Probiotics in Fish Nutrition—Long-Standing Household Remedy or Native Nutraceuticals?
Abstract
1. Introduction
2. The “Living Gut” and Its Core Microbiome
3. Species-Specific, Native Probiotics
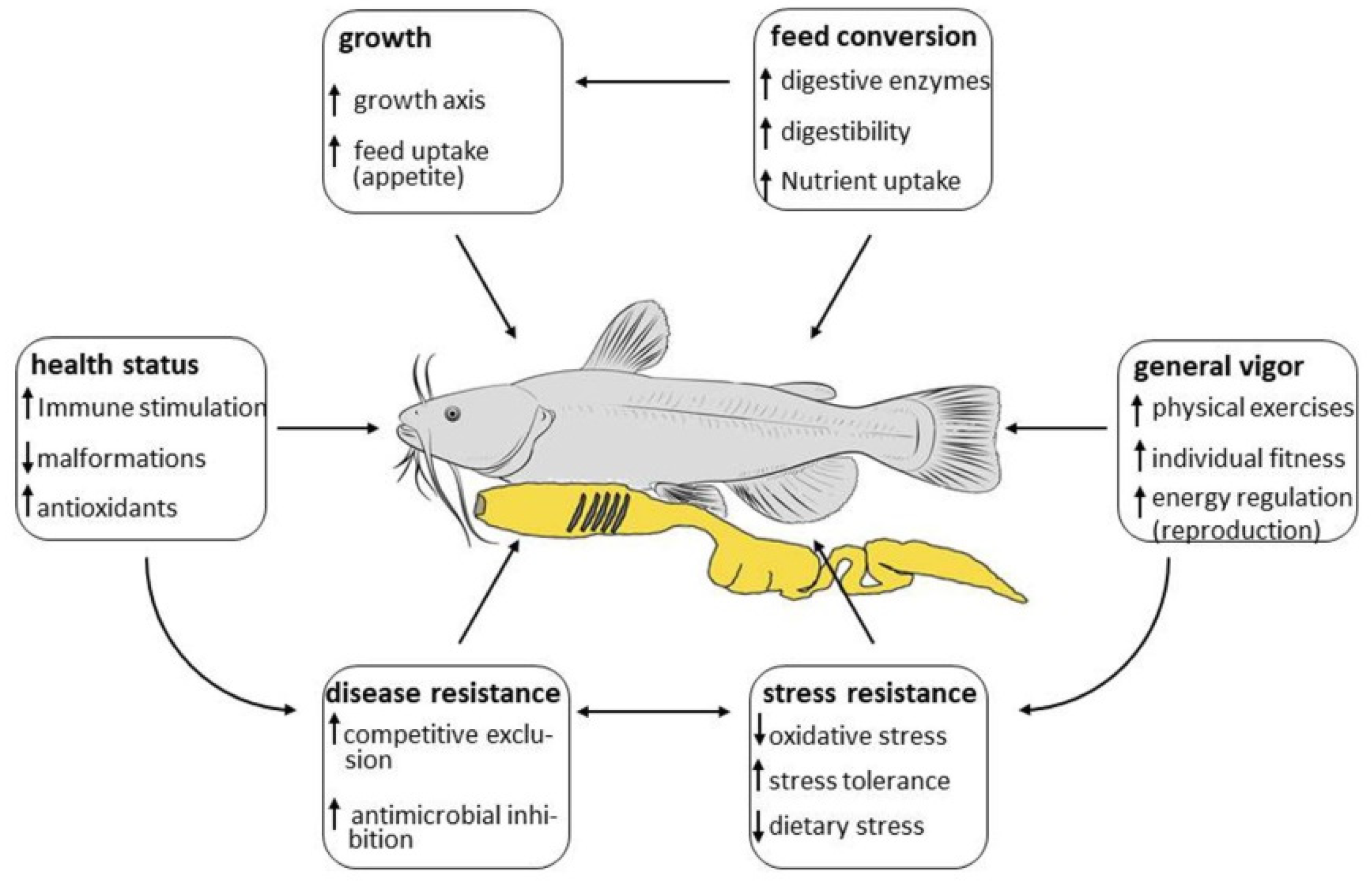
4. Modes of Action
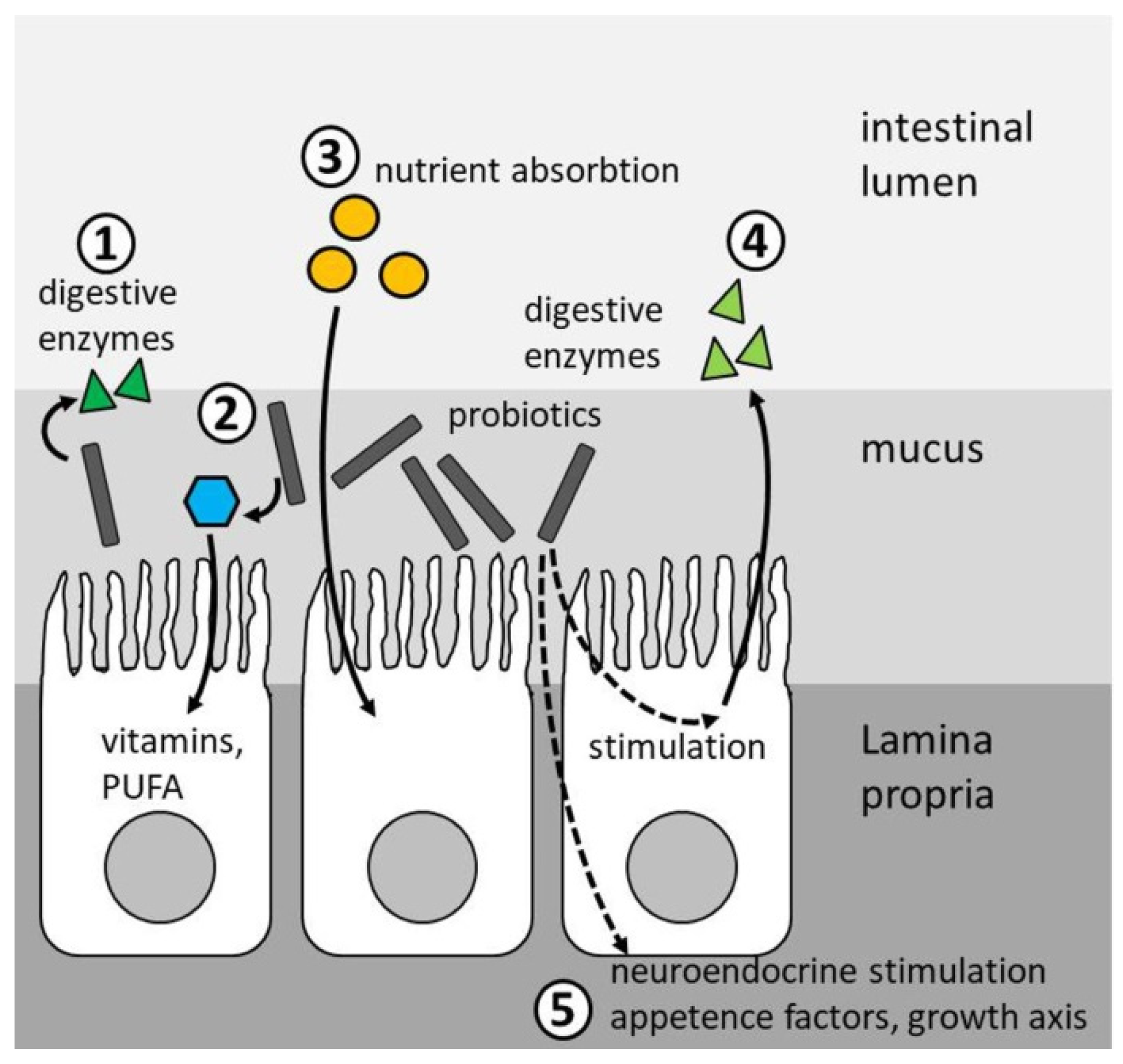
4.1. Growth
4.2. Feed Conversion
4.3. Stress Resistance
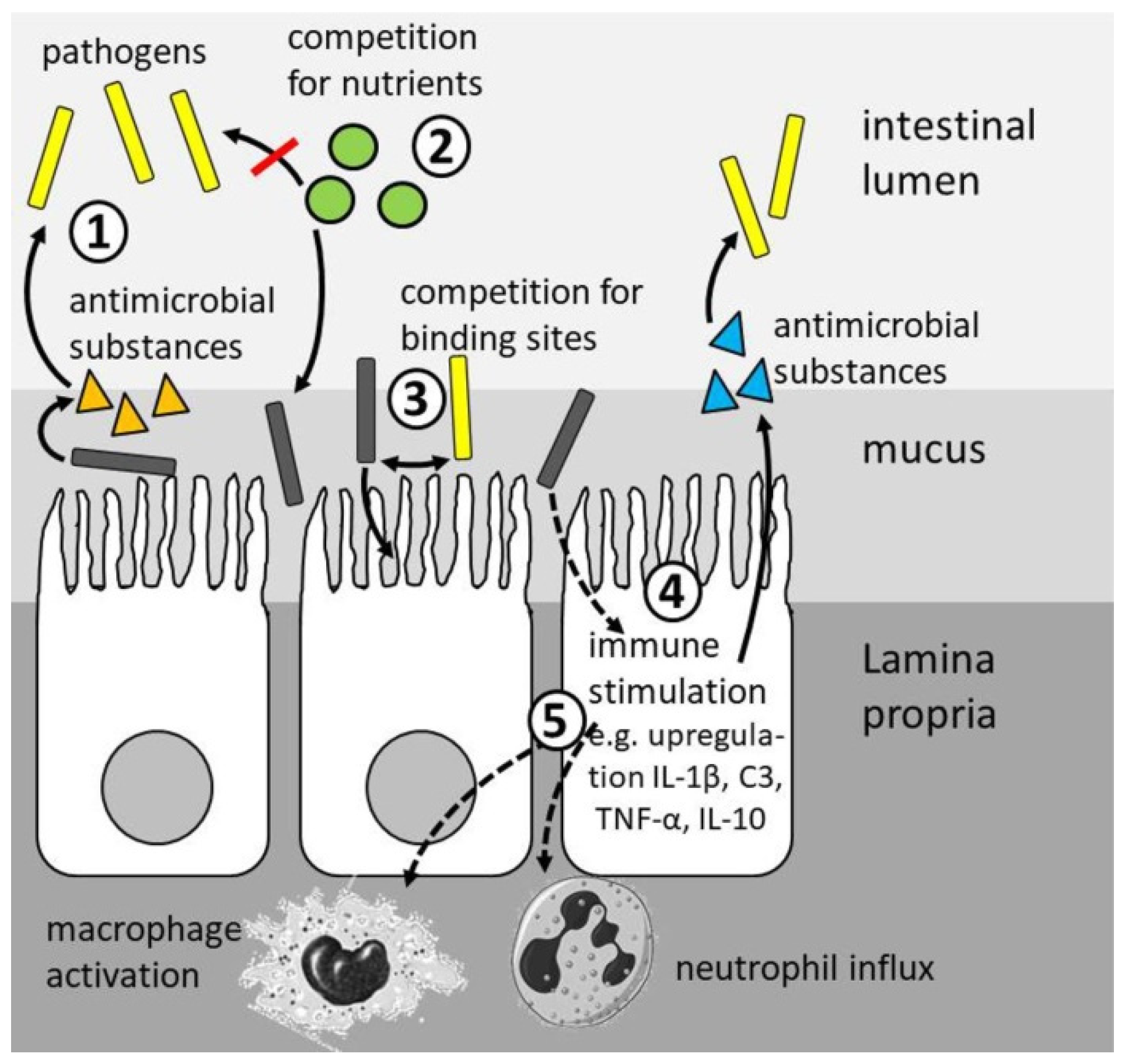
4.4. Health Status
4.5. Disease Resistance
4.6. General Vigor (Fitness)
5. In Vitro Screening for Candidate Probiotics
6. Administration
7. Conclusions
Author Contributions
Funding
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- FAO. The State of World Fisheries and Aquaculture 2020. Sustainability in Action; FAO: Rome, Italy, 2020; pp. 2–36. [Google Scholar]
- Turnbull, J.F. Stress and resistance to infectious diseases in fish. In Infectious Disease in Aquaculture; Woodhead Publishing: Amsterdam, The Netherlands, 2012; pp. 111–125. [Google Scholar] [CrossRef]
- Kroeckel, S.; Harjes, A.G.E.; Roth, I.; Katz, H.; Wuertz, S.; Susenbeth, A.; Schulz, C. When a turbot catches a fly: Evaluation of a pre-pupae meal of the Black Soldier Fly (Hermetia illucens) as fish meal substitute—Growth performance and chitin degradation in juvenile turbot (Psetta maxima). Aquaculture 2012, 364, 345–352. [Google Scholar] [CrossRef]
- Nasr, M.; Reda, R.; Ismail, T.; Moustafa, A. Growth, hemato-biochemical parameters, body composition, and myostatin gene expression of Clarias gariepinus fed by replacing fishmeal with plant protein. Animals 2021, 11, 889. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Liu, Z.-L.; Niu, J. Growth performance, intestinal histomorphology, body composition, hematological and antioxidant parameters of Oncorhynchus mykiss were not detrimentally affected by replacement of fish meal with concentrated dephenolization cottonseed protein. Aquac. Rep. 2021, 19, 100557. [Google Scholar] [CrossRef]
- Tusche, K.; Nagel, F.; Arning, S.; Wuertz, S.; Susenbeth, A.; Schulz, C. Effect of different dietary levels of potato protein concentrate supplemented with feed attractants on growth performance of rainbow trout (Oncorhynchus mykiss). Anim. Feed. Sci. Technol. 2013, 183, 202–209. [Google Scholar] [CrossRef]
- Azeredo, R.; Machado, M.; Kreuz, E.; Wuertz, S.; Oliva-Teles, A.; Enes, P.; Costas, B. The European seabass (Dicentrarchus labrax) innate immunity and gut health are modulated by dietary plant-protein inclusion and prebiotic supplementation. Fish Shellfish. Immunol. 2017, 60, 78–87. [Google Scholar] [CrossRef]
- Estruch, G.; Collado, M.C.; Monge-Ortiz, R.; Tomás-Vidal, A.; Jover-Cerdá, M.; Peñaranda, D.S.; Martínez, G.P.; Martínez-Llorens, S. Long-term feeding with high plant protein based diets in gilthead seabream (Sparus aurata, L.) leads to changes in the inflammatory and immune related gene expression at intestinal level. BMC Veter Res. 2018, 14, 302. [Google Scholar] [CrossRef]
- Piazzon, M.C.; Calduch-Giner, J.A.; Fouz, B.; Estensoro, I.; Simó-Mirabet, P.; Puyalto, M.; Karalazos, V.; Palenzuela, O.; Sitjà-Bobadilla, A.; Pérez-Sánchez, J. Under control: How a dietary additive can restore the gut microbiome and proteomic profile, and improve disease resilience in a marine teleostean fish fed vegetable diets. Microbiome 2017, 5, 1–23. [Google Scholar] [CrossRef]
- Kollath, W. The increase of the diseases of civilization and their prevention. Münch Med. Wochenschr. 1953, 95, 2. [Google Scholar]
- Fuller, R. Probiotics in man and animals. J. Appl. Bacteriol. 1989, 66, 365–378. [Google Scholar]
- Verschuere, L.; Rombaut, G.; Sorgeloos, P.; Verstraete, W. Probiotic Bacteria as Biological Control Agents in Aquaculture. Microbiol. Mol. Biol. Rev. 2000, 64, 655–671. [Google Scholar] [CrossRef] [PubMed]
- Merrifield, D.L.; Dimitroglou, A.; Foey, A.; Davies, S.J.; Baker, R.T.; Bøgwald, J.; Castex, M.; Ringø, E. The current status and future focus of probiotic and prebiotic applications for salmonids. Aquaculture 2010, 302, 1–18. [Google Scholar] [CrossRef]
- Hoseinifar, S.H.; Sun, Y.-Z.; Wang, A.; Zhou, Z. Probiotics as Means of Diseases Control in Aquaculture, a Review of Current Knowledge and Future Perspectives. Front. Microbiol. 2018, 9, 2429. [Google Scholar] [CrossRef] [PubMed]
- Van Hai, N. The use of probiotics in aquaculture. J. Appl. Microbiol. 2015, 119, 917–935. [Google Scholar] [CrossRef]
- Ringø, E.; Van Doan, H.; Lee, S.H.; Soltani, M.; Hoseinifar, S.H.; Harikrishnan, R.; Song, S.K. Probiotics, lactic acid bacteria and bacilli: Interesting supplementation for aquaculture. J. Appl. Microbiol. 2020, 129, 116–136. [Google Scholar] [CrossRef] [PubMed]
- Ringø, E.; Hoseinifar, S.H.; Ghosh, K.; Van Doan, H.; Beck, B.R.; Song, S.K. Lactic Acid Bacteria in Finfish—An Update. Front. Microbiol. 2018, 9, 1818. [Google Scholar] [CrossRef]
- Soltani, M.; Ghosh, K.; Hoseinifar, S.H.; Kumar, V.; Lymbery, A.J.; Roy, S.; Ringø, E. Genusbacillus, promising probiotics in aquaculture: Aquatic animal origin, bio-active components, bioremediation and efficacy in fish and shellfish. Rev. Fish. Sci. Aquac. 2019, 27, 331–379. [Google Scholar] [CrossRef]
- Li, X.; Ringø, E.; Hoseinifar, S.H.; Lauzon, H.L.; Birkbeck, H.; Yang, D. The adherence and colonization of microorganisms in fish gastrointestinal tract. Rev. Aquac. 2019, 11, 603–618. [Google Scholar] [CrossRef]
- Zhou, S.; Sun, Y.; Yu, M.; Shi, Z.; Zhang, H.; Peng, R.; Li, Z.; Cui, J.; Luo, X. Linking Shifts in Bacterial Community Composition and Function with Changes in the Dissolved Organic Matter Pool in Ice-Covered Baiyangdian Lake, Northern China. Microorganisms 2020, 8, 883. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wang, P.; Wang, C.; Chen, J.; Miao, L.; Yuan, Q.; Liu, S.; Feng, T. Do bacterioplankton respond equally to different river regulations? A quantitative study in the single-dammed Yarlung Tsangpo River and the cascade-dammed Lancang River. Environ. Res. 2020, 191, 110194. [Google Scholar] [CrossRef]
- Schulhof, M.A.; Allen, A.E.; Allen, E.E.; Mladenov, N.; McCrow, J.P.; Jones, N.T.; Blanton, J.; Cavalheri, H.B.; Kaul, D.; Symons, C.C.; et al. Sierra Nevada mountain lake microbial communities are structured by temperature, resources and geographic location. Mol. Ecol. 2020, 29, 2080–2093. [Google Scholar] [CrossRef]
