Photocatalytic Bactericidal Performance of LaFeO3 under Solar Light in the Presence of Natural Organic Matter: Spectroscopic and Mechanistic Evaluation
Abstract
:1. Introduction
2. Materials and Methods
2.1. Materials
2.1.1. Photocatalyst Specimen: LaFeO3
2.1.2. NOM Representatives: SRNOM, SRHA, and SRFA
2.1.3. Indicator Organism: E. coli
2.2. Methodology
2.2.1. E. coli Enumeration
2.2.2. Spectroscopic Characterization
2.2.3. Detection of Cellular Contents upon Cell Lysis
2.2.4. Additional Analysis
3. Results
3.1. SPCI in the Presence of Various NOM Analogues Using LF
3.1.1. Bacterial Inactivation Profiles
Mono System Properties
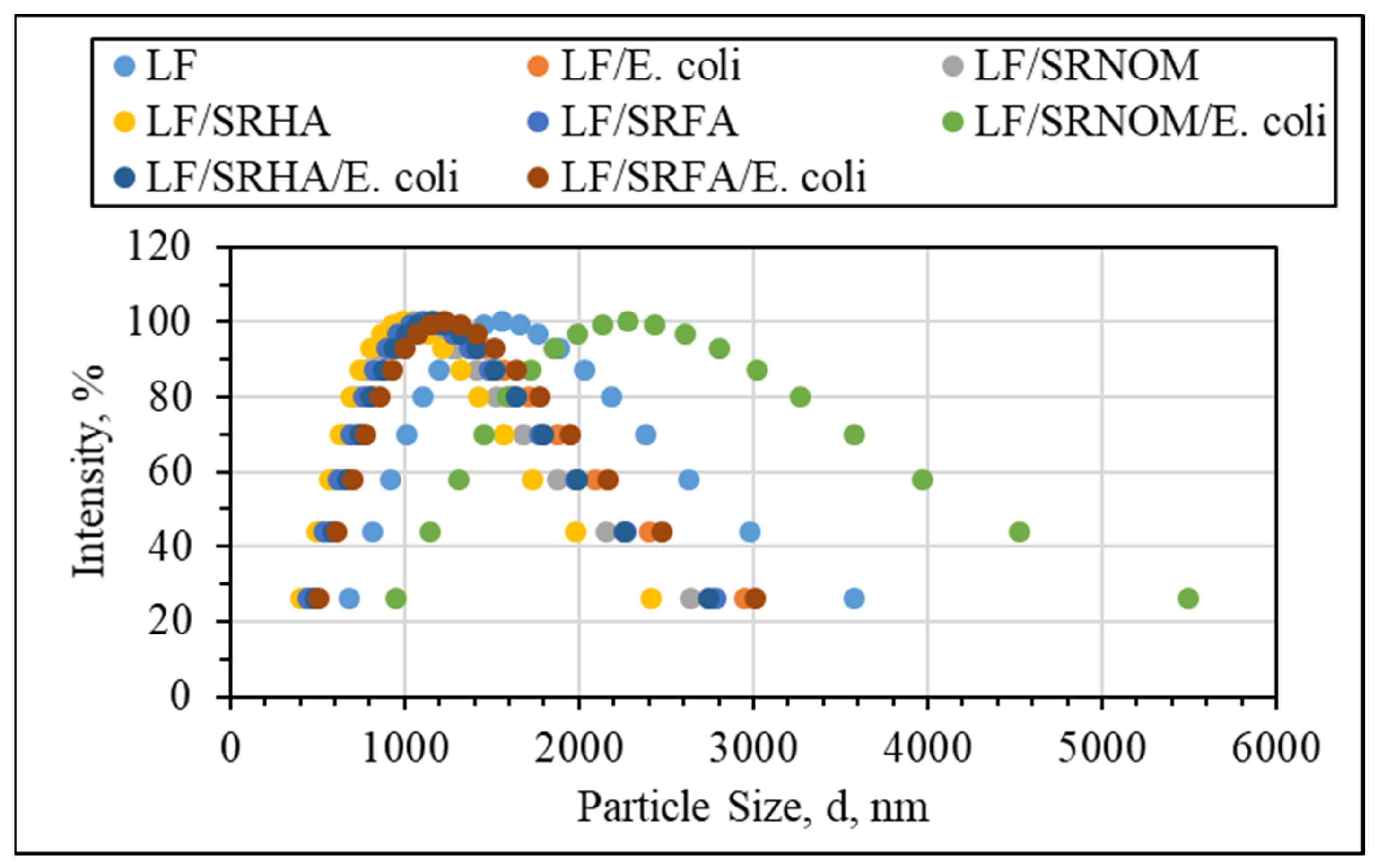
Binary System Interactions
Ternary Interactions Prevailing between E. coli, NOM Analogue Compounds, and LF
3.1.2. Spectroscopic Detection of Cellular Constituents Released upon SPCI
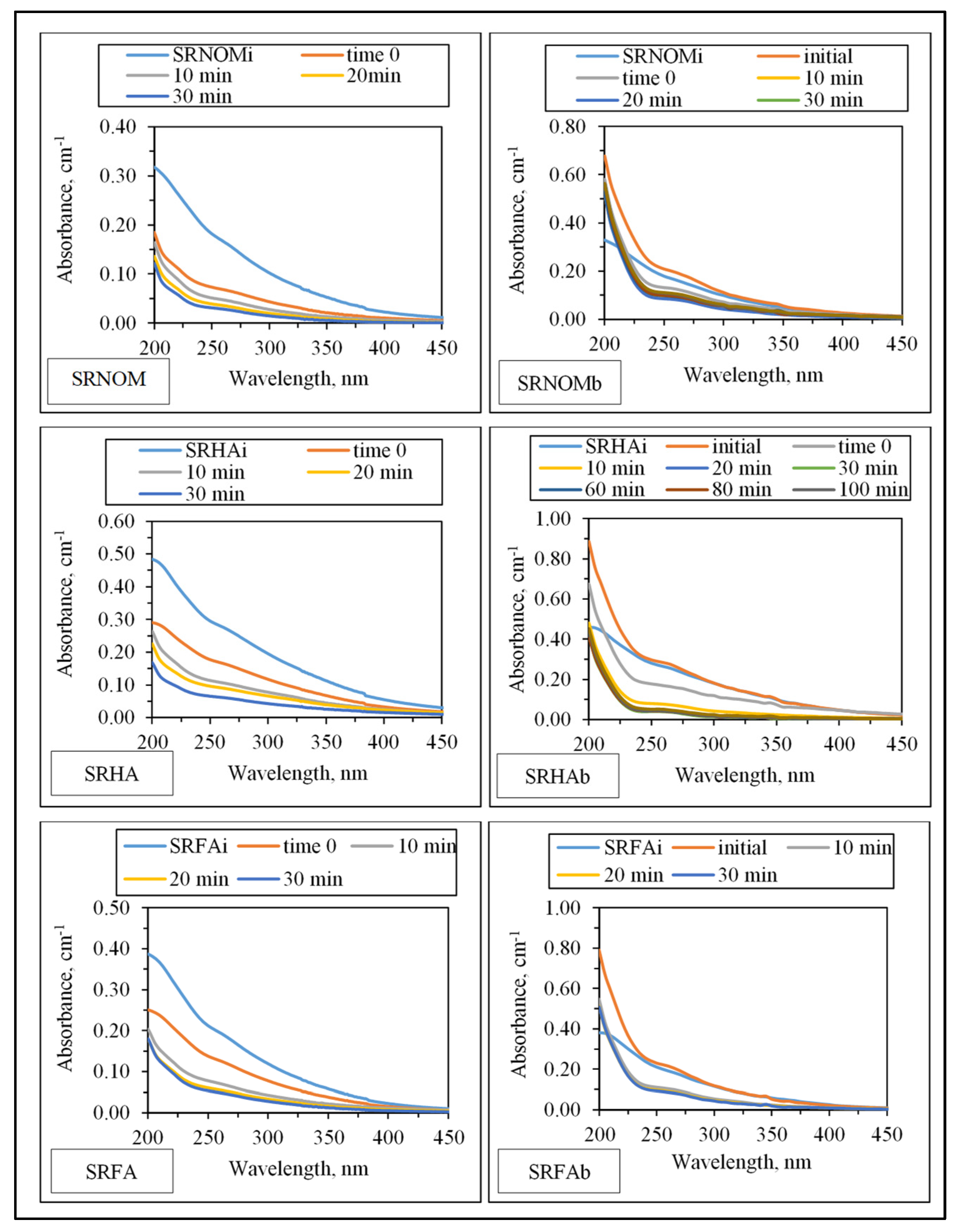
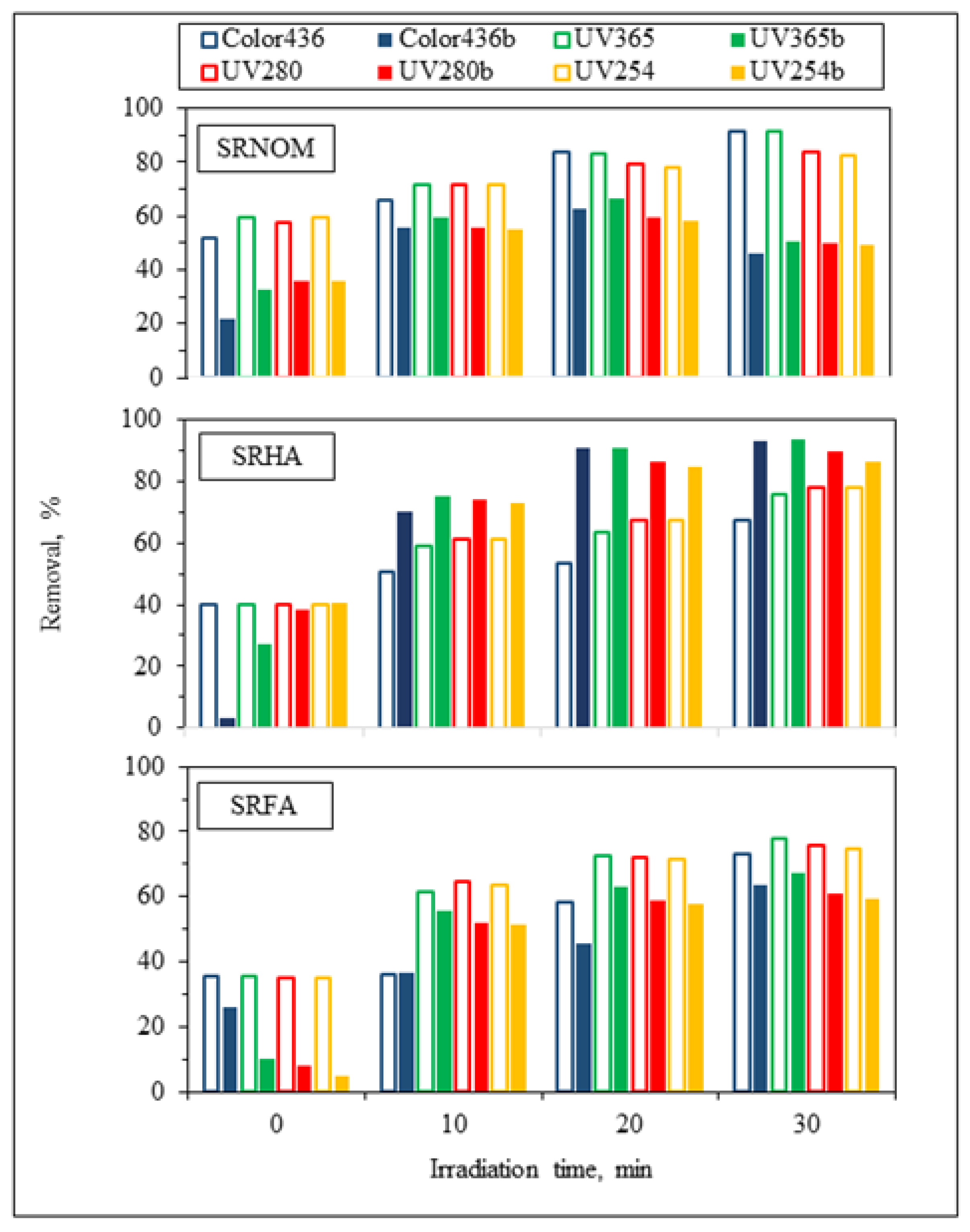
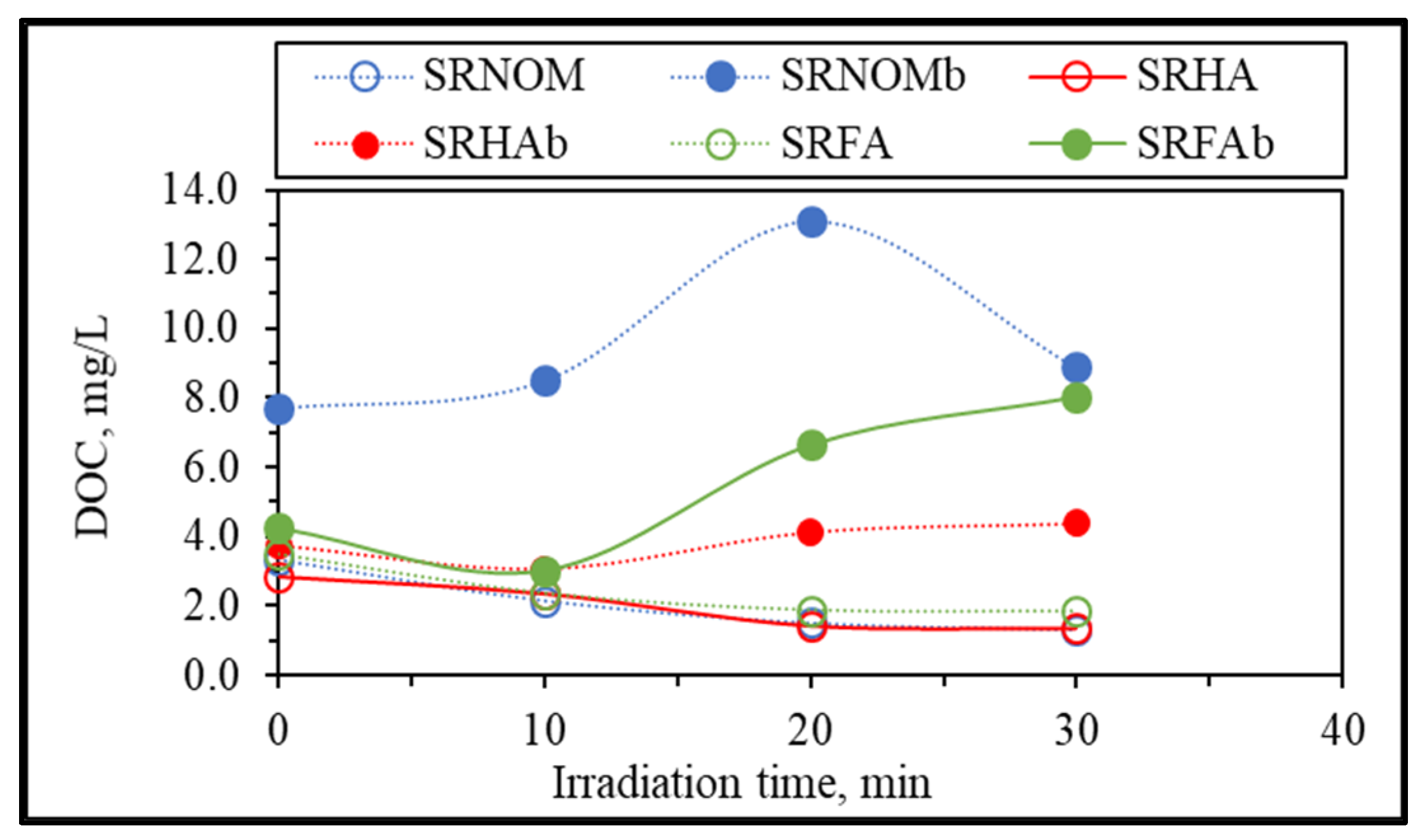
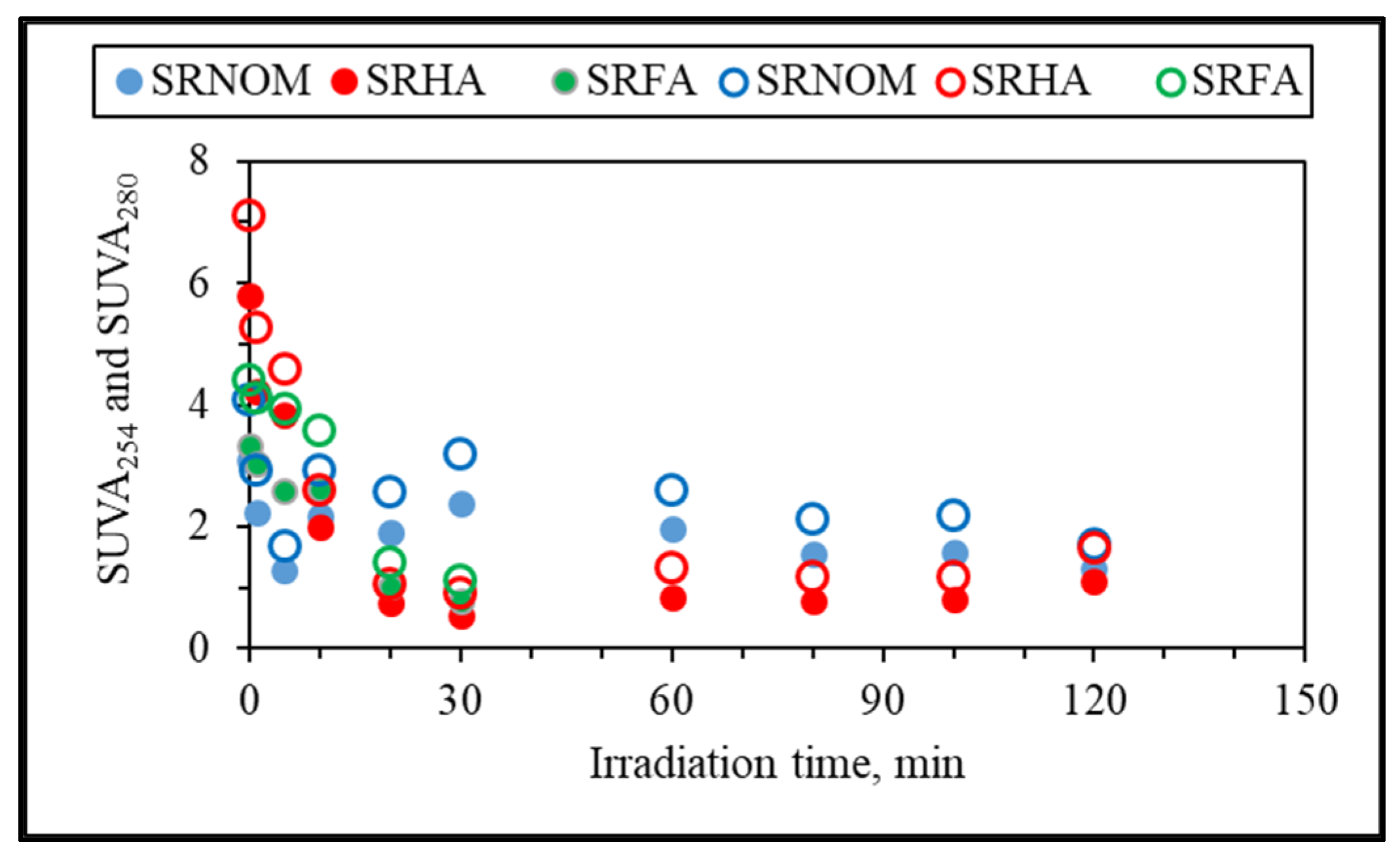
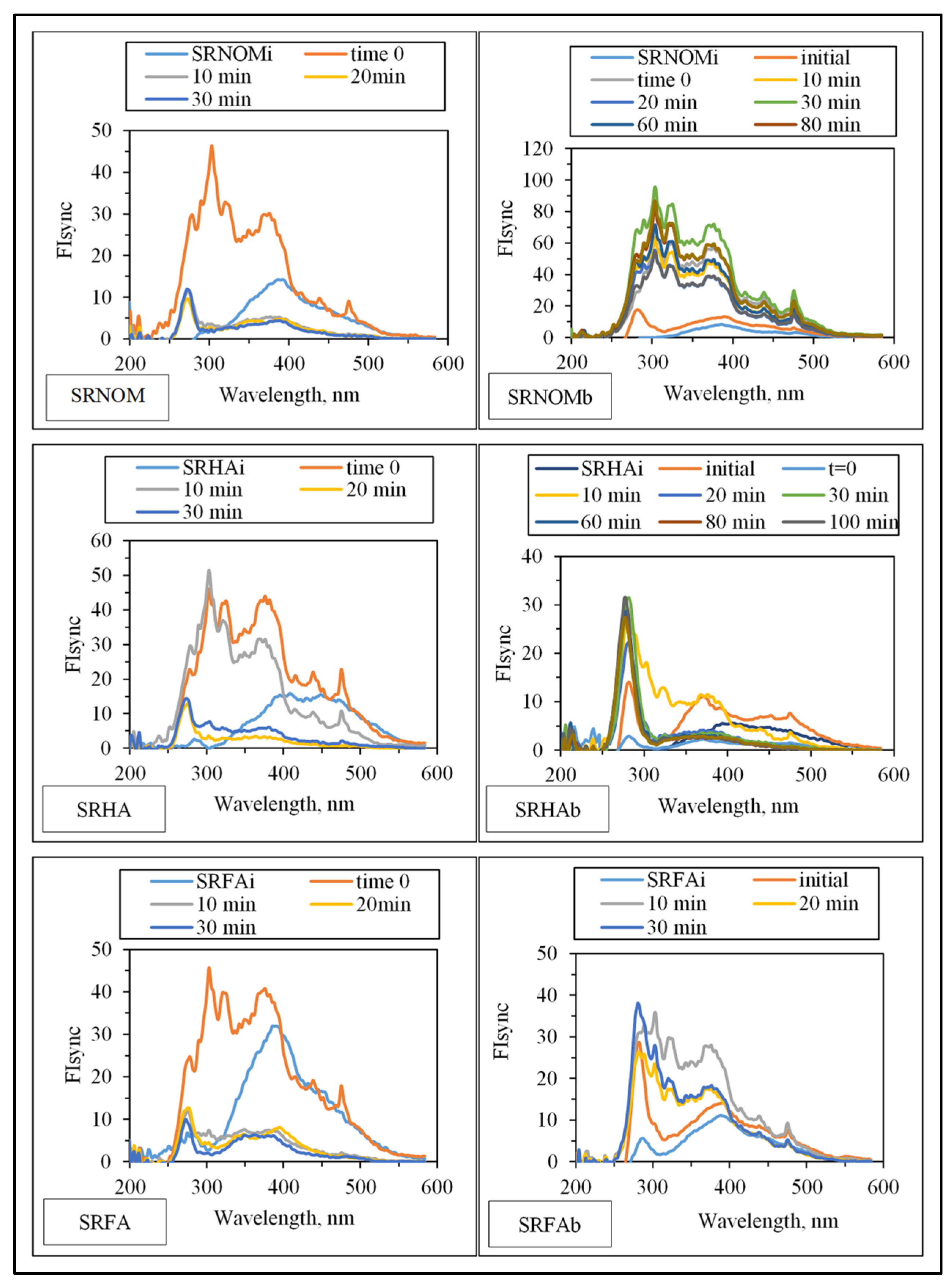
