Design and Evaluation of a Millifluidic Insulator-Based Dielectrophoresis (DEP) Retention Device to Separate Bacteria from Tap Water
Abstract
1. Introduction
2. Materials and Methods
2.1. Device Design and Fabrication
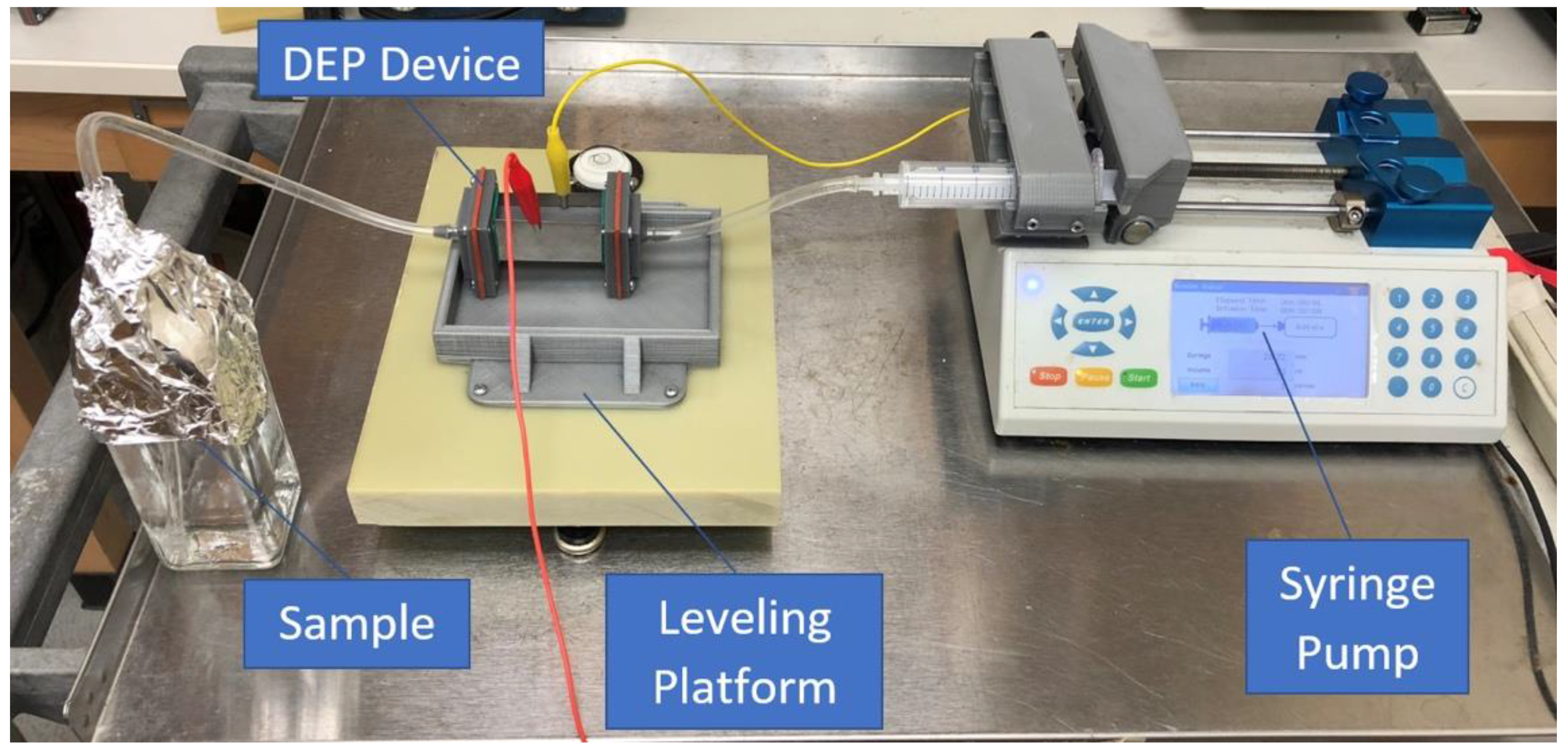
2.2. Experimental Set-Up
2.3. Microbial Preparation
2.4. Experimental Design and Quantification of Removal Efficiency
3. Results and Discussion
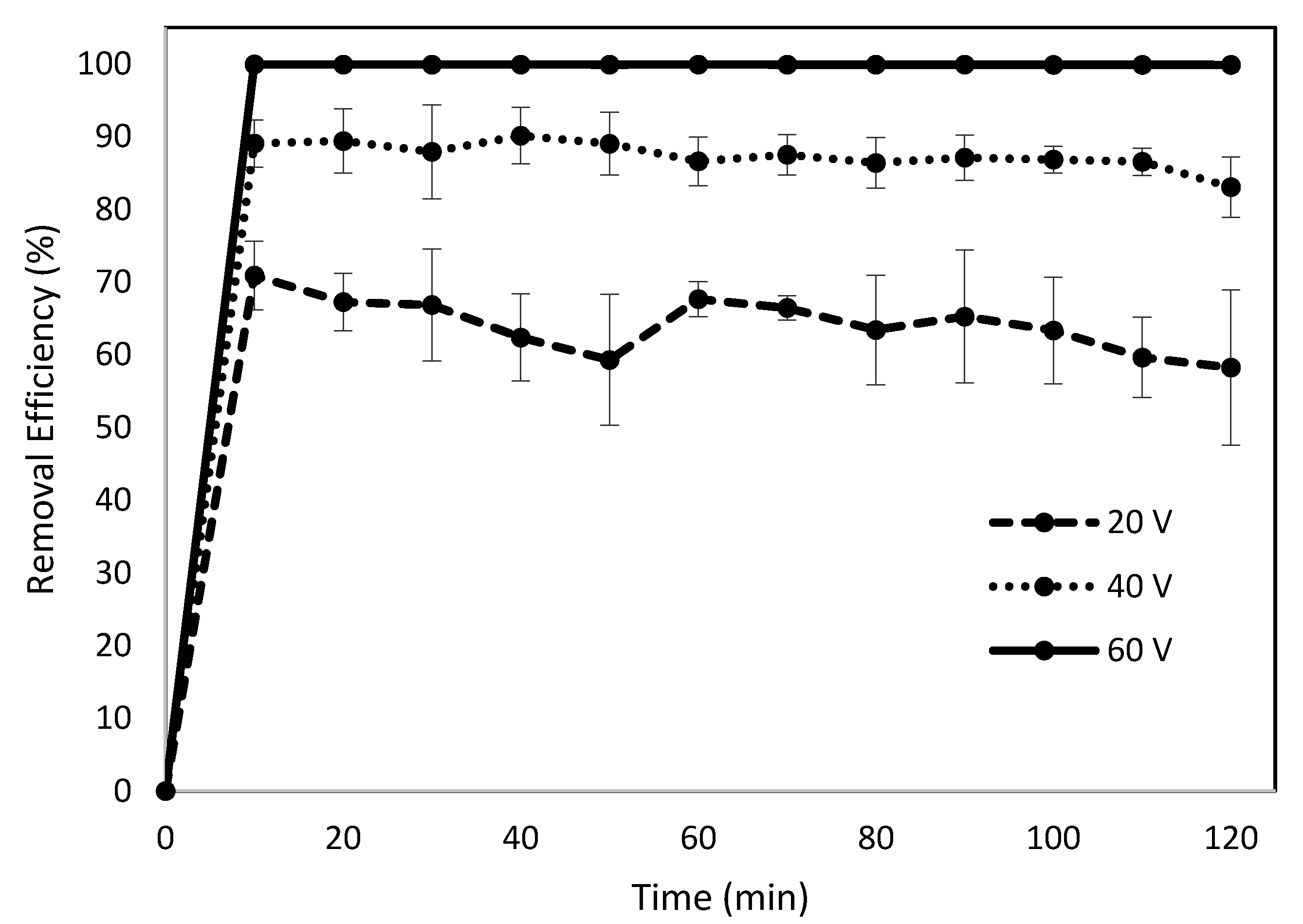
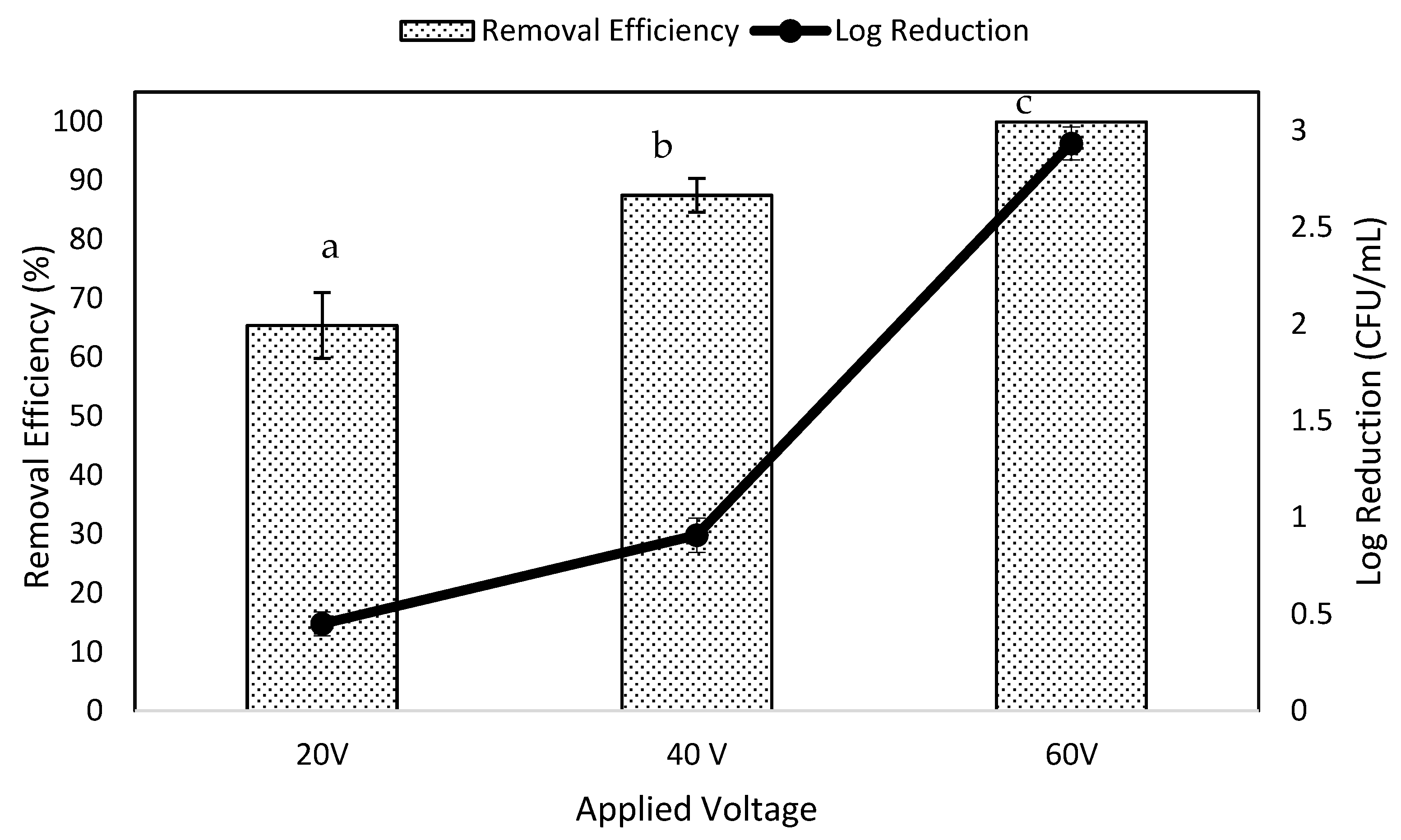
3.1. Effects of Voltage on Removal Efficiency
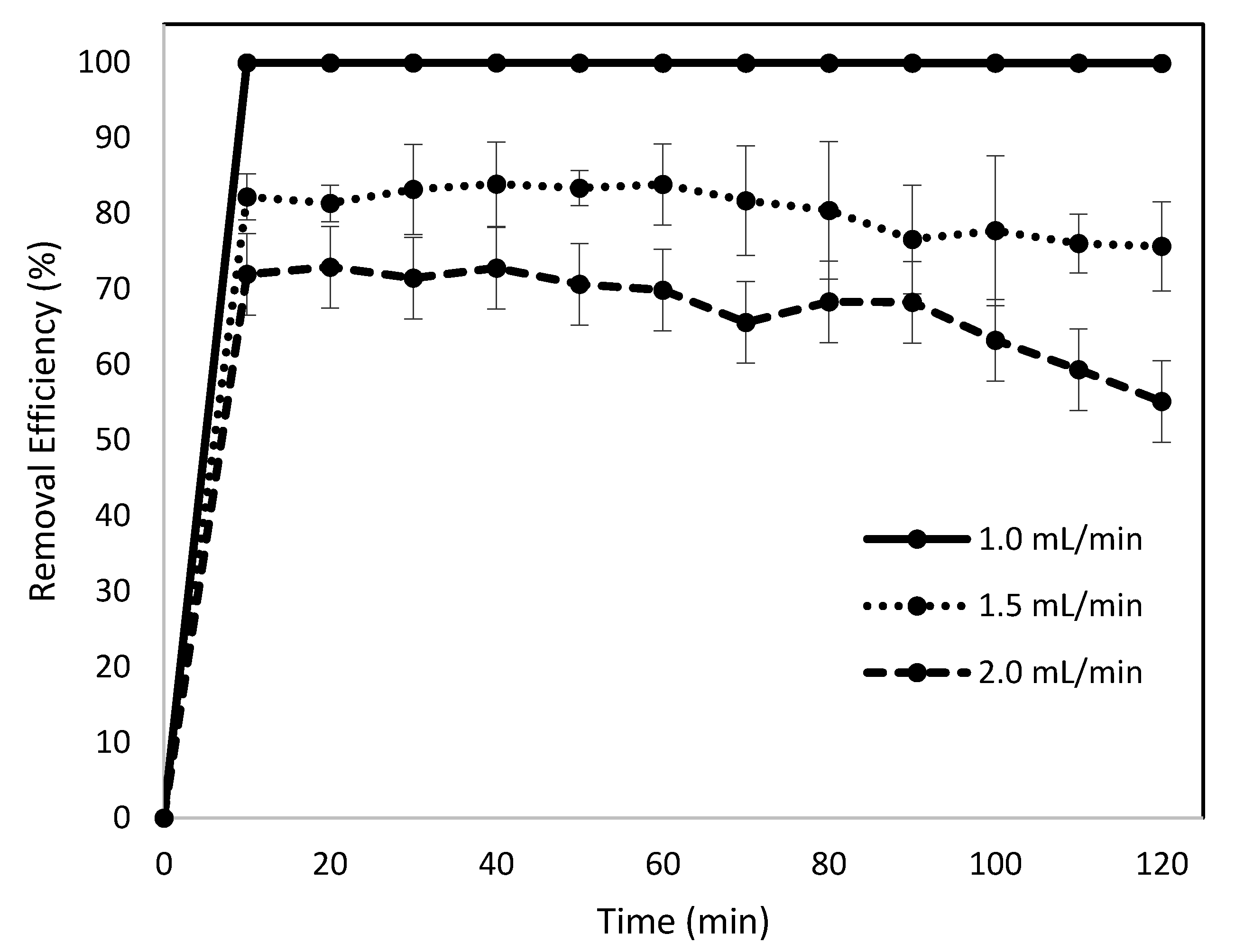
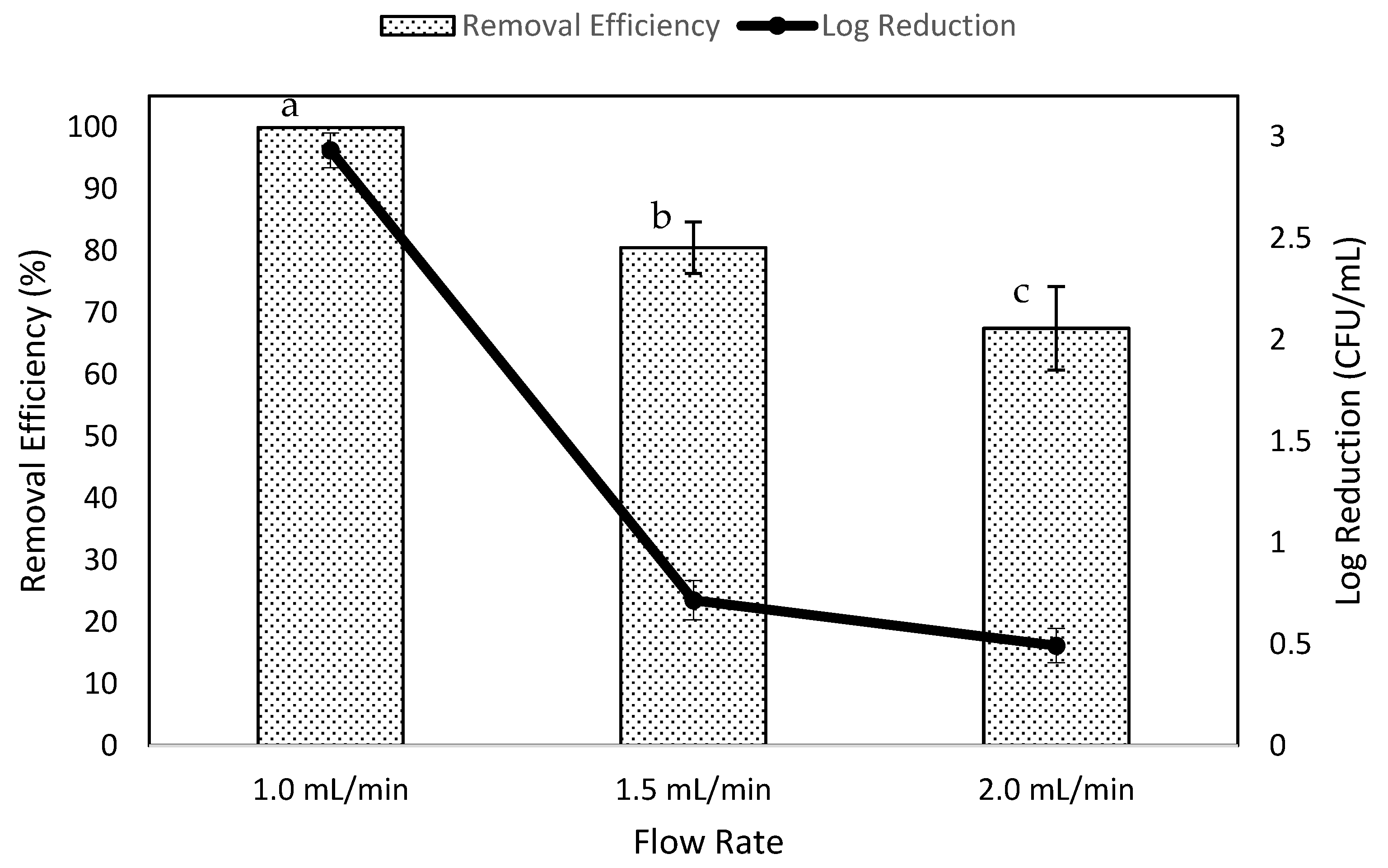
3.2. Effects of Flow Rate on Removal Efficiency
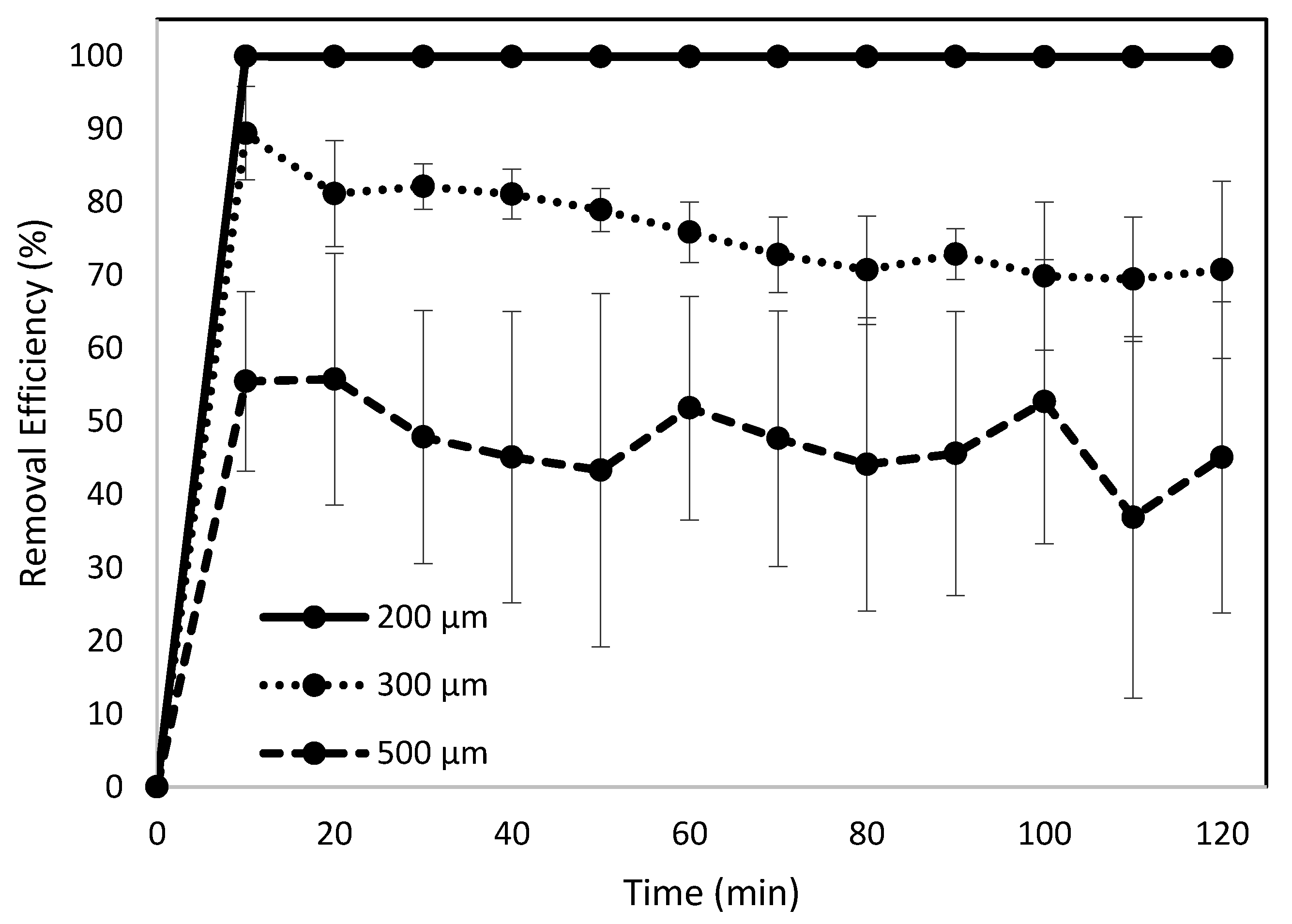
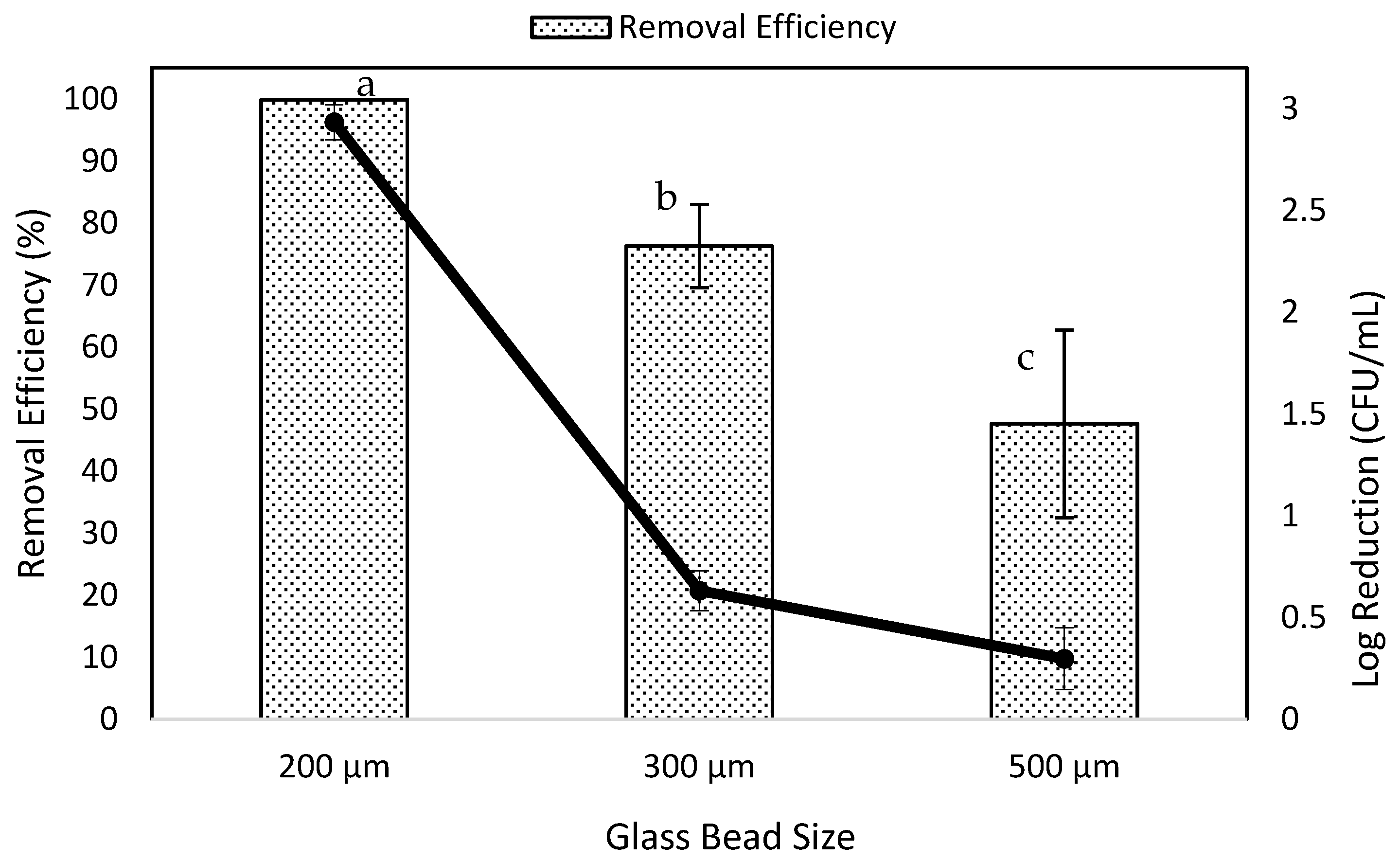
3.3. Effects of Bead Size on Removal Efficiency
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Gleick, P.H. The Human Right to Water. Water Policy 1998, 1, 487–503. [Google Scholar] [CrossRef]
- World Health Organization. Progress on Drinking Water, Sanitation and Hygiene: 2017 Update and SDG Baselines. 2017. Available online: https://www.who.int/mediacentre/news/releases/2017/launch-version-report-jmp-water-sanitation-hygiene.pdf (accessed on 20 February 2021).
- Grey, D.; Sadoff, C.W. Sink or Swim? Water Security for Growth and Development. Water Policy 2007, 9, 545–571. [Google Scholar] [CrossRef]
