Abstract
Wastewater contamination and urbanization contribute to the spread of antibiotic resistance in aquatic environments. This is a particular concern in areas receiving chronic pollution of untreated waste via combined sewer overflow (CSO) events. The goal of this study was to expand knowledge of CSO impacts, with a specific focus on multidrug resistance. We sampled a CSO-impacted segment of the James River (Virginia, USA) during both clear weather and an active overflow event and compared it to an unimpacted upstream site. Bacteria resistant to ampicillin, streptomycin, and tetracycline were isolated from all samples. Ampicillin resistance was particularly abundant, especially during the CSO event, so these isolates were studied further using disk susceptibility tests to assess multidrug resistance. During a CSO overflow event, 82% of these isolates were resistant to five or more antibiotics, and 44% were resistant to seven or more. The latter statistic contrasts starkly with the upstream reference site, where only 4% of isolates displayed resistance to more than seven antibiotics. DNA sequencing (16S rRNA gene) revealed that ~35% of our isolates were opportunistic pathogens, comprised primarily of the genera Stenotrophomonas, Pseudomonas, and Chryseobacterium. Together, these results demonstrate that CSOs can be a significant source of viable clinically-relevant bacteria to the natural environment and that multidrug resistance is an important understudied component of the environmental spread of antibiotic resistance.
1. Introduction
Widespread, indiscriminate use of antibiotics in recent decades has stimulated a proliferation of antibiotic-resistant (AR) strains of bacteria, which are a major public health threat to millions of people worldwide. While antibiotic resistance is most often studied in medical settings, there is growing concern about spread to the natural environment. There has been a particular focus on freshwater ecosystems because they are a vital source of drinking water and widely used for recreation. Sources of AR to these ecosystems include municipal wastewater systems, effluent from hospitals, pharmaceutical manufacturing, and upstream agricultural runoff. Wastewater treatment plants are well-known hotspots for AR [1] due to high densities of bacteria in close proximity as well as the presence of contaminants, such as pharmaceuticals and heavy metals, that select for resistant organisms. Because these wastewater treatment facilities do not completely remove antibiotics, antibiotic-resistant bacteria, or antibiotic resistance genes [2], effluent causes chronic pollution of receiving water bodies (e.g., see [3,4,5,6,7]).
In many urban aquatic ecosystems, the potential for spreading AR is further exacerbated by reliance on combined sewer systems and the unintentional release of untreated sewage directly into the environment. Used by more than 40 million people in the United States [8], these systems collect stormwater and sewage for treatment at a single facility. During large precipitation events, the volume of water that needs to be treated exceeds the capacity of the facility, and the combined untreated wastewater is released into the receiving water body in what is called a CSO (“combined sewer overflow”) event. Urban rivers and streams that are heavily impacted by CSO discharge have been found to contain elevated concentrations of AR bacteria and AR genes [9,10,11,12,13], contributed to primarily by storm-driven transport [14]. Many of these prior studies have utilized molecular techniques and focused on assessing the abundance of AR genes, but it remains unclear whether the genes are present in viable clinically-relevant organisms. Moreover, as CSO events occur only occasionally, it remains uncertain whether there is a significant risk of AR exposure at impacted sites during clear weather (i.e., when there is no overflow). A recent study by Brown et al. [15] suggests that the high levels of AR in rivers are due to continuous wastewater and CSO inputs but would not persist if these sources were removed.
Given the diversity of AR bacteria and genes associated with wastewater, as well as selection for AR traits due to sublethal concentrations of antibiotics and heavy metals, the potential for the development of multidrug-resistance (MDR) is high [16,17,18,19]. This can occur when an organism accumulates multiple genes that each code for resistance to a single antibiotic, collectively conferring resistance to multiple drugs. The phenomena can also manifest due to increased expression of individual genes that code for multidrug efflux pumps. Genes associated with MDR have been found in several metagenomic studies of surface water [20,21,22], including wastewater-impacted rivers [23]. In addition, MDR organisms have been isolated from both clinical [24,25,26] and municipal wastewater effluent [27] and from receiving water bodies [28,29]. This has led to serious concerns about the cycling of MDR between clinical settings, the human microbiome, and the water system, but the potential effects of CSOs have not been fully explored.
This study examined the role that CSOs may play in disseminating viable MDR bacteria to urban rivers. The research was conducted in Richmond, Virginia (USA), which is a moderately-sized city that relies on a combined sewer system that overflows into the James River (Figure 1). We utilized culture-based methods to isolate viable resistant bacteria from a CSO-impacted section of the river during both clear weather conditions and an active overflow event, then compared this to an upstream site with no proximal CSOs. We initially focused on determining what fraction of the bacterial community was resistant to ampicillin, streptomycin, and tetracycline. Results indicate ampicillin-resistant bacteria were particularly abundant, especially during a CSO overflow event, so these isolates were identified (16S rRNA gene sequencing) and screened for MDR. We hypothesized that bacterial communities associated with CSO discharge would exhibit higher levels of MDR and contain more human-associated and potentially pathogenic species. Further, we predicted that the chronic inputs at the at the CSO site would result in a community with elevated MDR and pathogenicity even during clear weather (i.e., when the CSO was not discharging).
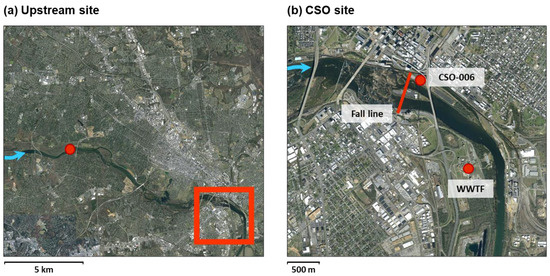
Figure 1.
(a) Map of study area showing the direction of water flow (blue arrows) and the location of the upstream sampling site (red circle). The red inset box is expanded in panel (b) to show the sampling site at the CSO discharge point, the location of the city′s wastewater treatment facility (WWTF), and the approximate fall line of the James River.
2. Materials and Methods
2.1. Study Sites
Richmond, Virginia (USA) has approximately 227,000 residents and a population density of ~9800 people per km2 (United States Census Bureau, 2017). The city has the most extensive CSO system in the state of Virginia, servicing approximately 50 km2. This study considered two sites along the James River as it flows through Richmond. The first site (37.560471, −77.545801) was ~10 km “upstream” from the city center and before all known CSO outfalls; as such, it allowed us to characterize AR in the absence of the urban and CSO inputs. The second site was downtown (37.529486–77.429382), adjacent to the city′s largest CSO discharge point (identified as CSO-006 by the Richmond Department of Public Utilities). Long-term water quality monitoring indicates consistently elevated levels of Escherichia coli (E. coli) at this “CSO site” (Supplemental Figure S1), and prior research has documented an increased abundance of AR genes in the CSO-affected area of the river [30]. The CSO site is approximately 500 m downstream of the James′ fall line—the point up to which the river becomes tidal—and about 2.5 km upstream from the outflow of the Richmond wastewater treatment plant (Figure 1). Owing to strong tidal and fluvial forces, the James in and around the city is considered to be well-mixed both vertically and laterally.
2.2. Sampling
Surface water samples were collected from the river during an active CSO overflow event (28 June 2016) and again a week later following clear weather and 5+ days without CSO discharge (5 July 2016). The upstream location was sampled on the same date (5 July 2016) during the clear weather condition. Specific conductance, temperature, dissolved oxygen, turbidity, and pH of each water sample were measured in the field using a YSI 6600 water quality sonde (YSI Environmental, Inc., Yellow Springs, OH, USA). Water used for microbiological analyses was collected from the surface (~30 cm depth) of the main channel of the river using sterile plastic containers (~5 L), transported back to the laboratory on ice, and processed within 2 h. River discharge data were obtained from the United States Geological Survey (USGS) gaging station at site 02037500 (37.563055, −77.547222), which is near the upstream sampling site. Precipitation data was provided by the National Climatic Data Center using the Richmond International Airport “KRIC” station.
2.3. Microbiological Analyses
E. coli abundance was determined using Modified mTEC agar (BD Difco, Sparks, MD, USA) following EPA Method #1603 [31]. In addition, plate counts were performed using R2A agar (BD Difco). Dilutions were prepared in sterile phosphate-buffered saline (PBS) and plated in triplicate onto four types of media: (i) unamended R2A, (ii) R2A amended with ampicillin (100 μg mL−1), (iii) R2A with streptomycin (100 μg mL−1), and (iv) R2A with tetracycline (50 μg mL−1). These antibiotic concentrations are near the maximum of the range recommended by major antibiotic retailers (e.g., ATCC (Manassas, VA, USA), Thermo Fisher Scientific (Waltham, MA, USA), and Sigma Aldrich (St. Louis, MO, USA)) and were chosen in an effort to limit the growth of environmental bacteria, which often have low levels of natural resistance [32]. Plates were incubated in the dark at 27 °C for 72 h, after which time any containing 20–200 colony forming units (CFU) were counted and moved to 4 °C for storage (<2 weeks). The total abundance of heterotrophic bacteria was quantified from unamended plates, and the fraction of the community resistant to each antibiotic was calculated.