3.1.2.1. UV–VIS and Synchronous Scan Fluorescence Spectroscopic Evaluation
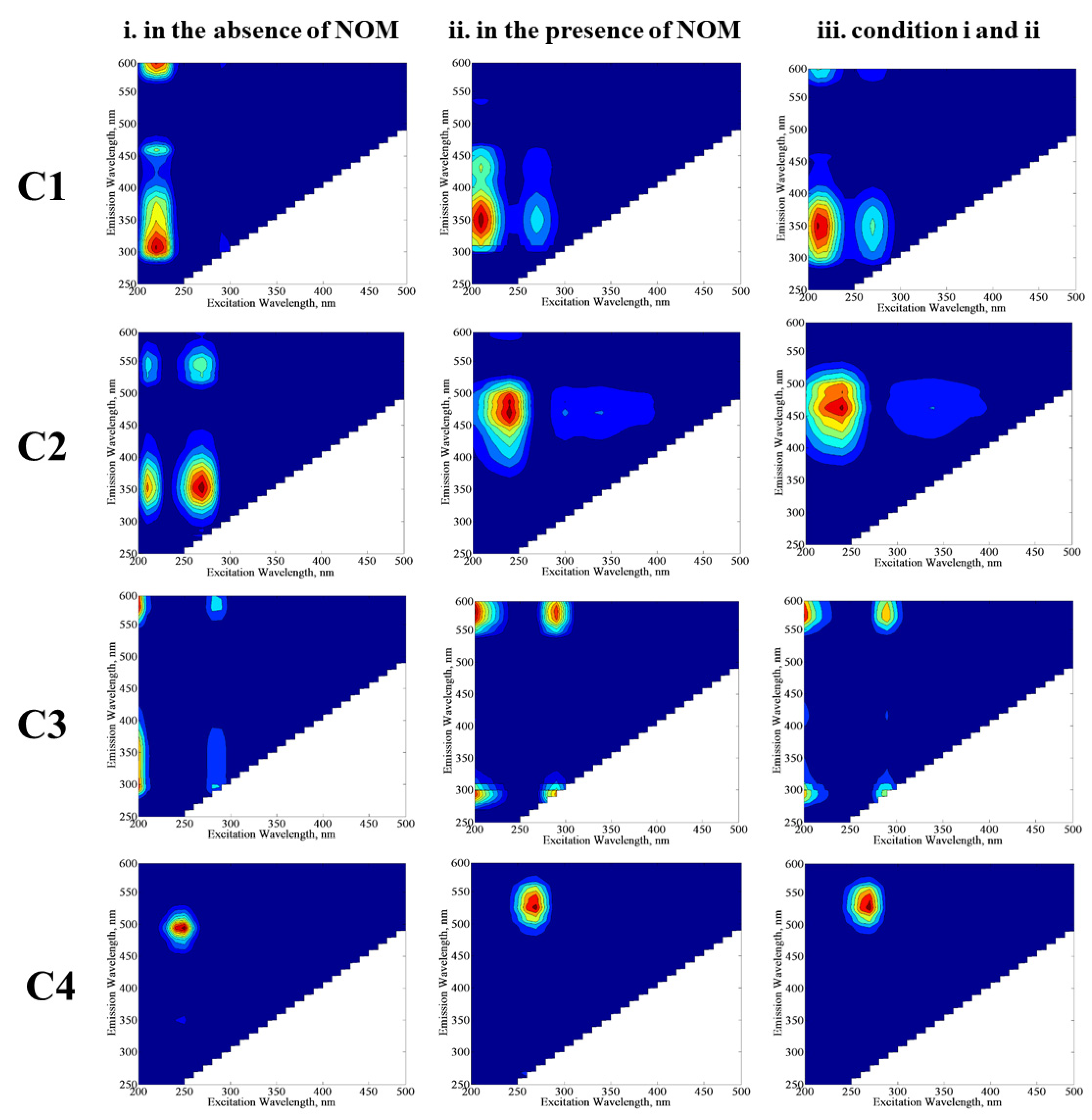
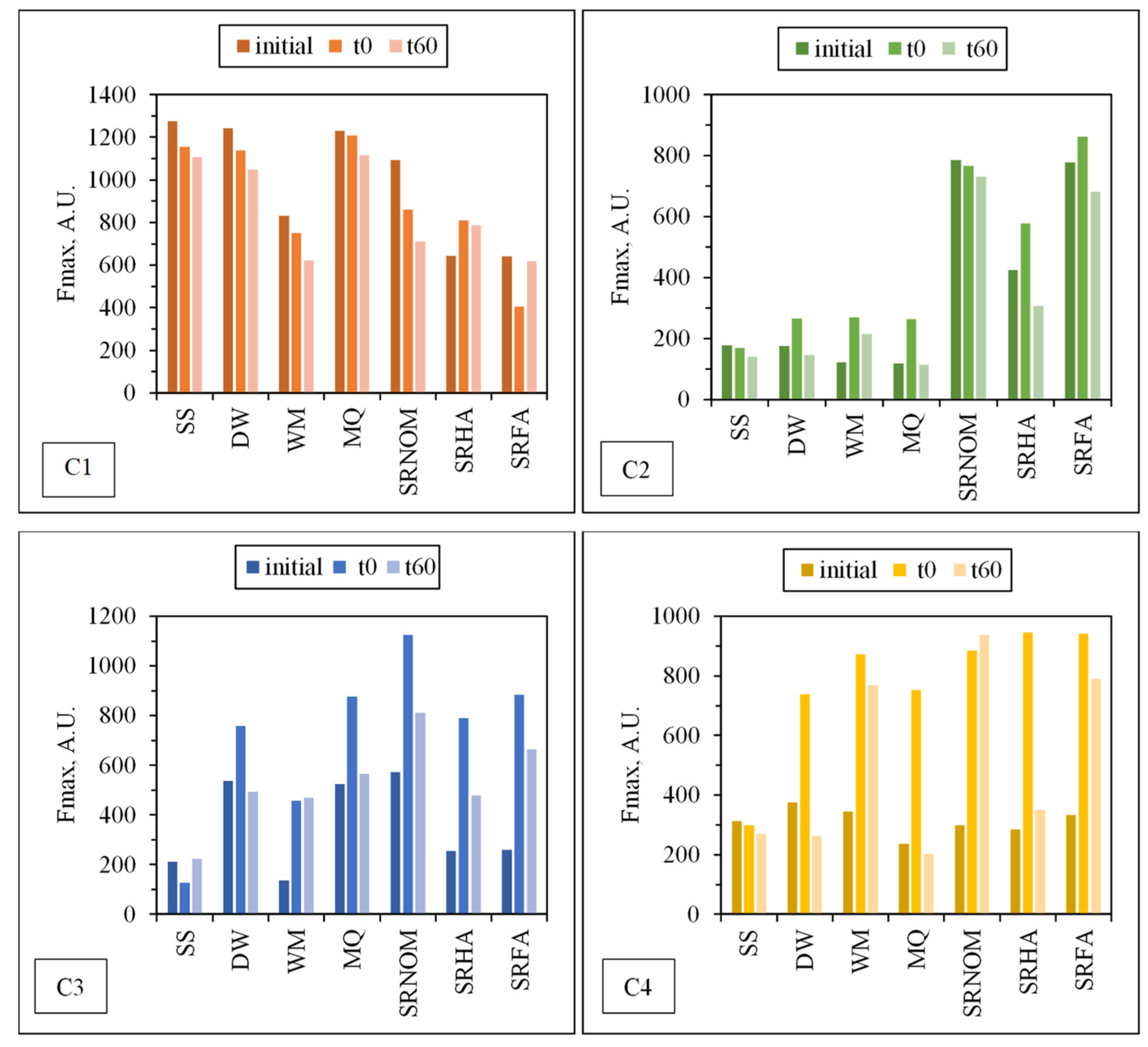
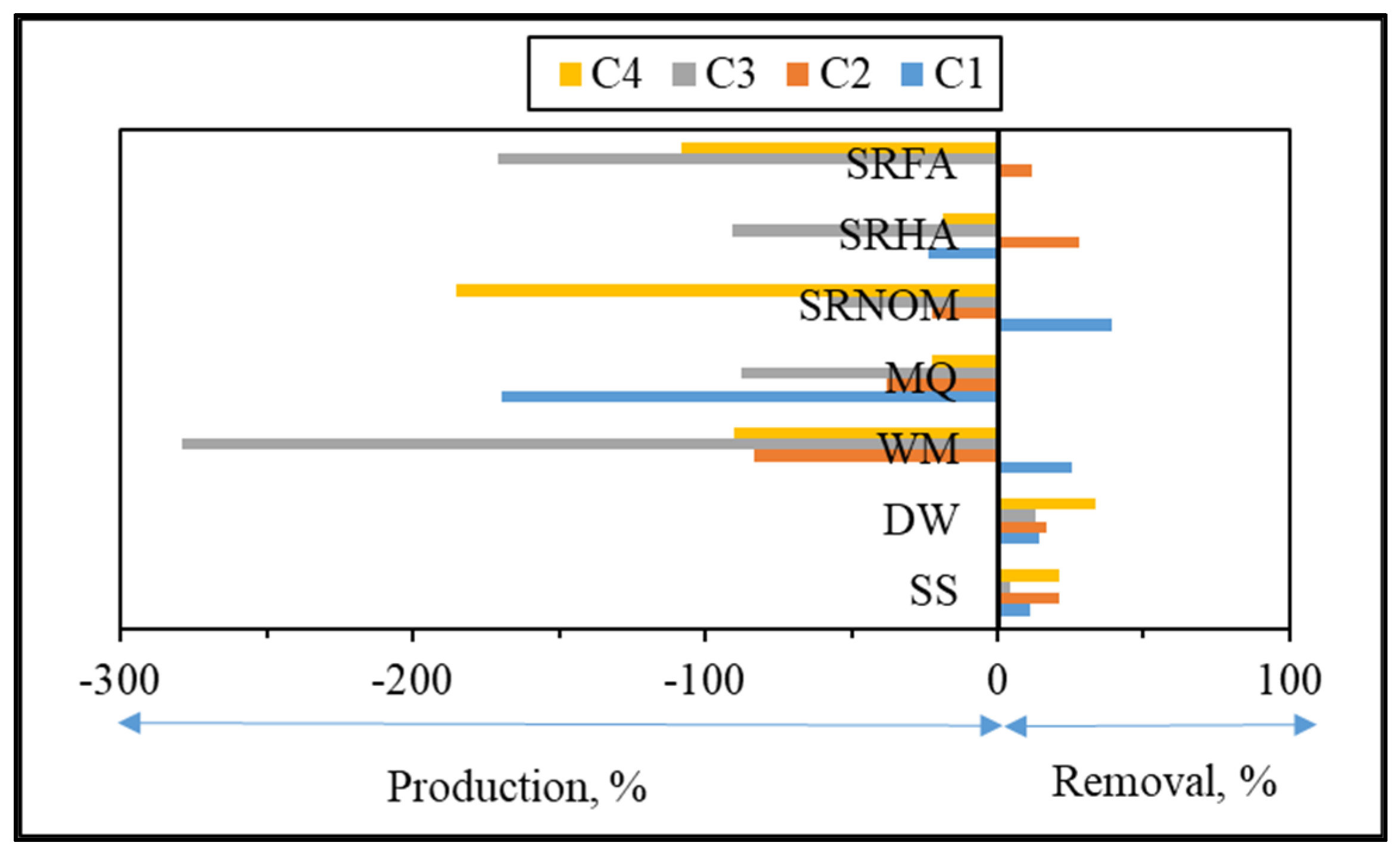
3.1.2.2. EEM Fluorescence Features in Combination with PARAFAC Modeling
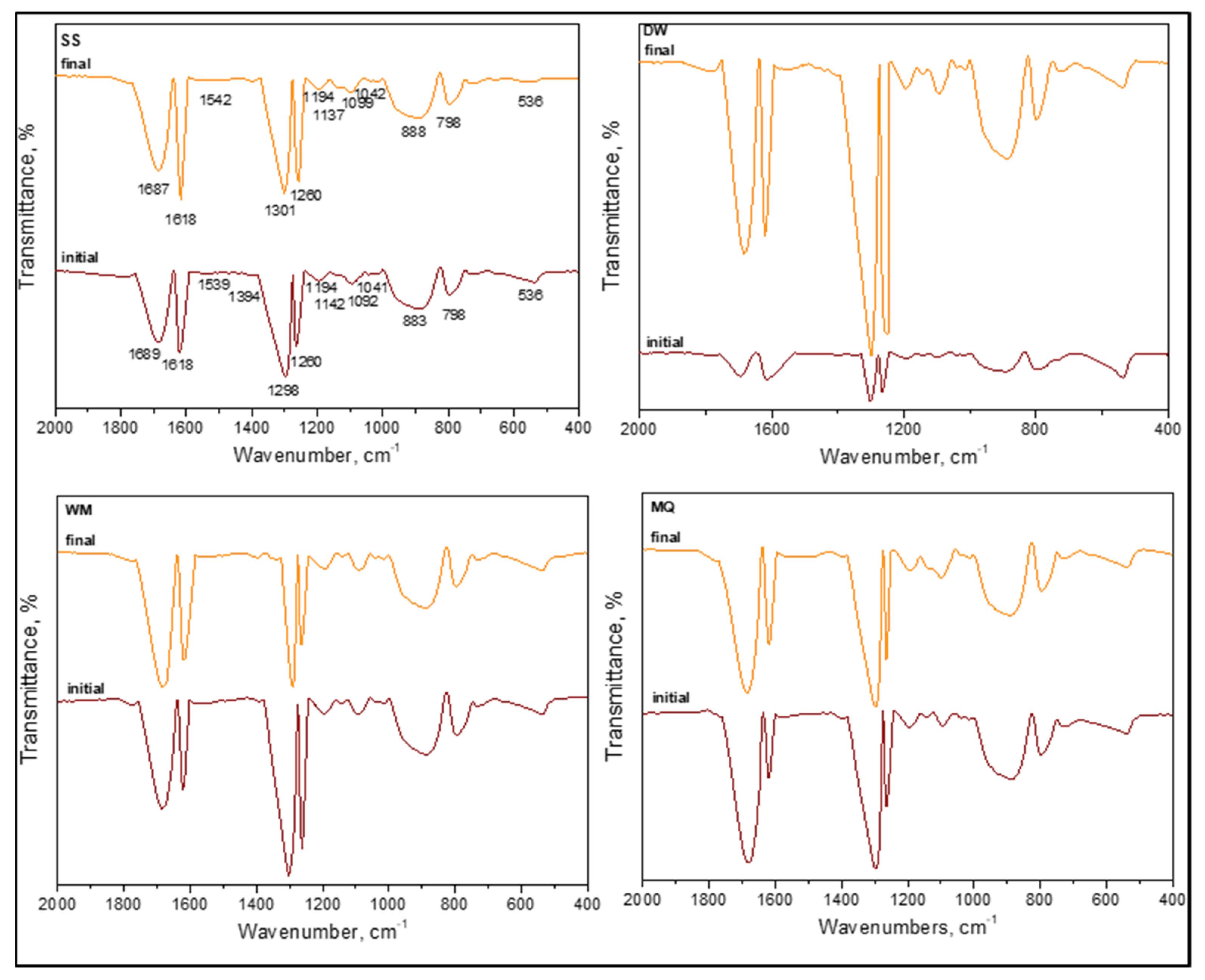
3.1.2.3. FTIR Spectroscopy
3.1.2.4. FTIR Spectral Features of E. coli in the Absence of Organic Matter
3.1.2.5. FTIR Spectral Features of E. coli in the Presence of Organic Matter
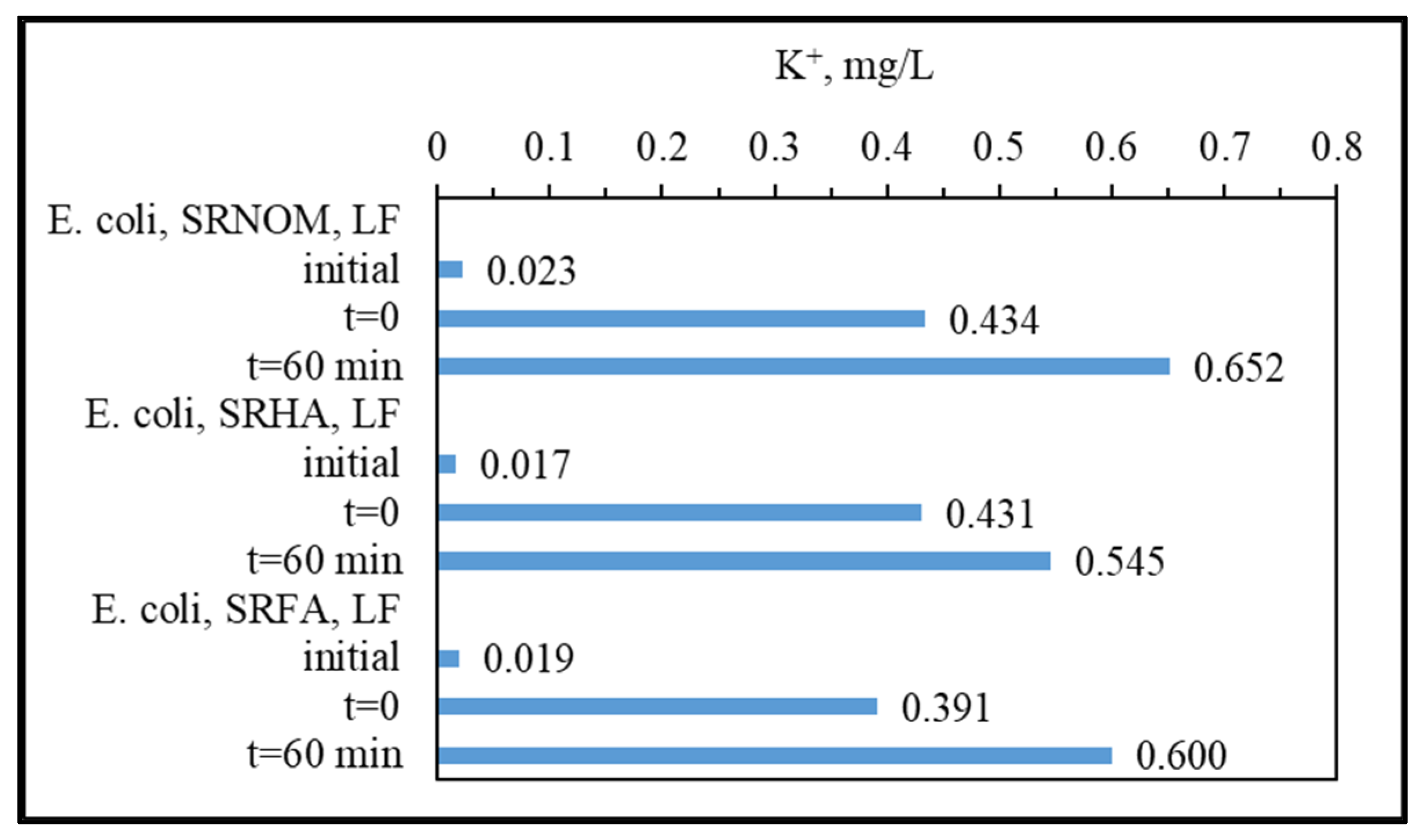
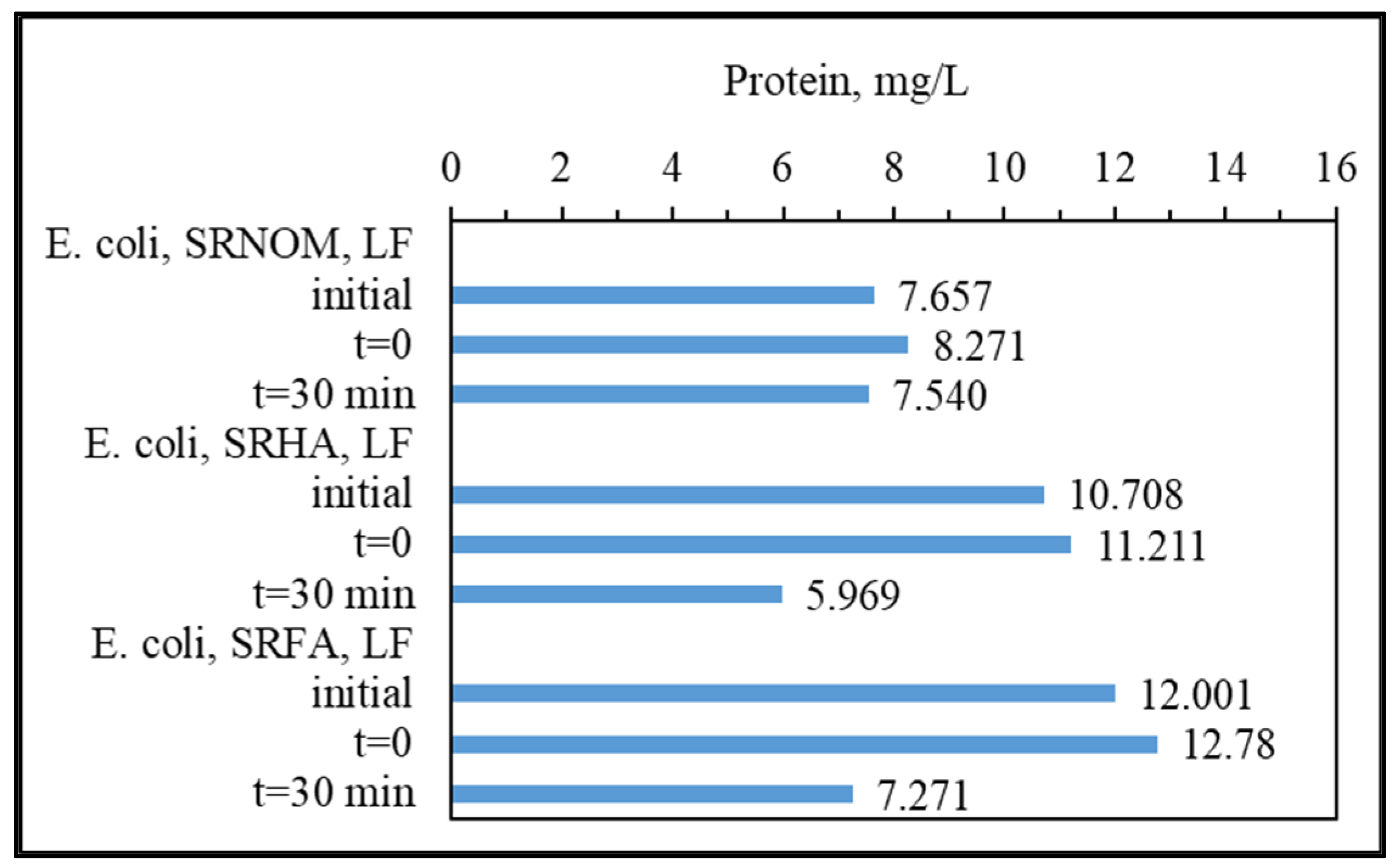
3.1.3. Analytical Detection of Cellular Constituents Released upon SPCI
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AOPs | Advanced oxidation processes |
| C1, C2, C3, and C4 | The excitation and emission maxima values of the components determined by PARAFAC model. |
| CFU | Colony forming units |
| DOC | Dissolved organic carbon |
| DW | Distilled/deionized water |
| E. coli | Escherichia coli |
| EEM | Excitation emission matrix |
| EPS | Extracellular polymeric substance |
| FA | Fulvic acid |