- Breen, P.; Winters, A.D.; Nag, D.; Ahmad, M.M.; Theis, K.R.; Withey, J.H. Internal Versus External Pressures: Effect of Housing Systems on the Zebrafish Microbiome. Zebrafish 2019, 16, 388–400. [Google Scholar] [CrossRef]
- Riiser, E.S.; Haverkamp, T.H.A.; Varadharajan, S.; Borgan, Ø.; Jakobsen, K.S.; Jentoft, S.; Star, B. Metagenomic Shotgun Analyses Reveal Complex Patterns of Intra- and Interspecific Variation in the Intestinal Microbiomes of Codfishes. Appl. Environ. Microbiol. 2020, 86. [Google Scholar] [CrossRef] [PubMed]
- Xiao, F.; Zhu, W.; Yu, Y.; He, Z.; Wu, B.; Wang, C.; Shu, L.; Li, X.; Yin, H.; Wang, J.; et al. Host development overwhelms environmental dispersal in governing the ecological succession of zebrafish gut microbiota. NPJ Biofilms Microbiomes 2021, 7, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Zha, Y.; Lindström, E.S.; Eiler, A.; Svanbäck, R. Different Roles of Environmental Selection, Dispersal, and Drift in the Assembly of Intestinal Microbial Communities of Freshwater Fish With and Without a Stomach. Front. Ecol. Evol. 2020, 8, 8. [Google Scholar] [CrossRef]
- Fogarty, C.; Burgess, C.M.; Cotter, P.D.; Cabrera-Rubio, R.; Whyte, P.; Smyth, C.; Bolton, D.J. Diversity and composition of the gut microbiota of Atlantic salmon (Salmo salar) farmed in Irish waters. J. Appl. Microbiol. 2019, 127, 648–657. [Google Scholar] [CrossRef] [PubMed]
- Webster, T.M.U.; Consuegra, S.; Hitchings, M.; De Leaniz, C.G. Interpopulation Variation in the Atlantic Salmon Microbiome Reflects Environmental and Genetic Diversity. Appl. Environ. Microbiol. 2018, 84, 84. [Google Scholar] [CrossRef]
- Dulski, T.; Kujawa, R.; Godzieba, M.; Ciesielski, S. Effect of Salinity on the Gut Microbiome of Pike Fry (Esox lucius). Appl. Sci. 2020, 10, 2506. [Google Scholar] [CrossRef]
- Hassenrück, C.; Reinwald, H.; Kunzmann, A.; Tiedemann, I.; Gärdes, A. Effects of Thermal Stress on the Gut Microbiome of Juvenile Milkfish (Chanos chanos). Microorganisms 2020, 9, 5. [Google Scholar] [CrossRef]
- Zhang, Z.; Li, D.; Refaey, M.M.; Xu, W. High Spatial and Temporal Variations of Microbial Community along the Southern Catfish Gastrointestinal Tract: Insights into Dynamic Food Digestion. Front. Microbiol. 2017, 8, 1531. [Google Scholar] [CrossRef]
- Kuebutornye, F.K.A.; Wang, Z.W.; Lu, Y.S.; Abarike, E.D.; Sakyi, M.E.; Li, Y.; Xie, C.X.; Hlordzi, V. Effects of three host-associated Bacillus species on mucosal immunity and gut health of Nile tilapia, Oreochromis niloticus and its resistance against Aeromonas hydrophila infection. Fish Shellfish. Immunol. 2020, 97, 83–95. [Google Scholar] [CrossRef]
- Zhang, C.; Derrien, M.; Levenez, F.; Brazeilles, R.; Ballal, S.A.; Kim, J.; Degivry, M.-C.; Quéré, G.; Garault, P.; Vlieg, J.E.T.V.H.; et al. Ecological robustness of the gut microbiota in response to ingestion of transient food-borne microbes. ISME J. 2016, 10, 2235–2245. [Google Scholar] [CrossRef]
- Aerts, J.; Schaeck, M.; De Swaef, E.; Ampe, B.; Decostere, A. Vibrio lentus as a probiotic candidate lowers glucocorticoid levels in gnotobiotic sea bass larvae. Aquaculture 2018, 492, 40–45. [Google Scholar] [CrossRef]
- Pérez-Ramos, A.; Mohedano, M.L.; Pardo, M.Á.; López, P. β-Glucan-Producing Pediococcus parvulus 2.6: Test of Probiotic and Immunomodulatory Properties in Zebrafish Models. Front. Microbiol. 2018, 9, 1684. [Google Scholar] [CrossRef]
- Zhang, Z.; Li, D.; Refaey, M.M.; Xu, W.; Tang, R.; Li, L. Host Age Affects the Development of Southern Catfish Gut Bacterial Community Divergent From That in the Food and Rearing Water. Front. Microbiol. 2018, 9, 495. [Google Scholar] [CrossRef] [PubMed]
- Rawls, J.F.; Samuel, B.S.; Gordon, J.I. Gnotobiotic zebrafish reveal evolutionarily conserved responses to the gut microbiota. Proc. Natl. Acad. Sci. USA 2004, 101, 4596–4601. [Google Scholar] [CrossRef]
- Rawls, J.F.; Mahowald, M.A.; Ley, R.E.; Gordon, J.I. Reciprocal Gut Microbiota Transplants from Zebrafish and Mice to Germ-free Recipients Reveal Host Habitat Selection. Cell 2006, 127, 423–433. [Google Scholar] [CrossRef] [PubMed]
- Roeselers, G.; Mittge, E.K.; Stephens, W.Z.; Parichy, D.M.; Cavanaugh, C.M.; Guillemin, K.; Rawls, J.F. Evidence for a core gut microbiota in the zebrafish. ISME J. 2011, 5, 1595–1608. [Google Scholar] [CrossRef]
- Etyemez, M.; Balcázar, J.L. Bacterial community structure in the intestinal ecosystem of rainbow trout (Oncorhynchus mykiss) as revealed by pyrosequencing-based analysis of 16S rRNA genes. Res. Veter Sci. 2015, 100, 8–11. [Google Scholar] [CrossRef]
- Larsen, A.M.; Mohammed, H.H.; Arias, C.R. Comparison of DNA extraction protocols for the analysis of gut microbiota in fishes. FEMS Microbiol. Lett. 2014, 362, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Long, M.; Gatesoupe, F.-J.; Zhang, Q.; Li, A.; Gong, X. Comparative Analysis of the Intestinal Bacterial Communities in Different Species of Carp by Pyrosequencing. Microb. Ecol. 2015, 69, 25–36. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Gong, Y.-X.; Zhu, B.; Liu, G.-L.; Wang, G.-X.; Ling, F. Effect of a new recombinant Aeromonas hydrophila vaccine on the grass carp intestinal microbiota and correlations with immunological responses. Fish Shellfish. Immunol. 2015, 45, 175–183. [Google Scholar] [CrossRef]
- Narrowe, A.B.; Albuthi-Lantz, M.; Smith, E.P.; Bower, K.J.; Roane, T.M.; Vajda, A.M.; Miller, C.S. Perturbation and restoration of the fathead minnow gut microbiome after low-level triclosan exposure. Microbiome 2015, 3, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Llewellyn, M.S.; Boutin, S.; Hoseinifar, S.H.; Derome, N. Teleost microbiomes: The state of the art in their characterization, manipulation and importance in aquaculture and fisheries. Front. Microbiol. 2014, 5, 207. [Google Scholar] [CrossRef]
- Huang, Q.; Sham, R.C.; Deng, Y.; Mao, Y.; Wang, C.; Zhang, T.; Leung, K.M.Y. Diversity of gut microbiomes in marine fishes is shaped by host-related factors. Mol. Ecol. 2020, 29, 5019–5034. [Google Scholar] [CrossRef]
