Effectiveness of Permeable Reactive Bio-Barriers for Bioremediation of an Organohalide-Polluted Aquifer by Natural-Occurring Microbial Community
Abstract
:1. Introduction
2. Materials and Methods
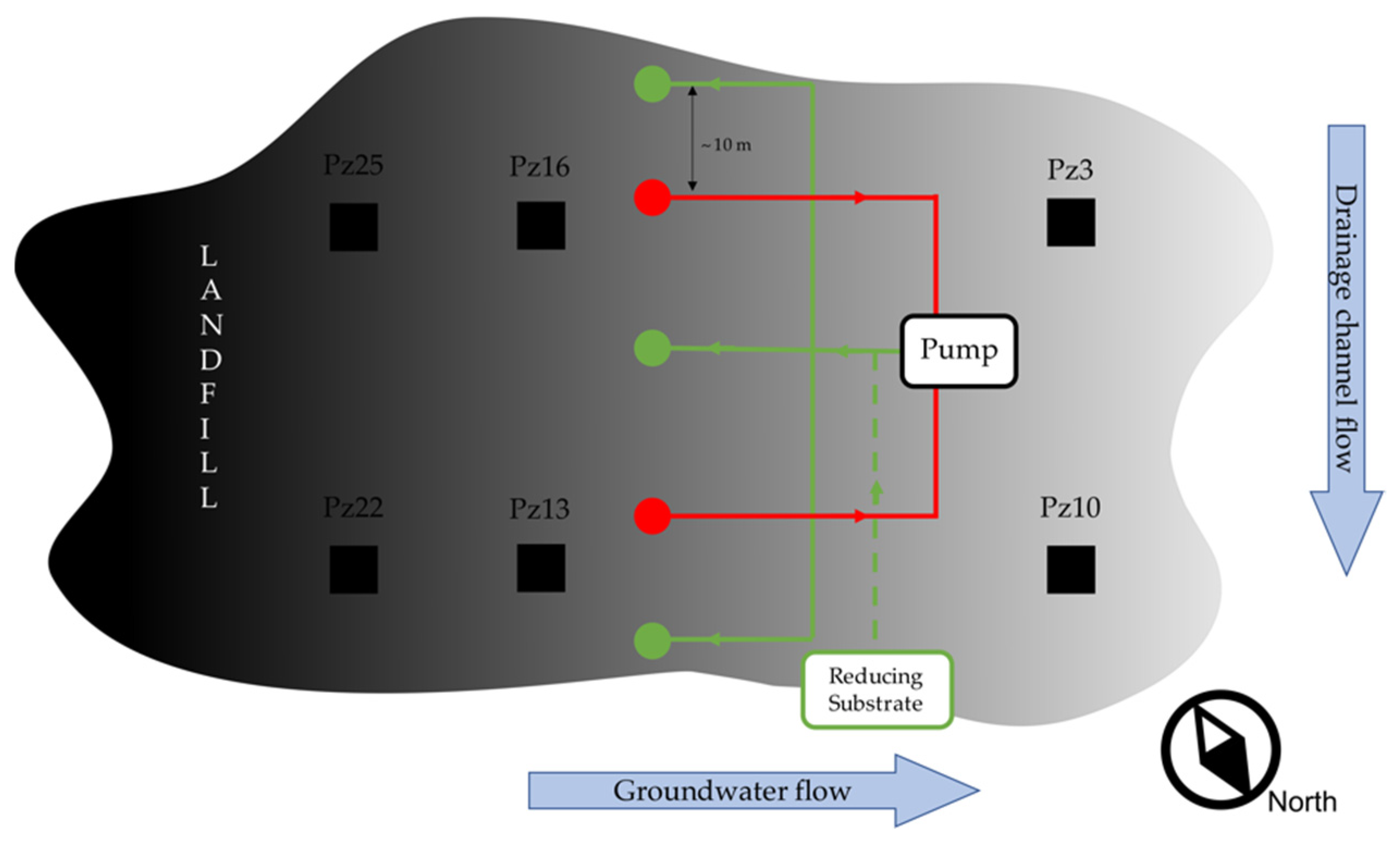
2.1. Site Description and Pilot Scale Experiment
2.2. Microcosm Experiments
2.3. Chemical Methods
2.4. Nucleic Acid Extraction from Groundwater Samples
2.5. Real Time Quantitative PCR
2.6. Illumina MiSeq Sequencing of 16S rRNA Genes
2.7. Statistical Analyses
3. Results
3.1. Landfill Groundwater: Chemical Characterization and OHR Biomarkers