| Fmax | Maximum fluorescence intensity at PARAFAC modeling |
| Fmaxt/Fmaxi | Normalized Fmax intensities of each component. |
| ATR-FTIR | Fourier transform infrared spectroscopy equipped with an attenuated total reflection |
| HA | Humic acid |
| IHSS | International Humic Substances Society |
| LaFeO3 | LF |
| LPS | Lipopolysaccharide |
| MQ | MilliQ water, ultra pure water |
| NOM | Natural organic matter |
| NOM analogues | SRNOM, SRHA, and SRFA |
| PARAFAC | Fluorescence spectra with parallel factor multiway analysis |
| pHzpc | pH of zero point of charge |
| ROS | Reactive oxygen species |
| SEM | Scanning Electron Microscopy |
| SPCI | Solar photocatalytic inactivation |
| SRFA | Suwannee River Fulvic Acid |
| SRFAi | Suwannee River Fulvic Acid referring initial conditions |
| SRHA | Suwannee River Humic Acid |
| SRHAi | Suwannee River Humic Acid referring initial conditions |
| SRNOM | Suwannee River Natural Organic matter |
| SRNOMi | Suwannee River Natural Organic matter referring initial conditions |
| SS | Saline solution |
| SUVA254 | Specific UV–vis parameter represents UV254/DOC |