2.4. Isolations and MDR Testing
Because plate count data revealed an elevated abundance of ampicillin-resistant bacteria, as well as a strong increase associated with CSO discharge (Figure 2), we elected to isolate colonies from the ampicillin-amended plates for identification and additional resistance screening. For each sample, we selected 50–60 morphologically distinct colonies and streaked them onto ampicillin-amended R2A agar (100 μg mL−1). Colonies were re-streaked onto new plates a minimum of three times to ensure pure cultures. Isolates were then transferred to R2A broth (27 °C for 72 h). A small aliquot of each turbid liquid culture was archived (15% glycerol, stored at −80 °C).
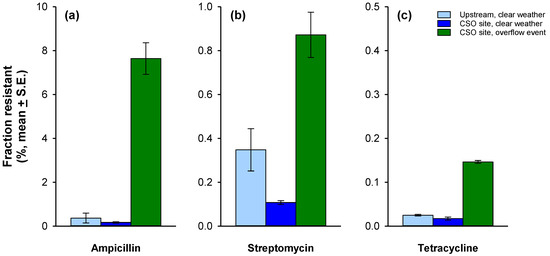
Figure 2.
Samples collected from the upstream site during clear weather (left bar, light blue), the CSO-impacted site during clear weather (center bar, dark blue), and the CSO-impacted site during an overflow event (right bar, green) were analyzed to determine the fraction of the culturable bacterial community resistant to either: (a) ampicillin, (b) streptomycin, or (c) tetracycline. Note the difference in scale on the y-axis.
OxoidTM Antimicrobial Susceptibility Disk (Thermo Scientific, Waltham, MA, USA) tests were used to screen isolates for multidrug resistance (MDR). As our main research foci were (i) CSO events and fecal contamination and (ii) obtaining clinically-relevant isolates, we used the recommendations for screening Enterobacteriaceae outlined in the Clinical and Laboratory Standards Institute (CLSI) manual [33]. This decision was supported by the fact that ~50% of clinically significant bacteria are Enterobacteriaceae [33]. We chose 1–2 drugs per antibiotic class and used the recommended disk concentrations. A total of 10 were screened, including two penicillins (ampicillin (10 μg) and piperacillin (100 μg)), 1 β-lactam/β-lactamase inhibitor combination (amoxicillin clavulanate (20 μg amoxicillin with 10 μg clavulanic acid), also known as Augmentin), two cephems (cefepime (30 μg) and cefotaxime (30 μg)), one aminoglycoside (streptomycin (10 μg)), 1 tetracycline (tetracycline (30 μg)), one fluoroquinolone (ciprofloxacin (5 μg)), and one folate pathway inhibitor (co-trimoxazole (1.25 μg sulfamethoxazole with 23.75 μg trimethoprim), also known as Bactrim). We also tested vancomycin (30 μg), a glycopeptide antibiotic that was historically the last resort for the treatment of multidrug-resistant Gram-positive bacterial infections [34]. Acquired resistance to vancomycin is now an increasing problem in pathogenic bacteria [35], especially Enterococci [36].
Following disk diffusion methods in the Manual of Antimicrobial Susceptibility Testing [37], archived isolates were tested using the log phase method, with the inoculum reaching the 0.5 McFarland turbidity standard in no longer than 24 h. Once cultures reached the 0.5 McFarland standard, they were spread on Mueller–Hinton agar plates (BD Difco) and incubated at 35 °C for 16–18 h, after which time the zone of inhibition surrounding each antibiotic disk was measured. E. coli (ATCC 25922, Manassas, VA, USA) was used as a control and tested with each batch of samples.
Isolates were classified as being resistant or susceptible based on CLSI reference tables [33]. We then calculated the total number of antibiotics to which each isolate was resistant (MDR index) and the fraction of isolates resistant to each individual antibiotic.
2.5. DNA Extraction and Sequencing
For each isolate, an aliquot (2 mL) of turbid broth culture was centrifuged (1 min at 10,000× g), and DNA was extracted from the resulting cell pellet using the Dneasy UltraClean Microbial DNA Isolation Kit (Qiagen, Germantown, MD, USA). Successful extraction was confirmed using agarose gel electrophoresis (1% gel), and extracted DNA was shipped to Fasteris SA (Geneva, Switzerland) for sequencing of the full 16S rRNA gene. PCR was performed using their proprietary primers: P0 (5′-AGAGTTTGATCCTGGCTCAG-3′) and P6 (5′-GTACGGCTACCTTGTTACGA-3′); the sequence for P0 matches the universal primer 8F and P6 is a slightly longer version of 1492 [38]. Sanger sequencing was performed on purified PCR products using 3730XL DNA analyzer (Applied Biosystems, Foster City, CA, USA) set with a 50 cm 96-capillaries array and a LongSeq-POP7 run module (800–1000 bp in general) with the Z-BigDyeV3 Dye set. Base calling was performed automatically using KB Basecaller Software Version 1.4.1.8 (Applied Biosystems). Forward and reverse sequences were assembled into contigs using BioEdit [39] and quality checked using Sequence Analysis Version 6 (Applied Biosystems). Sequences were deposited into GenBank under accession numbers MZ490613 through MZ490746. Sequencing results were obtained for a total of 134 isolates (45 for the CSO site during the overflow event, 44 for the CSO site during clear weather, and 45 for the upstream site during clear weather).
2.6. Statistical Analyses
All statistical analyses were performed using the PAST 4.0 statistical package [40].
3. Results
3.1. Water Quality
When sampled during clear weather, water quality of the two sites was quite similar except for a small increase in turbidity, bacterial abundance, and E. coli at the CSO site compared to upstream (Table 1). During the wet-weather CSO event, water quality and hydrological parameters changed considerably, including a 2.5-fold increase in river discharge, a 16-fold increase in turbidity, and a doubling of E. coli abundance. The elevated E. coli abundance exceeded the Virginia State law of primary contact-230 CFU 100-mL−1 (Code of Virginia; Clean Water Act). § 62.1-44.15 of the Code of Virginia; Clean Water Act (33 USC § 1251 et seq.); 40 CFR Part 131.

Table 1.
Hydrologic and water quality data for each site and set of sampling conditions.
3.2. Antibiotic Resistance
Across all samples, the fraction of the community resistant to ampicillin was highest (range: 0.2–7.6%), followed by streptomycin (0.1–0.9%), then by tetracycline (0.02–0.15%) (Figure 2). For all three antibiotics, the percentage of bacteria harboring resistance was much greater in conjunction with the active CSO event. Resistance to ampicillin was most affected. The fraction of the community displaying resistance to this antibiotic increased almost 50-fold during the overflow event compared to clear weather conditions; the corresponding change for streptomycin and tetracycline was only 8-fold.
Antibiotic disk susceptibility tests were performed on all ampicillin-resistant isolates to determine MDR to 10 antibiotics in total. Consistent with our findings from the antibiotic-amended plate counts, greater resistance was associated with the CSO site during the overflow event (Figure 3). Approximately 65% of the CSO event isolates were resistant to 6 or more antibiotics. In contrast, this level of MDR was found in only 50% of isolates from the CSO site during clear weather and fewer than 20% of the isolates from the upstream site. All of the isolates obtained during the active CSO event were found to have resistance to at least four of the tested antibiotics.
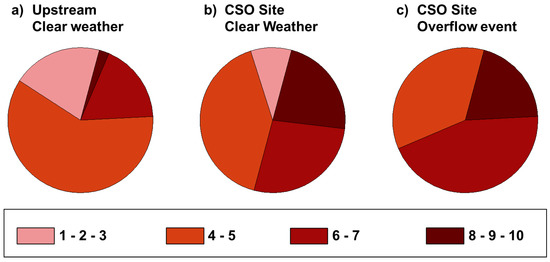
Figure 3.
Fraction of isolates resistant to varying numbers of antibiotics.
A Kruskal–Wallis test revealed a statistically significant difference in MDR between the samples (H = 22.1, p < 0.0001). Isolates from the upstream site had significantly lower levels of MDR (4.5 ± 0.2 S.E.) compared to the CSO site during either clear weather (5.8 ± 0.3) or the overflow event (6.2 ± 0.2) (Dunn′s post hoc, p < 0.001). The difference in MDR between the two sampling events at the CSO site was not significant (p = 0.16).