- Dehler, C.E.; Secombes, C.J.; Martin, S.A. Environmental and physiological factors shape the gut microbiota of Atlantic salmon parr (Salmo salar L.). Aquaculture 2017, 467, 149–157. [Google Scholar] [CrossRef]
- Dulski, T.; Kozłowski, K.; Ciesielski, S. Habitat and seasonality shape the structure of tench (Tinca tinca L.) gut microbiome. Sci. Rep. 2020, 10, 1–11. [Google Scholar] [CrossRef]
- Wang, A.R.; Ran, C.; Ringø, E.; Zhou, Z.G. Progress in fish gastrointestinal microbiota research. Rev. Aquac. 2018, 10, 626–640. [Google Scholar] [CrossRef]
- Liu, H.; Guo, X.; Gooneratne, S.R.; Lai, R.; Zeng, C.; Zhan, F.; Wang, W. The gut microbiome and degradation enzyme activity of wild freshwater fishes influenced by their trophic levels. Sci. Rep. 2016, 6, 24340. [Google Scholar] [CrossRef] [PubMed]
- Bajinka, O.; Tan, Y.; Abdelhalim, K.A.; Özdemir, G.; Qiu, X. Extrinsic factors influencing gut microbes, the immediate consequences and restoring eubiosis. AMB Express 2020, 10, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Vasemägi, A.; Visse, M.; Kisand, V. Effect of Environmental Factors and an Emerging Parasitic Disease on Gut Microbiome of Wild Salmonid Fish. mSphere 2017, 2, e00418-17. [Google Scholar] [CrossRef]
- Stephens, W.Z.; Burns, A.R.; Stagaman, K.; Wong, S.; Rawls, J.F.; Guillemin, K.; Bohannan, B.J.M. The composition of the zebrafish intestinal microbial community varies across development. ISME J. 2016, 10, 644–654. [Google Scholar] [CrossRef]
- Michl, S.C.; Ratten, J.-M.; Beyer, M.; Hasler, M.; Laroche, J.; Schulz, C. The malleable gut microbiome of juvenile rainbow trout (Oncorhynchus mykiss): Diet-dependent shifts of bacterial community structures. PLoS ONE 2017, 12, e0177735. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Wang, G.; Angert, E.R.; Wang, W.; Li, W.; Zou, H. Composition, Diversity, and Origin of the Bacterial Community in Grass Carp Intestine. PLoS ONE 2012, 7, e30440. [Google Scholar] [CrossRef]
- Xing, M.; Hou, Z.; Yuan, J.; Liu, Y.; Qu, Y.; Liu, B. Taxonomic and functional metagenomic profiling of gastrointestinal tract microbiome of the farmed adult turbot (Scophthalmus maximus). FEMS Microbiol. Ecol. 2013, 86, 432–443. [Google Scholar] [CrossRef]
- Murall, C.L.A.; Touzel, M.P.; Allen-Vercoe, E.; Alizon, S.; Froissart, R.; McCann, K. Chapter five—Invasions of host-associated microbiome networks. In Advances in Ecological Research; Bohan, D.A., Dumbrell, A.J., Massol, F., Eds.; Academic Press: Cambridge, MA, USA, 2017; Volume 57, p. 80. [Google Scholar]
- Ibrahem, M.D. Evolution of probiotics in aquatic world: Potential effects, the current status in Egypt and recent prospectives. J. Adv. Res. 2015, 6, 765–791. [Google Scholar] [CrossRef] [PubMed]
- Nikoskelainen, S.; Ouwehand, A.; Salminen, S.; Bylund, G. Protection of rainbow trout (Oncorhynchus mykiss) from furunculosis by Lactobacillus rhamnosus. Aquaculture 2001, 198, 229–236. [Google Scholar] [CrossRef]
- Nikoskelainen, S.; Salminen, S.; Bylund, G.; Ouwehand, A.C. Characterization of the Properties of Human- and Dairy-Derived Probiotics for Prevention of Infectious Diseases in Fish. Appl. Environ. Microbiol. 2001, 67, 2430–2435. [Google Scholar] [CrossRef]
- Rinkinen, M.; Mättö, J.; Salminen, S.; Westermarck, E.; Ouwehand, A.C. In vitro adhesion of lactic acid bacteria to canine small intestinal mucus. J. Anim. Physiol. Anim. Nutr. 2000, 84, 43–47. [Google Scholar] [CrossRef]
- Rinkinen, M.; Westermarck, E.; Salminen, S.; Ouwehand, A.C. Absence of host specificity for in vitro adhesion of probiotic lactic acid bacteria to intestinal mucus. Veter Microbiol. 2003, 97, 55–61. [Google Scholar] [CrossRef]
- Gildberg, A.; Mikkelsen, H. Effects of supplementing the feed to Atlantic cod (Gadus morhua) fry with lactic acid bacteria and immuno-stimulating peptides during a challenge trial with Vibrio anguillarum. Aquaculture 1998, 167, 103–113. [Google Scholar] [CrossRef]
- Askarian, F.; Kousha, A.; Salma, W.; Ringø, E. The effect of lactic acid bacteria administration on growth, digestive enzyme activity and gut microbiota in Persian sturgeon (Acipenser persicus) and beluga (Huso huso) fry. Aquac. Nutr. 2011, 17, 488–497. [Google Scholar] [CrossRef]
- Salma, W.; Zhou, Z.; Wang, W.; Askarian, F.; Kousha, A.; Ebrahimi, M.T.; Myklebust, R.; Ringø, E. Histological and bacteriological changes in intestine of beluga (Huso huso) following ex vivo exposure to bacterial strains. Aquaculture 2011, 314, 24–33. [Google Scholar] [CrossRef][Green Version]
- Ringø, E.; Birkbeck, T.; Munro, P.; Vadstein, O.; Hjelmeland, K. The effect of early exposure to Vibrio pelagiuson the aerobic bacterial flora of turbot, Scophthalmus maximus (L.) larvae. J. Appl. Bacteriol. 1996, 81, 207–211. [Google Scholar] [CrossRef]
- Bansil, R.; Turner, B.S. Mucin structure, aggregation, physiological functions and biomedical applications. Curr. Opin. Colloid Interface Sci. 2006, 11, 164–170. [Google Scholar] [CrossRef]
- Van Tassell, M.L.; Miller, M.J. Lactobacillus adhesion to mucus. Nutrients 2011, 3, 613–636. [Google Scholar] [CrossRef]
- Buck, B.L.; Altermann, E.; Svingerud, T.; Klaenhammer, T.R. Functional Analysis of Putative Adhesion Factors in Lactobacillus acidophilus NCFM. Appl. Environ. Microbiol. 2005, 71, 8344–8351. [Google Scholar] [CrossRef]
- Etzold, S.; Juge, N. Structural insights into bacterial recognition of intestinal mucins. Curr. Opin. Struct. Biol. 2014, 28, 23–31. [Google Scholar] [CrossRef]
- Johnson, B.; Selle, K.; O’Flaherty, S.; Goh, Y.J.; Klaenhammer, T. Identification of extracellular surface-layer associated proteins in Lactobacillus acidophilus NCFM. Microbiol-Sgm 2013, 159, 2269–2282. [Google Scholar] [CrossRef]
- Bautista-Gallego, J.; Arroyo-López, F.; Rantsiou, K.; Jiménez-Díaz, R.; Garrido-Fernández, A.; Cocolin, L. Screening of lactic acid bacteria isolated from fermented table olives with probiotic potential. Food Res. Int. 2013, 50, 135–142. [Google Scholar] [CrossRef]
- Son, S.-H.; Jeon, H.-L.; Yang, S.-J.; Lee, N.-K.; Paik, H.-D. In vitro characterization of Lactobacillus brevis KU15006, an isolate from kimchi, reveals anti-adhesion activity against foodborne pathogens and antidiabetic properties. Microb. Pathog. 2017, 112, 135–141. [Google Scholar] [CrossRef]