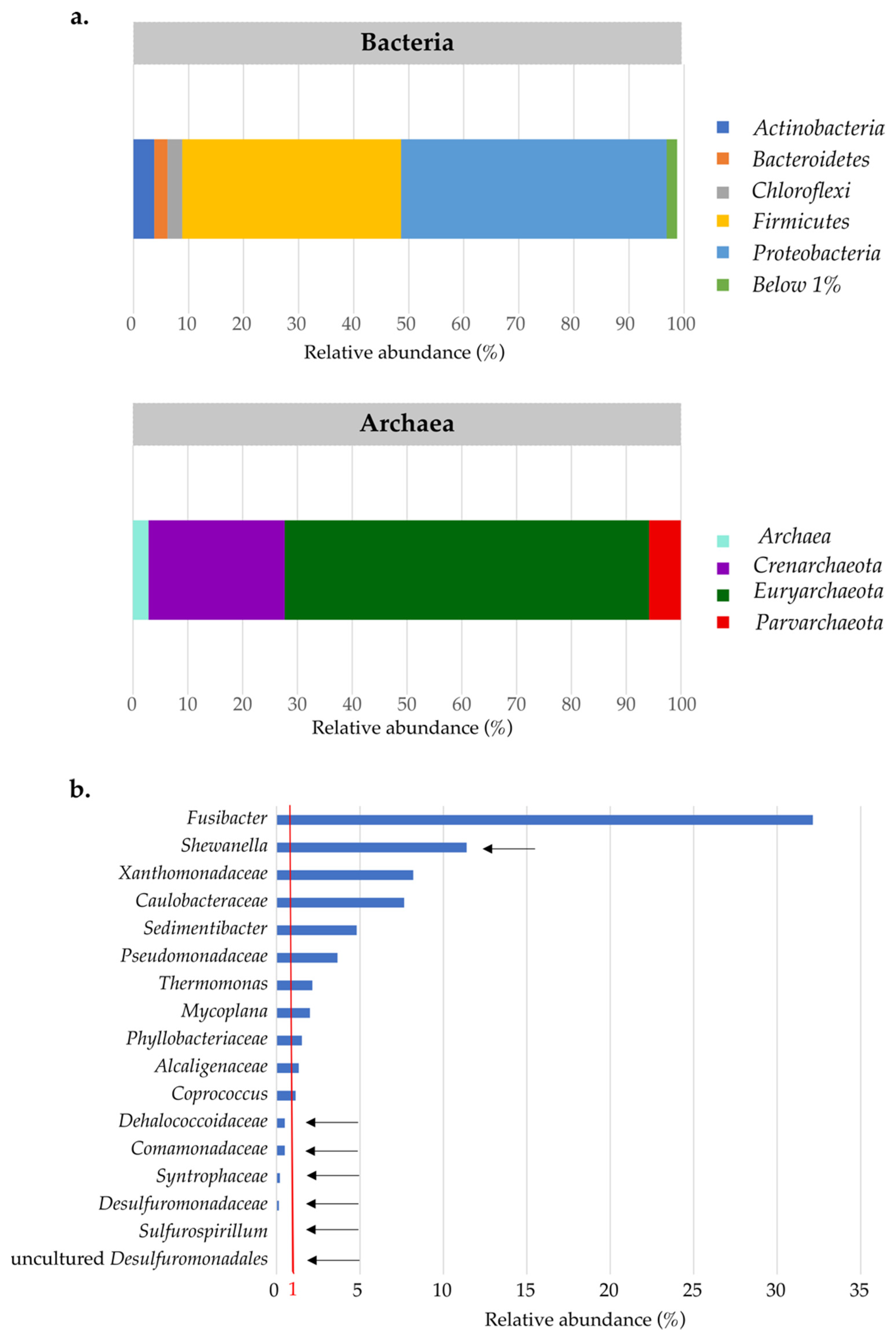
3.2. Active Microbial Community of Landfill Groundwater
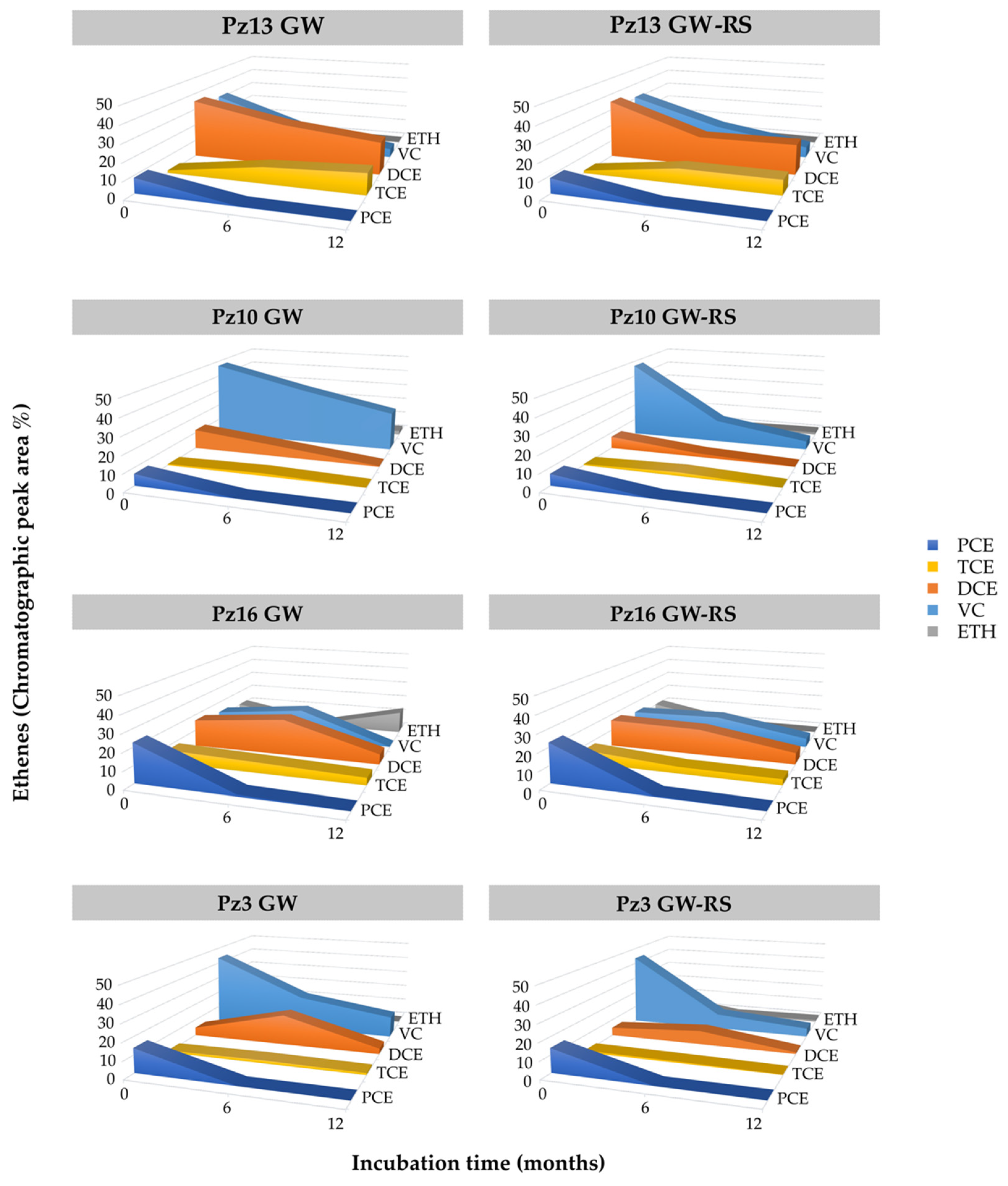
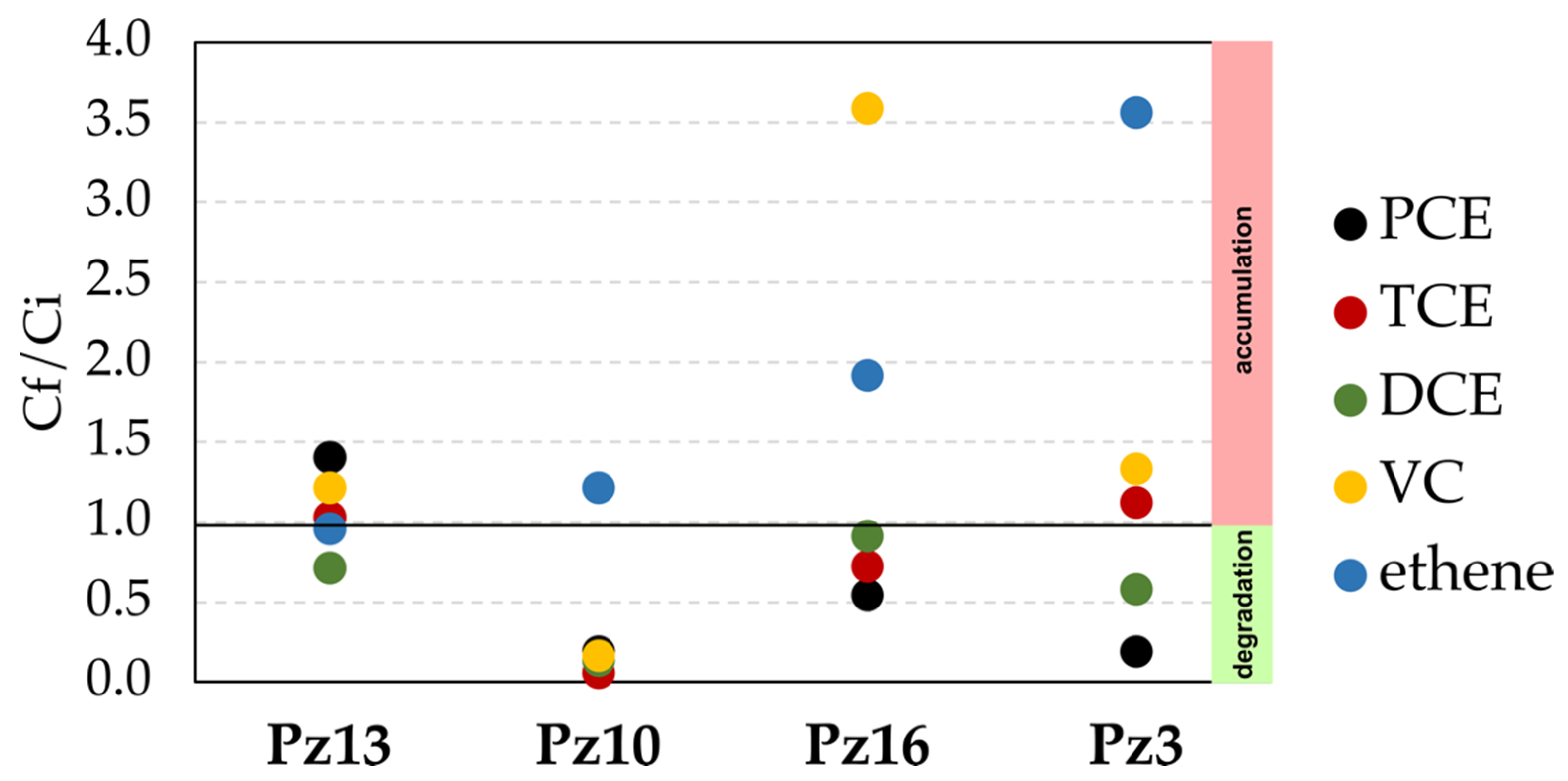
3.3. Determination of Natural OHR in the Aquifer by Microcosm Incubations
3.4. CE Field Monitoring in Pilot-Scale Experiment
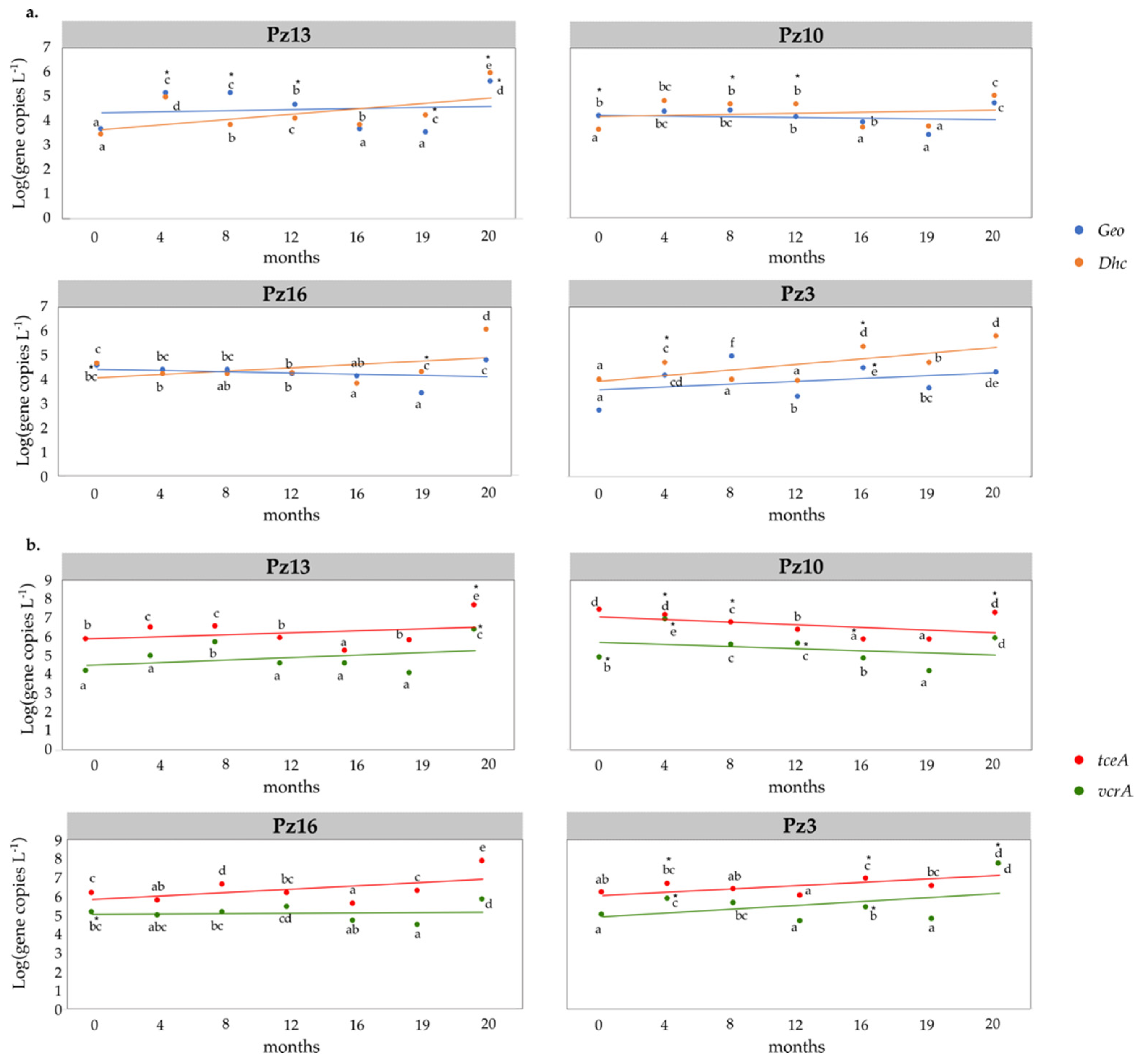
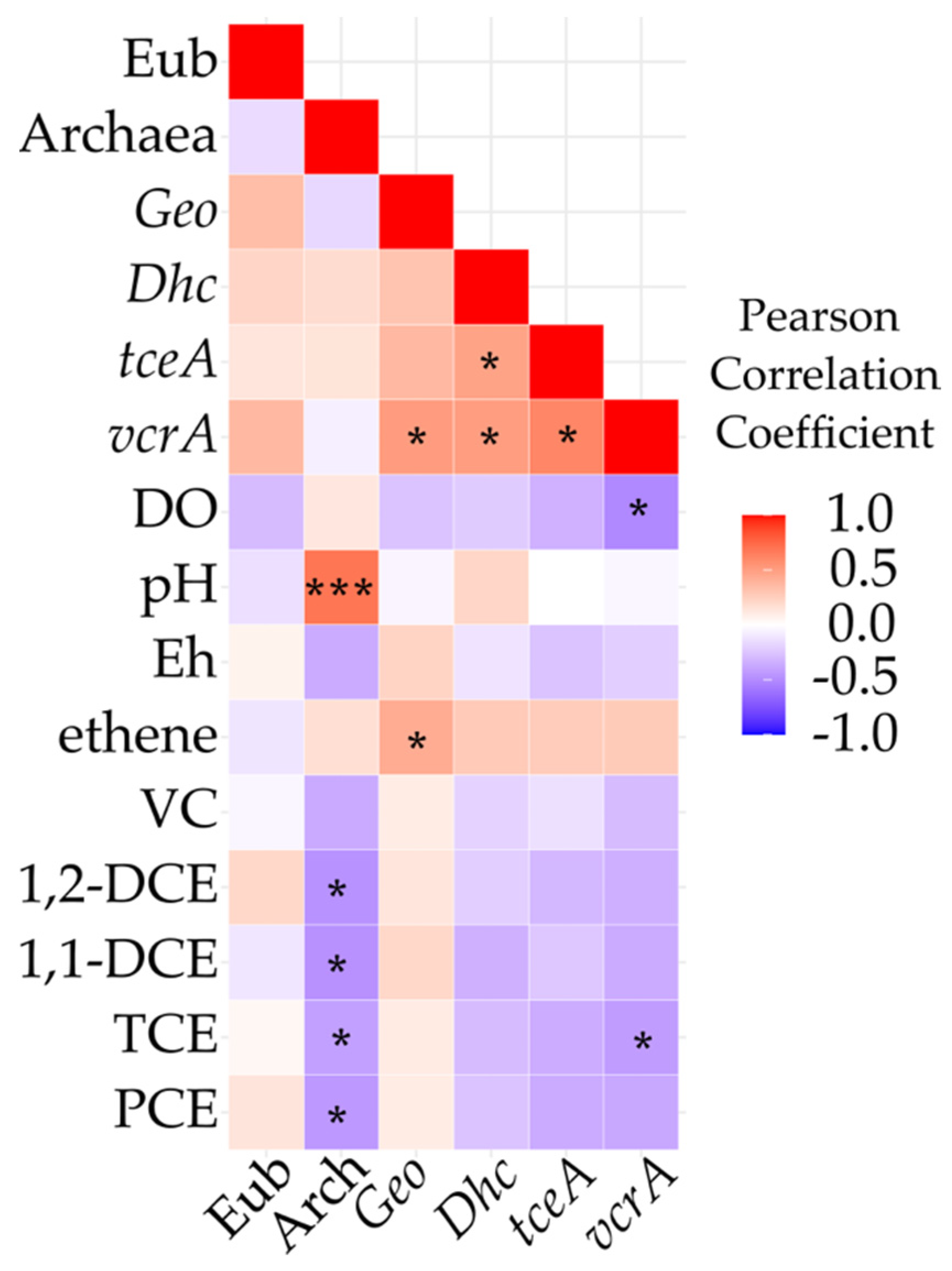
3.5. Field Monitoring of Microbial Populations Involved in OHR
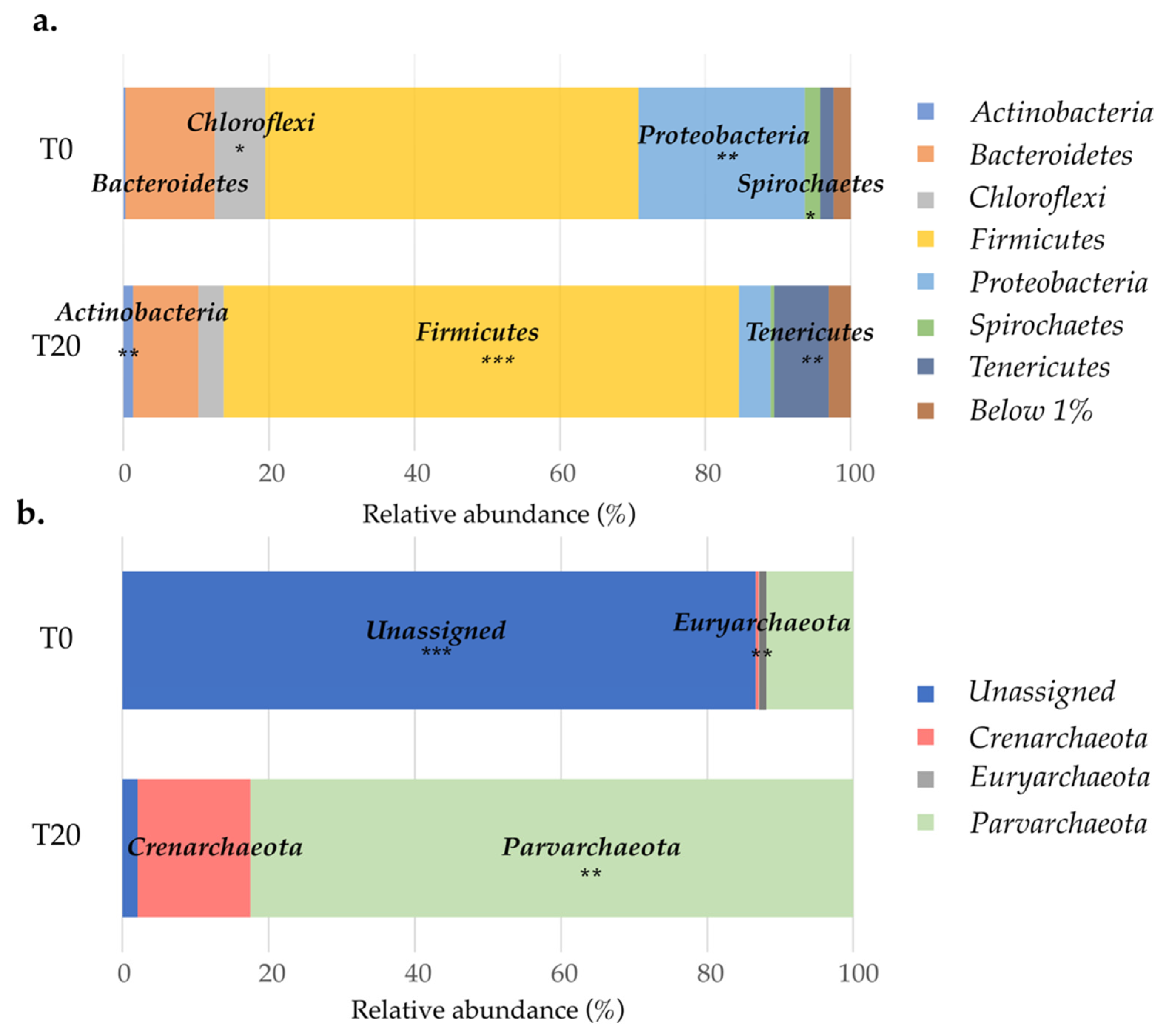
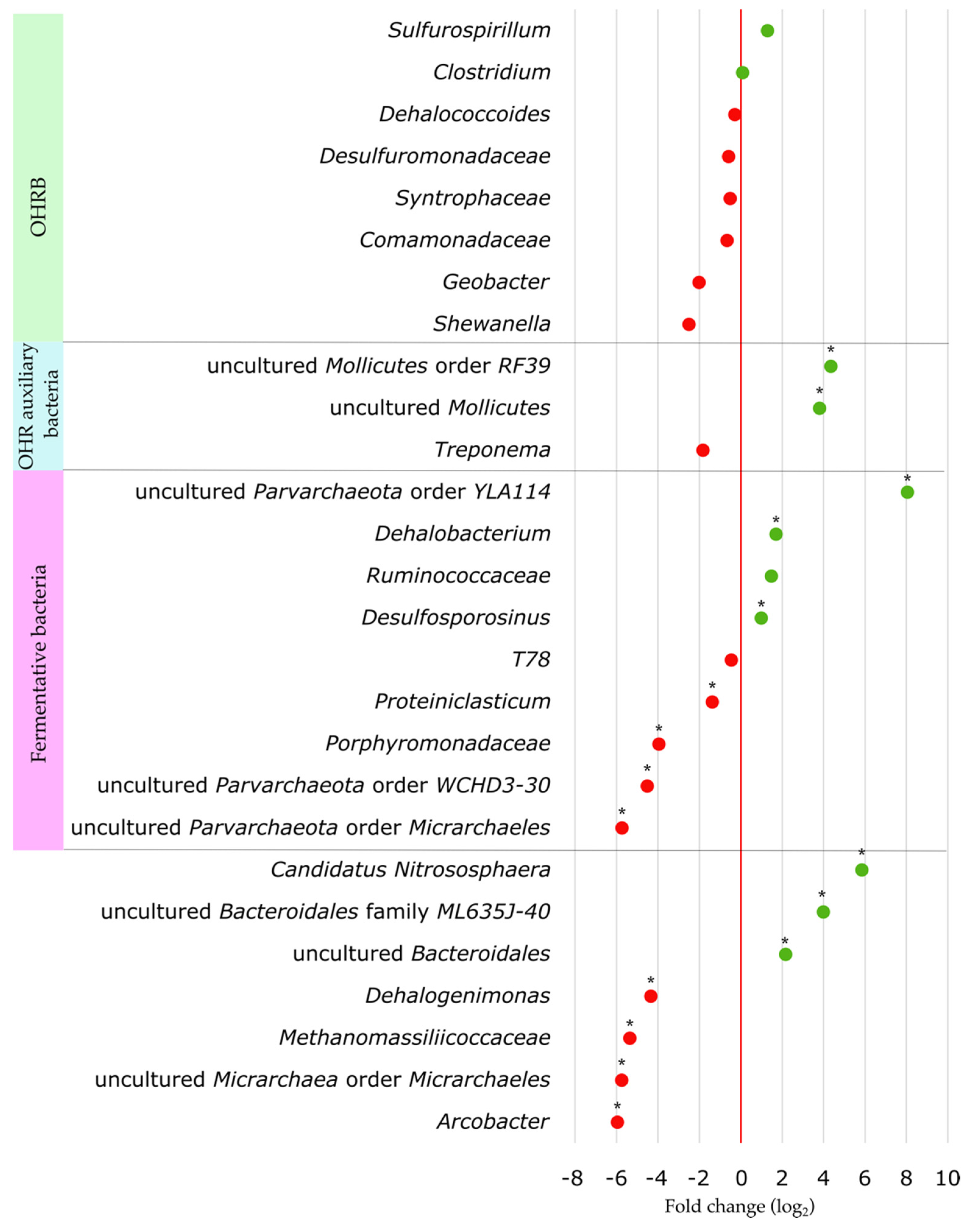
3.6. Effect of Substrate Addition on the Microbial Community of the Aquifer
4. Discussion
4.1. Site Characterization
4.2. Organohalide Respiration Affected by Reducing Substrate
4.3. Impact of the Biostimulation Treatment on the Aquifer Microbial Community
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Moran, M.J.; Zogorski, J.S.; Squillace, P.J. Chlorinated solvents in groundwater of the United States. Environ. Sci. Technol. 2007, 41, 74–81. [Google Scholar] [CrossRef]
- Beamer, P.I.; Luik, C.E.; Abrell, L.; Camposc, S.; Martínez, M.E.; Sáez, A.E. Concentration of Trichloroethylene in Breast Milk and Household Water from Nogales, Arizona. Environ. Microbiol. 2012, 46, 9055–9061. [Google Scholar] [CrossRef] [Green Version]
- Linge, E.; Anttila, A.; Hemminki, K. Organic solvents and cancer. Cancer Causes Control 1992, 55, 1353–1395. [Google Scholar]
- USA EPA. Toxicity and exposure assessment for children’s health. Trichloroethylene—TEACH chemical summary. 2007. Available online: http://www.epa.gov/teach/chem_summ/TCE_summary.pdf (accessed on 15 May 2021).