Cross-site differences were also evident when the results for each antibiotic were reviewed separately (Figure 4). The fraction of isolates resistant was always greatest for the CSO site, and isolates obtained during the overflow event had a further increased likelihood of being resistant to cefotaxime, cefepime, piperacillin, and tetracycline compared to CSO isolates obtained during clear weather. Across all three samples, resistance to vancomycin was highest (82% all isolates) followed by amoxicillin clavulanate (Augmentin) (75%), cefotaxime (69%), and streptomycin (66%). Resistance to co-trimoxazole (Bactrim) (21% of all isolates) and ciprofloxacin (15%) was relatively rare.
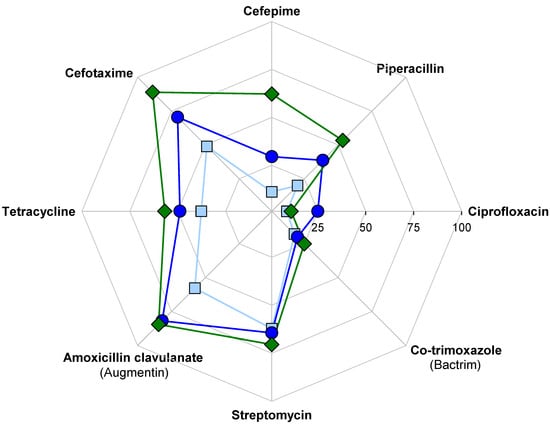
Figure 4.
Fraction of isolates resistant to each antibiotic, with gray lines indicating 25 (innermost), 50, 75, and 100%. Samples were collected from the upstream site during clear weather (light blue squares), the CSO-impacted site during clear weather (dark blue circles), and the CSO-impacted site during an overflow event (green diamonds). Resistance to vancomycin (not shown) was 82% in all three samples.
3.3. Identification of Bacterial Isolates
Representative matches with >99% sequence identity at the species level were found for 118 of the 134 isolates. For the other 16 isolates, representative matches were all >98.3% similar. Classifications comprised 20 families and 30 genera. Diversity was highest for both samples collected during clear weather, and significantly reduced when the CSO site was sampled during overflow conditions (Figure 5). Rarefaction indicates the sampling effort was adequate to capture the diversity of culturable ampicillin-resistant organisms (Figure 5).
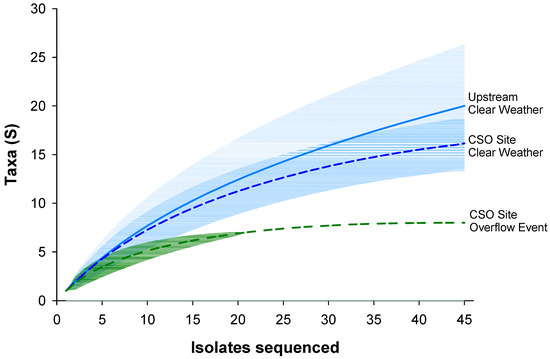
Figure 5.
Rarefaction curves for genus-level classification of isolates. Shading represents 95% confidence intervals.
A complete table of organism identification, percent matching value, MDR profile results, and pathogenicity can be found in the Supplemental Table S2. For subsequent discussion, we rely primarily on the genus-level classifications with a few exceptions: (i) Well-known human pathogens are discussed as “species”. (ii) We aggregated at the family level for Comamondaceae, Enterobacteriaceae, and Sphingomonadaceae. (iii) The category “Other Proteobacteria” was created to aggregate results for five families that had few representatives, specifically Colwelliaceae (only one isolate classified in this family), Erythrobacteraceae (one), Pseudoalteromonadaceae (one), Rhodobacteraceae (one), and Oxalobacteraceae (two).
During the CSO overflow event, the greatest proportion of organisms were Stenotrophomonas (37.8%), of which 28.9% were Stenotrophomonas maltophilia (Figure 6). Fisher’s exact test (Supplemental Table S1) indicated that Stenotrophomonas were found significantly less often in clear weather samples. Mean MDR of Stenotrophomonas was similar at the CSO site during both overflow (6.9 antibiotics) and clear (7.0) weather conditions. No Stenotrophomonas were found in the upstream sample.
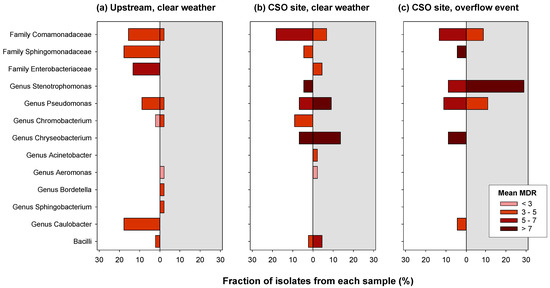
Figure 6.
The fraction of isolates at each site that fall within each classification group. Bar color represents the mean MDR for all isolates in the group, with darker bars indicating resistance to a greater number of antibiotics. Within some groups, both pathogenic and non-pathogenic strains were identified. These are presented separately: bars extending left against the white background represent non-pathogenic isolates, while bars extending right against the gray background represent potentially pathogenic isolates. The graph does not include 6 isolates classified as “Other Proteobacteria” (all non-pathogenic).
Pseudomonas comprised 22.2% of isolates from the CSO site during the overflow event and were in lower abundance during clear weather sampling events (Upstream: 11.1%, CSO site: 15.9%), although chi-squared tests indicate this difference was not significant. In contrast, Sphingomonadaceae were found in much greater abundance upstream (17.8%) compared to either sample from the CSO site (4.4% of each). The family Comamonadaceae had the greatest number of isolates overall, with a similar frequency at each site (Supplemental Table S1). On average, members of this group were resistant to five antibiotics.
Of the 134 total isolates sequenced, 47 could be classified as opportunistic pathogens or potentially pathogenic based on their best match species-level classification (Table 2). Of those 47, only 5 (9%) were obtained from the upstream site; the remaining 42 were found exclusively at the CSO site. Approximately one-third (31.0%) of these pathogens were Stenotrophomonas maltophilia. These were only isolated from the CSO site during an overflow event and had an average MDR of 7.7. Potentially pathogenic Pseudomonas isolates were the next most abundant group (21.3%). One potentially pathogenic Pseudomonas isolate was obtained from the upstream site, and the remainder were from the CSO site. The third most common potentially pathogenic bacteria were from the genus Chryseobacterium (specifically Chryseobacterium indologenes, Chryseobacterium gleum, and Chryseobacterium timonianum), which were found primarily at the CSO site during clear weather.

Table 2.
Distribution of potentially pathogenic taxa. Data are presented as the % in each classification/group relative to the total number of potentially pathogenic isolates for each sample. Mean MDR for each group is shown in parentheses.
4. Discussion
In this study, we focused on CSO discharge as a potential reservoir of pathogenic bacteria and MDR to an urbanized river. For all three of the antibiotics that we initially considered, we found that a much greater fraction of the culturable bacterial community was resistant during the CSO event (Figure 2). The increase was particularly striking for ampicillin. Young et al. [41] observed similar increases associated with wet-weather sampling of a CSO-impacted section of the Hudson River Estuary, and the fraction of culturable heterotrophs that were resistant was also similar to ours (~0.7% for tetracycline and ~10% for ampicillin). Ash et al. [42] also found widespread resistance to ampicillin when considering 77 samples from 16 U.S. rivers over the span of 3 years. High prevalence of ampicillin resistance relative to other common antibiotics was also evident in studies of cultured fecal coliforms by Guardabassi et al. [43] and West et al. [44]. Together, these findings motivated our subsequent focus on identifying the ampicillin-resistant isolates and screening for MDR.
4.1. Multidrug Resistance
Though there have been several recent studies examining the prevalence of MDR in aquatic ecosystems, most have focused on enteropathogenic microorganisms [45,46] and the dissemination of MDR to the broader microbial community is less often considered. These studies all found higher levels of MDR associated with polluted waterways and speculated that wastewater/sewage is the likely contamination source. Our study is unique in its consideration of MDR associated with point source wastewater inputs. We found MDR was elevated in isolates obtained from the CSO site. This was especially true during the CSO overflow event, where 82% of isolates were resistant to five or more antibiotics, and 44% of isolates were resistant to seven or more. The latter statistic contrasts starkly with data for the upstream site, where only 4% of isolates had this level of MDR (≥7). The observed increase in MDR at the CSO site is most likely due to the release of resistant organisms associated with the wastewater overflow and possibly sloughing of biofilms within the sewer system [27,47]. It is also possible that the antibiotics, disinfectants, and metals commonly found in wastewater and urban runoff exert selective pressure on the microorganisms at this site, as these pollutants have been correlated with increased prevalence of AR [13,48], even at low concentrations [49]. The selective pressure of antimicrobial residues creates an environment where bacteria with AR will have an increased survival rate in comparison to bacteria without AR [50,51,52], and thus comprise a greater fraction of the total community.