- Wanka, K.M.; Damerau, T.; Costas, B.; Krueger, A.; Schulz, C.; Wuertz, S. Isolation and characterization of native probiotics for fish farming. BMC Microbiol. 2018, 18, 119. [Google Scholar] [CrossRef]
- Chowdhury, A.; Roy, N.C. Probiotic supplementation for enhanced growth of striped catfish (Pangasianodon hypophthalmus) in cages. Aquac. Rep. 2020, 18, 100504. [Google Scholar] [CrossRef]
- Wanka, K.M.; Schulz, C.; Kloas, W.; Wuertz, S. Administration of host-derived probiotics does not affect utilization of soybean meal enriched diets in juvenile turbot (Scophthalmus maximus). J. Appl. Ichthyol. 2019, 35, 1004–1015. [Google Scholar] [CrossRef]
- Chen, X.; Xie, J.; Liu, Z.; Yin, P.; Chen, M.; Liu, Y.; Tian, L.; Niu, J. Modulation of growth performance, non-specific immunity, intestinal morphology, the response to hypoxia stress and resistance to Aeromonas hydrophila of grass carp (Ctenopharyngodon idella) by dietary supplementation of a multi-strain probiotic. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2020, 231, 108724. [Google Scholar] [CrossRef] [PubMed]
- El-Haroun, E.R.; Goda, A.M.A.-S.; Chowdhury, M.A.K. Effect of dietary probiotic Biogen(R) supplementation as a growth promoter on growth performance and feed utilization of Nile tilapia Oreochromis niloticus (L.). Aquac. Res. 2006, 37, 1473–1480. [Google Scholar] [CrossRef]
- Kuebutornye, F.K.A.; Abarike, E.D.; Sakyi, M.E.; Lu, Y.; Wang, Z. Modulation of nutrient utilization, growth, and immunity of Nile tilapia, Oreochromis niloticus: The role of probiotics. Aquac. Int. 2020, 28, 277–291. [Google Scholar] [CrossRef]
- Silva, V.V.; Salomão, R.A.S.; Mareco, E.A.; Pai, M.D.; Santos, V.B. Probiotic additive affects muscle growth of Nile tilapia (Oreochromis niloticus). Aquac. Res. 2021, 52, 2061–2069. [Google Scholar] [CrossRef]
- Yi, C.-C.; Liu, C.-H.; Chuang, K.-P.; Chang, Y.-T.; Hu, S.-Y. A potential probiotic Chromobacterium aquaticum with bacteriocin-like activity enhances the expression of indicator genes associated with nutrient metabolism, growth performance and innate immunity against pathogen infections in zebrafish (Danio rerio). Fish Shellfish. Immunol. 2019, 93, 124–134. [Google Scholar] [CrossRef]
- Guidoli, M.G.; Mendoza, J.A.; Falcón, S.L.; Boehringer, S.I.; Sánchez, S.; Macías, M.E.F.N. Autochthonous probiotic mixture improves biometrical parameters of larvae of Piaractus mesopotamicus (Caracidae, Characiforme, Teleostei). Ciência Rural 2018, 48. [Google Scholar] [CrossRef]
- Chiu, K.-H.; Liu, W.-S. Dietary administration of the extract of Rhodobacter sphaeroides WL-APD911 enhances the growth performance and innate immune responses of seawater red tilapia (Oreochromis mossambicus×Oreochromis niloticus). Aquaculture 2014, 418–419, 32–38. [Google Scholar] [CrossRef]
- Hosseini, M.; Miandare, H.K.; Hoseinifar, S.H.; Yarahmadi, P. Dietary Lactobacillus acidophilus modulated skin mucus protein profile, immune and appetite genes expression in gold fish (Carassius auratus gibelio). Fish Shellfish. Immunol. 2016, 59, 149–154. [Google Scholar] [CrossRef]
- Giorgia, G.; Elia, C.; Andrea, P.; Cinzia, C.; Stefania, S.; Ana, R.; Daniel, M.L.; Ike, O.; Oliana, C. Effects of Lactogen 13, a New Probiotic Preparation, on Gut Microbiota and Endocrine Signals Controlling Growth and Appetite of Oreochromis niloticus Juveniles. Microb. Ecol. 2018, 76, 1063–1074. [Google Scholar] [CrossRef]
- Irianto, A.; Austin, B. Probiotics in aquaculture. J. Fish Dis. 2002, 25, 633–642. [Google Scholar] [CrossRef]
- Irianto, A.; Austin, B. Use of probiotics to control furunculosis in rainbow trout, Oncorhynchus mykiss (Walbaum). J. Fish Dis. 2002, 25, 333–342. [Google Scholar] [CrossRef]
- Bates, J.M.; Mittge, E.; Kuhlman, J.; Baden, K.N.; Cheesman, S.E.; Guillemin, K. Distinct signals from the microbiota promote different aspects of zebrafish gut differentiation. Dev. Biol. 2006, 297, 374–386. [Google Scholar] [CrossRef] [PubMed]
- Merrifield, D.; Bradley, G.; Harper, G.; Baker, R.; Munn, C.; Davies, S. Assessment of the effects of vegetative and lyophilized Pediococcus acidilactici on growth, feed utilization, intestinal colonization and health parameters of rainbow trout (Oncorhynchus mykiss Walbaum). Aquac. Nutr. 2009, 17, 73–79. [Google Scholar] [CrossRef]
- Merrifield, D.L.; Harper, G.M.; Dimitroglou, A.; Ringã, E.; Davies, S.J. Possible influence of probiotic adhesion to intestinal mucosa on the activity and morphology of rainbow trout (Oncorhynchus mykiss) enterocytes. Aquac. Res. 2009, 41, 1268–1272. [Google Scholar] [CrossRef]
- Tan, H.Y.; Chen, S.-W.; Hu, S.-Y. Improvements in the growth performance, immunity, disease resistance, and gut microbiota by the probiotic Rummeliibacillus stabekisii in Nile tilapia (Oreochromis niloticus). Fish Shellfish. Immunol. 2019, 92, 265–275. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, Q.; Xing, K.; Jiang, P.; Wang, J. Dietary cinnamaldehyde and Bacillus subtilis improve growth performance, digestive enzyme activity, and antioxidant capability and shape intestinal microbiota in tongue sole, Cynoglossus semilaevis. Aquaculture 2021, 531, 735798. [Google Scholar] [CrossRef]
- Santos, K.O.; Costa-Filho, J.; Spagnol, K.L.; Nornberg, B.F.; Lopes, F.M.; Tesser, M.B.; Marins, L.F. The inclusion of a transgenic probiotic expressing recombinant phytase in a diet with a high content of vegetable matter markedly improves growth performance and the expression of growth-related genes and other selected genes in zebrafish. Aquaculture 2020, 519, 734878. [Google Scholar] [CrossRef]
- Zhang, M.; Dong, B.; Lai, X.; Chen, Z.; Hou, L.; Shu, R.; Huang, Y.; Shu, H. Effects of Clostridium butyricum on growth, digestive enzyme activity, antioxidant capacity and gut microbiota in farmed tilapia (Oreochromis niloticus). Aquac. Res. 2021, 52, 1573–1584. [Google Scholar] [CrossRef]
- Wang, Y.; Al Farraj, D.A.; Vijayaraghavan, P.; Hatamleh, A.A.; Biji, G.D.; Rady, A.M. Host associated mixed probiotic bacteria induced digestive enzymes in the gut of tiger shrimp Penaeus monodon. Saudi J. Biol. Sci. 2020, 27, 2479–2484. [Google Scholar] [CrossRef]