- Falkenmark, M. The Greatest Water Problem: The Inability to Link Environmental Security, Water Security and Food Security. Int. J. Water Resour. Dev. 2001, 17, 539–554. [Google Scholar] [CrossRef]
- Mekonnen, M.M.; Hoekstra, A.Y. Four Billion People Facing Severe Water Scarcity. Sci. Adv. 2016, 2, e1500323. [Google Scholar] [CrossRef] [PubMed]
- Vörösmarty, C.J.; McIntyre, P.B.; Gessner, M.O.; Dudgeon, D.; Prusevich, A.; Green, P.; Glidden, S.; Bunn, S.; Sullivan, C.A.; Liermann, R.C.; et al. Global threats to human water security and river biodiversity. Nature 2010, 467, 555–561. [Google Scholar] [CrossRef] [PubMed]
- Boyce, T.G.; Swerdlow, D.L.; Patricia, M. Griffin Escherichia Coli O157: H7 and the Hemolytic–Uremic Syndrome. N. Engl. J. Med. 1995, 333, 364–368. [Google Scholar] [CrossRef]
- Wang, G.; Michael, P. Doyle Survival of Enterohemorrhagic Escherichia Coli O157: H7 in Water. J. Food Prot. 1998, 61, 662–667. [Google Scholar] [CrossRef]
- Al-Karaghouli, A.; Kazmerski, L.L. Energy Consumption and Water Production Cost of Conventional and Renewable-Energy-Powered Desalination Processes. Renew. Sustain. Energy Rev. 2013, 24, 343–356. [Google Scholar] [CrossRef]
- Shannon, M.A.; Bohn, P.W.; Elimelech, M.; Georgiadis, J.G.; Marinas, B.J.; Mayes, A.M. Science and Technology for Water Purification in the Coming Decades. Nanosci. Technol. A Collect. Rev. Nat. J. 2010, 337–346. [Google Scholar] [CrossRef]