| SUVA280 | Specific UV–vis parameter represents UV280/DOC |
| WM | Water matrix |
| XPS | X-ray photoelectron spectroscopy |
| XRD | X-ray diffraction |
References
- Oppenländer, T. Photochemical Purification of Water and Air: Advanced Oxidation Processes (AOPs): Principles, Reaction Mechanisms, Reactor Concepts; John Wiley & Sons: Hoboken, NJ, USA, 2007. [Google Scholar]
- Silanpää, M.; Matilainen, A.; Lahtinen, T. Characterization of NOM. In Natural Organic Matter in Water, Characterization and Treatment Methods; Silanpää, M., Ed.; IWA Publishing: London, UK, 2015. [Google Scholar]
- An, T.; Zhao, H.; Wong, P.K. Advances in Photocatalytic Disinfection; Springer: Berlin, Germany, 2017. [Google Scholar]
- Uyguner-Demirel, C.S.; Birben, N.C.; Bekbolet, M. A comprehensive review on the use of second generation TiO2 photocatalysts: Microorganism inactivation. Chemosphere 2018, 211, 420–448. [Google Scholar] [CrossRef]
- Daub, N.A.; Aziz, F.; Aziz, M.; Jaafar, J.; Wan Saleh, W.N.; Yusof, N.; Ismail, A.F. A mini review on parameters affecting the semiconducting oxide photocatalytic microbial disinfection. Water Air Soil Poll. 2020, 231, 1–13. [Google Scholar] [CrossRef]