- Tarkhani, R.; Imani, A.; Hoseinifar, S.H.; Ashayerizadeh, O.; Moghanlou, K.S.; Manaffar, R.; Van Doan, H.; Reverter, M. Comparative study of host-associated and commercial probiotic effects on serum and mucosal immune parameters, intestinal microbiota, digestive enzymes activity and growth performance of roach (Rutilus rutilus caspicus) fingerlings. Fish Shellfish. Immunol. 2020, 98, 661–669. [Google Scholar] [CrossRef]
- Niu, K.-M.; Kothari, D.; Lee, W.-D.; Zhang, Z.; Lee, B.-J.; Kim, K.-W.; Wu, X.; Han, H.-S.; Khosravi, S.; Lee, S.-M.; et al. Probiotic Potential of the Farmed Olive Flounder, Paralichthys olivaceus, Autochthonous Gut Microbiota. Probiotics Antimicro 2021. [Google Scholar] [CrossRef] [PubMed]
- Hmani, H.; Daoud, L.; Jlidi, M.; Jalleli, K.; Ben Ali, M.; Brahim, A.H.; Bargui, M.; Dammak, A.; Ben Ali, M. A Bacillus subtilis strain as probiotic in poultry: Selection based on in vitro functional properties and enzymatic potentialities. J. Ind. Microbiol. Biotechnol. 2017, 44, 1157–1166. [Google Scholar] [CrossRef]
- Kong, Y.; Li, M.; Chu, G.; Liu, H.; Shan, X.; Wang, G.; Han, G. The positive effects of single or conjoint administration of lactic acid bacteria on Channa argus: Digestive enzyme activity, antioxidant capacity, intestinal microbiota and morphology. Aquaculture 2021, 531, 735852. [Google Scholar] [CrossRef]
- Tarkhani, R.; Imani, A.; Hoseinifar, S.H.; Moghanlou, K.S.; Manaffar, R. The effects of host-associated Enterococcus faecium CGMCC1.2136 on serum immune parameters, digestive enzymes activity and growth performance of the Caspian roach (Rutilus rutilus caspicus) fingerlings. Aquaculture 2020, 519, 734741. [Google Scholar] [CrossRef]
- Nguyen, T.L.; Chun, W.-K.; Kim, A.; Kim, N.; Roh, H.J.; Lee, Y.; Yi, M.; Kim, S.; Park, C.-I.; Kim, D.-H. Dietary Probiotic Effect of Lactococcus lactis WFLU12 on Low-Molecular-Weight Metabolites and Growth of Olive Flounder (Paralichythys olivaceus). Front. Microbiol. 2018, 9, 2059. [Google Scholar] [CrossRef]
- Ray, A.; Ghosh, K.; Ringø, E. Enzyme-producing bacteria isolated from fish gut: A review. Aquac. Nutr. 2012, 18, 465–492. [Google Scholar] [CrossRef]
- Ray, A.K.R.E. Fishes—Digestive organs. In Aquaculture Nutrition—Gut Health, Probiotics, and Prebiotics; Merrifield, D., Ringo, E., Eds.; John Wiley & Sons Ltd.: West Sussex, UK, 2014. [Google Scholar]
- Eck, P.; Friel, J. Should probiotics be considered as vitamin supplements? Vitam. Miner. 2013, 2, e124. [Google Scholar] [CrossRef]
- Ringø, E.; Jelstensen, J.P.; Olsen, R.E. Production of eicosapentaenoic acid by freshwaterVibrio. Lipids 1992, 27, 564–566. [Google Scholar] [CrossRef]
- Ringø, E.; Sinclair, P.D.; Birkbeck, H.; Barbour, A. Production of Eicosapentaenoic Acid (20:5 n-3) by Vibrio pelagius Isolated from Turbot (Scophthalmus maximus (L.)) Larvae. Appl. Environ. Microbiol. 1992, 58, 3777–3778. [Google Scholar] [CrossRef]
- Yano, Y.; Nakayama, A.; Saito, H.; Ishihara, K. Production of docosahexaenoic acid by marine bacteria isolated from deep sea fish. Lipids 1994, 29, 527–528. [Google Scholar] [CrossRef]
- Yano, Y.; Nakayama, A.; Yoshida, K. Distribution of polyunsaturated Fatty acids in bacteria present in intestines of deep-sea fish and shallow-sea poikilothermic animals. Appl. Environ. Microbiol. 1997, 63, 2572–2577. [Google Scholar] [CrossRef]
- Estupiñán, M.; Hernández, I.; Saitua, E.; Bilbao, M.E.; Mendibil, I.; Ferrer, J.; Alonso-Sáez, L. Novel Vibrio spp. Strains Producing Omega-3 Fatty Acids Isolated from Coastal Seawater. Mar. Drugs 2020, 18, 99. [Google Scholar] [CrossRef] [PubMed]
- Nichols, D.S.; Davies, N.W. Improved detection of polyunsaturated fatty acids as phenacyl esters using liquid chromatography-ion trap mass spectrometry. J. Microbiol. Methods 2002, 50, 103–113. [Google Scholar] [CrossRef]
- Nichols, D.S.; McMeekin, T.A. Biomarker techniques to screen for bacteria that produce polyunsaturated fatty acids. J. Microbiol. Methods 2002, 48, 161–170. [Google Scholar] [CrossRef]
- Sutthi, N.; Van Doan, H. Saccharomyces crevices and Bacillus spp. effectively enhance health tolerance of Nile tilapia under transportation stress. Aquaculture 2020, 528, 735527. [Google Scholar] [CrossRef]
- Rollo, A.; Sulpizio, R.; Nardi, M.; Silvi, S.; Orpianesi, C.; Caggiano, M.; Cresci, A.; Carnevali, O. Live microbial feed supplement in aquaculture for improvement of stress tolerance. Fish Physiol. Biochem. 2006, 32, 167–177. [Google Scholar] [CrossRef]
- Abass, D.A.; Obirikorang, K.A.; Campion, B.B.; Edziyie, R.E.; Skov, P.V. Dietary supplementation of yeast (Saccharomyces cerevisiae) improves growth, stress tolerance, and disease resistance in juvenile Nile tilapia (Oreochromis niloticus). Aquac. Int. 2018, 26, 843–855. [Google Scholar] [CrossRef]
- Carnevali, O.; de Vivo, L.; Sulpizio, R.; Gioacchini, G.; Olivotto, I.; Silvi, S.; Cresci, A. Growth improvement by probiotic in European sea bass juveniles (Dicentrarchus labrax, L.), with particular attention to IGF-1, myostatin and cortisol gene expression. Aquaculture 2006, 258, 430–438. [Google Scholar] [CrossRef]
- Dawood, M.A.; Eweedah, N.M.; Moustafa, E.M.; Farahat, E.M. Probiotic effects of Aspergillus oryzae on the oxidative status, heat shock protein, and immune related gene expression of Nile tilapia (Oreochromis niloticus) under hypoxia challenge. Aquaculture 2020, 520, 734669. [Google Scholar] [CrossRef]
- Lin, Y.-S.; Saputra, F.; Chen, Y.-C.; Hu, S.-Y. Dietary administration of Bacillus amyloliquefaciens R8 reduces hepatic oxidative stress and enhances nutrient metabolism and immunity against Aeromonas hydrophila and Streptococcus agalactiae in zebrafish (Danio rerio). Fish Shellfish. Immunol. 2019, 86, 410–419. [Google Scholar] [CrossRef]
- Gioacchini, G.; Giorgini, E.; Olivotto, I.; Maradonna, F.; Merrifield, D.L.; Carnevali, O. The Influence of Probiotics on Zebrafish Danio Rerio Innate Immunity and Hepatic Stress. Zebrafish 2014, 11, 98–106. [Google Scholar] [CrossRef]