- IARC. Vinyl Chloride; International Agency for Research on Cancer: Ottawa, ON, Canada, 1987. [Google Scholar]
- Mattes, T.E.; Alexander, A.K.; Coleman, N.V. Aerobic biodegradation of the chloroethenes: Pathways, enzymes, ecology, and evolution. Fems Microbiol. Rev. 2010, 34, 445–475. [Google Scholar] [CrossRef] [Green Version]
- Maymo-Gallett, X.; Chien, Y.; Gossett, J.M.; Zinder, S.H. Isolation of a bacterium that reductively dechlorinates tetrachloroethene to ethene. Science 1997, 276, 1568–1571. [Google Scholar] [CrossRef] [PubMed]
- Cichocka, D.; Nikolausz, M.; Haest, P.J.; Nijenhuis, I. Tetrachloroethene conversion to ethene by a Dehalococcoides-containing enrichment culture from Bitterfeld. FEMS Microbiol. Ecol. 2010, 72, 297–310. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Atashgahi, S.; Lu, Y.; Smidt, H. Overview of known organohaliderespiring bacteria—Phylogenetic diversity and environmental distribution. In Organohalide-Respiring Bacteria; Adrian, L., Löffler, F., Eds.; Springer: Berlin/Heidelberg, Germany, 2016; pp. 63–105. [Google Scholar] [CrossRef]
- DiStefano, T.D.; Gossett, J.M.; Zinder, S.H. Reductive Dechlorination of High Concentrations of Tetrachloroethene to Ethene by an Anaerobic Enrichment Culture in the Absence of Methanogenesis. Appl. Environ. Microbiol. 1991, 57, 2287–2292. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maillard, J.; Schumacher, W.; Vazquez, F.; Regeard, C.; Hagen, W.R.; Holliger, C. Characterization of the Corrinoid Iron-Sulfur Protein Tetrachloroethene Reductive Dehalogenase of Dehalobacter restrictus. Appl. Environ. Microbiol 2003, 69, 4628–4638. [Google Scholar] [CrossRef] [Green Version]
- Pöritz, M.; Goris, T.; Wubet, T.; Tarkka, M.T.; Buscot, F.; Nijenhuis, I.; Lechner, U.; Adrian, L. Genome sequences of two dehalogenation specialists—Dehalococcoides mccartyi strains BTF08 and DCMB5 enriched from the highly polluted Bitterfeld region. Fems Microbiol. Lett. 2013, 343, 101–104. [Google Scholar] [CrossRef] [Green Version]
- Krajmalnik-Brown, R.; Hölscher, T.; Thomson, I.N.; Saunders, F.M.; Ritalahti, K.M.; Löffler, F.E. Genetic identification of a putative vinyl chloride reductase in Dehalococcoides sp. strain BAV1. Appl. Environ. Microbiol. 2004, 70, 6347–6351. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Müller, J.A.; Rosner, B.M.; Von Abendroth, G.; Meshulam-Simon, G.; McCarty, P.L.; Spormann, A.M. Molecular identification of the catabolic vinyl chloride reductase from Dehalococcoides sp. strain VS and its environmental distribution. Appl. Environ. Microbiol. 2004, 70, 4880–4888. [Google Scholar] [CrossRef] [Green Version]
- Yang, Y.; McCarty, P.L. Competition for hydrogen within a chlorinated solvent dehalogenating anaerobic mixed culture. Environ. Sci. Technol. 1998, 32, 3591–3597. [Google Scholar] [CrossRef]
- Chen, L.X.; Méndez-García, C.; Dombrowski, N.; Servín-Garcidueñas, L.E.; Eloe-Fadrosh, E.A.; Fang, B.Z.; Luo, Z.H.; Tan, S.; Zhi, X.Y.; Hua, Z.S.; et al. Metabolic versatility of small archaea Micrarchaeota and Parvarchaeota. ISME J. 2018, 12, 756–775. [Google Scholar] [CrossRef] [PubMed]
- Parsons Corporation. Principles and Practices of Enhanced Anaerobic Bioremediation of Chlorinated Solvents; AFCEE: Brooks City, TX, USA; NFEC: Port Hueneme, CA, USA; ESTCP: Arlington, VA, USA, 2004; p. 457. [Google Scholar]
- Liang, S.H.; Kuo, Y.C.; Chen, S.H.; Chen, C.Y.; Kao, C.M. Development of a slow polycolloid-releasing substrate (SPRS) biobarrier to remediate TCE-contaminated aquifers. J. Hazard. Mater. 2013, 254, 107–115. [Google Scholar] [CrossRef]
- Fennell, D.E.; Gossett, J.M.; Zinder, S.H. Comparison of butyric acid, ethanol, lactic acid, and propionic acid as hydrogen donors for the reductive dechlorination of tetrachloroethene. Environ. Sci. Technol. 1997, 31, 918–926. [Google Scholar] [CrossRef]
- Yang, Y.; McCarty, P.L. Comparison between donor substrates for biologically enhanced tetrachloroethene DNAPL dissolution. Environ. Sci. Technol. 2002, 36, 3400–3404. [Google Scholar] [CrossRef]
- Matteucci, F.; Ercole, C.; Del Gallo, M. A study of chlorinated solvent contamination of the aquifers of an industrial area in central Italy: A possibility of bioremediation. Front. Microbiol. 2015, 6, 924. [Google Scholar] [CrossRef]
- Adrian, L.; Löffler, F. Organohalide-Respiring Bacteria; Springer: Berlin/Heidelberg, Germany, 2016; pp. 63–105. [Google Scholar] [CrossRef]
- Battelle Memorial Institute. Permeable Reactive Barrier Cost and Performance Report; NAVFAC: Washington, DC, USA, 2012. [Google Scholar]
- Wu, Y.-J.; Liu, P.-W.G.; Hsu, Y.-S.; Whang, L.-M.; Lin, T.-F.; Hung, W.-N.; Cho, K.-C. Application of molecular biological tools for monitoring efficiency of trichloroethylene remediation. Chemosphere 2019, 233, 697–704. [Google Scholar] [CrossRef]
- Fierer, N.; Jackson, J.A.; Vilgalys, R.; Jackson, R.B. Assessment of soil microbial community structure by use of taxon-specific quantitative PCR assays. Appl. Environ. Microbiol. 2005, 71, 4117–4120. [Google Scholar] [CrossRef] [Green Version]
- Rago, L.; Cristiani, P.; Villa, F.; Zecchin, S.; Colombo, A.; Cavalca, L.; Schievano, A. Influ- ences of dissolved oxygen concentration on biocathodic microbial communities in microbial fuel cells. Bioelectrochemistry 2017, 116, 39–51. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [Green Version]
- Rognes, T.; Flouri, T.; Nichols, B.; Quince, C.; Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 2016, 4, e2584. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Callahan, B.J.; McMurdie, P.J.; Holmes, S.P. Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. ISME J. 2017, 11, 2639–2643. [Google Scholar] [CrossRef] [Green Version]
- Spring, S.; Rosenzweig, F. The genera Desulfitobacterium and Desulfosporosinus: Taxonomy. Prokaryotes 2006, 4, 771–786. [Google Scholar]
- Zhang, K.; Song, L.; Dong, X. Proteiniclasticum ruminis gen. nov.; sp. nov.; a strictly anaerobic proteolytic bacterium isolated from yak rumen. Int. J. Syst. Evol. Microbiol. 2010, 60, 2221–2225. [Google Scholar] [CrossRef]
- Sakamoto, M. The Family Porphyromonadaceae. In The Prokaryotes—Other Major Lineages of Bacteria and the Archaea; Rosenberg, E., DeLong, R.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; pp. 811–824. [Google Scholar]
- Jiang, Y.; Dennehy, C.; Lawlor, P.G.; Hu, Z.; McCabe, M.; Cormican, P.; Zhan, X.; Gardiner, G.E. Exploring he roles of and interactions among microbes in dry co-digestion of food waste and pig manure using high throughput 16S rRNA gene amplicon sequencing. Biotechnol. Biofuels 2019, 12, 5. [Google Scholar] [CrossRef] [Green Version]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2015. [Google Scholar]
- Chambers, J.M.; Freeny, A.; Heiberger, R.M. Analysis of variance; designed experiments. In Statistical Models in S; Chapter 5; Chambers, S.J.M., Hastie, T.J., Eds.; Routledge: Abingdon, UK, 2017. [Google Scholar]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McCarthy, D.J.; Chen, Y.; Smyth, G.K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 2012, 40, 4288–4297. [Google Scholar] [CrossRef] [Green Version]