- Hua, G.; Reckhow, D.A. Comparison of Disinfection Byproduct Formation from Chlorine and Alternative Disinfectants. Water Res. 2007, 41, 1667–1678. [Google Scholar] [CrossRef]
- Boorman, G.A. Drinking Water Disinfection Byproducts: Review and Approach to Toxicity Evaluation. Environ. Health Perspect. 1999, 107, 207–217. [Google Scholar] [PubMed]
- van Reis, R.; Zydney, A. Bioprocess Membrane Technology. J. Membr. Sci. 2007, 297, 16–50. [Google Scholar] [CrossRef]
- Madaeni, S.S. The Application of Membrane Technology for Water Disinfection. Water Res. 1999, 33, 301–308. [Google Scholar] [CrossRef]
- Gao, W.; Liang, H.; Ma, J.; Han, M.; Chen, Z.; Han, Z.; Li, G. Membrane Fouling Control in Ultrafiltration Technology for Drinking Water Production: A Review. Desalination 2011, 272, 1–8. [Google Scholar] [CrossRef]
- Plappally, A.K. Energy Requirements for Water Production, Treatment, End Use, Reclamation, and Disposal. Renew. Sustain. Energy Rev. 2012, 16, 4818–4848. [Google Scholar] [CrossRef]
- Kimura, K.; Hane, Y.; Watanabe, Y.; Amy, G.; Ohkuma, N. Irreversible Membrane Fouling during Ultrafiltration of Surface Water. Water Res. 2004, 38, 3431–3441. [Google Scholar] [CrossRef]
- Pohl, H.A. Dielectrophoresis: The Behavior of Neutral Matter in Nonuniform Electric Fields (Cambridge Monographs on Physics); Cambridge University Press: Cambridge, UK, 1978. [Google Scholar]
