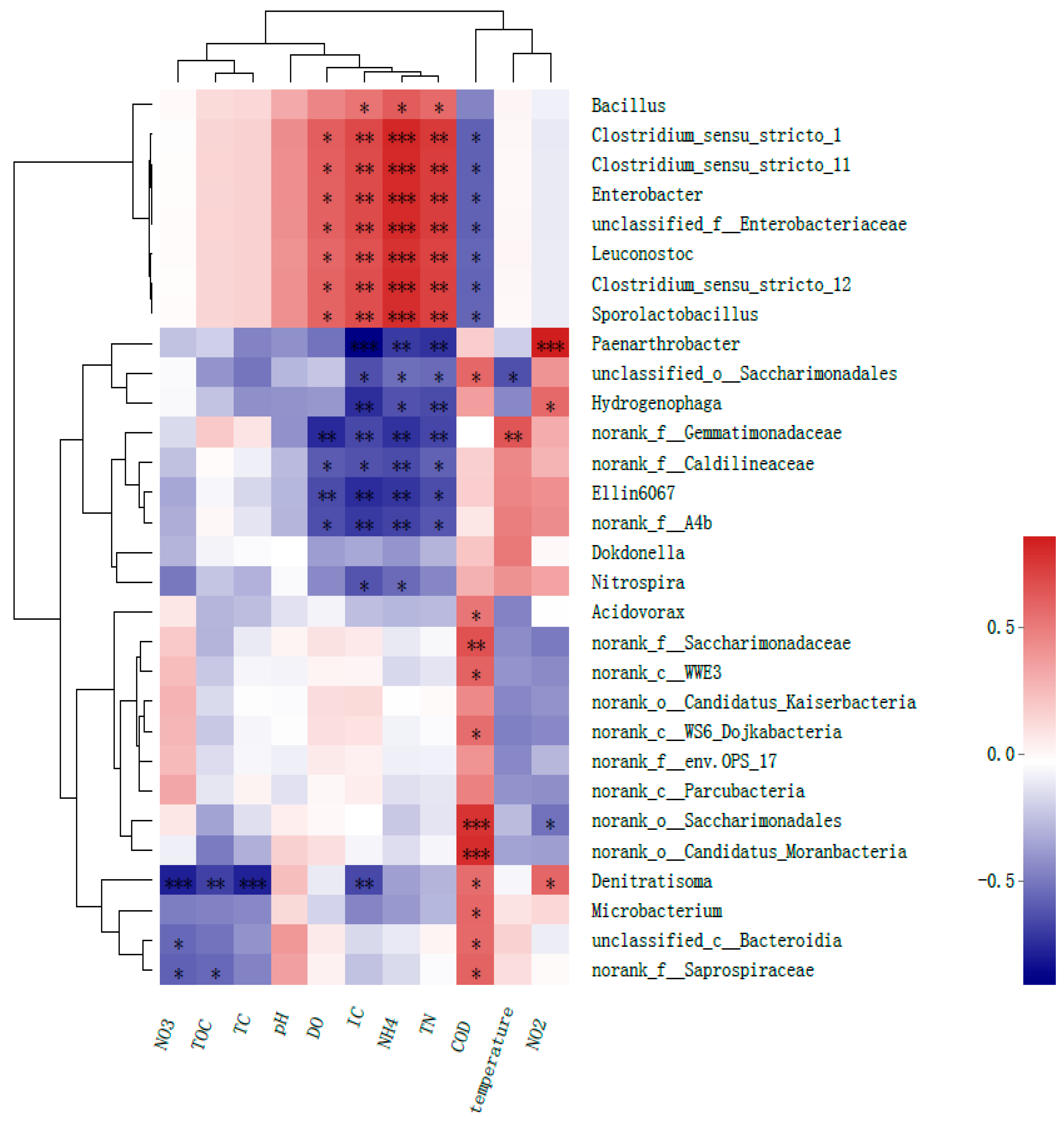
3.3.2. Correlation Heatmap Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bagchi, S.; Behera, M. Bioaugmentation using Pseudomonas aeruginosa with an approach of intermittent aeration for enhanced power generation in ceramic MFC. Sustain. Energy Technol. Assess. 2021, 45, 101138. [Google Scholar]
- Albina, P.; Durban, N.; Bertron, A.; Albrecht, A.; Robinet, J.-C.; Erable, B. Influence of hydrogen electron donor, alkaline pH, and high nitrate concentrations on microbial denitrification: A review. Int. J. Mol. Sci. 2019, 20, 5163. [Google Scholar] [CrossRef] [PubMed]
- Crini, G.; Lichtfouse, E. Advantages and disadvantages of techniques used for wastewater treatment. Environ. Chem. Lett. 2019, 17, 145–155. [Google Scholar] [CrossRef]
- Di Bella, G.; Mannina, G. Intermittent aeration in a hybrid moving bed biofilm-reactor for carbon and nutrient biological removal. Water 2020, 12, 492. [Google Scholar] [CrossRef]
- Velkova, Z.; Kirova, G.; Stoytcheva, M.; Kostadinova, S.; Todorova, K.; Gochev, V. Immobilized microbial biosorbents for heavy metals removal. Eng. Life Sci. 2018, 18, 871–881. [Google Scholar] [CrossRef]
- Skoczko, I.; Puzowski, P.; Szatyłowicz, E. Experience from the Implementation and Operation of the Biological Membrane Reactor (MBR) at the Modernized Wastewater Treatment Plant in Wydminy. Water 2020, 12, 3410. [Google Scholar] [CrossRef]
- Wang, J.; Chu, L. Biological nitrate removal from water and wastewater by solid phase denitrification process. Biotechnol. Adv. 2016, 34, 1103–1112. [Google Scholar] [CrossRef]
- Hu, Z.; Lotti, T.; De Kreuk, M.; Kleerebezem, R.; Loosdrecht, M.V.; Kruit, J.; Jetten, M.S.M.; Kartal, B. Nitrogen removal by a nitritation-anammox bioreactor at low temperature. Appl. Environ. Microbiol. 2013, 79, 2807–2818. [Google Scholar] [CrossRef]
- Zhao, B.; Ran, X.C.; Tian, M. Assessing the performance of a sequencing batch biofilm reactor bioaugmented with P. stutzeri strain XL-2 treating ammonium-rich wastewater. Bioresour. Technol. 2018, 270, 70–79. [Google Scholar] [CrossRef]
- Liao, R.H.; Miao, Y.; Li, J.; Li, Y.; Zhu, W.; Du, J.; Li, Y.M.; Li, A.M.; Shen, H.J. Temperature dependence of denitrification microbial communities and functional genes in an expanded granular sludge bed reactor treating nitrate-rich wastewater. RSC Adv. 2018, 8, 42087–42094. [Google Scholar] [CrossRef]
- Schlegel, S.; Koeser, H. Wastewater treatment with submerged fixed bed biofilm reactor systems—Design rules, operating experiences and ongoing developments. Water Sci. Technol. 2009, 55, 83–89. [Google Scholar] [CrossRef]