The mechanism driving increased MDR of isolates from the CSO site during clear weather is less obvious. One possible explanation is a legacy effect wherein the persistent exposure to antibiotics and inputs of MDR bacteria from CSO overflows have created a community that remains enriched even during clear weather. It is also possible that the urbanization of the catchment between the two sampling locations is a factor [9]. Of particular interest may be the elevated turbidity at the CSO site (Table 1), as previous studies indicate that suspended material can be important for the accumulation and dissemination of AR genes and microorganisms in rivers [53]. It is also possible that tidal flow carries MDR bacteria upstream to the CSO site from the discharge point of the wastewater treatment plant (Figure 1). Further research that considers multiple environmental compartments (e.g., sediment vs. water column) and additional sites is needed to distinguish these possibilities.
4.2. Resistance to Individual Antibiotics
We saw no difference in the frequency of resistance across sampling locations or conditions for three of the antibiotics that we considered (Figure 4): co-trimoxazole (overall average: 21%), streptomycin (66%), and vancomycin (82%). Low environmental resistance to co-trimoxazole is expected, given its pattern of limited clinical use [54]. Likewise, because streptomycin use is widespread in the agricultural and veterinary industries and the compound is among the more chemically-stable antibiotics [55], its higher frequency of resistance makes sense. In contrast, the high frequency of resistance for vancomycin was somewhat surprising; vancomycin resistance has previously been found in wastewater biofilms, surface water, and drinking water [12,56], but at much lower levels than in our study. It is important to keep in mind that vancomycin is generally ineffective against Gram-negative bacteria and unable to penetrate their outer membrane barrier. Approximately 85% of our isolates were Proteobacteria, which are Gram negative, so our high frequency of vancomycin resistance is primarily due to intrinsic natural resistance associated with cell wall and membrane structure. When only Gram-positive organisms are considered, the fraction displaying vancomycin resistance decreases to 50%.
For five of the six remaining antibiotics, resistance increased with wastewater exposure. Specifically, the number of isolates resistant to piperacillin, cefepime, cefotaxime, tetracycline, and amoxicillin clavulanate was always lowest for the upstream site, intermediate for the CSO site during clear weather, and highest for the CSO site during an overflow event (Figure 4). This difference was particularly high for cefepime, which is a fourth-generation cephalosporin with broad-spectrum antibacterial activity. Because resistance to cefepime is rare in environmental samples [57], the marked increase we saw at the CSO site is strong evidence of an anthropogenic source. Tetracycline and amoxicillin clavulanate (Augmentin) are also broad-spectrum antibiotics but with a longer history of use in both human and veterinary medicine [26,58]. Thus, our finding of higher baseline levels of resistance to these compounds at the upstream site, compared to cefepime, is very reasonable.
In addition to looking at the overall frequency of resistance across isolates, we examined the data for co-occurrence patterns following the approach outlined in Martins da Costa et al. [59] (Supplemental Figure S2). The strongest association among the resistance profiles was for vancomycin and ampicillin. Sanderson et al. [60] and Martins da Costa et al. [61] also observed a link between resistance to these two antibiotics, but further research is necessary to ascertain the mechanism of the relationship. We also observed a strong association of the resistance profiles for cefotaxime and amoxicillin clavulanate. Of the 93 isolates displaying resistance to cefotaxime, 84 (90%) also showed resistance to amoxicillin clavulanate, which suggests co-selection associated with resistance to these two compounds. Prior research by Tacao et al. [62] has also identified cefotaxime co-selection for bacteria isolated from polluted rivers. Co-selection is being increasingly recognized as a major driving force in the development of MDR [63]. These processes can be driven by direct selective pressure imposed by the antibiotics or indirectly via exposure to pollutants such as heavy metals, disinfectants, and elevated nutrients [64,65]. These processes may be especially important in aquatic ecosystems and wastewater, which integrate chemical and microbial contaminants from diverse industrial, agricultural, and domestic sources, creating incubation conditions that promote genetic exchange and contribute to the spread of antibiotic resistance.
4.3. Identification and Diversity of Resistant Bacteria
The diversity of MDR isolates was highest at the upstream site and significantly lower at the CSO site during the overflow event (Figure 5). This is consistent with several prior studies that have documented reduced diversity in environments experiencing wastewater pollution [66,67,68]. In our study, nearly all (97%) isolates were Proteobacteria and Bacteroidetes, which are among the phyla most commonly reported in rivers, tap water, and wastewater [69,70,71]. These phyla are recognized as major reservoirs of AR among environmental bacteria and include both antibiotic producers and degraders [72,73,74,75]. The majority of isolates were included in classes of ubiquitous Proteobacteria, specifically Alpha-, Beta-, and Gammaproteobacteria (87%) which includes many species found in the human microbiome [69,76]. Interestingly, 75% percent of the isolates classified as Gammaproteobacteria were obtained from the CSO site. This class includes several medically important groups, including Enterobacteriaceae and Pseudomonadaceae. Well-known pathogens such as Pseudomonas aeruginosa, Escherichia coli, and Salmonella are part of these groups. As hypothesized, potentially pathogenic isolates were more common at the CSO site and exhibited a greater level of MDR (Table 2 and Figure 6). In contrast, the upstream site included a greater abundance of the family Sphingomonadaceae and the genus Caulobacter. Sphingomonadaceae are commonly isolated from soils, freshwater, and marine habitats [77], and Caulobacter have been reported from a wide variety of aquatic habitats, mainly freshwater [78,79].
Our decision to use culture-based approaches in this study was motivated by a desire to understand the complexities associated with AR in clinically-relevant and viable organisms, but does present some limitations. Several prior studies have demonstrated that the abundance and diversity of organisms recovered during water quality assessment can be affected by cultivation conditions such as media type and incubation temperature. The general consensus is that R2A incubated at lower temperatures (20–25 °C) tends to yield higher levels of diversity [80,81], so we elected to use this approach to gain broad insight into environmental resistance. This choice of incubation conditions is likely the reason there were no fecal Enterobacteriaceae among our isolates from the CSO event sample, even though the E. coli data (Table 1) give clear evidence of fecal contamination. This result is consistent with previous studies showing that Enterobacteriaceae are not often detected in studies involving bacterial diversity unless selective media is used [69]. It is also important to keep in mind that our focus on ampicillin-resistant organisms further restricted the types of bacteria we isolated. Approximately half (54%) of our isolates were members of the orders of Enterobacteriales, Pseudomonadales, Burkholderiales, Bacteroidales, and Flavobacteriales, which have previously been selected for in ampicillin-enriched cultures from urban aquatic ecosystems [82].
When MDR of the individual microbial groups was considered, there were clear differences across samples (Figure 6). For example, though the abundance of the family Comamonadaceae was reasonably consistent (~20%), pathogenic species were primarily isolated from the CSO site, and MDR increased with the level of wastewater exposure. In the case of Stenotrophomonas, all of these isolates were obtained from the CSO site, and potentially pathogenic strains were only found during the CSO overflow event. Of particular interest was the appearance of Stenotrophomonas maltophilia, which was only found during the CSO overflow event and constituted nearly 30% of those isolates. While Stenotrophomonas maltophilia has occasionally been found in aquatic habitats, including plant rhizospheres, it is most well-known as a multidrug-resistant opportunistic pathogen that causes respiratory infections. For example, S. maltophilia has been found to be a co-colonizer with Pseudomonas aeruginosa in samples recovered from the respiratory tract of cystic fibrosis patients [83]. Members of the genus Pseudomonas were found in all samples, but abundance and MDR were higher for the CSO site. Numerous species of Pseudomonas were isolated in our samples, including P. fluorescens, P. mosselli, and P. putida, which have all been found to be opportunistic pathogens in humans [84]. Additionally, P. fluorescens has been reported to cause infection from contaminated drinking water [85].
We found MDR members of the genus Chryseobacterium at the CSO site during both sampling conditions. Although species of Chryseobacterium are often known as environmental bacteria, our samples included C. gleum and C. indologenes, which have been reported to cause a wide variety of infections in immunocompromised individuals (e.g., respiratory tract and urinary tract infections) and are intrinsically resistant to several antibiotics including those used in this study [86,87]. All of our Chryseobacterium isolates were resistant to tetracycline, streptomycin, cefotaxime, and amoxicillin clavulanate (Augmentin). This contributed to the high average resistance observed (8.3) (Table 2). The remaining pathogens observed were Acinetobacter, Aeromonas, Bordetella, Paenibacillus, Chromobacterium violaceum, Sphingopyxis witflariensis, D. acidovorans, C. testeroni, and P. alcalifaciens, which are all uncommon, but have been shown to cause infections in humans [88,89,90,91,92,93,94,95,96].