- Van Doan, H.; Hoseinifar, S.H.; Khanongnuch, C.; Kanpiengjai, A.; Unban, K.; Van Kim, V.; Srichaiyo, S. Host-associated probiotics boosted mucosal and serum immunity, disease resistance and growth performance of Nile tilapia (Oreochromis niloticus). Aquaculture 2018, 491, 94–100. [Google Scholar] [CrossRef]
- Veiga, P.T.D.N.; Owatari, M.S.; Nunes, A.L.; Rodrigues, R.A.; Kasai, R.Y.D.; Fernandes, C.E.; De Campos, C.M. Short communication: Bacillus subtilis C-3102 improves biomass gain, innate defense, and intestinal absorption surface of native Brazilian hybrid Surubim (Pseudoplatystoma corruscans x P. reticulatum). Aquac. Int. 2020, 28, 1183–1193. [Google Scholar] [CrossRef]
- Nunes, A.L.; Owatari, M.S.; Rodrigues, R.A.; Fantini, L.E.; Kasai, R.Y.D.; Martins, M.L.; Mouriño, J.L.P.; De Campos, C.M. Effects of Bacillus subtilis C-3102-supplemented diet on growth, non-specific immunity, intestinal morphometry and resistance of hybrid juvenile Pseudoplatystoma sp. challenged with Aeromonas hydrophila. Aquac. Int. 2020, 28, 2345–2361. [Google Scholar] [CrossRef]
- Meidong, R.; Nakao, M.; Sakai, K.; Tongpim, S. Lactobacillus paraplantarum L34b-2 derived from fermented food improves the growth, disease resistance and innate immunity in Pangasius bocourti. Aquaculture 2021, 531, 735878. [Google Scholar] [CrossRef]
- Zhou, S.; Song, D.; Zhou, X.; Mao, X.; Zhou, X.; Wang, S.; Wei, J.; Huang, Y.; Wang, W.; Xiao, S.-M.; et al. Characterization of Bacillus subtilis from gastrointestinal tract of hybrid Hulong grouper (Epinephelus fuscoguttatus × E. lanceolatus) and its effects as probiotic additives. Fish Shellfish. Immunol. 2019, 84, 1115–1124. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.-X.; Kang, Y.-H.; Zhan, S.; Zhao, Z.-L.; Jin, S.-N.; Chen, C.; Zhang, L.; Shen, J.-Y.; Wang, C.-F.; Wang, G.-Q.; et al. Effect of Bacillus velezensis on Aeromonas veronii-Induced Intestinal Mucosal Barrier Function Damage and Inflammation in Crucian Carp (Carassius auratus). Front. Microbiol. 2019, 10, 2663. [Google Scholar] [CrossRef] [PubMed]
- Tachibana, L.; Telli, G.S.; Dias, D.D.C.; Gonçalves, G.S.; Guimarães, M.C.; Ishikawa, C.M.; Cavalcante, R.B.; Natori, M.M.; Alarcon, M.F.F.; Tapia-Paniagua, S.; et al. Bacillus subtilis and Bacillus licheniformis in diets for Nile tilapia (Oreochromis niloticus): Effects on growth performance, gut microbiota modulation and innate immunology. Aquac. Res. 2021, 52, 1630–1642. [Google Scholar] [CrossRef]
- Arani, M.M.; Salati, A.P.; Keyvanshokooh, S.; Safari, O. The effect of Pediococcus acidilactici on mucosal immune responses, growth, and reproductive performance in zebrafish (Danio rerio). Fish Physiol. Biochem. 2021, 47, 153–162. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.-P.; Dong, W.-L.; Chen, L.; Wang, Y.-M.; Muhammad, I.; Ju, A.-Q.; Shan, X.-F.; Ma, H.-X.; Kong, L.-C. Effects of dietary Lactobacillus plantarum C20015 on growth, immunity, and disease resistance in koi carp. Aquac. Int. 2020, 28, 1797–1809. [Google Scholar] [CrossRef]
- Abarike, E.D.; Jian, J.; Tang, J.; Cai, J.; Yu, H.; Lihua, C.; Jun, L. Influence of traditional Chinese medicine and Bacillus species (TCMBS) on growth, immune response and disease resistance in Nile tilapia, Oreochromis niloticus. Fish Shellfish. Immunol. 2019, 91, 417. [Google Scholar] [CrossRef]
- Midhun, S.J.; Neethu, S.; Arun, D.; Vysakh, A.; Divya, L.; Radhakrishnan, E.; Jyothis, M. Dietary supplementation of Bacillus licheniformis HGA8B improves growth parameters, enzymatic profile and gene expression of Oreochromis niloticus. Aquaculture 2019, 505, 289–296. [Google Scholar] [CrossRef]
- Abarike, E.D.; Cai, J.; Lu, Y.; Yu, H.; Chen, L.; Jian, J.; Tang, J.; Jun, L.; Kuebutornye, F.K. Corrigendum to “Effects of a commercial probiotic BS containing Bacillus subtilis and Bacillus licheniformis on growth, immune response and disease resistance in Nile tilapia, Oreochromis niloticus” [Fish Shellfish Immunol. 82 (2018) 229–238]. Fish Shellfish. Immunol. 2019, 84, 1180. [Google Scholar] [CrossRef] [PubMed]
- Aly, S.M.; Mohamed, M.F.; John, G. Effect of probiotics on the survival, growth and challenge infection in Tilapia nilotica (Oreochromis niloticus). Aquac. Res. 2008, 39, 647–656. [Google Scholar] [CrossRef]
- Thy, H.T.T.; Tri, N.N.; Quy, O.M.; Fotedar, R.; Kannika, K.; Unajak, S.; Areechon, N. Effects of the dietary supplementation of mixed probiotic spores of Bacillus amyloliquefaciens 54A, and Bacillus pumilus 47B on growth, innate immunity and stress responses of striped catfish (Pangasianodon hypophthalmus). Fish Shellfish. Immunol. 2017, 60, 391–399. [Google Scholar] [CrossRef]
- Ha, C.W.Y.; Lam, Y.Y.; Holmes, A.J. Mechanistic links between gut microbial community dynamics, microbial functions and metabolic health. World J. Gastroenterol. 2014, 20, 16498–16517. [Google Scholar] [CrossRef]
- Liu, W.; Yang, Y.; Zhang, J.; Gatlin, D.M.; Ringø, E.; Zhou, Z. Effects of dietary microencapsulated sodium butyrate on growth, intestinal mucosal morphology, immune response and adhesive bacteria in juvenile common carp (Cyprinus carpio) pre-fed with or without oxidised oil. Br. J. Nutr. 2014, 112, 15–29. [Google Scholar] [CrossRef]
- Montalban-Arques, A.; De Schryver, P.; Bossier, P.; Gorkiewicz, G.; Mulero, V.; Gatlin, D.M.I.; Galindo-Villegas, J. Selective Manipulation of the Gut Microbiota Improves Immune Status in Vertebrates. Front. Immunol. 2015, 6, 512. [Google Scholar] [CrossRef]
- Louis, P.; Hold, G.L.; Flint, H.J. The gut microbiota, bacterial metabolites and colorectal cancer. Nat. Rev. Genet. 2014, 12, 661–672. [Google Scholar] [CrossRef]
- Shapiro, H.; Thaiss, C.A.; Levy, M.; Elinav, E. The cross talk between microbiota and the immune system: Metabolites take center stage. Curr. Opin. Immunol. 2014, 30, 54–62. [Google Scholar] [CrossRef]
- Gomez-Gil, B.; Roque, A.; Turnbull, J.F. The use and selection of probiotic bacteria for use in the culture of larval aquatic organisms. Aquaculture 2000, 191, 259–270. [Google Scholar] [CrossRef]
- Picchietti, S.; Fausto, A.M.; Randelli, E.; Carnevali, O.; Taddei, A.R.; Buonocore, F.; Scapigliati, G.; Abelli, L. Early treatment with Lactobacillus delbrueckii strain induces an increase in intestinal T-cells and granulocytes and modulates immune-related genes of larval Dicentrarchus labrax (L.). Fish Shellfish. Immunol. 2009, 26, 368–376. [Google Scholar] [CrossRef]