- Birben, N.C.; Lale, E.; Pelosato, R.; Cozzarini, L.; Uyguner-Demirel, C.S.; Schmid, C.; Natali Sora, I.; Bekbolet, M. Photocatalytic bactericidal performance of LaFeO3 under solar light: Kinetics, spectroscopic and mechanistic evaluation. Water 2021, 13, 1135. [Google Scholar] [CrossRef]
- Turkten, N.; Natali Sora, I.; Tomruk, A.; Bekbolet, M. Photocatalytic degradation of humic acids using LaFeO3. Catalysts 2018, 8, 630. [Google Scholar] [CrossRef] [Green Version]
- Caronna, T.; Fontana, F.; Sora, I.N.; Pelosato, R. Chemical synthesis and structural characterization of the substitution compound LaFe1-xCuxO3 (x = 0 − 0.40). Mater. Chem. Phys. 2009, 116, 645–648. [Google Scholar] [CrossRef]
- Coble, P.G. Characterization of marine and terrestrial NOM in seawater using excitation-emission matrix spectroscopy. Mar. Chem. 1996, 51, 325–346. [Google Scholar] [CrossRef]
- Baker, A.; Tipping, E.; Thacker, A.; Gondar, D. Relating dissolved organic matter fluorescence and functional properties. Chemosphere 2008, 73, 1765–1772. [Google Scholar] [CrossRef] [PubMed]
- Sen-Kavurmaci, S.; Bekbolet, M. Tracing TiO2 photocatalytic degradation of humic acid in the presence of clay particles by excitation–emission matrix (EEM) fluorescence spectra. J. Photoch. Photobio. A 2014, 82, 53–61. [Google Scholar] [CrossRef]
- Stedmon, C.A.; Bro, R. Characterizing dissolved organic matter fluorescence with parallel factor analysis: A tutorial. Limnol. Oceanogr. Meth. 2008, 6, 572–579. [Google Scholar] [CrossRef]
- Darabdhara, G.; Boruah, P.K.; Hussain, N.; Borthakur, P.; Sharma, B.; Sengupta, P.; Das, M.R. Magnetic nanoparticles towards efficient adsorption of Gram positive and Gram negative bacteria: An investigation of adsorption parameters and interaction mechanism. Colloid Surface A 2017, 516, 161–170. [Google Scholar] [CrossRef]
- Harden, V.P.; Harris, J.O. The isoelectric point of bacterial cells. J. Bacteriol. 1953, 65, 198–202. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- IHSS. Available online: https://humic-substances.org/ (accessed on 3 March 2021).