- Hamdani, A.; Mountadar, M.; Assobhei, O. Comparative study of the efficacy of three coagulants in treating dairy factory waste water. Int. J. Dairy Technol. 2005, 58, 83–88. [Google Scholar] [CrossRef]
- Ibrahim, R.S.; Sabeen, A.H.; Noor, Z.Z.; Ahmad Mutamim, N.S. Comparative study of suspended and attached growth in membrane bioreactors for wastewater treatment. Water Environ. J. 2020, 34, 273–289. [Google Scholar] [CrossRef]
- Hamdani, A.; Mountadar, M.; Assobhei, O. Simultaneous nitrate and organic matter removal from a dairy effluent by biodenitrification. Rev. Sci. Eau. 2018, 3, 97–107. [Google Scholar] [CrossRef][Green Version]
- Sun, L.; Tian, Y.; Zhang, J.; Cui, H.; Zuo, W.; Li, J. A novel symbiotic system combining algae and sludge membrane bioreactor technology for wastewater treatment and membrane fouling mitigation: Performance and mechanism. Chem. Eng. J. 2018, 344, 246–253. [Google Scholar] [CrossRef]
- Hobbie, J.E.; Daley, R.J.; Jasper, S. Use of nuclepore filters for counting bacteria by fluorescence microscopy. Appl. Environ. Microbio. 1977, 33, 1225–1228. [Google Scholar] [CrossRef]
- Kargi, F.; Pamukoglu, M.Y. Aerobic biological treatment of pre-treated landfill leachate by fed-batch operation. Enzym. Microb. Technol. 2003, 33, 588–595. [Google Scholar] [CrossRef]
- APHA. Standard Methods for the Examination of Water and Wastewater, 20th ed.; AWWA, WEF, IPB University: Bragança, Portugal, 2012. [Google Scholar]
- Huang, H.; Peng, C.; Peng, P.C.; Lin, Y.; Zhang, X.X.; Ren, H.Q. Towards the biofilm characterization and regulation in biological wastewater treatment. Appl. Microbiol. Biotechnol. 2019, 103, 1115–1129. [Google Scholar] [CrossRef] [PubMed]
- Echevarria, C.; Valderrama, C.; Cortina, J.L.; Martin, I.; Arnaldos, M.; Bernat, X.; Boleda, M.R.; Vega, A.; Teuler, A.; Castellvi, E. Technoeconomic evaluation and comparison of PAC-MBR and ozonation-UV revamping for organic micropollutants removal from urban reclaimed wastewater. Sci. Total. Environ. 2019, 671, 288–298. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wang, Y.; Xie, F.L.; Chen, S.W.; Li, J.H.; Li, D.S.; Chen, X. Improvement of acetoin reductase activity enhances bacitracin production by Bacillus licheniformis. Process. Biochem. 2014, 49, 2039–2043. [Google Scholar] [CrossRef]
- Ding, L.J.; Xu, P.; Zhang, W.Y.; Yuan, Y.; He, X.P.; Su, D.Q.; Shi, Y.T.; Naman, C.B.; Yan, X.J.; Wu, B.; et al. Three New Diketopiperazines from the Previously Uncultivable Marine Bacterium Gallaecimonas mangrovi HK-28 Cultivated by iChip. Chem. Biodiversity 2018, 17, e2000221. [Google Scholar] [CrossRef]