- Chu, T.-W.; Chen, C.-N.; Pan, C.-Y. Antimicrobial status of tilapia (Oreochromis niloticus) fed Enterococcus avium originally isolated from goldfish intestine. Aquac. Rep. 2020, 17, 100397. [Google Scholar] [CrossRef]
- Feng, J.; Chang, X.; Zhang, Y.; Yan, X.; Zhang, J.; Nie, G. Effects of Lactococcus lactis from Cyprinus carpio L. as probiotics on growth performance, innate immune response and disease resistance against Aeromonas hydrophila. Fish Shellfish. Immunol. 2019, 93, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Chauhan, A.; Singh, R. Isolation and evaluation of putative probiotic strains from different teleost to prevent Pseudomonas aeruginosa infection in Cyprinus carpio. Aquac. Res. 2019, 50, 3616–3627. [Google Scholar] [CrossRef]
- Ghosh, B.; Cain, K.D.; Nowak, B.F.; Bridle, A.R. Microencapsulation of a putative probiotic Enterobacter species, C6-6, to protect rainbow trout, Oncorhynchus mykiss (Walbaum), against bacterial coldwater disease. J. Fish Dis. 2016, 39, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Puvanendran, V.; Rud, I.; Msw, B.; Arnesen, J.; Axelsson, L. Probiotic Carnobacterium divergens increase growth parameters and disease resistance in farmed Atlantic cod (Gadus morhua) larvae without influencing the microbiota. Aquaculture 2021, 532, 736072. [Google Scholar] [CrossRef]
- Salyers, A.A.; Whitt, D.D.; Whitt, D.D. Bacterial Pathogenesis: A Molecular Approach; American Society for Microbiology (ASM): Washington, DC, USA, 2011. [Google Scholar]
- Brunt, J.; Austin, B. Use of a probiotic to control lactococcosis and streptococcosis in rainbow trout, Oncorhynchus mykiss (Walbaum). J. Fish Dis. 2005, 28, 693–701. [Google Scholar] [CrossRef]
- Kesarcodi-Watson, A.; Kaspar, H.; Lategan, M.J.; Gibson, L. Probiotics in aquaculture: The need, principles and mechanisms of action and screening processes. Aquaculture 2008, 274, 1–14. [Google Scholar] [CrossRef]
- Kuebutornye, F.K.A.; Abarike, E.D.; Lu, Y.; Hlordzi, V.; Sakyi, M.E.; Afriyie, G.; Wang, Z.; Li, Y.; Xie, C.X. Mechanisms and the role of probiotic Bacillus in mitigating fish pathogens in aquaculture. Fish Physiol. Biochem. 2020, 46, 819–841. [Google Scholar] [CrossRef]
- Stein, T. Bacillus subtilis antibiotics: Structures, syntheses and specific functions. Mol. Microbiol. 2005, 56, 845–857. [Google Scholar] [CrossRef]
- Nadal, A.L.; Ikeda-Ohtsubo, W.; Sipkema, D.; Peggs, D.; McGurk, C.; Forlenza, M.; Wiegertjes, G.F.; Brugman, S. Feed, Microbiota, and Gut Immunity: Using the Zebrafish Model to Understand Fish Health. Front. Immunol. 2020, 11, 11. [Google Scholar] [CrossRef]
- Lee, M.-C.; Hsu, Y.-J.; Ho, H.-H.; Hsieh, S.-H.; Kuo, Y.-W.; Sung, H.-C.; Huang, C.-C. Lactobacillus salivarius Subspecies salicinius SA-03 is a New Probiotic Capable of Enhancing Exercise Performance and Decreasing Fatigue. Microorganisms 2020, 8, 545. [Google Scholar] [CrossRef] [PubMed]
- Wosinska, L.; Cotter, P.D.; O’Sullivan, O.; Guinane, C. The Potential Impact of Probiotics on the Gut Microbiome of Athletes. Nutrients 2019, 11, 2270. [Google Scholar] [CrossRef]
- Chen, Y.M.; Wei, L.; Chiu, Y.S.; Hsu, Y.J.; Tsai, T.Y.; Wang, M.F.; Huang, C.C. Lactobacillus plantarum TWK10 supplementation improves exercise performance and increases muscle mass in mice. Nutrients 2016, 8, 205. [Google Scholar] [CrossRef]
- Lee, M.-C.; Hsu, Y.-J.; Chuang, H.-L.; Hsieh, P.-S.; Ho, H.-H.; Chen, W.-L.; Chiu, Y.-S. In Vivo Ergogenic Properties of the Bifidobacterium longum OLP-01 Isolated from a Weightlifting Gold Medalist. Nutrients 2019, 11, 2003. [Google Scholar] [CrossRef] [PubMed]
- Barton, W.; Penney, N.C.; Cronin, O.; Garcia-Perez, I.; Molloy, M.G.; Holmes, E.; Shanahan, F.; Cotter, P.D.; O’Sullivan, O. The microbiome of professional athletes differs from that of more sedentary subjects in composition and particularly at the functional metabolic level. Gut Microbiota 2018, 67, 625–633. [Google Scholar] [CrossRef]
- Petersen, L.M.; Bautista, E.J.; Nguyen, H.; Hanson, B.M.; Chen, L.; Lek, S.H.; Sodergren, E.; Weinstock, G.M. Community characteristics of the gut microbiomes of competitive cyclists. Microbiome 2017, 5, 1–13. [Google Scholar] [CrossRef]
- Tovar-Ramírez, D.; Mazurais, D.; Gatesoupe, J.; Quazuguel, P.; Cahu, C.; Zambonino-Infante, J. Dietary probiotic live yeast modulates antioxidant enzyme activities and gene expression of sea bass (Dicentrarchus labrax) larvae. Aquaculture 2010, 300, 142–147. [Google Scholar] [CrossRef]
- Lamari, F.; Castex, M.; Larcher, T.; Ledevin, M.; Mazurais, D.; Bakhrouf, A.; Gatesoupe, F.-J. Comparison of the effects of the dietary addition of two lactic acid bacteria on the development and conformation of sea bass larvae, Dicentrarchus labrax, and the influence on associated microbiota. Aquaculture 2013, 376-379, 137–145. [Google Scholar] [CrossRef]
- Carnevali, O.; Maradonna, F.; Gioacchini, G. Integrated control of fish metabolism, wellbeing and reproduction: The role of probiotic. Aquaculture 2017, 472, 144–155. [Google Scholar] [CrossRef]
- Gioacchini, G.; Ciani, E.; Pessina, A.; Cecchini, C.; Silvi, S.; Rodiles, A.; Merrifield, D.L.; Olivotto, I.; Carnevali, O. Correction to: Effects of Lactogen 13, a New Probiotic Preparation, on Gut Microbiota and Endocrine Signals Controlling Growth and Appetite of Oreochromis niloticus Juveniles. Microb. Ecol. 2018, 76, 1075. [Google Scholar] [CrossRef]
- Mehrim, A.I.; Khalil, F.F.; Hassan, M.E. Hydroyeast Aquaculture®® as a reproductive enhancer agent for the adult Nile tilapia (Oreochromis niloticus Linnaeus, 1758). Fish Physiol. Biochem. 2014, 41, 371–381. [Google Scholar] [CrossRef] [PubMed]
- Gioacchini, G.; Giorgini, E.; Merrifield, D.L.; Hardiman, G.; Borini, A.; Vaccari, L.; Carnevali, O. Probiotics Can Induce Follicle Maturational Competence: The Danio rerio Case. Biol. Reprod. 2012, 86, 65. [Google Scholar] [CrossRef] [PubMed]
- Gioacchini, G.; Maradonna, F.; Lombardo, F.; Bizzaro, D.; Olivotto, I.; Carnevali, O. Increase of fecundity by probiotic administration in zebrafish (Danio rerio). Reproduction 2010, 140, 953–959. [Google Scholar] [CrossRef] [PubMed]