- Parrino, F.; García-López, E.; Marcì, G.; Palmisano, L.; Felice, V.; Sora, I.N.; Armelao, L. Cu-substituted lanthanum ferrite perovskites: Preparation, characterization and photocatalytic activity in gas-solid regime under simulated solar light irradiation. J. Alloy. Comp. 2016, 682, 686–694. [Google Scholar] [CrossRef]
- Chowdhury, I.; Cwiertny, D.M.; Walker, S.L. Combined factors influencing the aggregation and deposition of nano-TiO2 in the presence of humic acid and bacteria. Environ. Sci. Technol. 2012, 46, 6968–6976. [Google Scholar] [CrossRef] [PubMed]
- Elbourne, A.; Chapman, J.; Gelmi, A.; Cozzolino, D.; Crawford, R.J.; Truong, V.K. Bacterial-nanostructure interactions: The role of cell elasticity and adhesion forces. J. Colloid Interf. Sci. 2019, 546, 192–210. [Google Scholar] [CrossRef] [PubMed]
- Natali Sora, I.; Fontana, F.; Passalacqua, R.; Ampelli, C.; Perathoner, S.; Centi, G.; Parrino, F.; Palmisano, L. Photoelectrochemical properties of doped lanthanum orthoferrites. Electrochim. Acta 2013, 109, 710–715. [Google Scholar] [CrossRef] [Green Version]
- Lin, N.; Berton, P.; Moraes, C.; Rogers, R.D.; Tufenkji, N. Nanodarts, nanoblades, and nanospikes: Mechano-bactericidal nanostructures and where to find them. Adv. Colloid Interfac. 2018, 252, 55–68. [Google Scholar] [CrossRef] [Green Version]
- Cantwell, R.E.; Hofmann, R.; Templeton, M.R. Interactions between humic matter and bacteria when disinfecting water with UV light. J. Appl. Microbiol. 2008, 105, 25–35. [Google Scholar] [CrossRef] [Green Version]
- Albanese, A.; Walkey, C.D.; Olsen, J.B.; Guo, H.; Emili, A.; Chan, W.C.W. Secreted biomolecules alter the biological identity and cellular interactions of nanoparticles. ACS Nano 2014, 8, 5515–5526. [Google Scholar] [CrossRef]
- Wang, Q.; Kang, F.; Gao, Y.; Mao, X.; Hu, X. Sequestration of nanoparticles by an EPS matrix reduces the particle specific bactericidal activity. Sci. Rep. 2016, 6, 1–10. [Google Scholar] [CrossRef] [Green Version]
- Baalousha, M.; Afshinnia, K.; Guo, L. Natural organic matter composition determines the molecular nature of silver nanomaterial-NOM corona. Environ. Sci. Nano 2018, 5, 868–881. [Google Scholar] [CrossRef]
- Muela, A.; Garcia-Bringas, J.M.; Arana, I.; Barcina, I. Humic materials offer photoprotective effect to Escherichia coli exposed to damaging luminous radiation. Microb. Ecol. 2000, 40, 336–344. [Google Scholar] [CrossRef]
- Ateia, M.; Ran, J.; Fujii, M.; Yoshimura, C. The relationship between molecular composition and fluorescence properties of humic substances. Int. J. Environ. Sci. Technol. 2017, 14, 867–880. [Google Scholar] [CrossRef]
- Awfa, D.; Ateia, M.; Fujii, M.; Yoshimura, C. Photocatalytic degradation of organic micropollutants: Inhibition mechanisms by different fractions of natural organic matter. Water Res. 2020, 174, 115643. [Google Scholar] [CrossRef]
- Guerard, J.J.; Miller, P.L.; Trouts, T.D.; Chin, Y.-P. The role of fulvic acid composition in the photosensitized degradation of aquatic contaminants. Aquat. Sci. 2009, 71, 160–169. [Google Scholar] [CrossRef]
- Uyguner-Demirel, C.S.; Bekbolet, M. Significance of analytical parameters for the understanding of natural organic matter in relation to photocatalytic oxidation. Chemosphere 2011, 84, 1009–1031. [Google Scholar] [CrossRef] [PubMed]
- Nelson, D.L.; Cox, M.M. Lehninger Principles of Biochemistry, 4th ed.; W.H. Freeman: New York, NY, USA, 2004. [Google Scholar]
- Madigan, M.T.; Martinko, J.M. Brock Biology of Microorganisms, 11th ed.; Pearson Education, Inc: Upper Saddle River, NJ, USA, 2006. [Google Scholar]
- Murphy, K.R.; Hambly, A.; Singh, S.; Henderson, R.K.; Baker, A.; Stuetz, R.; Khan, S.J. Organic matter fluorescence in municipal water recycling schemes: Toward a unified PARAFAC model. Environ. Sci. Technol. 2011, 45, 2909–2916. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Hur, J.; Zhuang, W. Occurrence and behaviors of fluorescence EEM-PARAFAC components in drinking water and wastewater treatment systems and their applications: A review. Environ. Sci. Poll. Res. 2015, 22, 6500–6510. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Wang, Z.; Wang, C.; Li, X.; Wang, C.-C. Photocatalytic degradation of NOM in urban stormwater runoff with TiO2 nanoparticles under UV light irradiation: EEM-PARAFAC analysis and influence of co-existing inorganic ions. Environ. Pollut. 2018, 243, 177–188. [Google Scholar] [CrossRef]
- Turong, H.B.; Huy, B.T.; Ly, Q.V.; Lee, Y.-I.; Hur, J. Visible light-activated degradation of natural organic matter (NOM) using zinc-bismuth oxides-graphitic carbon nitride (ZBO-CN) photocatalyst: Mechanistic insights from EEM-PARAFAC. Chemosphere 2019, 224, 597–606. [Google Scholar] [CrossRef]
- Phong, D.D.; Hur, J. Insight into photocatalytic degradation of dissolved organic matter in UVA/TiO2 systems revealed by fluorescence EEM-PARAFAC. Water Res. 2015, 87, 119–126. [Google Scholar] [CrossRef]
- Ly, Q.V.; Kim, H.-C.; Hur, J. Tracking fluorescent dissolved organic matter in hybrid ultrafiltration systems with TiO2/UV oxidation via EEM-PARAFAC. J. Membrane Sci. 2018, 549, 275–282. [Google Scholar] [CrossRef]