- Huang, H.K.; Tseng, S.K. Nitrate reduction by Citrobacter diversus under aerobic environment. Appl. Microbiol. Biotechnol. 2001, 55, 90–94. [Google Scholar] [CrossRef]
- Xuan, L.; Sheng, Z.; Lu, J.; Qiu, Q.; Chen, J.; Xiong, J. Bacterioplankton community responses and the potential ecological thresholds along disturbance gradients. Sci.Total Environ. 2019, 696, 134015–134024. [Google Scholar] [CrossRef]
- Vijay, A.; Chhabra, M.; Vincent, T. Microbial community modulates electrochemical performance and denitrification rate in a biocathodic autotrophic and heterotrophic denitrifying microbial fuel cell. Bioresour. Technol. 2019, 272, 217–225. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Liu, Y.; Lv, W.; Xia, S.; Han, J.; Wang, Z.; Yu, X.; Cai, L. Enhancement of nitrogen removal by supplementing fluidized-carriers into the aerobic tank in a full-scale A(2)/O system. Sci. Total Environ. 2019, 660, 817–825. [Google Scholar] [CrossRef] [PubMed]
- Ji, B.; Yang, K.; Zhu, L.; Jiang, Y.; Wang, H.Y.; Zhou, J.; Zhang, H.N. Aerobic denitrification: A review of important advances of the last 30 years. Biol. process. Eng. 2019, 20, 643–651. [Google Scholar] [CrossRef]
- Chen, C.J.; Huang, X.X.; Lei, C.X. Effect of organic matter strength on anammox for modified greenhouse turtle breeding wastewater treatment. Bioresour. Technol. 2013, 148, 172–179. [Google Scholar] [CrossRef]
- Kindaichi, T.; Yuri, S.; Ozaki, N.; Ohashi, A. Ecophysiological role and function of uncultured Chloroflexi in an anammox reactor. Water Sci. Technol. 2012, 66, 2256–2261. [Google Scholar] [CrossRef]
- Zhan, X.; Yang, Y.Q.; Chen, F.R. Treatment of secondary effluent by a novel tidal-integrated vertical flow constructed wetland using raw sewage as a carbon source: Contribution of partial denitrification-anammox. Chem. Eng. J. 2020, 395, 125165. [Google Scholar] [CrossRef]
- Yang, X.; Lai, J.L.; Li, J.; Zhang, Y.; Luo, X.G.; Han, M.W.; Zhu, Y.B.; Zhao, S.P. Biodegradation and physiological response mechanism of Bacillus aryabhattai to cyclotetramethylenete-tranitramine (HMX) contamination. J. Environ. Manag. 2021, 288, 112247. [Google Scholar] [CrossRef]
- Li, Y.; Yang, Y.; Feng, Y.R.; Pu, J.; Hou, L.A. Combined effects of Pseudomonas quinolone signal-based quorum quenching and graphene oxide on the mitigation of biofouling and improvement of the application potential for the thin-film composite membrane. Sci. Total Environ. 2021, 760, 143348. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; He, X.; Jia, G. Simultaneous nitrification and denitrification process using novel surface-modified suspended carriers for the treatment of real domestic wastewater. Chemosphere 2021, 247, 125831. [Google Scholar] [CrossRef]