- Chen, Y.; Lun, A.T.L.; Smyth, G.K. Differential expression analysis of complex RNA-seq experiments using edgeR. In Statistical Analysis of Next Generation Sequence Data; Datta, S., Nettleton, D.S., Eds.; Springer: New York, NY, USA, 2014. [Google Scholar]
- Robinson, M.D.; Smyth, G.K. Small-sample estimation of negative binomial dispersion, with applications to SAGE data. Biostatistics 2008, 9, 321–332. [Google Scholar] [CrossRef] [Green Version]
- Leri, A.C.; Mayer, L.M.; Thornton, K.R.; Northrup, P.A.; Dunigan, M.R.; Ness, K.J.; Gellis, A.B. A marine sink for chlorine in natural organic matter. Nat. Geosci. 2015, 8, 620–624. [Google Scholar] [CrossRef]
- Atashgahi, S.; Häggblom, M.M.; Smidt, H. Organohalide respiration in pristine environments: Implications for the natural halogen cycle. Environ. Microbiol. 2018, 20, 934–948. [Google Scholar] [CrossRef] [Green Version]
- Nobre, R.C.M.; Nobre, M.M.M.; Campos, T.M.P.; Ogles, D. In-situ biodegradation potential of 1,2-DCA and VC at sites with different hydrogeological settings. J. Hazard. Mater. 2017, 340, 417–426. [Google Scholar] [CrossRef]
- Lee, P.K.; Macbeth, T.W.; Sorenson, K.S.; Deeb, R.A.; Alvarez-Cohen, L. Quantifying genes and transcripts to assess the in situ physiology of “Dehalococcoides” spp. in a trichloroethene-contaminated groundwater site. Appl. Environ. Microbiol. 2008, 74, 2728–2739. [Google Scholar] [CrossRef] [Green Version]
- Courbet, C.; Rivière, A.; Jeannottat, S.; Rinaldi, S.; Hunkeler, D.; Bendjoudi, H.; De Marsily, G. Complementing approaches to demonstrate chlorinated solvent biodegradation in a complex pollution plume: Mass balance, PCR and compound-specific stable isotope analysis. J. Contam. Hydrol. 2011, 126, 315–329. [Google Scholar] [CrossRef] [PubMed]
- Lendvay, J.M.; Loffler, F.E.; Dollhopf, M.; Aiello, M.R.; Daniels, G.; Fathepure, B.Z.; Gebhard, M.; Heine, R.; Helton, R.; Shi, J.; et al. Bioreactive barriers: A comparison of bioaugmentation and biostimulation for chlorianted solvent remediation. Environ. Sci. Technol. 2003, 37, 1422–1431. [Google Scholar] [CrossRef]
- Matturro, B.; Pierro, L.; Frascadore, E.; Petrangeli Papini, M.; Rossetti, S. Microbial community changes in a chlorinated solvents polluted aquifer over the field scale treatment with poly-3-hydroxybutyrate as amendment. Front. Microbiol. 2018, 9, 1664. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, Y.; Cápiro, N.L.; Marcet, T.F.; Yan, J.; Pennell, K.D.; Löffler, F.E. Organohalide respiration with chlorinated ethenes under low pH conditions. Environ. Sci. Technol. 2017, 51, 8579–8588. [Google Scholar] [CrossRef]
- Jing, R.; Kjellerup, B.V. Predicting the potential for organohalide respiration in wastewater: Comparison of intestinal and wastewater microbiomes. Sci. Total Env. 2020, 705, 135833. [Google Scholar] [CrossRef]
- Qiu, L.; Fang, W.; He, H.; Liang, Z.; Zhan, Y.; Lu, Q.; Liang, D.; Wang, S. Organohalide-respiring bacteria in polluted urban rivers employ novel bifunctional reductive dehalogenases to dechlorinate polychlorinated biphenyls and tetrachloroethene. Environ. Sci. Technol. 2020, 54, 8791–8800. [Google Scholar] [CrossRef]
- Comeau, A.; Harding, T.; Galand, P.; Vincent, W.F.; Lovejoy, C. Vertical distribution of microbial communities in a perennially stratified Arctic lake with saline, anoxic bottom waters. Sci. Rep. 2020, 2, 604. [Google Scholar] [CrossRef] [Green Version]
- Kubo, K.; Lloyd, K.G.; Biddle, J.F.; Amann, R.; Teske, A.; Knittel, K. Archaea of the Miscellaneous Crenarchaeotal Group are abundant, diverse and widespread in marine sediments. ISME J 2012, 6, 1949–1965. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Z.; Pan, J.; Wang, F.; Gu, J.D.; Li, M. Bathyarchaeota: Globally distributed metabolic generalists in anoxic environments. Fems Microbiol. Rev. 2018, 42, 639–655. [Google Scholar] [CrossRef]
- Meng, J.; Xu, J.; Qin, D.; He, Y.; Xiao, X.; Wang, F. Genetic and functional properties of uncultivated MCG archaea assessed by metagenome and gene expression analyses. ISME J. 2014, 8, 650–659. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vanwonterghem, I.; Evans, P.N.; Parks, D.H.; Jensen, P.D.; Woodcroft, B.J.; Hugenholtz, P.; Tyson, G.W. Methylotrophic methanogenesis discovered in the archaeal phylum Verstraetearchaeota. Nat. Microbiol. 2016, 1, 1–9. [Google Scholar] [CrossRef] [Green Version]
- Maphosa, F.; de Vos, W.M.; Smidt, H. Exploiting the ecogenomics toolbox for environmental diagnostics of organohalide-respiring bacteria. Trends Biotechnol. 2010, 28, 308–316. [Google Scholar] [CrossRef] [PubMed]
- Temme, H.R.; Carlson, A.; Novak, P.J. Presence, Diversity, and Enrichment of Respiratory Reductive Dehalogenase and Non-respiratory Hydrolytic and Oxidative Dehalogenase Genes in Terrestrial Environments. Front. Microbiol. 2019, 10, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Türkowsky, D.; Jehmlich, N.; Diekert, G.; Adrian, L.; von Bergen, M.; Goris, T. An integrative overview of genomic, transcriptomic and proteomic analyses in organohalide respiration research. FEMS Microbiol. Ecol. 2018, 94. [Google Scholar] [CrossRef] [Green Version]
- Lohner, S.T.; Spormann, A.M. Identification of a reductive tetrachloroethene dehalogenase in Shewanella sediminis. Philos. Trans. R. Soc. B Biol. Sci. 2013, 368, 20120326. [Google Scholar] [CrossRef] [Green Version]
- Kuribayashi, T.A.; Fujii, S.; Janda, J.M.; Abbott, S.L. The genus Shewanella: From the briny depths below to human pathogen. Crit. Rev. Microbiol. 2014, 40, 293–312. [Google Scholar]
- Sucharita, K.; Sasikala, C.; Park, S.C.; Baik, K.S.; Seong, C.N.; Ramana, C.V. Shewanella chilikensis sp. nov.; a moderately alkaliphilic gammaproteobacterium isolated from a lagoon. Int. J. Syst. Evol. Microbiol. 2009, 59, 3111–3115. [Google Scholar] [CrossRef] [Green Version]
- Jiang, Z.; Shi, M.; Shi, L. Degradation of organic contaminants and steel corrosion by the dissimilatory metal-reducing microorganisms Shewanella and Geobacter spp. Int. Biodeterior. Biodegrad. 2020, 147, 104842. [Google Scholar] [CrossRef]
- Viñas, M.; Sabaté, J.; Espuny, M.J.; Solanas, A.M. Bacterial community dynamics and polycyclic aromatic hydrocarbon degradation during bioremediation of heavily creosote-contaminated soil. Appl. Environ. Microbiol. 2005, 71, 7008–7018. [Google Scholar] [CrossRef] [Green Version]
- Farber, R.; Rosenberg, A.; Rozenfeld, S.; Banet, G.; Cahan, R. Bioremediation of artificial diesel-contaminated soil using bacterial consortium immobilized to plasma-pretreated wood waste. Microorganisms 2019, 7, 497. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Obi, C.C.; Adebusoye, S.A.; Amund, O.O.; Ugoji, E.O.; Ilori, M.O.; Hedman, C.J.; Hickey, W.J. Structural dynamics of microbial communities in polycyclic aromatic hydrocarbon-contaminated tropical estuarine sediments undergoing simulated aerobic biotreatment. Appl. Microbiol. Biotechnol 2017, 101, 4299–4314. [Google Scholar] [CrossRef]
- Hatayama, K.; Sakihama, Y.; Matsumiya, Y.; Kubo, M. Construction of an Efficient Bioremediation System for Petroleum Hydrocarbon-Contaminated Soils Using Specific Hydrocarbon-Degrading Bacteria and Bacterial Biomass Monitoring. Soil Contam. Res. Trends 2008, 6, 143–160. [Google Scholar]