- Zahra, Z.; Maqbool, T.; Arshad, M.; Badshah, M.A.; Choi, H.-K.; Hur, J. Changes in fluorescent dissolved organic matter and their association with phytoavailable phosphorus in soil amended with TiO2 nanoparticles. Chemosphere 2019, 227, 17–25. [Google Scholar] [CrossRef] [PubMed]
- Phong, D.D.; Hur, J. Using two-dimensional correlation size exclusion chromatography (2D-CoSEC) and EEM-PARAFAC to explore the heterogeneous adsorption behavior of humic substances on nanoparticles with respect to molecular sizes. Environ. Sci. Technol. 2018, 16, 427–435. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.Z.; Peleato, N.M.; Legge, R.L.; Andrews, R.C. Towards real-time detection of wastewater in surface waters using fluorescence spectroscopy. J. Environ. Sci. 2019, 86, 195–202. [Google Scholar] [CrossRef]
- Tang, J.; Wu, J.; Li, Z.; Cheng, C.; Liu, B.; Chai, Y.; Wang, Y. Novel insights into variation of fluorescent dissolved organic matters during antibiotic wastewater treatment by excitation emission matrix coupled with parallel factor analysis and cosine similarity assessment. Chemosphere 2018, 210, 843–848. [Google Scholar] [CrossRef] [PubMed]
- Hedges, J.I. Formation and clay mineral reactions of melanoidins. Geochim. Cosmochim. Ac. 1978, 42, 69–76. [Google Scholar] [CrossRef]
- Nagata, T. Production mechanisms of dissolved organic matter. In Microbial Ecology of the Oceans; Kirchman, D.L., Ed.; Wiley Series in Ecological and Applied Microbiology: Hoboken, NJ, USA, 2000; pp. 121–152. [Google Scholar]
- Harvey, G.R.; Boran, D.A.; Chesal, L.A.; Tokar, J.M. The structure of marine fulvic and humic acids. Mar. Chem. 1983, 12, 119–132. [Google Scholar] [CrossRef]
- Harvey, G.R.; Boran, D.A.; Piotrowicz, S.R.; Weisel, C.P. Synthesis of marine humic substances from unsaturated lipids. Nature 1984, 309, 244–246. [Google Scholar] [CrossRef]
- Sharon, M.; Datta, S.; Shah, S.; war Sharon, M.; Soga, T.; Afre, R. Photocatalytic degradation of E. coli and S. aureus by multi walled carbon nanotubes. Carbon Lett. 2007, 8, 184–190. [Google Scholar] [CrossRef]
- Carré, G.; Hamon, E.; Ennahar, S.; Estner, M.; Lett, M.-C.; Horvatovich, P.; Gies, J.-P.; Keller, V.; Keller, N.; Andrea, P. TiO2 photocatalysis damages lipids and proteins in Escherichia coli. Appl. Environ. Microb. 2014, 80, 2573–2581. [Google Scholar] [CrossRef] [Green Version]
- Kiwi, J.; Nadtochenko, V. Evidence for the mechanism of photocatalytic degradation of the bacterial wall membrane at the TiO2 interface by ATR-FTIR and Laser Kinetic Spectroscopy. Langmuir 2005, 21, 4631–4641. [Google Scholar] [CrossRef] [PubMed]
- Nadtochenko, V.A.; Rincon, A.G.; Stanca, S.E.; Kiwi, J. Dynamics of E. coli membrane cell peroxidation during TiO2 photocatalysis studied by ATR-FTIR spectroscopy and AFM microscopy. J. Photoch. Photobio. A 2005, 169, 131–137. [Google Scholar] [CrossRef]
- Rtimi, S.; Sanjines, R.; Andrzejczuk, M.; Pulgarin, C.; Kulik, A.; Kiwi, J. Innovative transparent non-scattering TiO2 bactericide thin films inducing increased E. coli cell wall fluidity. Surf. Coat. Tech. 2014, 254, 333–343. [Google Scholar] [CrossRef]
- Abazari, R.; Sanati, S.; Saghatforoush, L.A. A unique and facile preparation of lanthanum ferrite nanoparticles in emulsion nanoreactors: Morphology, structure, and efficient photocatalysis. Mat. Sci. Semicon. Proc. 2014, 25, 301–306. [Google Scholar] [CrossRef]
- Foster, H.A.; Ditta, I.B.; Varghese, S.; Steele, A. Photocatalytic disinfection using titanium dioxide: Spectrum and mechanism of antimicrobial activity. Appl. Microbiol. Biot. 2011, 90, 1847–1868. [Google Scholar] [CrossRef] [PubMed]
- FTIR, 13C-NMR and Fluorescence Spectra of IHSS Standards. Available online: https://humic-substances.org/ftir-13c-nmr-and-fluorescence-spectra/ (accessed on 3 March 2021).
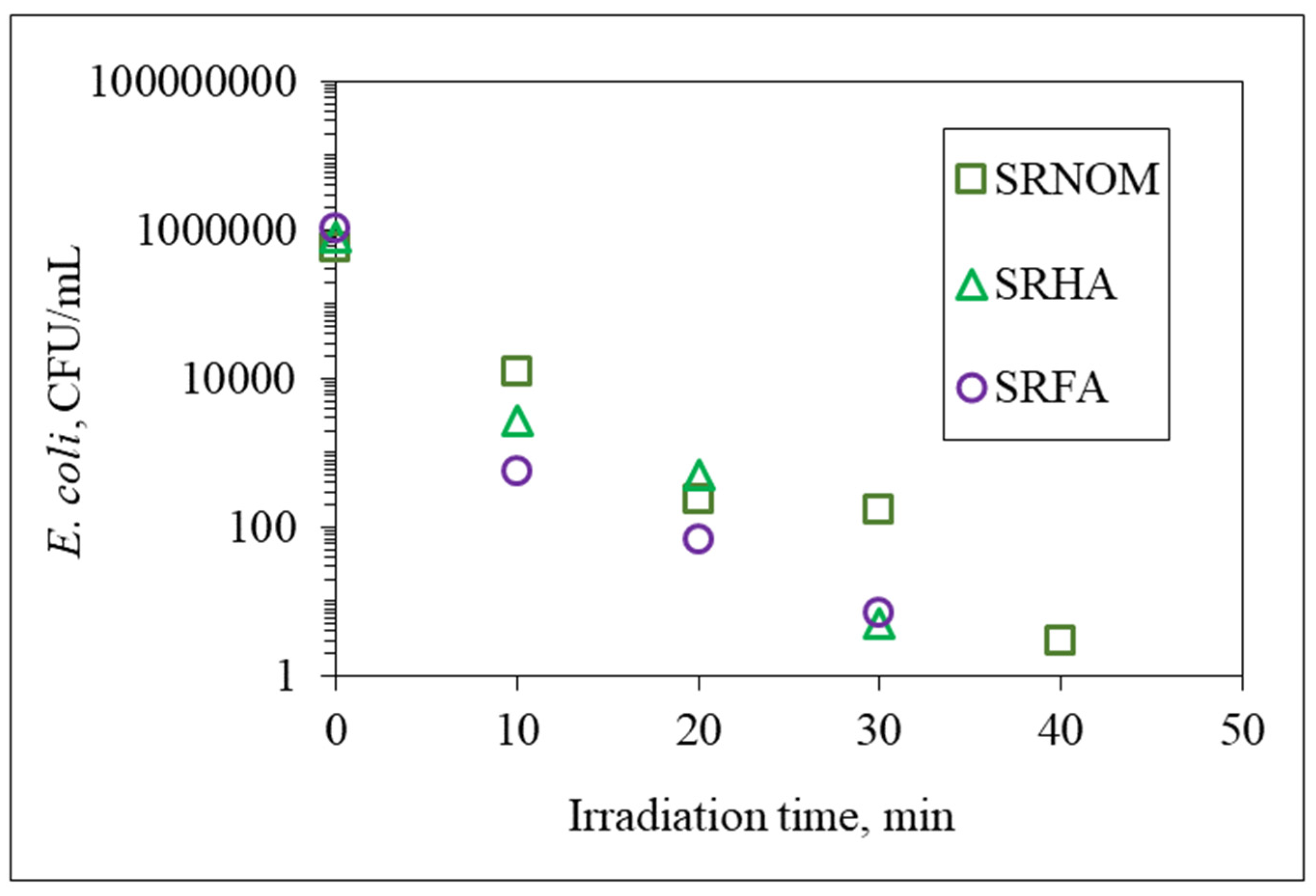

| NOM | k, min−1 | t1/2, min | Rate, CFU/mL min |
|---|---|---|---|
| SRNOM | 0.287 | 2.42 | 2.66 × 105 |
| SRHA | 0.376 | 1.84 | 3.61 × 105 |
| SRFA | 0.379 | 1.83 | 3.98 × 105 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Birben, N.C.; Lale, E.; Pelosato, R.; Turkten, N.; Natali Sora, I.; Bekbolet, M. Photocatalytic Bactericidal Performance of LaFeO3 under Solar Light in the Presence of Natural Organic Matter: Spectroscopic and Mechanistic Evaluation. Water 2021, 13, 2785. https://doi.org/10.3390/w13192785
Birben NC, Lale E, Pelosato R, Turkten N, Natali Sora I, Bekbolet M. Photocatalytic Bactericidal Performance of LaFeO3 under Solar Light in the Presence of Natural Organic Matter: Spectroscopic and Mechanistic Evaluation. Water. 2021; 13(19):2785. https://doi.org/10.3390/w13192785
Chicago/Turabian StyleBirben, Nazmiye Cemre, Ezgi Lale, Renato Pelosato, Nazli Turkten, Isabella Natali Sora, and Miray Bekbolet. 2021. "Photocatalytic Bactericidal Performance of LaFeO3 under Solar Light in the Presence of Natural Organic Matter: Spectroscopic and Mechanistic Evaluation" Water 13, no. 19: 2785. https://doi.org/10.3390/w13192785
APA StyleBirben, N. C., Lale, E., Pelosato, R., Turkten, N., Natali Sora, I., & Bekbolet, M. (2021). Photocatalytic Bactericidal Performance of LaFeO3 under Solar Light in the Presence of Natural Organic Matter: Spectroscopic and Mechanistic Evaluation. Water, 13(19), 2785. https://doi.org/10.3390/w13192785