- Pethig, R. Dielectrophoresis: Status of the Theory, Technology, and Applications. Biomicrofluidics 2010, 4, 22811. [Google Scholar] [CrossRef] [PubMed]
- Pethig, R.; Markx, G.H. Applications of Dielectrophoresis in Biotechnology. Trends Biotechnol. 1997, 15, 426–432. [Google Scholar] [CrossRef]
- Hawkins, B.G.; Smith, A.E.; Syed, Y.A.; Kirby, B.J. Continuous-Flow Particle Separation by 3D Insulative Dielectrophoresis Using Coherently Shaped, Dc-Biased, Ac Electric Fields. Anal. Chem. 2007, 79, 7291–7300. [Google Scholar] [CrossRef]
- Doh, I.; Cho, Y.-H. A Continuous Cell Separation Chip Using Hydrodynamic Dielectrophoresis (DEP) Process. Sens. Actuators A Phys. 2005, 121, 59–65. [Google Scholar] [CrossRef]
- Pamme, N. Continuous Flow Separations in Microfluidic Devices. Lab A Chip 2007, 7, 1644–1659. [Google Scholar] [CrossRef]
- Cheng, I.-F.; Chang, H.-C.; Hou, D.; Chang, H.-C. An Integrated Dielectrophoretic Chip for Continuous Bioparticle Filtering, Focusing, Sorting, Trapping, and Detecting. Biomicrofluidics 2007, 1, 21503. [Google Scholar] [CrossRef] [PubMed]
- Pohl, H.A.; Crane, J.S. Dielectrophoresis of Cells. Biophys. J. 1971, 11, 711–727. [Google Scholar] [CrossRef]
- Srivastava, S.K.; Gencoglu, A.; Minerick, A.R. DC Insulator Dielectrophoretic Applications in Microdevice Technology: A Review. Anal. Bioanal. Chem. 2011, 399, 301–321. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Zheng, Y.; Akin, D.; Bashir, R. Characterization and Modeling of a Microfluidic Dielectrophoresis Filter for Biological Species. J. Microelectromech. Syst. 2005, 14, 103–112. [Google Scholar] [CrossRef]