- Kaplan, C.W.; Kitts, C.L. Bacterial succession in a petroleum land treatment unit. Appl. Environ. Microbiol. 2004, 70, 1777–1786. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tan, X.; Liao, H.; Shu, L.; Yao, H. Effect of different substrates on soil microbial community structure and the mechanisms of reductive soil disinfestation. Front. Microbiol. 2019, 10, 2851. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Lee, T.K.; Löffler, F.E.; Park, J. Characterization of microbial community structure and population dynamics of tetrachloroethene-dechlorinating tidal mudflat communities. Biodegradation 2011, 22, 687–698. [Google Scholar] [CrossRef]
- Macbeth, T.W.; Cummings, D.E.; Spring, S.; Petzke, L.M.; Sorenson, K.S. Molecular characterization of a dechlorinating community resulting from in situ biostimulation in a trichloroethene-contaminated deep, fractured basalt aquifer and comparison to a derivative laboratory culture. Appl. Environ. Microbiol. 2004, 70, 7329–7341. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Militon, C.; Boucher, D.; Vachelard, C.; Perchet, G.; Barra, V.; Troquet, J.; Peyretaillade, E.; Peyret, P. Bacterial community changes during bioremediation of aliphatic hydrocarbon-contaminated soil. FEMS Microbiol. Ecol. 2010, 74, 669–681. [Google Scholar] [CrossRef]
- Yang, S.; Wen, X.; Zhao, L.; Shi, Y.; Jin, H. Crude oil treatment leads to shift of bacterial communities in soils from the deep active layer and upper permafrost along the China-Russia crude oil pipeline route. PLoS ONE 2014, 9, e96552. [Google Scholar] [CrossRef]
- Koo, H.; Mojib, N.; Huang, J.P.; Donahoe, R.J.; Bej, A.K. Bacterial community shift in the coastal Gulf of Mexico salt-marsh sediment microcosm in vitro following exposure to the Mississippi Canyon Block 252 oil (MC252). 3 Biotech 2015, 5, 379–392. [Google Scholar] [CrossRef] [Green Version]
- Yang, S.; Wen, X.; Shi, Y.; Liebner, S.; Jin, H.; Perfumo, A. Hydrocarbon degraders establish at the costs of microbial richness, abundance and keystone taxa after crude oil contamination in permafrost environments. Sci. Rep. 2016, 6, 37473. [Google Scholar] [CrossRef] [PubMed]
- DiStefano, T.D.; Baral, R.; Duran, M.; Speece, R.E. A comparison of complex electron donors for anaerobic dechlorination of PCE. Bioremediation J. 2001, 5, 131–143. [Google Scholar] [CrossRef]
- Kao, C.M.; Chen, Y.L.; Chen, S.C.; Yeh, T.Y.; Wu, W.S. Enhanced PCE dechlorination by biobarrier systems under different redox conditions. Water Res. 2003, 37, 4885–4894. [Google Scholar] [CrossRef]
- Aulenta, F.; Bianchi, A.; Majone, M.; Papini, M.P.; Potalivo, M.; Tandoi, V. Assessment of natural or enhanced in situ bioremediation at a chlorinated solvent-contaminated aquifer in Italy: A microcosm study. Environ. Int. 2005, 31, 185–190. [Google Scholar] [CrossRef] [PubMed]
- Imfeld, G.; Pieper, H.; Shani, N.; Rossi, P.; Nikolausz, M.; Nijenhuis, I.; Paschke, H.; Weiss, H.; Richnow, H.H. Characterization of groundwater microbial communities, dechlorinating bacteria, and in situ biodegradation of chloroethenes along a vertical gradient. Water Air Soil Pollut. 2011, 221, 107–122. [Google Scholar] [CrossRef]
- Fennell, D.E.; Carroll, A.B.; Gossett, J.M.; Zinder, S.H. Assessment of indigenous reductive dechlorinating potential at a TCE-contaminated site using microcosms, polymerase chain reaction analysis, and site data. Environ. Sci. Technol. 2001, 35, 1830–1839. [Google Scholar] [CrossRef]
- Lorah, M.M.; Voytek, M.A. Degradation of 1,1,2,2- tetrachloroethane and accumulation of vinyl chloride in wetland sediment microcosms and in situ porewater: Biogeochemical controls and associations with microbial communities. J. Contam. Hydrol. 2004, 70, 117–145. [Google Scholar] [CrossRef]
- Aeppli, C.; Hofstetter, T.B.; Amaral, H.I.; Kipfer, R.; Schwarzenbach, R.P.; Berg, M. Quantifying in situ transformation rates of chlorinated ethenes by combining compound-specific stable isotope analysis, groundwater dating, and carbon isotope mass balances. Environ. Sci. Technol. 2010, 44, 3705–3711. [Google Scholar] [CrossRef] [Green Version]
- Matturro, B.; Presta, E.; Rossetti, S. Reductive dechlorination of tetrachloroethene in marine sediments: Biodiversity and dehalorespiring capabilities of the indigenous microbes. Sci. Total Env. 2016, 545, 445–452. [Google Scholar] [CrossRef]
- Hermon, L.; Hellal, J.; Denonfoux, J.; Vuilleumier, S.; Imfeld, G.; Urien, C.; Ferreira, S.; Joulian, C. Functional genes and bacterial communities during organohalide respiration of chloroethenes in microcosms of multi-contaminated groundwater. Front. Microbiol. 2019, 10, 89. [Google Scholar] [CrossRef] [Green Version]
- Futagami, T.; Goto, M.; Furukawa, K. Biochemical and genetic bases of dehalorespiration. Chem. Rec. 2008, 8, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Abe, Y.; Aravena, R.; Zopfi, J.; Parker, B.; Hunkeler, D. Evaluating the fate of chlorinated ethenes in streambed sediments by combining stable isotope, geochemical and microbial methods. J. Contam. Hydrol. 2009, 107, 10–21. [Google Scholar] [CrossRef] [Green Version]
- Tiehm, A.; Schmidt, K.R. Sequential anaerobic/aerobic biodegradation of chloroethenes-aspects of field application. Curr. Opin. Biotechnol. 2011, 22, 415–421. [Google Scholar] [CrossRef] [PubMed]
- Yoshikawa, M.; Zhang, M.; Toyota, K. Biodegradation of volatile organic compounds and their effects on biodegradability under co-existing conditions. Microbes Env. 2017, 32, 188–200. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Weatherill, J.J.; Atashgahi, S.; Schneidewind, U.; Krause, S.; Ullah, S.; Cassidy, N.; Rivett, M.O. Natural attenuation of chlorinated ethenes in hyporheic zones: A review of key biogeochemical processes and in-situ transformation potential. Water Res. 2018, 128, 362–382. [Google Scholar] [CrossRef] [Green Version]
- Perez-de-Mora, A.; Zila, A.; McMaster, M.L.; Edwards, E.A. Bioremediation of chlorinated ethenes in fractured bedrock and associated changes in dechlorinating and nondechlorinating microbial populations. Environ. Sci. Technol. 2014, 48, 5770–5779. [Google Scholar] [CrossRef] [PubMed]
- Van der Zaan, B.; Hannes, F.; Hoekstra, N.; Rijnaarts, H.; de Vos, W.M.; Smidt, H.; Gerritse, J. Correlation of Dehalococcoides 16S rRNA and chloroethene-reductive dehalogenase genes with geochemical conditions in chloroethene-contaminated groundwater. Appl. Environ. Microbiol. 2010, 76, 843–850. [Google Scholar] [CrossRef] [Green Version]
- Liang, Y.; Liu, X.; Singletary, M.A.; Wang, K.; Mattes, T.E. Relationships between the abundance and expression of functional genes from vinyl chloride (VC)-degrading bacteria and geochemical parameters at VC-contaminated sites. Environ. Sci. Technol. 2017, 51, 12164–12174. [Google Scholar] [CrossRef]
- Dugat-Bony, E.; Biderre-Petit, C.; Jaziri, F.; David, M.M.; Denonfoux, J.; Lyon, D.Y.; Richard, J.Y.; Curvers, C.; Boucher, D.; Vogel, T.M.; et al. In situ TCE degradation mediated by complex dehalorespiring communities during biostimulation processes. Microb. Biotechnol. 2012, 5, 642–653. [Google Scholar] [CrossRef]
- Yoshikawa, M.; Zhang, M. Constraints in Anaerobic Microbial Dechlorination, Fermentation, and Sulfate-Reduction Induced by High Concentrations of Tetrachloroethylene. Water Air Soil Pollut. 2020, 231, 1–11. [Google Scholar] [CrossRef]