In addition to these potential pathogens, we isolated several groups of non-pathogenic human-associated bacteria, including Sphingomonas, Pseudomonas, Acinetobacter, and non-fecal Enterobacteriaceae. These organisms are common residents of the human microbiome and capable of resisting several different antibiotics [97,98,99,100,101]. We also isolated several members of the family Comamonadaceae from all sites. While Comamonadaceae is most commonly viewed as a large and diverse group of environmental bacteria, it is worth noting that they have recently been found to be abundant in urban runoff [102] and urban wastewater treatment plants [103], and can be relatively sensitive to antimicrobials [104,105].
5. Conclusions
Our results demonstrate that surface waters contaminated with wastewater discharge from CSOs can contain a diverse group of MDR bacteria, including several viable pathogenic species. Many MDR organisms were present at the CSO site even during periods of clear weather, which could represent a significant public health concern to local residents. Exposure to contaminated surface water used for drinking and recreation has previously been linked to adverse health outcomes due to bacterial exposure [106,107,108], and transmission of MDR further elevates the risk. Understanding the impact of persistent episodic contamination such as urban CSOs is especially important given increasing evidence that wastewater organisms can survive and even regrow in downstream rivers. Our findings complement many prior studies that have focused on the spread of antibiotic residuals and AR genes to aquatic ecosystems via wastewater inputs and urbanization, and raise unique concerns about multidrug resistance of clinically-relevant organisms that transition between the human and environmental microbiomes. The results also highlight the need for a holistic approach to wastewater management that includes not only consideration of alternate treatment approaches within WWTF (e.g., expanded tertiary treatment or incorporation of antibiotic removal technologies [2]) but also of further updates to existing infrastructure to reduce urban runoff and the frequency of CSO events. Reducing AR associated with episodic discharge will become increasingly important as storm intensity and frequency increase due to global change.
Supplementary Materials
The following are available online at https://www.mdpi.com/article/10.3390/w13152122/s1, Figure S1: Weekly E. coli data from the two years prior to sampling, Figure S2: Resistance profiles showing associations among antibiotics, Table S1: Number of isolates in each taxonomic group, Table S2: Taxonomic classifications and resistance profiles for all isolates.
Author Contributions
Conceptualization, E.S.L. and R.B.F.; Investigation, G.B. and E.S.L.; Methodology, G.B., E.S.L., J.M.B. and R.B.F.; Validation, G.B. and R.B.F.; Formal Analysis, R.B.F.; Data Curation, G.B. and R.B.F.; Visualization, R.B.F.; Writing—Original Draft, G.B. and R.B.F.; Writing—Review and Editing, G.B., J.M.B. and R.B.F.; Supervision, J.M.B. and R.B.F.; Project Administration, R.B.F.; Funding Acquisition, E.S.L. and R.B.F. All authors have read and agreed to the published version of the manuscript.
Funding
The project was supported by a VCU Rice Center Student Research Grant to Levengood and a VCU College of Humanities and Sciences Scholarship Catalyst Award to Franklin.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
MDR data and a summary of the sequencing results presented in this study are available in Supplementary Material (Table S1). All DNA sequences have been deposited into GenBank under accession numbers MZ490613 through MZ490746.
Acknowledgments
We thank Sunauz Moezzi and Shira Lanyi for assistance with lab work and William Mac Lee for help with field sampling and water quality data. This paper is contribution #93 from the VCU Rice Rivers Center.
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
References
- Rizzo, L.; Manaia, C.; Merlin, C.; Schwartz, T.; Dagot, C.; Ploy, M.C.; Michael, I.; Fatta-Kassinos, D. Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: A review. Sci. Total Environ. 2013, 447, 345–360. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Chu, L.; Wojnárovits, L.; Takács, E. Occurrence and fate of antibiotics, antibiotic resistant genes (ARGs) and antibiotic resistant bacteria (ARB) in municipal wastewater treatment plant: An overview. Sci. Total Environ. 2020, 744, 140997. [Google Scholar] [CrossRef] [PubMed]
- Chu, B.T.T.; Petrovich, M.L.; Chaudhary, A.; Wright, D.; Murphy, B.; Wells, G.; Poretsky, R. Metagenomics reveals the impact of wastewater treatment plants on the dispersal of microorganisms and genes in aquatic sediments. Appl. Environ. Microbiol. 2018, 84, e02168-17. [Google Scholar]
- Michael, I.; Rizzo, L.; McArdell, C.S.; Manaia, C.M.; Merlin, C.; Schwartz, T.; Dagot, C.; Fatta-Kassinos, D. Urban wastewater treatment plants as hotspots for the release of antibiotics in the environment: A review. Water Res. 2013, 47, 957–995. [Google Scholar] [CrossRef]
- Raza, S.; Jo, H.; Kim, J.; Shin, H.; Hur, H.G.; Unno, T. Metagenomic exploration of antibiotic resistome in treated wastewater effluents and their receiving water. Sci. Total Environ. 2021, 765, 142755. [Google Scholar]
- Rodríguez-Mozaz, S.; Chamorro, S.; Marti, E.; Huerta, B.; Gros, M.; Sànchez-Melsió, A.; Borrego, C.; Barcelo, D.; Balcázar, J.L. Occurrence of antibiotics and antibiotic resistance genes in hospital and urban wastewaters and their impact on the receiving river. Water Res. 2015, 69, 234–242. [Google Scholar] [CrossRef]
- Singh, R.; Singh, A.P.; Kumar, S.; Giri, B.S.; Kim, K.H. Antibiotic resistance in major rivers in the world: A systematic review on occurrence, emergence, and management strategies. J. Clean. Prod. 2019, 234, 1484–1505. [Google Scholar] [CrossRef]
- Tibbetts, J. Combined sewer systems: Down, dirty, and out of date. Environ. Health Perspect. 2005, 113, A464–A467. [Google Scholar] [CrossRef][Green Version]
- Honda, R.; Watanabe, T.; Sawaittayotin, V.; Masago, Y.; Chulasak, R.; Tanong, K.; Chaminda, G.; Wongsila, K.; Sienglum, C.; Sunthonwatthanaphong, V.; et al. Impacts of urbanization on the prevalence of antibiotic-resistant Escherichia coli in the Chaophraya River and its tributaries. Water Sci. Technol. 2015, 73, 362–374. [Google Scholar] [CrossRef]
- Port, J.A.; Cullen, A.C.; Wallace, J.C.; Smith, M.N.; Faustman, E.M. Metagenomic frameworks for monitoring antibiotic resistance in aquatic environments. Environ. Health Perspect. 2014, 122, 222–228. [Google Scholar] [CrossRef]
- Kumar, M.; Ram, B.; Honda, R.; Poopipattana, C.; Canh, V.; Chaminda, T.; Furumai, H. Concurrence of antibiotic resistant bacteria (ARB), viruses, pharmaceuticals and personal care products (PPCPs) in ambient waters of Guwahati, India: Urban vulnerability and resilience perspective. Sci. Total Environ. 2019, 693, 133640. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, T.; Kohnen, W.; Jansen, B.; Obst, U. Detection of antibiotic-resistant bacteria and their resistance genes in wastewater, surface water, and drinking water biofilms. FEMS Microbiol. Ecol. 2003, 43, 325–335. [Google Scholar] [CrossRef]
- Berendonk, T.U.; Manaia, C.M.; Merlin, C.; Fatta-Kassinos, D.; Cytryn, E.; Walsh, F.; Burgmann, H.; Sorum, H.; Norstrom, M.; Pons, M.N.; et al. Tackling antibiotic resistance: The environmental framework. Nat. Rev. Microbiol. 2015, 13, 310–317. [Google Scholar] [CrossRef] [PubMed]