- Cummings, E.B.; Singh, A.K. Dielectrophoresis in Microchips Containing Arrays of Insulating Posts: Theoretical and Experimental Results. Anal. Chem. 2003, 75, 4724–4731. [Google Scholar] [CrossRef]
- Baylon-Cardiel, J.L.; Lapizco-Encinas, B.H.; Reyes-Betanzo, C.; Chávez-Santoscoy, A.V.; MartÌnez-Chapa, S.O. Prediction of Trapping Zones in an Insulator-Based Dielectrophoretic Device. Lab A Chip 2009, 9, 2896–2901. [Google Scholar] [CrossRef]
- Lapizco-Encinas, B.H.; Davalos, R.V.; Simmons, B.A.; Cummings, E.B.; Fintschenko, Y. An Insulator-Based (Electrodeless) Dielectrophoretic Concentrator for Microbes in Water. J. Microbiol. Methods 2005, 62, 317–326. [Google Scholar] [CrossRef]
- Masuda, S.; Itagaki, T.; Kosakada, M. Detection of Extremely Small Particles in the Nanometer and Ionic Size Range. IEEE Trans. Ind. Appl. 1988, 24, 740–744. [Google Scholar] [CrossRef]
- Cummings, E.B.; Singh, A.K. Dielectrophoretic Trapping without Embedded Electrodes. In Microfluidic Devices and Systems III; SPIE: Bellingham, WA, USA, 2000; Volume 4177, pp. 151–160. [Google Scholar]
- Chou, C.-F.; Tegenfeldt, J.O.; Bakajin, O.; Chan, S.S.; Cox, E.C.; Darnton, N.; Duke, T.; Austin, R.H. Electrodeless Dielectrophoresis of Single-and Double-Stranded DNA. Biophys. J. 2002, 83, 2170–2179. [Google Scholar] [CrossRef]