- Falcinelli, S.; Picchietti, S.; Rodiles, A.; Cossignani, L.; Merrifield, D.L.; Taddei, A.R.; Maradonna, F.; Olivotto, I.; Gioacchini, G.; Carnevali, O. Lactobacillus rhamnosus lowers zebrafish lipid content by changing gut microbiota and host transcription of genes involved in lipid metabolism. Sci. Rep. 2015, 5, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Geraylou, Z.; Vanhove, M.P.M.; Souffreau, C.; Rurangwa, E.; Buyse, J.; Ollevier, F. In vitroselection and characterization of putative probiotics isolated from the gut of Acipenser baerii (Brandt, 1869). Aquac. Res. 2012, 45, 341–352. [Google Scholar] [CrossRef]
- Kavitha, M.; Raja, M.; Perumal, P. Evaluation of probiotic potential of Bacillus spp. isolated from the digestive tract of freshwater fish Labeo calbasu (Hamilton, 1822). Aquac. Rep. 2018, 11, 59–69. [Google Scholar] [CrossRef]
- Linh, N.T.H.; Sakai, K.; Taoka, Y. Screening of lactic acid bacteria isolated from fermented food as potential probiotics for aquacultured carp and amberjack. Fish. Sci. 2018, 84, 101–111. [Google Scholar] [CrossRef]
- EFSA. 2008 Annual report on pesticide residues according to Article 32 of Regulation (EC) No 396/2005. EFSA J. 2010, 8, 1646. [Google Scholar] [CrossRef]
- Pisano, M.B.; Viale, S.; Conti, S.; Fadda, M.E.; Deplano, M.; Melis, M.P.; Deiana, M.; Cosentino, S. Preliminary Evaluation of Probiotic Properties of Lactobacillus Strains Isolated from Sardinian Dairy Products. BioMed Res. Int. 2014, 2014, 1–9. [Google Scholar] [CrossRef]
- Harty, D.; Oakey, H.; Patrikakis, M.; Hume, E.; Knox, K. Pathogenic potential of lactobacilli. Int. J. Food Microbiol. 1994, 24, 179–189. [Google Scholar] [CrossRef]
- Jena, P.K.; Trivedi, D.; Thakore, K.; Chaudhary, H.; Giri, S.S.; Seshadri, S. Isolation and characterization of probiotic properties of Lactobacilli isolated from rat fecal microbiota. Microbiol. Immunol. 2013, 57, 407–416. [Google Scholar] [CrossRef] [PubMed]
- Kaushik, J.K.; Kumar, A.; Duary, R.K.; Mohanty, A.K.; Grover, S.; Batish, V.K. Functional and Probiotic Attributes of an Indigenous Isolate of Lactobacillus plantarum. PLoS ONE 2009, 4, e8099. [Google Scholar] [CrossRef]
- Papadimitriou, K.; Zoumpopoulou, G.; Folign, B.; Alexandraki, V.; Kazou, M.; Pot, B.; Tsakalidou, E. Discovering probiotic microorganisms: In vitro, in vivo, genetic and omics approaches. Front. Microbiol. 2015, 6, 58. [Google Scholar] [CrossRef]
- Laparra, J.M.; Sanz, Y. Comparison ofin vitromodels to study bacterial adhesion to the intestinal epithelium. Lett. Appl. Microbiol. 2009, 49, 695–701. [Google Scholar] [CrossRef] [PubMed]
- Mosso, A.; Jimenez, M.; Vignolo, G.; Leblanc, J.; Samman, N. Increasing the folate content of tuber based foods using potentially probiotic lactic acid bacteria. Food Res. Int. 2018, 109, 168–174. [Google Scholar] [CrossRef] [PubMed]
- Solopova, A.; Bottacini, F.; Degli Esposti, E.V.; Amaretti, A.; Raimondi, S.; Rossi, M.; Van Sinderen, D. Riboflavin Biosynthesis and Overproduction by a Derivative of the Human Gut Commensal Bifidobacterium longum subsp. infantis ATCC. Front. Microbiol. 2020, 11, 11. [Google Scholar] [CrossRef] [PubMed]
- Page, J.W.; Andrews, J.W.; Murai, T.; Murray, M.W. Hydrogen ion concentration in the gastrointestinal tract of channel catfish. J. Fish Biol. 1976, 8, 225–228. [Google Scholar] [CrossRef]
- Hoehne-Reitan, K.; Kjorsvik, E.; Reitan, K.I. Development of the pH in the intestinal tract of larval turbot. Mar. Biol. 2001, 139, 1159–1164. [Google Scholar]
- Kaktcham, P.M.; Temgoua, J.-B.; Zambou, F.N.; Diaz-Ruiz, G.; Wacher, C.; Pérez-Chabela, M.D.L. In Vitro Evaluation of the Probiotic and Safety Properties of Bacteriocinogenic and Non-Bacteriocinogenic Lactic Acid Bacteria from the Intestines of Nile Tilapia and Common Carp for Their Use as Probiotics in Aquaculture. Probiotics Antimicrob. Proteins 2018, 10, 98–109. [Google Scholar] [CrossRef]
- Gunzburg, W.H.; Aung, M.M.; Toa, P.; Ng, S.; Read, E.; Tan, W.J.; Brandtner, E.M.; Dangerfield, J.; Salmons, B. Efficient protection of microorganisms for delivery to the intestinal tract by cellulose sulphate encapsulation. Microb. Cell Factories 2020, 19, 1–14. [Google Scholar] [CrossRef]
- Gharibzahedi, S.M.T.; Smith, B. Legume proteins are smart carriers to encapsulate hydrophilic and hydrophobic bioactive compounds and probiotic bacteria: A review. Compr. Rev. Food Sci. Food Saf. 2021, 20, 1250–1279. [Google Scholar] [CrossRef] [PubMed]
- Kwiecień, I.; Kwiecień, M. Application of Polysaccharide-Based Hydrogels as Probiotic Delivery Systems. Gels 2018, 4, 47. [Google Scholar] [CrossRef] [PubMed]
- Tan, E.W.; Tan, K.Y.; Phang, L.V.; Kumar, P.V.; In, L.L.A. Enhanced gastrointestinal survivability of recombinant Lactococcus lactis using a double coated mucoadhesive film approach. PLoS ONE 2019, 14, e0219912. [Google Scholar] [CrossRef]
- Rosas-Ledesma, P.; León-Rubio, J.M.; Alarcón, F.J.; Moriñigo, M.A.; Balebona, M.C. Calcium alginate capsules for oral administration of fish probiotic bacteria: Assessment of optimal conditions for encapsulation. Aquac. Res. 2011, 43, 106–116. [Google Scholar] [CrossRef]
- Lee, H.; Ko, G. Antiviral effect of vitamin A on norovirus infection via modulation of the gut microbiome. Sci. Rep. 2016, 6, 25835. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wuertz, S.; Schroeder, A.; Wanka, K.M. Probiotics in Fish Nutrition—Long-Standing Household Remedy or Native Nutraceuticals? Water 2021, 13, 1348. https://doi.org/10.3390/w13101348
Wuertz S, Schroeder A, Wanka KM. Probiotics in Fish Nutrition—Long-Standing Household Remedy or Native Nutraceuticals? Water. 2021; 13(10):1348. https://doi.org/10.3390/w13101348
Chicago/Turabian StyleWuertz, Sven, Arne Schroeder, and Konrad M. Wanka. 2021. "Probiotics in Fish Nutrition—Long-Standing Household Remedy or Native Nutraceuticals?" Water 13, no. 10: 1348. https://doi.org/10.3390/w13101348
APA StyleWuertz, S., Schroeder, A., & Wanka, K. M. (2021). Probiotics in Fish Nutrition—Long-Standing Household Remedy or Native Nutraceuticals? Water, 13(10), 1348. https://doi.org/10.3390/w13101348