- Garner, E.; Benitez, R.; von Wagoner, E.; Sawyer, R.; Schaberg, E.; Hession, W.; Krometis, L.; Badgley, B.; Pruden, A. Stormwater loadings of antibiotic resistance genes in an urban stream. Water Res. 2017, 123, 144–152. [Google Scholar] [CrossRef]
- Brown, P.C.; Borowska, E.; Peschke, R.; Schwartz, T.; Horn, H. Decay of elevated antibiotic resistance genes in natural river sediments after sedimentation of wastewater particles. Sci. Total Environ. 2020, 705, 135861. [Google Scholar] [CrossRef]
- Tennstedt, T.; Szczepanowski, R.; Braun, S.; Pühler, A.; Schlüter, A. Occurrence of integron-associated resistance gene cassettes located on antibiotic resistance plasmids isolated from a wastewater treatment plant. FEMS Microbiol. Ecol. 2003, 45, 239–252. [Google Scholar] [CrossRef]
- Von Wintersdorff, C.J.H.; Penders, J.; van Niekerk, J.M.; Mills, N.D.; Majumder, S.; van Alphen, L.B.; Savelkoul, P.H.M.; Wolffs, P.F.G. Dissemination of antimicrobial resistance in microbial ecosystems through horizontal gene transfer. Front. Microbiol. 2016, 7, 173. [Google Scholar] [CrossRef]
- Kim, S.; Aga, D.S. Potential ecological and human health impacts of antibiotics and antibiotic-resistant bacteria from wastewater treatment plants. J. Toxicol. Environ. Health Part B Crit. Rev. 2007, 10, 559–573. [Google Scholar] [CrossRef]
- Alekshun, M.N.; Levy, S.B. Molecular mechanisms of antibacterial multidrug resistance. Cell 2007, 128, 1037–1050. [Google Scholar] [CrossRef]
- Chen, B.; Liang, X.; Huang, X.; Zhang, T.; Li, X. Differentiating anthropogenic impacts on ARGs in the Pearl River Estuary by using suitable gene indicators. Water Res. 2013, 47, 2811–2820. [Google Scholar] [CrossRef]
- Garner, E.; Wallace, J.; Argoty, G.; Wilkinson, C.; Fahrenfeld, N.; Heath, L.; Zhang, L.; Arabi, M.; Aga, D.; Pruden, A. Metagenomic profiling of historic Colorado Front Range flood impact on distribution of riverine antibiotic resistance genes. Sci. Rep. 2016, 6, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Imchen, M.; Kumavath, R. Shotgun metagenomics reveals a heterogeneous prokaryotic community and a wide array of antibiotic resistance genes in mangrove sediment. FEMS Microbiol. Ecol. 2020, 96, 173. [Google Scholar] [CrossRef] [PubMed]
- Reichert, G.; Hilgert, S.; Alexander, J.; de Azevedo, J.C.R.; Morck, T.; Fuchs, S.; Schwartz, T. Determination of antibiotic resistance genes in a WWTP-impacted river in surface water, sediment, and biofilm: Influence of seasonality and water quality. Sci. Total Environ. 2021, 768, 144526. [Google Scholar] [CrossRef]
- Korzeniewska, E.; Korzeniewska, A.; Harnisz, M. Antibiotic resistant Escherichia coli in hospital and municipal sewage and their emission to the environment. Ecotoxicol. Environ. Saf. 2013, 91, 96–102. [Google Scholar] [CrossRef] [PubMed]
- Müller, H.; Sib, E.; Gajdiss, M.; Klanke, U.; Lenz-Plet, F.; Barabasch, V.; Albert, C.; Schallenberg, A.; Timm, C.; Zacharias, N.; et al. Dissemination of multi-resistant Gram-negative bacteria into German wastewater and surface waters. FEMS Microbiol. Ecol. 2018, 94, fiy057. [Google Scholar] [CrossRef]
- Vaz-Moreira, I.; Varela, A.R.; Pereira, T.V.; Fochat, R.C.; Manaia, C.M. Multidrug resistance in quinolone-resistant gram-negative bacteria isolated from hospital effluent and the municipal wastewater treatment plant. Microb. Drug Resist. 2016, 22, 155–163. [Google Scholar] [CrossRef]
- Osińska, A.; Harnisz, M.; Korzeniewska, E. Prevalence of plasmid-mediated multidrug resistance determinants in fluoroquinolone-resistant bacteria isolated from sewage and surface water. Environ. Sci. Pollut. Res. 2016, 23, 10818–10831. [Google Scholar] [CrossRef]
- Ho, J.Y.; Jong, M.C.; Acharya, K.; Liew, S.S.X.; Smith, D.R.; Noor, Z.Z.; Goodson, M.L.; Werner, D.; Graham, D.W.; Eswaran, J. Multidrug-resistant bacteria and microbial communities in a river estuary with fragmented suburban waste management. J. Hazard. Mater. 2021, 405, 124687. [Google Scholar] [CrossRef]
- Yewale, P.P.; Lokhande, K.B.; Sridhar, A.; Vaishnav, M.; Khan, F.A.; Mandal, A.; Swamy, K.V.; Jass, J.; Nawani, N. Molecular profiling of multidrug-resistant river water isolates: Insights into resistance mechanism and potential inhibitors. Environ. Sci. Pollut. Res. Int. 2020, 27, 27279–27292. [Google Scholar] [CrossRef]
- Levengood, E.S. Effect of combined sewer overflow on the abundance of antibiotic resistant bacteria in the James River. Master’s Thesis, Virginia Commonwealth University, Richmond, VA, USA, 2017. [Google Scholar]
- United States Environmental Protection Agency [EPA]. Method 1603: Escherichia Coli (E. coli) in Water by Membrane Filtration Using Modified Membrane-Thermotolerant Escherichia Coli Agar (Modified mTEC); United States Environmental Protection Agency: Washington, DC, USA, 2009. [Google Scholar]
- Sandegren, L. Selection of antibiotic resistance at very low antibiotic concentrations. Ups. J. Med. Sci. 2014, 119, 103–107. [Google Scholar] [CrossRef]
- Patel, J.B.; Cockerill, F.; Alder, J.; Bradford, P.; Eliopoulos, G.; Hardy, D.; Hindler, J.; Jenkins, S.; Lewis, J.; Miller, L.; et al. Performance Standards for Antimicrobial Susceptibility Testing: Twenty-Fourth Informational Supplement; CLSI: Wayne, PA, USA, 2014. [Google Scholar]
- Dhanda, G.; Sarkar, P.; Samaddar, S.; Haldar, J. Battle against vancomycin-resistant bacteria: Recent developments in chemical strategies. J. Med. Chem. 2019, 62, 3184–3205. [Google Scholar] [CrossRef] [PubMed]
- Werner, G.; Strommenger, B.; Witte, W. Acquired vancomycin resistance in clinically relevant pathogens. Future Microbiol. 2008, 3, 547–562. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, M.O.; Baptiste, K.E. Vancomycin-resistant Enterococci: A review of antimicrobial resistance mechanisms and perspectives of human and animal health. Microb. Drug Resist. 2018, 24, 590–606. [Google Scholar] [CrossRef] [PubMed]
- Ortez, J. Disk diffusion testing. In Manual of Antimicrobial Susceptibility Testing; Coyle, M.B., Ed.; American Society for Microbiology: Washington, DC, USA, 2005; pp. 39–52. [Google Scholar]
- Turner, S.; Pryer, K.M.; Miao, V.P.W.; Palmer, J.D. Investigating deep phylogenetic relationships among cyanobacteria and plastids by small subunit rRNA sequence analysis. J. Eukaryot. Microbiol. 1999, 46, 327–338. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.A. BioEdit: A User-Friendly Biological Sequence Alignment Editor and Analysis Program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 1999, 41, 95–98. [Google Scholar]
- Hammer, Ø.; Harper, D.A.T.; Ryan, P.D. PAST: Paleontological statistics software package for education and data analysis. Palaeontol. Electron. 2001, 4, 9. [Google Scholar]
- Young, S.; Juhl, A.; O’Mullan, G. Antibiotic-resistant bacteria in the Hudson River Estuary linked to wet weather sewage contamination. J. Water Health 2013, 11, 297–310. [Google Scholar] [CrossRef] [PubMed]
- Ash, R.J.; Mauck, B.; Morgan, M. Antibiotic resistance of gram-negative bacteria in rivers, United States. Emerg. Infect. Dis. 2002, 8, 713–716. [Google Scholar] [CrossRef]
- Guardabassi, L.; Lo Fo Wong, D.M.A.; Dalsgaard, A. The effects of tertiary wastewater treatment on the prevalence of antimicrobial resistant bacteria. Water Res. 2002, 36, 1955–1964. [Google Scholar] [CrossRef]