- Zhou, G.; Imamura, M.; Suehiro, J.; Hara, M. A Dielectrophoretic Filter for Separation and Collection of Fine Particles Suspended in Liquid. In Proceedings of the Conference Record of the 2002 IEEE Industry Applications Conference, 37th IAS Annual Meeting (Cat. No. 02CH37344), Pittsburgh, PA, USA, 13–18 October 2002; Volume 2, pp. 1404–1411. [Google Scholar]
- McGraw, G.J.; Davalos, R.V.; Brazzle, J.D.; Hachman, J.; Hunter, M.; Chames, J.; Fiechtner, G.J.; Cummings, E.B.; Fintschenko, Y.; Simmons, B.A. Polymeric Insulator-Based Dielectrophoresis (IDEP) for the Monitoring of Water-Borne Pathogens. In Proceedings of the ASPE Spring Topical Meeting on Precision Micro/Nano Scale Polymer Based Component and Device Fabrication, ASPE 2005, Cambridge, UK, 18–29 April 2005. [Google Scholar]
- Lapizco-Encinas, B.H.; Simmons, B.A.; Cummings, E.B.; Fintschenko, Y. Insulator-Based Dielectrophoresis for the Selective Concentration and Separation of Live Bacteria in Water. Electrophoresis 2004, 25, 1695–1704. [Google Scholar] [CrossRef] [PubMed]
- Lapizco-Encinas, B.H.; Simmons, B.A.; Cummings, E.B.; Fintschenko, Y. Dielectrophoretic Concentration and Separation of Live and Dead Bacteria in an Array of Insulators. Anal. Chem. 2004, 76, 1571–1579. [Google Scholar] [CrossRef]