- Trueba-Santiso, A.; Parladé, E.; Rosell, M.; Lliros, M.; Mortan, S.H.; Martínez-Alonso, M.; Gaju, N.; Martín-González, L.; Vicent, T.; Marco-Urrea, E. Molecular and carbon isotopic characterization of an anaerobic stable enrichment culture containing Dehalobacterium sp. during dichloromethane fermentation. Sci. Total Environ. 2017, 581, 640–648. [Google Scholar] [CrossRef]
- Siliakus, M.F.; van der Oost, J.; Kengen, S.W. Adaptations of archaeal and bacterial membranes to variations in temperature, pH and pressure. Extremophiles 2017, 21, 651–670. [Google Scholar] [CrossRef] [PubMed]
- Boone, D.R.; Castenholz, R.W. Bergey’s manual of systematic bacteriology. In The Archaea and the Deeply Branching and Phototrophic Bacteria; Garrity, G.M., Ed.; Springer: New York, NY, USA, 2001. [Google Scholar]
- Rinke, C.; Schwientek, P.; Sczyrba, A.; Ivanova, N.N.; Anderson, I.J.; Cheng, J.F.; Darling, A.; Malfatti, S.; Swan, B.K.; Gies, E.A.; et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature 2013, 499, 431–437. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shelton, J.L.; Akob, D.M.; McIntosh, J.C.; Fierer, N.; Spear, J.R.; Warwick, P.D.; McCray, J.E. Environmental drivers of differences in microbial community structure in crude oil reservoirs across a methanogenic gradient. Front. Microbiol. 2016, 7, 1535. [Google Scholar] [CrossRef] [Green Version]
- Delgado, A.G.; Fajardo-Williams, D.; Kegerreis, K.L.; Parameswaran, P.; Krajmalnik-Brown, R. Impact of ammonium on syntrophic organohalide-respiring and fermenting microbial communities. mSphere 2016, 1, e00053-16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ziv-El, M.; Delgado, A.G.; Yao, Y.; Kang, D.W.; Nelson, K.G.; Halden, R.U.; Krajmalnik-Brown, R. Development and characterization of DehaloR^ 2, a novel anaerobic microbial consortium performing rapid dechlorination of TCE to ethene. Appl. Microbiol. Biotech. 2011, 92, 1063–1071. [Google Scholar] [CrossRef]
- Liu, N.; Li, H.; Li, M.; Ding, L.; Weng, C.H.; Dong, C.D. Oxygen exposure effects on the dechlorinating activities of a trichloroethene-dechlorination microbial consortium. Bioresour. Technol. 2017, 240, 98–105. [Google Scholar] [CrossRef]
- Němeček, J.; Dolinová, I.; Macháčková, J.; Špánek, R.; Ševců, A.; Lederer, T.; Černík, M. Stratification of chlorinated ethenes natural attenuation in an alluvial aquifer assessed by hydrochemical and biomolecular tools. Chemosphere 2017, 184, 1157–1167. [Google Scholar] [CrossRef]
- Hug, L.A.; Beiko, R.G.; Rowe, A.R.; Richardson, R.E.; Edwards, E.A. Comparative metagenomics of three Dehalococcoides-containing enrichment cultures: The role of the non-dechlorinating community. BMC Genom. 2012, 13, 327. [Google Scholar] [CrossRef] [Green Version]
- Wei, K.; Grostern, A.; Chan, W.W.; Richardson, R.E.; Edwards, E.A. Electron acceptor interactions between organohalide-respiring bacteria: Cross-feeding, competition, and inhibition. In Organohalide-Respiring Bacteria; Springer: Berlin/Heidelberg, Germany, 2016; pp. 283–308. [Google Scholar]
- Maphosa, F.; van Passel, M.W.; de Vos, W.M.; Smidt, H. Metagenome analysis reveals yet unexplored reductive dechlorinating potential of Dehalobacter sp. E 1 growing in co-culture with Sedimentibacter sp. Environ. Microbiol. Rep. 2012, 4, 604–616. [Google Scholar]
- Ding, C.; Ma, T.; Hu, A.; Dai, L.; He, Q.; Cheng, L.; Zhang, H. Enrichment and characterization of a psychrotolerant consortium degrading crude oil alkanes under methanogenic conditions. Microb. Ecol. 2015, 70, 433–444. [Google Scholar] [CrossRef]
- Gao, W.; Yin, X.; Mi, T.; Zhang, Y.; Lin, F.; Han, B.; Zhao, X.; Cui, Z.; Zheng, L. Microbial diversity and ecotoxicity of sediments 3 years after the Jiaozhou Bay oil spill. AMB Express 2018, 8, 1–10. [Google Scholar] [CrossRef]
- Jung, J.; Choi, S.; Hong, H.; Sung, J.S.; Park, W. Effect of red clay on diesel bioremediation and soil bacterial community. Microb. Ecol. 2014, 68, 314–323. [Google Scholar] [CrossRef]
- Campeão, M.E.; Reis, L.; Leomil, L.; de Oliveira, L.; Otsuki, K.; Gardinali, P.; Pelz, O.; Valle, R.; Thompson, F.L.; Thompson, C.C. The deep-sea microbial community from the Amazonian basin associated with oil degradation. Front. Microbiol. 2017, 8, 1019. [Google Scholar] [CrossRef] [Green Version]
- Li, J.; Luo, C.; Zhang, D.; Cai, X.; Jiang, L.; Zhao, X.; Zhang, G. Diversity of the active phenanthrene degraders in PAH-polluted soil is shaped by ryegrass rhizosphere and root exudates. Soil Biol. Biochem. 2019, 128, 100–110. [Google Scholar] [CrossRef]
- Sun, W.; Sun, X.; Cupples, A.M. Identification of Desulfosporosinus as toluene-assimilating microorganisms from a methanogenic consortium. Int. Biodeterior. Biodegrad. 2014, 88, 13–19. [Google Scholar] [CrossRef]
- Sette, L.D.; Simioni, K.C.; Vasconcellos, S.P.; Dussan, L.J.; Neto, E.V.; Oliveira, V.M. Analysis of the composition of bacterial communities in oil reservoirs from a southern offshore Brazilian basin. Antonie Van Leeuwenhoek 2007, 91, 253–266. [Google Scholar] [CrossRef]
| Parameter/Target | Unit | Law Limits Directive 2000/60/EC | Pz22 | St. Dv. | Pz25 | St. Dv. |
|---|---|---|---|---|---|---|
| PCE | µg L−1 | 1.1 | 7300.00 | 802.1 | 2860.00 | 622.3 |
| TCE | µg L−1 | 1.5 | 43,000.00 | 15,143.8 | 7300.00 | 1850.2 |
| 1,1 DCE | µg L−1 | 0.05 | 22,800.00 | 16,607.9 | 1370.00 | 640.9 |
| 1,2 DCE | µg L−1 | 60 | 17,000.00 | 6455.0 | 5640.00 | 1683.6 |
| VC | µg L−1 | 0.5 | 72,000.00 | 62,649.8 | 6200.00 | 4346.6 |
| ethene | µg L−1 | - | 19,800.00 | 282.8 | 980.00 | 195.2 |
| Eh * | mV | - | −126.00 | −85.00 | ||
| pH | - | 7.4 | 7.5 | |||
| DO ** | mg L−1 | - | 1.20 | 0.9 | ||
| Eub | log(gene copies L−1) | - | 9.42 ± 0.08 | 9.79 ± 0.12 | ||
| Arc | log(gene copies L−1) | - | 5.70 ± 0.06 | 7.27 ± 0.15 | ||
| Geo | log(gene copies L−1) | - | 3.34 ± 0.21 | 3.89 ± 0.05 | ||
| Dhc | log(gene copies L−1) | - | 6.42 ± 0.02 | 5.17 ± 0.11 | ||
| tcrA | log(gene copies L−1) | - | 7.80 ± 0.10 | 7.05 ± 0.14 | ||
| vcrA | log(gene copies L−1) | - | 6.77 ± 0.05 | 6.01 ± 0.04 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bertolini, M.; Zecchin, S.; Beretta, G.P.; De Nisi, P.; Ferrari, L.; Cavalca, L. Effectiveness of Permeable Reactive Bio-Barriers for Bioremediation of an Organohalide-Polluted Aquifer by Natural-Occurring Microbial Community. Water 2021, 13, 2442. https://doi.org/10.3390/w13172442
Bertolini M, Zecchin S, Beretta GP, De Nisi P, Ferrari L, Cavalca L. Effectiveness of Permeable Reactive Bio-Barriers for Bioremediation of an Organohalide-Polluted Aquifer by Natural-Occurring Microbial Community. Water. 2021; 13(17):2442. https://doi.org/10.3390/w13172442
Chicago/Turabian StyleBertolini, Martina, Sarah Zecchin, Giovanni Pietro Beretta, Patrizia De Nisi, Laura Ferrari, and Lucia Cavalca. 2021. "Effectiveness of Permeable Reactive Bio-Barriers for Bioremediation of an Organohalide-Polluted Aquifer by Natural-Occurring Microbial Community" Water 13, no. 17: 2442. https://doi.org/10.3390/w13172442
APA StyleBertolini, M., Zecchin, S., Beretta, G. P., De Nisi, P., Ferrari, L., & Cavalca, L. (2021). Effectiveness of Permeable Reactive Bio-Barriers for Bioremediation of an Organohalide-Polluted Aquifer by Natural-Occurring Microbial Community. Water, 13(17), 2442. https://doi.org/10.3390/w13172442