- West, B.M.; Liggit, P.; Clemans, D.L.; Francoeur, S.N. Antibiotic resistance, gene transfer, and water quality patterns observed in waterways near CAFO farms and wastewater treatment facilities. Water Air Soil Pollut. 2011, 217, 473–489. [Google Scholar] [CrossRef]
- Delgado-Gardea, M.; Tamez-Guerra, P.; Gomez-Flores, R.; Zavala-Díaz de la Serna, F.; Eroza-de la Vega, G.; Nevárez-Moorillón, G.; Pérez-Recoder, M.; Sánchez-Ramírez, B.; González-Horta, M.; Infante-Ramírez, R. Multidrug-resistant bacteria isolated from surface water in Bassaseachic Falls National Park, Mexico. Int. J. Environ. Res. Public Health 2016, 13, 597. [Google Scholar] [CrossRef] [PubMed]
- Odonkor, S.; Addo, K. Prevalence of multidrug-resistant Escherichia coli isolated from drinking water sources. Int. J. Microbiol. 2018, 2018, 7204013. [Google Scholar] [CrossRef] [PubMed]
- Auguet, O.; Pijuan, M.; Borrego, C.M.; Rodriguez-Mozaz, S.; Triadó-Margarit, X.; Giustina, S.V.D.; Gutierrez, O. Sewers as potential reservoirs of antibiotic resistance. Sci. Total Environ. 2017, 605, 1047–1054. [Google Scholar] [CrossRef] [PubMed]
- Almakki, A.; Jumas-Bilak, E.; Marchandin, H.; Licznar-Fajardo, P. Antibiotic resistance in urban runoff. Sci. Total Environ. 2019, 667, 64–76. [Google Scholar] [CrossRef]
- Karkman, A.; Do, T.T.; Walsh, F.; Virta, M.P.J. Antibiotic-resistance genes in waste water. Trends Microbiol. 2018, 3, 220–228. [Google Scholar] [CrossRef]
- Andersson, D.I.; Hughes, D. Persistence of antibiotic resistance in bacterial populations. FEMS Microbiol. Rev. 2011, 35, 901–911. [Google Scholar] [CrossRef]
- Barbosa, T.M.; Levy, S.B. The impact of antibiotic use on resistance development and persistence. Drug Resist. Updates 2000, 3, 303–311. [Google Scholar] [CrossRef]
- Martinez, J.L. Environmental pollution by antibiotics and by antibiotic resistance determinants. Environ. Pollut. 2009, 157, 2893–2902. [Google Scholar] [CrossRef]
- Barrón, M.D.L.C.; Merlin, C.; Guilloteau, H.; Montargès-Pelletier, E.; Bellanger, X. Suspended materials in river waters differentially enrich Class 1 Integron- and IncP-1 plasmid-carrying bacteria in sediments. Front. Microbiol. 2018, 9, 1443. [Google Scholar] [CrossRef]
- Kim, S.; Thiessen, P.A.; Bolton, E.E.; Chen, J.; Fu, G.; Gindulyte, A.; Han, L.; He, J.; He, S.; Shoemaker, B.A.; et al. PubChem Substance and Compound databases. Nucleic Acids Res. 2016, 44, D1202–D1213. [Google Scholar] [CrossRef]
- Shen, Y.; Zhao, W.; Zhang, C.; Shi, Y.S. Degradation of streptomycin in aquatic environment: Kinetics, pathway, and antibacterial activity analysis. Environ. Sci. Pollut. Res. 2017, 24, 14337. [Google Scholar] [CrossRef]
- Morris, D.; Galvin, S.; Boyle, F.; Hickey, P.; Mulligan, M.; Cormican, M. Enterococcus faecium of the vanA genotype in rural drinking water, effluent, and the aqueous environment. Appl. Environ. Microbiol. 2011, 78, 596–598. [Google Scholar] [CrossRef] [PubMed]
- Carsenti-Etesse, H.; Cavallo, J.; Roger, P.; Ziha-Zarifi, I.; Plesiat, P.; Garrabé, E.; Dellamonica, P. Effect of β-lactam antibiotics on the in vitro development of resistance in Pseudomonas aeruginosa. Clin. Microbiol. Infect. 2001, 7, 144–151. [Google Scholar] [CrossRef]
- Pezzella, C.; Ricci, A.; DiGiannatale, E.; Luzzi, I.; Carattoli, A. Tetracycline and streptomycin resistance genes, transposons, and plasmids in Salmonella enterica isolates from animals in Italy. Antimicrob. Agents Chemother. 2004, 48, 903–908. [Google Scholar] [CrossRef] [PubMed]
- Martins da Costa, P.M.; Vaz-Pires, P.M.; Bernardo, F.M. Antibiotic resistance of Enterococcus spp. isolated from wastewater and sludge of poultry slaughterhouses. J. Environ. Sci. Health Part B 2006, 41, 1393–1403. [Google Scholar] [CrossRef] [PubMed]
- Sanderson, H.; Ortega-Polo, R.; McDermott, K.; Hall, G.; Zaheer, R.; Brown, R.; Majury, A.; McAllister, T.; Liss, S. Quantification and multidrug resistance profiles of vancomycin-resistant Enterococci isolated from two wastewater treatment plants in the same municipality. Microorganisms 2019, 7, 626. [Google Scholar] [CrossRef]
- Martins da Costa, P.; Vaz-Pires, P.; Bernardo, F. Antimicrobial resistance in Enterococcus spp. isolated in inflow, effluent and sludge from municipal sewage water treatment plants. Water Res. 2006, 40, 1735–1740. [Google Scholar] [CrossRef]
- Tacão, M.; Moura, A.; Correia, A.; Henriques, I. Co-resistance to different classes of antibiotics among ESBL-producers from aquatic systems. Water Res. 2014, 48, 100–107. [Google Scholar] [CrossRef]
- Baindara, P. Mechanism of Bacterial Co-resistance. In Bacterial Adaptation to Co-Resistance; Mandal, S., Paul, D., Eds.; Springer: Singapore, 2019; pp. 191–210. [Google Scholar]
- Grenni, P.; Corno, G. Knowledge gaps and research needs in bacterial co-resistance in the environment. In Bacterial Adaptation to Co-Resistance; Mandal, S., Paul, D., Eds.; Springer: Singapore, 2019; pp. 39–59. [Google Scholar]
- Squadrone, S. Water environments: Metal-tolerant and antibiotic-resistant bacteria. Environ. Monit. Assess. 2020, 192, 1–12. [Google Scholar] [CrossRef]
- Marathe, N.P.; Shetty, S.A.; Shouche, Y.S.; Larsson, D.G.J. Limited bacterial diversity within a treatment plant receiving antibiotic-containing waste from bulk drug production. PLoS ONE 2016, 11, e0165914. [Google Scholar] [CrossRef] [PubMed]
- Calderon, O.; Porter-Morgan, H.; Jacob, J.; Elkins, W. Bacterial diversity impacts as a result of combined sewer overflow in a polluted waterway. Glob. J. Environ. Sci. Manag. 2017, 3, 437–446. [Google Scholar]
- Wang, X.; Gu, J.; Gao, H.; Qian, X.; Li, H. Abundances of clinically relevant antibiotic resistance genes and bacterial community diversity in the Weihe River, China. Int. J. Environ. Res. Public Health 2018, 15, 708. [Google Scholar] [CrossRef]
- Vaz-Moreira, I.; Nunes, O.C.; Manaia, C.M. Bacterial diversity and antibiotic resistance in water habitats: Searching the links with the human microbiome. FEMS Microbiol. Rev. 2014, 38, 761–778. [Google Scholar] [CrossRef] [PubMed]
- Bai, Y.; Qi, W.; Liang, J.; Qu, J. Using high-throughput sequencing to assess the impacts of treated and untreated wastewater discharge on prokaryotic communities in an urban river. Appl. Microbiol. Biotechnol. 2013, 98, 1841–1851. [Google Scholar] [CrossRef] [PubMed]
- Ibekwe, A.M.; Leddy, M.; Murinda, S.E. Potential human pathogenic bacteria in a mixed urban watershed as revealed by pyrosequencing. PLoS ONE 2013, 8, e79490. [Google Scholar] [CrossRef]
- Dantas, G.; Sommer, M.O.A.; Oluwasegun, R.D.; Church, G.M. Bacteria subsisting on antibiotics. Science 2008, 320, 100–103. [Google Scholar] [CrossRef] [PubMed]
- Forsberg, K.J.; Reyes, A.; Wang, B.; Selleck, E.M.; Sommer, M.O.A.; Dantas, G. The shared antibiotic resistome of soil bacteria and human pathogens. Science 2012, 337, 1107–1111. [Google Scholar] [CrossRef]
- D’costa, V.M.; Mcgrann, K.M.; Hughes, D.W.; Wright, G.D. Sampling the antibiotic resistome. Science 2006, 311, 374–377. [Google Scholar] [CrossRef] [PubMed]
- Riesenfeld, C.S.; Goodman, R.M.; Handelsman, J. Uncultured soil bacteria are a reservoir of new antibiotic resistance genes. Environ. Microbiol. 2004, 6, 981–989. [Google Scholar] [CrossRef]
- Hugon, P.; Dufour, J.-C.; Colson, P.; Fournier, P.-E.; Sallah, K.; Raoult, D. A comprehensive repertoire of prokaryotic species identified in human beings. Lancet Infect. Dis. 2015, 15, 1211–1219. [Google Scholar] [CrossRef]