- Suehiro, J.; Zhou, G.; Imamura, M.; Hara, M. Dielectrophoretic Filter for Separation and Recovery of Biological Cells in Water. IEEE Trans. Ind. Appl. 2003, 39, 1514–1521. [Google Scholar] [CrossRef]
- Cebricos, J.; Hoptowit, R.; Jun, S. Separation of Escherichia Coli K12 from Contaminated Tap Water Using a Single-Stage, Continuous Flow Dielectrophoresis (DEP) Device. LWT 2017, 80, 185–192. [Google Scholar] [CrossRef]
- Wang, Z.; Hansen, O.; Petersen, P.K.; Rogeberg, A.; Kutter, J.P.; Bang, D.D.; Wolff, A. Dielectrophoresis Microsystem with Integrated Flow Cytometers for On-Line Monitoring of Sorting Efficiency. Electrophoresis 2006, 27, 5081–5092. [Google Scholar] [CrossRef]
- Abidin, Z.Z.; Baik, D.A.; Hafifudin, N.; Markx, G. FACTORS AFFECTING DIELECTROPHORETIC SEPARATION OF CELLS USING HIGH-GRADIENT ELECTRIC FIELD STRENGTH SYSTEM. J. Eng. Sci. Technol. 2008, 3, 30–39. [Google Scholar]
- Sankaranarayanan, A.; Sankarlal, K.M.P.; Raja, D. Impact of Different Frequencies in the Entrapment of Bacterial Pathogens from Drinking Water Using Dielectrophoretic Phenomena. J. Water Chem. Technol. 2016, 38, 117–122. [Google Scholar] [CrossRef]
- Kang, K.H.; Kang, Y.; Xuan, X.; Li, D. Continuous Separation of Microparticles by Size with Direct Current-Dielectrophoresis. Electrophoresis 2006, 27, 694–702. [Google Scholar] [CrossRef] [PubMed]
- Markx, G.H.; Dyda, P.A.; Pethig, R. Dielectrophoretic Separation of Bacteria Using a Conductivity Gradient. J. Biotechnol. 1996, 51, 175–180. [Google Scholar] [CrossRef]