- Liu, T.; Mao, Y.J.; Shi, Y.P.; Xie, Q. Start-up and bacterial community compositions of partial nitrification in moving bed biofilm reactor. Appl. Microbiol. Biotechnol. 2017, 101, 2563–2574. [Google Scholar] [CrossRef] [PubMed]
- Su, L.; Aga, D.; Chandran, K.; Khunjar, W.O. Factors impacting biotransformation kinetics of trace organic compounds in lab-scale activated sludge systems performing nitrification and denitrification. J. Hazard. Mater. 2015, 282, 116–124. [Google Scholar] [CrossRef] [PubMed]
- Wu, P.; Zhang, X.X.; Wang, C.C. Feasibility of applying intermittent aeration and baffles for achieving granular nitritation in a continuous short-cut denitrifying phosphorus removal system. Sci. Total Environ. 2020, 715, 137023. [Google Scholar] [CrossRef] [PubMed]
| Shannon | Simpson | Ace | Chao1 | |
|---|---|---|---|---|
| Day 1average | 2.04 | 0.18 | 42.57 | 40.25 |
| Day 7average | 6.13 | 0.01 | 2638.87 | 2640.29 |
| Day 21average | 5.52 | 0.02 | 2342.74 | 2316.22 |
| Day 30average | 5.78 | 0.02 | 2682.57 | 2667.27 |
| Day 45average | 6.09 | 0.01 | 2644.82 | 2631.45 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.; Lu, S.; Fang, Y.; Yang, K.; Ding, J.; Ye, X.; Zhang, H. Technology for Upgrading the Tailwater of Municipal Sewage Treatment Plants: The Efficacy and Mechanism of Microbial Coupling for Nitrogen and Carbon Removal. Water 2021, 13, 2850. https://doi.org/10.3390/w13202850
Zhang Y, Lu S, Fang Y, Yang K, Ding J, Ye X, Zhang H. Technology for Upgrading the Tailwater of Municipal Sewage Treatment Plants: The Efficacy and Mechanism of Microbial Coupling for Nitrogen and Carbon Removal. Water. 2021; 13(20):2850. https://doi.org/10.3390/w13202850
Chicago/Turabian StyleZhang, Yinan, Shihuan Lu, Yuxin Fang, Kexin Yang, Jiafeng Ding, Xueping Ye, and Hangjun Zhang. 2021. "Technology for Upgrading the Tailwater of Municipal Sewage Treatment Plants: The Efficacy and Mechanism of Microbial Coupling for Nitrogen and Carbon Removal" Water 13, no. 20: 2850. https://doi.org/10.3390/w13202850
APA StyleZhang, Y., Lu, S., Fang, Y., Yang, K., Ding, J., Ye, X., & Zhang, H. (2021). Technology for Upgrading the Tailwater of Municipal Sewage Treatment Plants: The Efficacy and Mechanism of Microbial Coupling for Nitrogen and Carbon Removal. Water, 13(20), 2850. https://doi.org/10.3390/w13202850