- Glaeser, S.P.; Kämpfer, P. The Family Sphingomonadaceae. In The Prokaryotes; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; pp. 641–707. [Google Scholar]
- Abraham, W.R.; Rohde, M.; Bennasar, A. The Family Caulobacteraceae. In The Prokaryotes; Rosenberg, E., DeLong, E.F., Lory, S., Stackebrandt, E., Thompson, F., Eds.; Springer: Berlin/Heidelberg, Germany, 2014; pp. 179–205. [Google Scholar]
- Ariskina, E.V.; Chernousova, E.Y.; Lapteva, N.A.; Akimov, V.N. Evaluation of the taxonomic diversity of prosthecate bacteria belonging to the genera Brevundimonas and Caulobacter isolated from various Eurasian ecosystems by analysis of the 16S rRNA genes. Microbiology 2011, 80, 403–410. [Google Scholar] [CrossRef]
- Gensberger, E.; Gössl, E.; Antonielli, L.; Sessitsch, A.; Kostić, T. Effect of different heterotrophic plate count methods on the estimation of the composition of the culturable microbial community. PeerJ 2015, 3, 862. [Google Scholar] [CrossRef]
- Reasoner, D.J. Heterotrophic plate count methodology in the United States. Int. J. Food Microbiol. 2004, 92, 307–315. [Google Scholar] [CrossRef] [PubMed]
- Coutinho, F.H.; Silveira, C.B.; Pinto, L.H.; Salloto, G.R.B.; Cardoso, A.M.; Martins, O.B.; Vieira, R.P.; Clementino, M.M. Antibiotic resistance is widespread in urban aquatic environments of Rio de Janeiro, Brazil. Microb. Ecol. 2014, 68, 441–452. [Google Scholar] [CrossRef]
- Brooke, J.S. Stenotrophomonas maltophilia: An emerging global opportunistic pathogen. Clin. Microbiol. Rev. 2012, 25, 2–41. [Google Scholar] [CrossRef] [PubMed]
- Leneveu-Jenvrin, C.; Madi, A.; Bouffartigues, E.; Biaggini, K.; Feuilloley, M.; Chevalier, S.; Connil, N. Cytotoxicity and inflammatory potential of two Pseudomonas mosselii strains isolated from clinical samples of hospitalized patients. BMC Microbiol. 2013, 13, 123. [Google Scholar] [CrossRef] [PubMed]
- Wong, V.; Levi, K.; Baddal, B.; Turton, J.; Boswell, T.C. Spread of Pseudomonas fluorescens due to contaminated drinking water in a bone marrow transplant unit. J. Clin. Microbiol. 2011, 49, 2093–2096. [Google Scholar] [CrossRef] [PubMed]
- Abdalhamid, B.; Elhadi, N.; Alsamman, K.; Aljindan, R. Chryseobacterium gleum pneumonia in an infant with nephrotic syndrome. IDCases 2016, 5, 34–36. [Google Scholar] [CrossRef] [PubMed]
- Loch, T.P.; Faisal, M. Emerging flavobacterial infections in fish: A review. J. Adv. Res. 2015, 6, 283–300. [Google Scholar] [CrossRef] [PubMed]
- Wen, H.; Wang, K.; Liu, Y.; Tay, M.; Lauro, F.M.; Huang, H.; Wu, H.; Liang, H.; Ding, Y.; Givskov, M.; et al. Population dynamics of an Acinetobacter baumannii clonal complex during colonization of patients. J. Clin. Microbiol. 2014, 52, 3200–3208. [Google Scholar] [CrossRef]
- Igbinosa, I.H.; Igumbor, E.U.; Aghdasi, F.; Tom, M.; Okoh, A.I. Emerging Aeromonas species infections and their significance in public health. Sci. World J. 2012, 2012, 625023. [Google Scholar] [CrossRef]
- Finger, H.; von Koenig, C.H.W. Chapter 31: Bordetella. In Medical Microbiology, 4th ed.; Baron, S., Ed.; University of Texas Medical Branch at Galveston: Galveston, TX, USA, 1996. [Google Scholar]
- Ouyang, J.; Pei, Z.; Lutwick, L.; Dalal, S.; Yang, L.; Cassai, N.; Sandhu, K.; Hanna, B.; Wieczorek, R.; Bluth, M.; et al. Case report: Paenibacillus thiaminolyticus: A new cause of human infection, inducing bacteremia in a patient on hemodialysis. Ann. Clin. Lab. Sci. 2008, 38, 393–400. [Google Scholar] [PubMed]
- Batista, J.H.; Neto, J.F.D.S. Chromobacterium violaceum pathogenicity: Updates and insights from genome sequencing of novel Chromobacterium species. Front. Microbiol. 2017, 8, 2213. [Google Scholar] [CrossRef] [PubMed]
- Kämpfer, P.; Denner, E.B.M.; Witzenberger, R.; Neef, A.; Busse, H.-J. Sphingopyxis witflariensis sp. nov., isolated from activated sludge. Int. J. Syst. Evol. Microbiol. 2002, 52, 2029–2034. [Google Scholar]
- Bilgin, H.; Sarmis, A.; Tigen, E.; Soyletir, G.; Mulazimoglu, L. Delftia Acidovorans: A rare pathogen in immunocompetent and immunocompromised patients. Can. J. Infect. Dis. Med. Microbiol. 2015, 26, 277–279. [Google Scholar] [CrossRef]
- Bayhan, G.I.; Tanir, G.; Karaman, I.; Ozkan, S. Comamonas testosteroni: An unusual bacteria associated with acute appendicitis. Balk. Med. J. 2013, 30, 447–448. [Google Scholar] [CrossRef] [PubMed]
- Haynes, J.; Hawkey, P.M. Providencia alcalifaciens and travellers diarrhoea. BMJ 1989, 299, 94. [Google Scholar] [CrossRef]
- Xi, C.; Zhang, Y.; Marrs, C.F.; Ye, W.; Simon, C.; Foxman, B.; Nriagu, J. Prevalence of antibiotic resistance in drinking water treatment and distribution systems. Appl. Environ. Microbiol. 2009, 75, 5714–5718. [Google Scholar] [CrossRef]
- Vaz-Moreira, I.; Nunes, O.C.; Manaia, C.M. Diversity and antibiotic resistance patterns of Sphingomonadaceae isolates from drinking water. Appl. Environ. Microbiol. 2011, 77, 5697–5706. [Google Scholar] [CrossRef] [PubMed]
- Vaz-Moreira, I.; Nunes, O.C.; Manaia, C.M. Diversity and antibiotic resistance in Pseudomonas spp. from drinking water. Sci. Total Environ. 2012, 426, 366–374. [Google Scholar] [CrossRef]
- Figueira, V.; Serra, E.A.; Vaz-Moreira, I.; Brandão, T.R.S.; Manaia, C.M. Comparison of ubiquitous antibiotic-resistant Enterobacteriaceae populations isolated from wastewaters, surface waters and drinking waters. J. Water Health 2011, 10, 1–10. [Google Scholar] [CrossRef]
- Narciso-Da-Rocha, C.; Vaz-Moreira, I.; Svensson-Stadler, L.; Moore, E.R.B.; Manaia, C.M. Diversity and antibiotic resistance of Acinetobacter spp. in water from the source to the tap. Appl. Microbiol. Biotechnol. 2012, 97, 329–340. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Suits, M.; Wituszynski, D.; Winston, R.; Martin, J.; Lee, J. Residential urban stormwater runoff: A comprehensive profile of microbiome and antibiotic resistance. Sci. Total Environ. 2020, 723, 138033. [Google Scholar] [CrossRef] [PubMed]
- Fernandes, T.; Vaz-Moreira, I.; Manaia, C.M. Neighbor urban wastewater treatment plants display distinct profiles of bacterial community and antibiotic resistance genes. Environ. Sci. Pollut. Res. 2019, 26, 11269–11278. [Google Scholar] [CrossRef] [PubMed]
- Saunders, A.M.; Albertsen, M.; Vollertsen, J.; Nielsen, P.H. The activated sludge ecosystem contains a core community of abundant organisms. ISME J. 2016, 10, 11–20. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Wan, Y.; Mao, Y.; Zhang, T. Partnership of Arthrobacter and Pimelobacter in aerobic degradation of sulfadiazine revealed by metagenomics analysis and isolation. Environ. Sci. Technol. 2018, 52, 2963–2972. [Google Scholar] [CrossRef]
- Curriero, F.C.; Patz, J.A.; Rose, J.B.; Lele, S. The association between extreme precipitation and waterborne disease outbreaks in the United States, 1948–1994. Am. J. Public Health 2001, 91, 1194–1199. [Google Scholar] [CrossRef] [PubMed]
- Nichols, G.; Lane, C.; Asgari, N.; Verlander, N.Q.; Charlett, A. Rainfall and outbreaks of drinking water related disease and in England and Wales. J. Water Health 2009, 7, 1–8. [Google Scholar] [CrossRef]
- Rose, J.; Daeschner, S.; Easterling, D.; Curriero, F.; Lele, S.; Patz, J.A. Climate and waterborne disease outbreaks. J. Am. Water Work. Assoc. 2000, 92, 77–87. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).