- Wang, X.-B.; Vykoukal, J.; Becker, F.F.; Gascoyne, P.R.C. Separation of Polystyrene Microbeads Using Dielectrophoretic/Gravitational Field-Flow-Fractionation. Biophys. J. 1998, 74, 2689–2701. [Google Scholar] [CrossRef]
- Shafiee, H.; Caldwell, J.L.; Sano, M.B.; Davalos, R.V. Contactless Dielectrophoresis: A New Technique for Cell Manipulation. Biomed. Microdevices 2009, 11, 997–1006. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jun, S.; Chun, C.; Ho, K.; Li, Y. Design and Evaluation of a Millifluidic Insulator-Based Dielectrophoresis (DEP) Retention Device to Separate Bacteria from Tap Water. Water 2021, 13, 1678. https://doi.org/10.3390/w13121678
Jun S, Chun C, Ho K, Li Y. Design and Evaluation of a Millifluidic Insulator-Based Dielectrophoresis (DEP) Retention Device to Separate Bacteria from Tap Water. Water. 2021; 13(12):1678. https://doi.org/10.3390/w13121678
Chicago/Turabian StyleJun, Shawn, Cherisse Chun, Kacie Ho, and Yong Li. 2021. "Design and Evaluation of a Millifluidic Insulator-Based Dielectrophoresis (DEP) Retention Device to Separate Bacteria from Tap Water" Water 13, no. 12: 1678. https://doi.org/10.3390/w13121678
APA StyleJun, S., Chun, C., Ho, K., & Li, Y. (2021). Design and Evaluation of a Millifluidic Insulator-Based Dielectrophoresis (DEP) Retention Device to Separate Bacteria from Tap Water. Water, 13(12), 1678. https://doi.org/10.3390/w13121678





